Abstract
This study aimed to increase maltose fermentation in industrial baker’s yeast to increase its leavening properties. To this end, we overexpressed MAL62 encoding alpha-glucosidase (maltase) and deleted MIG1 encoding a transcriptional repressor that regulates MAL gene expression. Strain overexpressing MAL62 showed 46.3 % higher alpha-glucosidase activity and enhanced leaving activity than the parental strain when tested in glucose–maltose low sugar model liquid dough (LSMLD). Deleting MIG1 was much less effective, but it could further strengthen leavening properties in a strain overexpressing MAL62. The relationship between maltose permease and alpha-glucosidase was further dissected by transforming the two genes. The results indicated that without increasing the maltose permease activity, maltose metabolism could also be enhanced by the increased alpha-glucosidase activity. Previous strategies for strain improvement have targeted the enhancement of alpha-glucosidase and maltose permease activities in concert. Our results suggest that increasing alpha-glucosidase activity is sufficient to improve maltose fermentation in lean dough.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The main component of lean dough is starch, which is hydrolyzed to maltose by amylases. Therefore, to improve the leavening ability in lean dough, it is essential to enhance maltose metabolism [2, 24]. Yeast cells require at least one of the five unlinked MAL loci (MAL1 through MAL4 and MAL6) in order to utilize maltose. Therefore, the key factor in increasing leavening ability is regulation of the MAL system [20]. A typical MAL locus consists of a MALx1 (MALxT) gene (where x is the locus), encoding maltose permease, a MALx2 (MALxS) gene, coding for alpha-glucosidase (maltase), and a MALx3 (MALxR) gene, encoding a positive regulatory protein [9]. In previous studies, maltose permease was considered to have a crucial role in both maltose uptake and maltose induction of the MAL genes [4, 6, 25]. Hence, many improvements intended to increase the leavening ability of yeast have concentrated on the modification of maltose permease [21].
The presence of glucose inhibits the transcription of genes specifically related to the consumption of other carbon sources, thereby delaying maltose uptake [18, 22]. An effective alleviation of glucose control would help to achieve a better process economy for bread-making. Several lines of evidence suggest that MIG1 encodes a repressor protein (MIG1p), which represses the transcription of all three MAL genes by binding upstream of these genes [3, 19]. Previous studies have shown that deleting MIG1 can enhance transcription of the MAL gene, thereby resulting in increased synthesis of proteins involved in maltose utilization [12].
Our objectives are (1) to gain better insight into the relationship between alpha-glucosidase and maltose permease at different extracellular maltose concentrations and (2) to study the effects of MAL62 overexpression and MIG1 deletion in industrial baker’s yeast.
Materials and methods
Strains, plasmids, and conditions of culture and fermentation
The genetic properties of all Saccharomyces cerevisiae strains and plasmids used in the present study are summarized in Table 1. Industrial baker’s yeast strain BY14 was obtained from the Yeast Collection Center of the Tianjin Key Laboratory of Industrial Microbiology.
Recombinant DNA was amplified in Escherichia coli DH5α. Transformants were grown in Luria–Bertani medium (10 g l−1 tryptone, 5 g l−1 yeast extract, and 10 g l−1 NaCl) with 100 μg ml−1 ampicillin. The plasmid was obtained using Plasmid Mini Kit II (D6945, Omega, USA).
