Abstract
Mung bean (Vigna radiata L.) is commonly grown in Asia as an important nutritional dry grain legume, as it can survive better in arid conditions than other crops. Abiotic stresses, such as drought and high-salt contents, negatively impact its growth and production. The dehydration-responsive element-binding protein 2 (DREB2) transcription factors play a significant role in the response to these stress stimuli via transcriptional regulation of downstream genes containing the cis-element dehydration-responsive element (DRE). However, the molecular mechanisms involved in the drought tolerance of this species remain elusive, with very few reported candidate genes. No DREB2 ortholog has been reported for mung bean, and the function of mung bean DREB2 is not clear. In this study, a novel VrDREB2A gene with conserved AP2 domains and transactivation ability was isolated from mung bean. A modified VrDREB2A protein lacking the putative negative regulatory domain encoded by nucleotides 394–543 was shown to be localized in the nucleus. Expression of the VrDREB2A gene was induced by drought, high salt concentrations and abscisic acid treatment. Furthermore, comparing with the wild type Arabidopsis, the overexpression of VrDREB2A activated the expression of downstream genes in transgenic Arabidopsis, resulting in enhanced tolerance to drought and high-salt stresses and no growth retardation. The results from this study indicate that VrDREB2A functions as an important transcriptional activator and may help increase the abiotic stress tolerance of the mung bean plant.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drought and high-salt concentrations are two common abiotic stresses that are severe threats to plant growth and food production around the world (Nakabayashi and Saito 2015). Many analyses have shown that transcriptional regulatory pathways are important to abiotic stress responses (Atkinson and Urwin 2012). Several transcription factors have been characterized with respect to their roles in the signaling network and transcriptional regulation, especially in abiotic stress responses (Yoshida et al. 2014). The DREB (dehydration-responsive element binding) transcription factor family, with the core sequence A/GCCGAC, was originally isolated in Arabidopsis (Liu et al. 1998) and shown to bind to DRE (dehydration-responsive element) core cis-acting sequences in the promoters of stress-responsive genes and regulate their expression via an abscisic acid (ABA)-independent pathway under conditions of drought, high-salt and cold stress in plants (Sakuma et al. 2002, 2006a). DREB transcription factors belong to the APETLA2/ethylene responsive element binding protein (AP2/EREBP) family, which is characterized by a conserved APETLA2/ethylene-responsive element binding factor (AP2/ERF) domain and is induced under abiotic stress conditions (Kizis et al. 2001; Mizoi et al. 2012).
Many DREB factors are induced by abiotic stresses, including drought (Liu et al. 1998; Nakashima et al. 2000; Qin et al. 2007b; Sakuma et al. 2006a), low temperatures (Gutha and Reddy 2008; Li et al. 2005; Qin et al. 2004, 2007b), high salt (Chen et al. 2009; Cong et al. 2008; Dubouzet et al. 2003; Huang et al. 2008; Nakashima et al. 2000; Wang et al. 2008) and extreme heat (Schramm et al. 2008). Many DREB genes have been identified, isolated and characterized in different plant species under various abiotic stresses, such as Arabidopsis (Sakuma et al. 2002), rice (Oryza sativa L.) (Cui et al. 2011), maize (Zea mays L.) (Qin et al. 2007a), soybean (Glycine max L.) (Chen et al. 2007; Mizoi et al. 2013) and cowpea (Vigna unguiculata L.) (Sadhukhan et al. 2014).
The roles of the DREB2A gene from Arabidopsis in abiotic stress tolerance are well characterized (Sakuma et al. 2006a). However, similar studies are very limited in legume plants, especially in mung bean, and the molecular mechanisms controlling plant tolerance to drought and salt remain largely unknown (Chung et al. 2013). In cowpea, VuDREB2A was induced by drought and salt stresses, and heterologous expression of VuDREB2A resulted in significant drought stress tolerance in Arabidopsis (Sadhukhan et al. 2014). Two GmDREB2A homologs (GmDREB2A;1 and GmDREB2A;2) have been identified in soybean, and their peptide sequences were very similar to each other, although the induction of GmDREB2A;2 was stronger than that of GmDREB2A;1. The two GmDREB2A homologs exhibited strong inducibility under abiotic condition. Overexpression of GmDREB2A;2 caused growth defects in the germination and seedling stages of the transgenic Arabidopsis (Mizoi et al. 2013). Moreover, DREB2A genes from different species, such as chrysanthemum (Dendranthema vestitum) (Liu et al. 2008), common wheat (Egawa et al. 2006), and Pennisetum glaucum (Agarwal et al. 2007), that result in physiological variations have been overexpressed in Arabidopsis thaliana and improved stress tolerance under drought, salt and freezing stresses in transgenic plants.
