Abstract
Breast cancer is the most prevalent malignancy in the US and the third highest cause of cancer-related mortality worldwide. Regular mammography screening has been attributed with doubling the rate of early cancer detection over the past three decades, yet estimates of mammographic accuracy in the hands of experienced radiologists remain suboptimal with sensitivity ranging from 62 to 87% and specificity from 75 to 91%. Advances in machine learning (ML) in recent years have demonstrated capabilities of image analysis which often surpass those of human observers. Here we present two novel techniques to address inherent challenges in the application of ML to the domain of mammography. We describe the use of genetic search of image enhancement methods, leading us to the use of a novel form of false color enhancement through contrast limited adaptive histogram equalization (CLAHE), as a method to optimize mammographic feature representation. We also utilize dual deep convolutional neural networks at different scales, for classification of full mammogram images and derivative patches combined with a random forest gating network as a novel architectural solution capable of discerning malignancy with a specificity of 0.91 and a specificity of 0.80. To our knowledge, this represents the first automatic stand-alone mammography malignancy detection algorithm with sensitivity and specificity performance similar to that of expert radiologists.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most prevalent malignancy in the US, accounting for over 230,000 new diagnoses and approximately 40,000 deaths annually [1]. Regular mammography screening has been attributed with doubling the rate of early cancer detection and has been credited with decreasing breast cancer mortality by up to 30% over the past three decades [2, 3]. Estimates of sensitivity and specificity in the hands of experienced radiologists currently range from 62 to 87% and 75 to 91%, respectively [4,5,6,7,8].
Computer aided detection (CAD) for mammography was first approved by the Food and Drug Administration (FDA) in 1998. CAD software functions essentially as a “second reader” to the interpreting radiologist. Early studies demonstrated increases of 19–23% in breast cancer detection rate with CAD utilization, resulting in reimbursement qualification and widespread adoption in the US [9,10,11]. Despite subsequent upgrades in traditional CAD algorithms, its clinical utility has remained controversial. The most definitive study to date pooled data from mammography registries of over 500,000 mammograms performed between 2003 and 2009 and found no added benefit of CAD in cancer detection or diagnostic accuracy for screening mammography [5].
Traditional CAD algorithms deploy conventional computer vision technologies based upon detection of hand-crafted imaging features broadly categorized into masses or micro-calcifications. In contrast, current machine learning (ML) methods are based upon feature discovery within samples of ground truth-validated images. ML has made substantial advances in feature recognition in natural-world images, often superseding that of human observers. Fundamental differences of data acquisition and content have limited the transferability of ML image algorithms to the domain of radiology. We present a ML based mammographic malignancy detection algorithm which utilizes novel data enhancement techniques and analytics architecture to achieve stand-alone accuracy similar to that reported for expert radiologists.
Data
Datasets were created using digital mammography images from The Digital Database for Screening Mammography (DDSM), which includes more than 6000 digital mammographic images evenly split between those with and without malignancy [12] and the proprietary Zebra Mammography Dataset (ZMDS), a dataset of 1739 full-sized mammograms divided into training, validation, and “untouched” test sets comprised of nearly equal numbers of malignant and nonmalignant samples.
Both databases represent a random variation of mammographic breast density. The “positive” samples contain malignancy and “negative” samples represent images of either normal breast tissue or tissue with definitively benign anomalies. Positive ground truth was defined by biopsy proven pathology. Negative samples were defined by pathology or at least 2 years of stable imaging follow-up.
The inclusion of a distinct benign class allowed for the generation of a tri-categorical network with the goal of differentiating not only normal from abnormal breast tissue, but further classifying abnormalities into those which are suspicious for malignancy and those which are not.
Using the DDSM and ZMDS, we trained experimental network designs, testing single image sensitivity, and specificity against a test set. Initial pathfinding experiments were designed to define the optimal preprocessing enhancement methods and parameters, testing combinations of parameter alterations as input to ensembles of shallow convolutional neural networks (CNN) [13] [14]. CNNs are a powerful means of extracting features from images (among other applications) and ultimately in performing automatic image classification. The depth of a CNN refers to the number of processing layers which input data must go through before the final layer of extracted features is passed to the discriminating classifier at the top of the network. We utilized a “shallow” network of six layers to identify optimal pre-processing measures. Pre-processed data was then input into the dual deep (greater than 20 layers) CNN networks. Searching the space of possible preprocessing methods directly against such a deep network would have prohibitively high computational cost.