Yeast strains were routinely cultivated in yeast extract peptone dextrose (YEPD) medium (10 g l−1 yeast extract, 20 g l−1 peptone, and 20 g l−1 glucose) in a shake-flask culture at 30 °C and shaken at 200 rpm in a rotary shaker. Cells were grown to stationary phase and inoculated into a LSMLD fermentation medium (2.5 g l−1 (NH4)2SO4, 5 g l−1 urea, 16 g l−1 KH2PO6, 5 g l−1 Na2HPO4, 0.6 g l−1 MgSO4, 0.0225 g l−1 nicotinic acid, 0.005 g l−1 Ca-pantothenate, 0.0025 g l−1 thiamine, 0.00125 g l−1 pyridoxine, 0.001 g l−1 riboflavin, and 0.0005 g l−1 folic acid) containing one of three specified carbon sources to further investigate the degree of glucose repression and determine the relationship between two MAL enzymes under different extracellular maltose concentrations. The C-mol of the three specified carbon sources (38 g l−1 maltose, 40 g l−1 glucose, and 33.25 g l−1 maltose mixed with 5 g l−1 glucose) was 1.33 C-mol. Cells were harvested by centrifugation (4 °C, 5,000 rpm, 5 min) and washed twice with sterile water at 4 °C for use in succeeding experiments. Episomal plasmids and yeast genomes were obtained using a yeast plasmid kit (PD1211-02, Omega, USA) and yeast DNA kit (D3370-01, Omega, USA), respectively.
Plasmid construction and yeast transformation
PCR primers used in the current work are listed in Table 2. Plasmid Yep-CPM, an episomal plasmid with MAL62 under the control of the PGK promoter, was constructed as follows: an XhoI fragment of MAL62 that was amplified with the primers MAL62-F and MAL62-R from genomes of the parent strain (BY14a) was cloned into vector pPGK1, resulting in plasmid pPGKM. A BamHI fragment of pPGKM containing the entire MAL62, the entire promoter, and the entire terminator of PGK1 was amplified with the primers PGK-F and PGK-R and then cloned into the BamHI linearized Yep-C plasmid, resulting in Yep-CPM plasmid. Yep-C is the control vector for Yep-CPM and contains the Cu resistant gene as the dominant selection marker when integrated into yeast. The pKAB plasmid used for deletion of MIG1 was created by the following process: an EcoRI upstream homologous fragment of the MIG1 gene, named A and amplified with A-F and A-R primers, was inserted into the multicopy episomal shuttle vector pUC19, resulting in the pUC19-A plasmid. The HindIII downstream homologous fragment of MIG1, named B and amplified with primers B–F and B-R, was also isolated from genomic DNA of the parent strain (BY14a) and subcloned into the HindIII site of pUC19-A, resulting in a plasmid named pUC-AB. The marker excised with primers Kan-F and Kan-R from pUG6 as a KpnI fragment was inserted into pUC-AB to obtain a plasmid called pKAB.
Yeast transformation was carried out using a lithium acetate procedure described previously [15]. Selection of strains BY-C, BY14a + MAL62, and BY14a + MAL62-MIG1 were performed in YEPD medium supplemented with 4 mmol l−1 CuSO4. Selection of BY14a-MIG1 strain was performed in YEPD medium supplemented with 0.8 mg ml−1 G418. After selection, recombinant strains were verified by primers Cu-F and Cu-R (BY-C), primers MIG1-U and MIG1-D (BY14a-MIG1), and primers PGK-F and PGK-R (BY14a + MAL62, BY14a + MAL62-MIG1).
Determination of cell dry weight and biomass yield
Nitrocellulose filters with a pore size of 0.45 mm (Gelman Sciences, Ann Arbor, MI, USA) were pre-dried in a microwave oven at 150 W for 10 min and subsequently weighed. Next, 5 ml of the cell culture was filtered, washed three times with 5 ml of distilled water, and dried at 110 °C for 24 h. Cell dry weight was determined with an accuracy of 2 %. Biomass yield was determined from the slopes of plots of biomass dry weight versus consumed sugar during exponential growth.
Analysis of sugars
For measurements of extracellular glucose and maltose, the cultivation medium was sampled, immediately filtered through a 0.45-μm-pore-size cellulose acetate filter (Millipore Corp., Danvers, MA, USA), and subsequently frozen and kept at −20 °C until analysis.