Mung bean (Vigna radiata L.) is an economically important grain legume widely grown in Asian countries for its protein-rich grains (Ahmad et al. 2011; Mishra et al. 2014). Drought and salt stresses are recognized as major constraints in the production and growth of mung beans (Hossain and Fujita 2010). Mung bean is moderately drought tolerant. Therefore, this distinctive characteristic makes it a valuable tropical legume for studying the molecular tolerance mechanisms of various abiotic stresses, including salinity and drought (Kim et al. 2004). However, no studies on the AP2/EREBP transcription factor gene in mung bean have been reported.
In the current study, we isolated and characterized the VrDREB2A gene in mung bean and demonstrated that the resulting protein is localized in the nucleus and exhibits transactivation ability. The expression of VrDREB2A was up-regulated under drought, high salt and ABA treatments. Furthermore, the overexpression of VrDREB2A enhanced the drought and high-salt tolerance of the transgenic Arabidopsis plants and did not cause growth retardation due to the activation of downstream genes.
Materials and methods
Plant growth
Mung bean (Vigna radiata L.) cultivar, VC2917, a high drought resistance accession used in this study was selected from the National Genebank of China, which is located at the Institute of Crop Science (ICS), in the Chinese Academy of Agricultural Sciences (CAAS), Beijing. The seeds were firstly germinated on moistened filter paper. When the roots had grown to approximately 1 cm, the seeds were transplanted into a soil/vermiculite (1:1, v/v) mix and grown in a greenhouse maintained at 22 °C under long days (16 h light, 8 h dark) with 50 % humidity. The 1-week old plants after germination were used for expression pattern analyses of VrDREB2A under different stress conditions.
For drought treatment, the seedlings were transferred to dry Whatman 3 MM paper for 1, 2 and 3 h. For salt treatment, the seedlings were watered with 1 M NaCl for 1, 6 and 24 h. For cold treatment, the seedlings were incubated at 4 °C for 1, 6 and 24 h. For ABA treatment, the leaves of seedlings were sprayed 200 μM ABA solution [0.05 % Tween-20 (v/v)] at 1, 6 and 24 h. After treatment, the whole seedlings were harvested, frozen in liquid nitrogen immediately, and stored at −80 °C for further analysis. Each treatment contained three biological repeats.
Isolation of VrDREB2A via degenerate PCR and RACE
RNA was isolated from 1-week old mung bean plants treated with 1 M NaCl for 3 h using the Trizol Reagent (Invitrogen, Carlsbad, CA, USA), and cDNA was amplified using reverse transcription PCR (RT-PCR) with MLV Reverse Transcriptase (Promega, Tokyo, Japan). A partial cDNA was obtained using PCR with degenerate primers designed from conserved regions of reported DREB2 sequences (see Table S1 in Supporting Information) and subcloned into pMD18-T simple (TaKaRa, Dalian, China). Then, the 3′ and 5′ regions were obtained using the RACE (rapid amplification of cDNA ends) PCR (Invitrogen, Carlsbad, CA, USA).
DNA/protein sequence and phylogenetic analyses
Nucleotide and protein sequences of DREB2A orthologs were retrieved from the NCBI Entrez database (http://www.ncbi.nlm.nih.gov). Multiple amino acid sequence alignment was performed using CLUSTALX 1.83 (Thompson et al. 1997). A phylogenetic tree was constructed using MEGA 6.06 (Tamura et al. 2007). The distances between the branches were calculated via the neighbor-joining method with 1,000 bootstrap replicates (Saitou and Nei 1987).