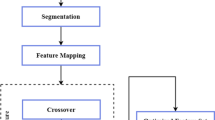
The parameter space (the set of all possible settings for the parameters) of the enhancement parameters, which described the combination of image processes, was searched by evolutionary selection of the ensemble members. Multiple functions were selected in the vast parameter space, each with a selection of inputs and arguments. The space was permutation dependent with specific function ordering, as the output of one function affected each subsequent function.
Most experiments used a maximum of eight possible preprocessing manipulations, including empty functions (NoOps), indicating a redundant portion to the “genome.” The optimal pre-processing combinations were selected by an evolutionary process: when a new shallow CNN with its associated input preprocessing method improved the ensemble’s top 1 precision, the network was added as a member. The genome-like description of its preprocessing method was added to a pool of parameter strings from which subsequent candidate members would be bred (the parameter string describing the input preprocessing method, constructed by splicing sections of previously successful preprocessing genomes).
Common computer vision methods were possible functions the “genome” could express. The genome also determining the parameters used for each function, whether to take the working memory buffer image, or the original image as input, and how to merge result to the working memory. Some methods available to the genetic preprocessing mechanism included the following:
-
gaussian blur
-
Otsus threshold masking
-
OpenCV 2’s Canny edge detection
-
CLAHE as implemented by OpenCV 2, with window and clipping parameters both chosen by the genome.
-
masking by hue matching (hue and threshold determined by the genome)
-
masking by shannon entropy threshold
Note that some functions were masking functions. These masks were then potentially used by other functions to apply processing to a selected region defined by that mask.
The resulting preprocessing evolved toward some common processing patterns. Initial experiments were performed on the less challenging task of classifying birds (Fig. 1). It was shown that sophisticated image manipulations could emerge from this process.
For our intended challenge of breast malignancy risk classification and localization, pathfinding experiments identified contrast limited adaptive histogram equalization (CLAHE) as consistently among the most useful enhancements during ensemble evolution. Unprocessed shallow network ensembles yielded accuracy of 86% in classifying malignant from non-malignant mammograms. In comparison, the addition of false color enhancement across the RGB spectrum by employing broad window resolution with low clipping value at the red channel, intermediate at the green and fine resolution with the highest limiting at the blue (see Scheme 1, Figs. 2 and 3). This resulted in substantially better accuracy of 92% in classifying malignant vs nonmalignant regions of interest with a shallow CNN.
We refer to the preprocessing method arrived at as spread parameter contrast limited adaptive histogram equalization. While more work could be carried out to search for the optimal hyper parameters of this method, we settled on a process that can be described by pseudocode snippet 1:
# pseudo code: red=CLAHE(sample_image, size=2, clipping=8) green=CLAHE(sample_image, size=4, clipping=4) blue=CLAHE(sample_image, size=8, clipping=2) input_image=merge(red, green, blue)
To enable lesion localization, we generated a separate dataset comprised of image patches from reader annotations. A sliding window approach would be taken to perform coarse localization of pathologically indicative regions. Each full breast image yielded approximately 100–400 sliding windows, with window traversal strides measuring 1/5 of the window’s width (see Fig. 4).
Inference would be run against these windows as regions of interest (RoIs). 512 × 512 pixel RoIs were downsampled and processed into 299 × 299 × 3 inputs, to suit the standard input for inception_v3 (described further below). Taking a larger window and downsampling were done simply to aid speed of processing, whilst retaining a good level of resolution.
Unlike the full images (which are more strongly isometric), the RoIs could be heavily augmented with flip and rotation, giving an eightfold augmentation for the detection window data. Two rounds of hard negative mining were performed to improve specificity of the RoI network. Hard negative mining refers to iterative training upon false-positive (FP) results with the aim of improving specificity. Specificity is particularly important for the RoI network as hundreds of inferences occur in the assessment of each full scale image.