The equipment used in the experiment included a differential refractometer detector (Waters 410 RI), Aminex HPX-87H columns (Bio-Rad, Hercules, CA, USA), and an HPLC pump (Waters 515). Samples were measured at 65 °C with 5 mM H2SO4 as the mobile phase at a flow rate of 0.6 ml min−1 [8]. Concentrations were determined with accuracy better than 5 %.
Determination of leavening ability of lean dough
The leavening ability of lean dough of yeast was determined according to the Chinese National Standards for yeast used in food processing. Lean dough was composed of 280 g of flour, 150 ml of water, 4 g of salt, and 9 g of fresh yeast. The dough was evenly and rapidly stirred for 5 min at (30 ± 0.2) °C and placed in a fermentograph box (Type JM451, Sweden). CO2 production was recorded for 60 min at 30 °C. Experiments were conducted three times.
Enzyme activity assays
Crude extracts were prepared using the Salema-Oom method to determine enzyme activities [24]. Alpha-glucosidase and maltose permease activities were determined following the Houghton-Larsen method [11]. Standard errors were less than 10 %.
Results
Overexpressed MAL62 gene results in enhanced maltose fermentation in three LSMLD
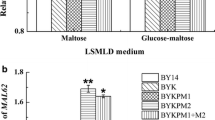
We tested the effects of MAL62 gene overexpression on maltose fermentation. Alpha-glucosidase and maltose permease activities were assayed in three LSMLD, and the amounts of CO2 produced in lean dough were measured. Alpha-glucosidase activities of strains BY14a + MAL62 and BY14a + MAL62-MIG1 were approximately 46.3 % and 82.2 %, respectively, higher than the parental strain (BY14a) in the glucose-maltose LSMLD (Fig. 1c), resulting in 33.9 and 39.5 % increases in 1-h CO2 production (Fig. 2) and 34 and 40 % enhancements in leavening ability (Table 3), respectively. Similar effects, but with higher levels of alpha-glucosidase activity, were observed in experiments utilizing the maltose LSMLD (Fig. 1b). In the glucose LSMLD, the activities of alpha-glucosidase in all strains were very low. However, an increase of at least 52.8 % (Fig. 1a) was observed in strains carrying the overexpressed MAL62. Both strains overexpressing the MAL62 gene showed enhanced alpha-glucosidase activities in the three LSMLD but displayed no increase in maltose permease activity (data not shown). These results suggest that, although it does not bring about increases in maltose permease activity, overexpression of the MAL62 gene could positively affect the level of alpha-glucosidase activity and production of gas.
Alpha-glucosidase activity and concentration of residual sugar in four yeast strains during shaking cultivation in three LSMLD: a glucose, b maltose, and c glucose-maltose. Filled square: residual sugar concentration of parent strain BY14a; filled circle: residual sugar concentration of BY14a-MIG1; Filled triangle: residual sugar concentration of BY14a + MAL62; Filled star: residual sugar concentration of BY14a + MAL62-MIG1; open square: alpha-glucosidase activity of parent strain BY14a; open circle: alpha-glucosidase activity of BY14a-MIG1; open triangle: alpha-glucosidase activity of BY14a + MAL62; and open star: alpha-glucosidase activity of BY14a + MAL62-MIG1. Data are averages from three independent experiments and error bars represent ± SD. BY-C was used as the blank control for the BY14a + MAL62 strain to avoid the possible effects of an empty vector on enzyme activities, fermentation characteristics, and leavening activity. Since no significant distinctions were observed (data not shown), BY14a was used as the control strain for the BY14a + MAL62 strain
MIG1 deletion was much less effective than MAL62 overexpression in enhancing maltose fermentation
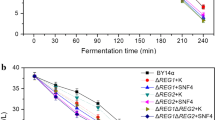
MIG1 is a major mediator of glucose repression in S. cerevisiae [1]. We measured alpha-glucosidase and maltose permease activities in the MIG1-deleted and parent yeast strains to examine whether MIG1 deletion is effective for MAL gene derepression. Loss of MIG1 in strain BY14a-MIG1 led to a slightly elevated level of alpha-glucosidase activity in three conditions (Fig. 1a–c) but no effect on maltose permease activity (data not shown). These findings confirm the role of MIG1 in the glucose repression of alpha-glucosidase expression in industrial baker’s yeast. In the glucose-maltose LSMLD, as glucose was consumed, a reduction in maltose permease activity was observed in the BY14a-MIG1 strain (Fig. 3). This result indicates that deletion of MIG1 in the industrial yeast strain does not entirely alleviate glucose control of the genes involved in maltose utilization.