Localization of the VrDREB2A-GFP fusion protein
The full length of VrDREB2A gene was amplified with two primers: 5′- CTCGAGCATGGGTGCTTATGATCAAGTTTCTG -3′ (XhoI site underlined) and 5′- GGTACCCTTCCTTGCTTGCTACCATTTC -3′ (KpnI site underlined). The CDS of a modified VrDREB2A (designated VrDREB2A-TR) gene was removed; the putative negative regulatory domain encoded by nucleotides 394–543 through megaprimer PCR (Ke and Madison 1997). In this study, the pE3025-GFP vector was derived from the pSATS-RFP-NI vector, which was replaced the red fluorescent protein (RFP) gene with the green fluorescent protein (GFP) gene under the control of the CaMV dual 35S promoter, as well as a TEV enhancer (Li et al. 2011). The two PCR products were cloned into the pE3025-GFP vector to generate the pE3025-VrDREB2A-GFP and pE3025-VrDREB2A-TR-GFP constructs. They were confirmed by sequencing and used for transient transformation of onion epidermal cells via a gene gun (Bio-Rad, California, USA). GFP fluorescence was observed under a confocal fluorescence microscopy system (Nikon, Tokyo, Japan). The pE3025-GFP empty vector was used as a control.
Transactivation analysis in yeast
The entire coding sequence of VrDREB2A was amplified using two primers: 5′- GAATTCATGGGTGCTTATGATCAAGTTTCTGTG -3′ (EcoRI site underlined) and 5′- GTCGACTCACTTCCTTGCTTGCTAGCATTTCCTTTG -3′ (SalI site underlined). The PCR product was sub-cloned into the DNA-binding domain vector pBD GAL4 with a DNA-binding domain which could activate transcription of the dual report genes His3 and LacZ both controlled by the GAL4 upstream activation sequence to construct plasmid pBD-VrDREB2A. The recombinant plasmid was then transferred into the yeast strain YRG-2 that had the reporter genes His3 and LacZ. The yeast strain YRG-2 cannot grow on the SD plates without histidine, and cannot induce LacZ (b-galactosidase) activity. The transformed yeast culture containing pBD-VrDREB2A, pBD GAL4 or pGAL4 vector was dropped onto SD plates lacking tryptophan or lacking both tryptophan and histidine. The plates were incubated at 30 °C for 3 d, and the resulting clones were used in a β-Gal assay to examine the transactivation ability of VrDREB2A. The pBD GAL4 empty vector (CK−) was used as the negative control, and pGAL4 vector (CK+) was used as the positive control.
Generation of VrDREB2A-overexpressing Arabidopsis
The VrDREB2A cDNA was cloned into the pMD18-T simple vector using the primers 5′- CCATGGGTGCTTATGATCAAGTTTCTGTG -3′ (Nco I site underlined) and 5′- ACTAGTTTCCTTGCTTGCTAGCATTTCCTTTG -3′ (SpeI site underlined) (TaKaRa, Dalian, China). An NcoI-SpeI fragment containing VrDREB2A cDNA was inserted into the pCAMBIA1302 vector containing the 35S CaMV promoter and a hygromycin (kanamycin) resistance marker. The plasmid was introduced into Agrobacterium EHA105 via heat shock. The pCAMBIA1302-VrDREB2A vector was transformed into Arabidopsis plants using the floral dip method (Clough and Bent 1998; Liu et al. 1998). Seeds of transformed Arabidopsis were selected on Murashige and Skoog (MS) medium containing 80 mg/ml) hygromycin. Genomic PCR and qRT-PCR confirmed the successful transfer of 35S:VrDREB2A. Three independent lines of the T3 generation were randomly chosen for further analysis.
Salt treatment
To determine the sensitivity of the seed germination process to NaCl, seeds from wild-type and transgenic plants were placed on MS agar plates or MS agar plates saturated with 210, 220 mM NaCl. The seeds were incubated at 4 °C for 72 h, and then transfered to a growth chamber with constant light (approximately 100 μmol m−2 s−1) at 22 °C for germination. After 14 d, the number of seed germination was recorded. Seeds were considered to have germinated when radicles were 1 mm long. Gernination rate statistics are presented as the number of seeds germinated VS the number of seeds tested.