Model
A model architecture was designed in light of the pathfinding experiments described above. Those experiments supporting the following assertions:
-
Preprocessing using SPCLAHE is beneficial to predictive performance.
-
Pathology can be discerned both at the small, window scale and at the large full image scale.
-
The shallow CNNs tested, alone or in ensembles, were not performant enough to achieve useful advances in performance, based on the experiments run.
The system architecture chosen is depicted in Fig. 5. Each mammogram underwent pre-processing and enhancement as both a single full image and as a set of patches cropped in a sliding window traversal of the full image, all standardized to 299 × 299 × 3, as described above, to suit the expected input dimensions for the chosen model. Full images and derived patches served as input for two respective independent deep CNN model instances, each based upon the Google inception_v3 model and pre-trained on ImageNet data.
Inception_v3 was chosen to optimize depth, stability, and availability of densely pretrained network snapshots. We employed a transfer-learning approach, benefiting from features learned by lower layers while fine tuning existing snapshots to the present mammographic challenge.
Network output from both inception_v3 instances was input to a random forest classification method in order to combine both the full scale and patch-based assessments into a single prediction of class: “suspicious” or “non-suspicious.” This approach permitted variations of the preprocessing methods for the full image vs the patches and also allowed independent hyper-parameter alterations to best suit each of the two image scales.
To find the highest performing network (which we refer to as the “ultimate classifier”), we concatenated the RoI network outputs and full image network outputs to train and run inference for a final prediction of risk. This final prediction from the ultimate classifier was thus dependent on both window scale and full image scale features.
To discover a suitable ultimate classifier, we applied each relevant classifier currently available in SciKit Learn, which was capable of giving a pseudo-probabilistic prediction (e.g., softmax score) of binary classification. While handcrafted logic produced similar results, we felt this logic was effective but unprincipled. We thus opted for a tried and tested general purpose discriminating classifier. This lead us to the random forest [15], as implemented in Scikit Learn.
The “validation set” (as opposed to the training set or final test set) was used to train the ultimate classifier, since the training set (used to train the full image and RoI networks) achieved near 100% accuracy after training when tested. The untouched test set remained unused for the final validation tests of the entire system.
Methods
Data was split as described above. Care was taken not to include images from any patient in more than one subset (e.g., for patient with images in the training set, no images existed in the test or validation sets).
It is worth noting that image samples were of standard mammogram screening studies, i.e., low dose x-ray imaging at width and high resolutions between 1 K and 5 K pixels. Sixteen-bit depth was available in the original images, which were reduced to eight-bit RGB during preprocessing and then presented to the receptive field as 32-bit floats, as the network model expects. This accepted loss in bit depth resolution was a compromise made to take advantage of the OpenCV CLAHE method, which at the time did not support 32-bit float grayscale images.
The full images were preprocessed as described above. The network was initiated with a checkpoint, pretrained on ImageNet [16]. The full image network then was trained on the training set until loss plateaued. Any parameter changes were made by retesting against the validation set (never against the test set) thereby avoiding the risk of information leakage to the ultimate test set.
The ROI images were preprocessed and, for the training set only, augmented, as described above. The ROI network then trained on the augmented, processed set until loss plateaued. This network also began with a checkpoint, pre-trained on ImageNet. The checkpoint being taken from the open source TensorFlow code repository.
Finally, the random forest classifier was trained, taking the outputs of the two deep CNNs and the common statistical attributes of the distribution of sliding window scores. This too was performed against the validation set. Final scores were validated for the untouched test set and defined for full image, RoI, and combined data.
Results
Each mammographic image was classified as “suspicious” or “non-suspicious” for malignancy based upon the random forest’s softmax score being above or below a chosen threshold. For the ROC curve, this threshold Θ is incremented from 0.0 to 1.0 by steps of 0.01. Results were assessed per image and per study separately—keeping in mind that a typical mammographic study includes two semi-orthogonal views of each breast. The study-level categories one, both or neither breast AS suspicious for malignancy.
Results are summarized in Table 1 and in the ROC curve (Scheme 2.). The overall stand-alone area under the curve for the ROC curve is 0.922. This is similar to those reported for contemporary single reader digital mammography and considerably above state of the art for stand-alone software performance [17].