Alpha-glucosidase activity, maltose permease activity, and concentration of residual sugar in parent strains BY14a and BY14a-MIG1 during shaking cultivation in the glucose-maltose LSMLD. Filled square: residual maltose concentration of parent strain BY14a; Filled triangle: residual maltose concentration of BY14a-MIG1; open square: alpha-glucosidase activity of parent strain BY14a; open triangle: alpha-glucosidase activity of BY14a-MIG1; open diamond: maltose permease activity of parent strain BY14a; and open inverted triangle: maltose permease activity of BY14a-MIG1. Data are averages from three independent experiments and error bars represent ± SD
Fermentation characteristics
We explored the physiological characteristics (specific growth rate and biomass yield) and leavening ability of four yeast strains to investigate their fermentation characteristics (Table 3). The physiological characteristics of strain BY14a-MIG1 were strongly affected by MIG1 deletion compared with the parent strain, BY14a. In the BY14a-MIG1 strain, less tightly controlled maltose metabolism increased the specific growth rate from 0.29 h−1 to 1.21 h−1 and the biomass yield from 0.62 g C-mol−1 to 0.68 g C-mol−1 in comparison to the parent strain. In contrast, overexpression of the MAL62 gene in strain BY14a + MAL62 had a slightly negative physiological effect and reduced the biomass yield from 0.62 g C-mol−1 to 0.58 g C-mol−1. Two strains overexpressing MAL62, BY14a + MAL62 and BY14a + MAL62-MIG1, showed significantly higher leavening abilities than strain BY14a-MIG1 (Table 3). These results suggest that only strains with a marked increase in alpha-glucosidase activity exhibit increased leavening ability.
Discussion
Previous studies have reported that maltose permease is the determining factor in leavening lean dough [6, 13]. In this study, we showed that maltose permease is less necessary than alpha-glucosidase, which is capable of enhancing maltose fermentation under any maltose concentration in LSMLD (i.e., glucose, maltose, or glucose-maltose). When cultivated in glucose medium and no extracellular maltose is available, intracellular maltose is sufficient to induce maltose-utilizing gene expression [25], rendering maltose transportation by maltose permease unnecessary. When yeast cells are grown in high extracellular maltose concentrations, alpha-glucosidase activity could also increase due to the presence of distinct maltose uptake systems that differ from maltose permease [5, 7, 23, 25]. Maltose permease does not play an exclusive role in either triggering MAL gene expression or maltose uptake. Hence, only overexpressed MAL62 could respond to maltose and bring about further increases in alpha-glucosidase activity. Furthermore, the increased alpha-glucosidase activity in yeast could have a strong correlation with the ability of the strain to leaven lean dough. These results support the conclusion of Jiang [15], who believed that alpha-glucosidase is the essential enzyme in maltose fermentation for high extracellular maltose concentrations. In this study, we verified that regardless of the extracellular maltose concentration, alpha-glucosidase, rather than maltose permease, is the most relevant feature in maltose fermentation.