To assess salt tolerance, 5-day-old seedlings were carefully transferred to MS media containing different concentrations of NaCl (0 or 150 mM) and incubated for 7 d. The position of the root tips was marked at the time of transfer, and the root growth was measured from this mark after 7 d. Three independent biological replicates were conducted for salt experiments.
Drought treatment
To assess drought tolerance, one-week-old seedlings were transferred to pots filled with a soil/vermiculite (1:1, v/v) mix for another 2 weeks in a greenhouse under conditions of continuous illumination of approximately 100 μmol m−2 s−1, 50 % relative humidity, a temperature of 22 °C, and long days (16 h light, 8 h dark), with regular watering every 4 d before water was withheld. After 16 d without water, all pots were watered simultaneously and plant recovery and survival rate were measured after 4 d. Survival statistics are presented as the number of plants survived.
The relative electrolyte leakage (EL) was determined as described (Hao et al. 2013). Fully expanded leaves (about 100 mg) of WT and transgenic lines were incubated in 15 ml ddH2O. The initial level of EL (Ci) was measured using a conductance meter (Thermo Scientific, Baverly, USA) after shaken for 24 h at room temperature. Leaf tissue was killed in an autoclave at 121 °C for 30 min. The conductance of the incubation solution (Cmax) was measured after 24 h incubation on a shaker. Relative EL was calculated as EL = (Ci/Cmax) × 100. Values are mean ± SD of twelve independent plants.
qRT-PCR analysis
RNA samples were isolated using Invitrogen reagents (Invitrogen, USA), and reverse transcription was performed on 2 µg of RNA using M-MLV Reverse Transcriptase (Promega, Tokyo, Japan). qRT-PCR was performed using a CFX-96 Real-Time System (Bio-Rad, Hercules, California, USA) with SYBR Premix Ex Taq (TaKaRa, Otsu, Shiga, Japan). The specificity of the primer pairs (see Table S2 in Supporting Information) was confirmed by sequencing the PCR amplicons. VrActin (GenBank: AF143208.1) or AtActin (GenBank: At5G09810) was used as an internal control to normalize the amount of cDNA in the samples.
Statistical analysis
The above described experiments were performed in three independent biological repetitions. The results are presented as the mean ± SD. Differences between groups were examined for statistical significance using Student’s t test (*P < 0.05, **P < 0.01) was regarded as statistically significant.
Results
Isolation and phylogenetic analysis of the VrDREB2A in mung bean
Firstly, a cDNA fragment (225 bp) was obtained using PCR with degenerate primers designed against the highly conserved regions of DREB2-type proteins. The full-length mRNA sequence was further isolated from total mung bean RNA using the 5′ and 3′ RACE procedures. The full-length mRNA was 1158 bp and had a complete open reading frame of 386 amino acids. Further analysis of the deduced amino acid sequence revealed that this protein contained conserved consensus amino acid residues in the unique AP/EREBP domain (the 14th valine and 19th glutamic acid at 79–142 amino acids) and was highly homologous to previously isolated DREB2 proteins (Fig. 1a). In addition, this protein included a conserved nuclear localization signal. Therefore, it was assumed that the encoding gene was mung bean DREB2A.
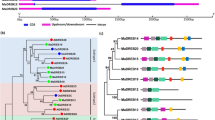
Multiple sequence alignment and phylogenetic relationship of DREB2A homologs. a The conserved DREB2A DNA-binding domain is indicated as the underlined segment. Stars and squares indicate the DREB2A signature sequences. b The phylogenetic relationship of DREB2A homologs constructed using MEGA 6.06 software. The bootstrapping value (out of 550 samples) for each node, obtained using the same software. The species designations are as follows: Mt, Medicago truncatula; At, Arabidopsis thaliana; Gm, Glycine max; Vu, Vigna unguiculata; Vr, Vigna radiata; Pv, Phaseolus vulgaris
To investigate the relationship between VrDREB2A and other DREB2A proteins, the deduced VrDREB2A amino acid sequence was compared with twelve other DREB2A proteins from Arabidopsis thaliana and other legume plants. Among these proteins, the reported VrDREB2A is closely related to cowpea VuDREB2A (GenBank: JN629045.3), with a 92 % match at the amino acid level, and has less similarity with the other DREB2A proteins from soybean (GmDREB2As, GenBank: AFU35562.1; GenBank: AFU35563.1) and the DREB proteins from Arabidopsis (AtDREB1A (GenBank: BAA33434.1) and AtDREB2A (GenBank: BAA36705.1)) (Fig. 1b). However, the characteristics and function of VrDREB2A remained unclear.