At a sensitivity of 0.91 (above the mean for expert radiologists), specificity by image was 0.80. This is also similar to those reported for expert radiologists with or without CAD [5].
Discussion and Conclusions
Digital mammography is the foundation of breast imaging practice and the only imaging modality to demonstrate mortality reduction with screening program [2, 3]. However, mammography continues to underperform with variable sensitivity and specificity, even with widespread CAD implementation [4,5,6,7,8]. Supplemental screening modalities, including digital breast tomosynthesis (DBT) demonstrate only modest improvement in cancer detection, with sensitivity ranging from 77 to 83% [18, 19]. Screening breast ultrasound also demonstrates modest improvement in cancer detection with variable sensitivity, but with low PPV (positive predictive value or precision) [20, 21]. While a recent study demonstrated breast MRI as high-performing in the screening setting for average-risk women, with specificity of 100%, implementation is hindered by cost, resources, and imaging time [22, 23].
Next generation CAD utilizing deep CNNs offers a promising approach for optimizing mammographic performance. Here we demonstrate stand-alone performance of a deep CNN-based algorithm which achieves AUC values superior to those described in expert interpretations of digital mammography and comparable to those described for digital breast tomosynthesis [17, 18, 23]. Our results were obtained by applying a novel false-color enhancement technique to CLAHE processed mammography images and utilizing a dual deep CNN engine capable of considering whole-image and ROI specific mammography features. This approach is intuitively aligned with how radiologists assess mammograms: first globally and then “zooming in” to analyze discrete regions. Indeed, some features of malignancy, such as regional architectural distortion or asymmetry are best revealed on the image level; whereas others, such as micro-calcifications or masses, are best seen in magnification.
Further investigation will assess additional features provided to the random forest as well as location dependent performance using metrics such as FROC (free-response ROC curve) and AFROC (alternative FROC). In addition, we hope to more closely examine CNN performance stratified by demographic attributes, such as breast density, age, and type of abnormality. Finally, we plan to perform more rounds of hard negative mining and consider the reliability and reproducibility of the deep CNN models.
References
American Cancer Society: Cancer Facts & Figures 2015. Cancer Facts Fig 2015:1–9, 2015
Tabar L, Vitak B, Chen T, Yen A, Cohen A, Tot T, Chiu S, Chen S, Fann J, Rosell J, Fohlin H, Smith R, Duffy S, Al E: Swedish two-county trial: impact of mammographic screening on breast cancer mortality during 3 decades - with comments. Radiology 260(3):658–663, 2011
Lauby-Secretan B, Scoccianti C, Loomis D, Benbrahim-Tallaa L, Bouvard V, Bianchini F, Straif K: Breast-cancer screening--viewpoint of the IARC Working Group. N. Engl. J. Med. 372(24):2353–2358, 2015
Kavanagh AM, Giles GG, Mitchell H, Cawson JN: The sensitivity, specificity, and positive predictive value of screening mammography and symptomatic status. J. Med. Screen. 7(2):105–110, 2000
Lehman CD, Wellman RD, Buist DSM, Kerlikowske K, Tosteson ANA, Miglioretti DL, Breast Cancer Surveillance Consortium: Diagnostic Accuracy of Digital Screening Mammography With and Without Computer-Aided Detection. JAMA Intern. Med. 175(11):1828–1837, 2015
Winkler NS, Raza S, Mackesy M, Birdwell RL: Breast density: clinical implications and assessment methods. Radiographics 35(2):316–324, 2015
Kolb TM, Lichy J, Newhouse JH: Comparison of the Performance of Screening Mammography, Physical Examination, and Breast US and Evaluation of Factors that Influence Them: An Analysis of 27,825 Patient Evaluations. Radiology 225(1):165–175, 2002