Glucose, which is most commonly present in industrial carbon sources, delays the uptake of sugars and prolongs the production process unnecessarily [16]. Effective alleviation of glucose repression, or the rapid transition from glucose to maltose metabolism, is essential to improve the leavening ability of yeast during bread-making. In laboratory strains, glucose repression could partially be relieved by deletion of MIG1 [3, 19], whereas in industrial strains, MIG1 deletion was much less effective to yields high levels of alpha-glucosidase and maltose permease activities [10, 14, 26, 27]. These results suggest that MAL gene control in industrial baker’s yeast occurs through a dual-level control mechanism (i.e., MIG1-dependent and MIG1-independent) [16]; thus, deletion of MIG1 in industrial yeast strains cannot completely eliminate glucose repression.
MAL62 overexpression negatively affected the specific growth rate and biomass yields. Deletion of MIG1 could compensate for the reduction in specific growth rate and biomass yield of the BY14a + MAL62 strain. BY14a + MAL62-MIG1 can readily ferment maltose and instantly produce CO2 when it enters the lean dough. These characteristics are consistent with what is required of a strain used commercially in terms of yield and leavening ability.
Based on the results of the present study, metabolic engineering of glucose derepression appears to be a greater challenge than maltose fermentation. Baker’s yeast containing overexpressed-MAL62 and featuring a deleted MIG1 genetic background that provides high levels of alpha-glucosidase activity is sufficient to enable the strain to rapidly ferment maltose and enhance its leavening properties. Such a strain could be useful for leavening lean dough.
References
Ahuatzi D, Riera A, Peláez R, Herrero P, Moreno F (2007) Hxk2 regulates the phosphorylation state of Mig1 and therefore its nucleocytoplasmic distribution. J Biol Chem 282:4485–4493
Bell PJ, Higgins VJ, Attfield PV (2001) Comparison of fermentative capacities of industrial baking and wild-type yeasts of the species Saccharomyces cerevisiae in different sugar media. Lett Appl Microbiol 32:224–229
Cao HL, Yue M, Li SG, Bai XF, Zhao XM, Du YG (2011) The impact of MIG1 and/or MIG2 disruption on aerobic metabolism of succinate dehydrogenase negative Saccharomyces cerevisiae. Appl Microbiol Biotechnol 89:733–738
Day RE, Higgins VJ, Rogers PJ, Dawes IW (2002) Characterization of the putative maltose transporters encoded by YDL247w and YJR160c. Yeast 19:1015–1027
Dietvorst J, Walsh MC, van Heusden GP, Steensma HY (2010) Comparison of the MTT1- and MAL31-like maltose transporter genes in lager yeast strains. FEMS Microbiol Lett 310:152–157
Goldenthal MJ, Vanoni M, Buchferer B, Marmur J (1987) Regulation of MAL gene expression in yeast: gene dosage effects. Mol Gen Genet 209:508–517
Hach A, Hon T, Zhang L (2000) The coiled coil dimerization element of the yeast transcriptional activator Hap1, a Gal4 family member, is dispensable for DNA binding but differentially affects transcriptional activation. J Biol Chem 275:248–254
Hauf J, Zimmermann FK, Müller S (2000) Simultaneous genomic overexpression of seven glycolytic enzymes in the yeast Saccharomyces cerevisiae. Enzyme Microb Technol 26:688–698
Higgins VJ, Braidwood M, Bell P, Bissinger P, Dawes IW, Attfield PV (1999) Genetic evidence that high noninduced maltase and maltose permease activities, governed by MALx3-encoded transcriptional regulators, determine efficiency of gas production by baker’s yeast in unsugared dough. Appl Environ Microbiol 65:680–685
Higgins VJ, Braidwood M, Bissinger P, Dawes IW, Attfield PV (1999) Leu343Phe substitution in the Malx3 protein of Saccharomyces cerevisiae increases the constitutivity and glucose insensitivity of MAL gene expression. Curr Genet 35:491–498