Localization and transactivation of the VrDREB2A protein
The nucleotides 394–543 of the VrDREB2A gene encode a putative negative regulatory domain, so we used a modified VrDREB2A construct (designated VrDREB2A-TR) to determine its subcellular localization. The full length of VrDREB2A and VrDREB2A-TR were fused in frame to the 5′ terminus of the GFP reporter gene under the control of the cauliflower mosaic virus dual 35S promoter (CaMV 35S) and a tobacco etch virus (TEV) enhancer. Recombinant constructs encoding the VrDREB2A-GFP, VrDREB2A-TR-GFP fusion gene and GFP alone were introduced into onion epidermal cells. The VrDREB2A-TR-GFP fusion protein accumulated primarily in the nucleus (Fig. 2g–i), whereas GFP was present throughout the entire cell (Fig. 2a–c). Interestingly, the VrDREB2A-GFP fusion protein didnot accumulate in the whole cell (Fig. 2d–f). Thus, VrDREB2A-TR was a nuclear-localized protein.
Subcellular localization and transcriptional activation analysis of VrDREB2A. a–c Subcellular localization of pE3025-GFP. d–f Subcellular localization of pE3025-VrDREB2A-GFP. g–i Subcellular localization of pE3025-VrDREB2A-TR-GFP. VrDREB2A-GFP, VrDREB2A-TR-GFP and GFP alone were bombarded into the onion epidermal cells using DNA-coated gold particles, and GFP expression was visualized after 20 h at room temperature in the dark. Cells expressing GFP alone were used as a control. a, d, g Bright field images; b, e, h GFP fluorescence (green); and c, f, i merged images. j All of the transformants containing pBD-VrDREB2A, pBD GAL4 or pGAL4 vector grew on SD/-Trp medium, SD/-Trp-His medium, or dyed b-galactosidase. The pBD GAL4 empty vector (CK−) was used as the negative control, and pGAL4 vector (CK+) was used as the positive control. Bars = 50 µm
The transactivation ability of VrDREB2A was analyzed using a yeast assay. The pBD-GAL4 had a DNA-binding domain which was controlled by the GAL4 upstream activation sequence and could activate transcription of the dual report genes His3 and LacZ. Yeast cells containing the fusion plasmid harboring VrDREB2A gene grew well on SD medium lacking histidine and could be stained blue using X-gal solution as well as yeast cells containing pGAL4 vector. On the contrary, yeast cells containing pBD GAL4 empty vector did not grow on SD medium lacking histidine, and could not stain blue (Fig. 2j). These results indicated that VrDREB2A showed transactivation activity. Taken together, these results reveal that VrDREB2A was consistent with its predicted function as a transcription factor.
Expression characteristics of the VrDREB2A in mung bean
qRT-PCR analysis showed that the transcription of the VrDREB2A gene was responsive to drought, high salt, and cold stresses as well as ABA treatment. Under drought conditions, VrDREB2A mRNA began to increase within 1 h and continued to increase after 3 h (Fig. 3a). Under high salt conditions, the expression pattern of VrDREB2A was similar to that during the drought treatment (Fig. 3b). Under cold conditions, the transcription of VrDREB2A was low and reduced its minimum level at 6 h (Fig. 3c). Interestingly, the expression of VrDREB2A was also induced by ABA treatment after 1 h and reached to its maximum at 24 h, suggesting that the gene was responsive to ABA and involved in ABA-dependent signal pathways (Fig. 3d). Taken together, these results indicate that VrDREB2A was induced by drought, salt, and cold stresses as well as ABA stimulation, indicating that it might play an important role in the response to abiotic stresses and ABA treatment.