Kerlikowske K, Hubbard RA, Miglioretti DL, Geller BM, Yankaskas BC, Lehman CD, Taplin SH, Sickles EA: Comparative effectiveness of digital versus film-screen mammography in community practice in the United States: A cohort study. Ann. Intern. Med. 155(8):493–502, 2011
Freer TW, Ulissey MJ: Screening mammography with computer-aided detection: prospective study of 12,860 patients in a community breast center. Radiology 220(3):781–786, 2001
Fenton JJ, Xing G, Elmore JG, Bang H, Chen SL, Lindfors KK, Baldwin LM: Short-term outcomes of screening mammography using computer-aided detection a population-based study of medicare enrollees. Ann. Intern. Med. 158(8):580–587, 2013
Rao VM, Levin DC, Parker L, Cavanaugh B, Frangos AJ, Sunshine JH: How widely is computer-aided detection used in screening and diagnostic mammography? J. Am. Coll. Radiol. 7(10):802–805, 2010
M Heath, K Bowyer, D Kopans, R Moore, P Kegelmeyer (2001) “The digital database for screening mammography,” Proc. Fifth Int. Work. Digit. Mammogr., pp. 212–218
Abdel-Zaher AM, Eldeib AM: Breast cancer classification using deep belief networks. Expert Syst. Appl. 46:139–144, 2016
J Arevalo, FA González, R Ramos-Pollán, JL Oliveira, MA Guevara Lopez (2015) “Representation learning for mammography mass lesion classification with convolutional neural networks,” Computer Methods and Programs in Biomedicine
Ho, Tin Kam (1995). Random Decision Forests (PDF). Proceedings of the 3rd International Conference on Document Analysis and Recognition, Montreal, QC, 14–16 August 1995
J Deng, W Dong, R Socher, L-J Li, K Li, L Fei-Fei (2009) “ImageNet: A large-scale hierarchical image database,” in 2009 I.E. Conference on Computer Vision and Pattern Recognition, pp. 248–255
Rafferty EA, Park JM, Philpotts LE, Poplack SP, Sumkin JH, Halpern EF, Niklason LT: Assessing radiologist performance using combined digital mammography and breast tomosynthesis compared with digital mammography alone: results of a multicenter, multireader trial. Radiology 266(1):104–113, 2013
Skaane P, Bandos AI, Gullien R, Eben EB, Ekseth U, Haakenaasen U, Izadi M, Jebsen IN, Jahr G, Krager M, Niklason LT, Hofvind S, Gur D: Comparison of digital mammography alone and digital mammography plus tomosynthesis in a population-based screening program. Radiology 267(1):47–56, 2013
Lee CI, Cevik M, Alagoz O, Sprague BL, Tosteson ANA, Miglioretti DL, Kerlikowske K, Stout NK, Jarvik JG, Ramsey SD, Lehman CD: Comparative effectiveness of combined digital mammography and tomosynthesis screening for women with dense breasts. Radiology 274(3):772–780, 2015
Berg WA, Blume JD, Cormack JB, Mendelson EB, Lehrer D, Böhm-Vélez M, Pisano ED, Jong RA, Evans WP, Morton MJ, Mahoney MC, Larsen LH, Barr RG, Farria DM, Marques HS, Boparai K, ACRIN 6666 Investigators: Combined screening with ultrasound and mammography vs mammography alone in women at elevated risk of breast cancer. JAMA 299(18):2151–2163, 2008
Brem RF, Lenihan MJ, Lieberman J, Torrente J: Screening breast ultrasound: past, present, and future. AJR. Am. J. Roentgenol. 204(2):234–240, 2015
CK Kuhl, K Strobel, H Bieling, C Leutner, HH Schild, S Schrading (2017) “Supplemental Breast MR Imaging Screening of Women with Average Risk of Breast Cancer,” Radiology, p. 161444
Helvie MA: Digital Mammography Imaging: Breast Tomosynthesis and Advanced Applications. Radiologic Clinics of North America 48(5):917–929, 2010
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
Philip Teare, Eyal Toledano, and Eldad Elnekave are employees of Zebra Medical Vision.
Oshra Benzaquen has no conflicts of interest to disclose.
Rights and permissions
About this article
Cite this article
Teare, P., Fishman, M., Benzaquen, O. et al. Malignancy Detection on Mammography Using Dual Deep Convolutional Neural Networks and Genetically Discovered False Color Input Enhancement. J Digit Imaging 30, 499–505 (2017). https://doi.org/10.1007/s10278-017-9993-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-017-9993-2