Houghton-Larsen J, Brandt A (2006) Fermentation of high concentrations of maltose by Saccharomyces cerevisiae is limited by the COMPASS methylation complex. Appl Environ Microbiol 72:7176–7182
Hu Z, Nehlin JO, Ronne H, Michels CA (1995) MIG1-dependent and MIG1-independent glucose regulation of MAL gene expression in Saccharomyces cerevisiae. Curr Genet 28:258–266
Hu Z, Yue Y, Jiang H, Zhang B, Sherwood PW, Michels CA (2000) Analysis of the mechanism by which glucose inhibits maltose induction of MAL gene expression in Saccharomyces. Genetics 154:121–132
Jiang H, Tatchell K, Liu S, Michels CA (2000) Protein phosphatase type-1 regulatory subunits Reg1p and Reg2p act as signal transducers in the glucose-induced inactivation of maltose permease in Saccharomyces cerevisiae. Mol Gen Genet 263:411–422
Jiang TX, Xiao DG, Gao Q (2008) Characterisation of maltose metabolism in lean dough by lagging and non-lagging baker’s yeast strains. Ann Microbiol 58:655–660
Klein CJL, Olsson L, Nielsen J (1998) Glucose control in Saccharomyces cerevisiae: the role of MIG1 in metabolic functions. Microbiol Sgm 144:13–24
Lilly M, Lambrechts MG, Pretorius IS (2000) Effect of increased yeast alcohol acetyltransferase activity on flavor profiles of wine and distillates. Appl Environ Microbiol 66:744–753
Lucero P, Herweijer M, Lagunas R (1993) Catabolite inactivation of the yeast maltose transporter is due to proteolysis. FEBS Lett 333:165–168
Nehlin JO, Carlberg M, Ronne H (1991) Control of yeast GAL genes by MIG1 repressor: a transcriptional cascade in the glucose response. EMBO J 10:3373–3377
Oda Y, Ouchi K (1990) Role of the yeast maltose fermentation genes in CO2 production rate from sponge dough. Food Microbiol 7:43–47
Peinado JM, Barbero A, van Uden N (1987) Repression and inactivation by glucose of the maltose transport system of Candida utilis. Appl Microbiol Biotechnol 26:154–157
Randez-Gil F, Sanz P (1994) Construction of industrial baker’s yeast strains able to assimilate maltose under catabolite repression conditions. Appl Microbiol Biotechnol 42:581–586
Richter K, Buchner J (2001) Hsp90: chaperoning signal transduction. J Cell Physiol 188:281–290
Salema-Oom M, de Sousa HR, Assuncao M, Goncalves P, Spencer-Martins I (2011) Derepression of a baker’s yeast strain for maltose utilization is associated with severe deregulation of HXT gene expression. J Appl Microbiol 110:364–374
Wang X, Bali M, Medintz I, Michels CA (2002) Intracellular maltose is sufficient to induce MAL gene expression in Saccharomyces cerevisiae. Eukaryot Cell 1:696–703
Wanke V, Vavassori M, Thevelein JM, Tortora P, Vanoni M (1997) Regulation of maltose utilization in Saccharomyces cerevisiae by genes of the RAS/protein kinase A pathway. FEBS Lett 402:251–255
Westergaarda SL, Broa C, Olssona L, Nielsena J (2004) Elucidation of the role of Grr1p in glucose sensing by Saccharomyces cerevisiae through genome-wide transcription analysis. FEMS Yeast Res 5:193–204
Acknowledgments
The current study was financially supported by the National Natural Science Foundation of China (31000043), Major Project of Research Program on Applied Fundamentals and Advanced Technologies of Tianjin (10JCZDJC16700), and program for Changjiang Scholars and Innovative Research Team in University (IRT1166).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, X., Zhang, C., Dong, J. et al. Enhanced leavening properties of baker’s yeast overexpressing MAL62 with deletion of MIG1 in lean dough. J Ind Microbiol Biotechnol 39, 1533–1539 (2012). https://doi.org/10.1007/s10295-012-1144-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-012-1144-7