Expression patterns of VrDREB2A in response to different abiotic stress inmung bean by qRT-PCR analyze. 1-week-old seedlings were treated with the following treatments: a for drought treatment, seedlings were transferred to dry Whatman 3 MM paper for 1, 2, or 3 h; b seedlings were subjected to 1 M NaCl for 1, 6, or 24 h; c seedlings were placed in a growth chamber at 4 °C for 1, 6, or 24 h; d the leaves were sprayed with 200 μl ABA solution containing 0.05 % Tween20 (v/v) for 1, 6, or 24 h. After treatment, the whole seedlings were harvested, frozen in liquid nitrogen immediately, and stored at −80 °C. VrActin was used as a reference gene to measure the relative quantification. Data are presented as mean ± SD (n = 3) and error bars represent SD
Overexpression of VrDREB2A improved the high salinity tolerance in transgenic Arabidopsis
The notable induction of VrDREB2A expression by multiple stresses indicated that this gene might be involved in stress resistance. Expression of VrDREB2A in transgenic Arabidopsis was detected via qRT-PCR (Fig. 4a). We randomly selected three independent T3 VrDREB2A overexpression lines (L2, L5 and L8) for drought and salinity resistance testing. The overexpression of VrDREB2A in these three lines did not cause significant growth retardation compared with the wild type line.
The overexpression of VrDREB2A increased the salt tolerance in Arabidopsis. a VrDREB2A expression in transgenic lines (L2, L5 and L8) and wild type (WT). The values were normalized against Atactin expression. Data represent the mean ± SD of three technical replicates. b Germination rate of wild type and VrDREB2A transgenic lines (L2, L5 and L8) with 0, 210 or 220 mM NaCl for 14 d. Mean germination and SD were calculated from the results of three replicated experiments each using more than 60 seeds. c Primary root elongation of wild type and VrDREB2A transgenic lines (L2, L5 and L8) seedlings in the presence of 0 mM or 150 mM NaCl for 7 d. All experiments included three replicates. Error bars represent the mean ± SD using 30 seeds. d, e Representative images of germination of wild type and VrDREB2A transgenic lines (L2, L5 and L8) with 0 mM or 210 mM NaCl for 14 d. f, g Representative images of root growth of wild type and VrDREB2A transgenic lines (L2, L5 and L8) with 0 mM or 150 mM NaCl for 7 d. Asterisks indicate significant difference from the wild type, Student’s t test, *P < 0.05, **P < 0.01)
To further evaluate the function of VrDREB2A in plants, we characterized the salinity tolerance phenotypes. In the germination stage, seed germination of the wild-type and transgenic plants did not differ under normal conditions (Fig. 4d). However, at a NaCl concentration of 210 or 220 mM, the seed of transgenic lines displayed significantly better germination rate than the wild type, and as the concentration of NaCl increased, the seed germination fulled down. At 220 mM NaCl seed germination of wild type was seriously inhibited (Fig. 4b). In the presence of 210 mM NaCl, 33.3 % of the wild-type seeds germinated, whereas the germination rates of the 35S:VrDREB2A L2, L5 and L8 transgenic lines were 70, 81.1 and 56.7 % (**P < 0.01, t test), respectively (Fig. 4e).
Similarly, when grown on MS plates containing 150 mM NaCl for 7 d, the primary root length of the wild-type plants was 1.2 cm, whereas the primary root lengths of the 35S:VrDREB2A L2, L5 and L8 transgenic lines were 1.7, 1.6 and 1.5 (*p < 0.05, t test), respectively (Fig. 4c, g). In conclusion, the transgenic plants exhibited longer primary roots than the WT plants, whereas no significant differences were observed under normal conditions (Fig. 4f, g). The results indicated that overexpression of VrDREB2A in Arabidopsis increased salt tolerance during germination and in the seedling stage.
Overexpression of VrDREB2A improved the drought tolerance of transgenic Arabidopsis
It was interesting that the expression of VrDREB2A was induced by drought stress; we further evaluated the function of VrDREB2A in plants response to drought stress. When without water for 16 d, 19.44 % of the wild-type plants survived, whereas the survival rates of the 35S:VrDREB2A L2, L5 and L8 transgenic plants were 49.31, 56.94 and 43.75 % (**P < 0.01, t test), respectively (Fig. 5b). Overall, the T3 transgenic Arabidopsis plants overexpressing VrDREB2A showed enhanced drought tolerance, whereas the wild-type plants wilted compared to the L2, L5 and L8 transgenic lines after drought treatment (Fig. 5a). Meanwhile, the relative electrolyte leakage of the 35S:VrDREB2A transgenic lines were lower than that of the wild-type line (Fig. 5c). These results indicated that overexpression of VrDREB2A could increase drought tolerance in Arabidopsis plants.
The overexpression of VrDREB2A increased the drought tolerance of Arabidopsis. a Representative images of drought tolerance of 3-week-old Arabidopsis plants from which water was withheld for 16 d. b Survival rate for wild type, L2, L5 and L8, respectively. Survival statistics are presented as the number of plants survived/number of plants tested. The experiment included three replicates. Error bars represent the mean ± SD and asterisks indicate a significant difference from the wild type (n = 48, Student’s t test, *P < 0.05, **P < 0.01). c The relative electrolyte leakage of wild type and transgenic plants (L2, L5 and L8) under normal condition was determined as described. d Expression analysis of AtCOR15A, AtCOR15B, AtKIN1, AtRD29A, AtRD29B and AtRD17, downstream genes of VrDREB2A in Arabidopsis. Abiotic stress-responsive genes in VrDREB2A transgenic and wild-type plants were analyzed by qRT-PCR. Total RNA was extracted from 3-week-old seedlings grown under normal conditions. The graphs indicate the induction levels of AtCOR15A (At2g42540), AtCOR15B (At2g42530), AtKIN1 (At5g15960), AtRD29A (At5g52310), AtRD29B (At5g52300) and AtRD17 (At1g20440) in the transgenic lines L2, L5 and L8 compared with those of wild-type plants (WT). Atactin was amplified as control. Data represent means and SE of three replications. Primers used are listed in Additional file (Supplementary Table S2)
Overexpression of the VrDREB2A in Arabidopsis activated the expression of downstream genes containing the DRE element
To understand the effects of VrDREB2A and the relationship between gene expression and stress tolerance in Arabidopsis, we analyzed the gene expression alterations of six genes containing DRE elements in their promoter regions (AtCOR15A, AtCOR15B, AtKIN1, AtRD17, AtRD29A and AtRD29B), all of which are downstream genes of DREBs in Arabidopsis (Li et al. 2011). In the transgenic plants, qRT-PCR analysis was performed using samples from WT and VrDREB2A transgenic plants. Expression levels of all six genes were enhanced in the VrDREB2A transgenic plants under normal growth conditions (Fig. 5d). These results indicated that VrDREB2A up-regulated the expression of downstream genes related to the drought and salt stress responses.
Discussion
Studying the regulation of stress-inducible genes may improve our understanding of the mechanisms by which plants maintain growth and thrive under abiotic stress conditions (Dolferus 2014). Identifying the transcription factors that mediate responses to abiotic stress is an important prerequisite for the use of stress-inducible genes in crop improvement. The DREB transcription factor is one of the most promising candidate genes conferring abiotic tolerance in several crops (Kizis et al. 2001; Roorkiwal et al. 2014). However, there have been no reports of mung bean DREB genes or the physiological processes regulated by these DREBs under abiotic stress conditions.
In this study, we successfully isolated VrDREB2A from mung bean, which is a drought-tolerant species (Kim et al. 2004), and demonstrated that it encoded a typical AP2/ERF domain. In addition, the phylogenetic analysis justified the classification of VrDREB2A in the DREB2 class of transcription factors, as its sequence was highly homologous to the cowpea protein VuDREB2A. Furthermore, VrDREB2A was a nuclear-localized protein and activated the transcription of dual reporter genes in yeast, which is characteristic of DREB2A from Arabidopsis and soybean (Mizoi et al. 2013). These data demonstrated that VrDREB2A was a new member of the DREB2 subclass and belonged to the AP2/EREBP family.
DREB2As play various roles in plants and participate in different pathways and crosstalk among pathways during the response to abiotic stresses. qRT-PCR analysis showed that the expression of the VrDREB2A gene was strongly induced by drought and salinity stresses. The rapid induction patterns of VrDREB2A in response to drought and high salt conditions were also characteristic of DREB2A proteins from Arabidopsis and other plant species. In particular, qRT-PCR data show that VrDREB2A is rapidly and strongly induced upon exposure to ABA treatment. This result indicated that VrDREB2A expression belongs to the ABA-dependent pathway in mung bean, which was completely different from the DREB2As isolated from other plants. This result confirmed that VrDREB2A is an abiotic stress response gene and may contribute to plant stress tolerance.
Many studies have indicated that transgenic methods are effective at improving the stress tolerance of crops. Earlier studies suggested that the overexpression of DREB2A proteins in plants increased abiotic stress, resulting in the induction of target stress-inducible genes in Arabidopsis (Kim et al. 2011; Sakuma et al. 2006b). Overexpression of VuDREB2A (Sadhukhan et al. 2014) and GmDREB2A:2 (Mizoi et al. 2013) in Arabidopsis resulted in enhanced tolerance to drought stress. Homologous expression of OsDREB2A in rice improved survival rates under drought and salt stress conditions compared with wild type plants (Cui et al. 2011). Our results indicated that overexpression of VrDREB2A improved tolerance to drought and salt stress. In addition, several unique genes containing DRE cis-elements were up-regulated exclusively in the DREB2A overexpressing transgenic plants. These results indicated that DREB2A was also an important transcription factor regulating the expression of other stress responsive genes via DRE cis-elements and that this protein might play a crucial role in mediating tolerance to multiple stresses.
Despite technological advances in the transformation of mung bean plants, the generation of transgenic mung bean plants is still difficult and time consuming (Somers et al. 2003). In our study, overexpression of VrDREB2A in Arabidopsis resulted in higher tolerance to drought and salt stress.
In conclusion, the VrDREB2A protein exhibited high transactivation activity and localized in the nucleus. Finally, the heterologous expression of VrDREB2A resulted in enhanced expression of DREB2A target, stress-inducible genes and improved the salt and drought stress tolerance of transgenic Arabidopsis. Therefore, our efforts in isolating mung bean VrDREB2A could provide a useful tool to increase plants tolerance against the increasing trend of droughts and/or salt restriction associated with global warming.
References
Agarwal P, Agarwal PK, Nair S, Sopory SK, Reddy MK (2007) Stress-inducible DREB2A transcription factor from Pennisetum glaucum is a phosphoprotein and its phosphorylation negatively regulates its DNA-binding activity. Mol Genet Genom 277:189–198
Ahmad M, Zahir ZA, Asghar HN, Asghar M (2011) Inducing salt tolerance in mung bean through coinoculation with rhizobia and plant-growth-promoting rhizobacteria containing 1-aminocyclopropane-1-carboxylate deaminase. Can J Microbiol 57:578–589
Atkinson NJ, Urwin PE (2012) The interaction of plant biotic and abiotic stresses: from genes to the field. J Exp Bot 63:3523–3543
Chen M, Wang QY, Cheng XG, Xu ZS, Li LC, Ye XG, Xia LQ, Ma YZ (2007) GmDREB2, a soybean DRE-binding transcription factor, conferred drought and high-salt tolerance in transgenic plants. Biochem Biophys Res Commun 353:299–305
Chen J, Xia X, Yin W (2009) Expression profiling and functional characterization of a DREB2-type gene from Populus euphratica. Biochem Biophys Res Commun 378:483–487
Chung E, Cho CW, So HA, Kang JS, Chung YS, Lee JH (2013) Overexpression of, a mung bean E2 ubiquitin-conjugating enzyme, enhances osmotic stress tolerance in Arabidopsis. PLoS One 8:e66056
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cong L, Zheng HC, Zhang YX, Chai TY (2008) Arabidopsis DREB1A confers high salinity tolerance and regulates the expression of GA dioxygenases in tobacco. Plant Sci 174:156–164
Cui M, Zhang W, Zhang Q, Xu Z, Zhu Z, Duan F, Wu R (2011) Induced over-expression of the transcription factor OsDREB2A improves drought tolerance in rice. Plant Physiol Biochem 49:1384–1391
Dolferus R (2014) To grow or not to grow: a stressful decision for plants. Plant Sci 229:247–261
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Egawa C, Kobayashi F, Ishibashi M, Nakamura T, Nakamura C, Takumi S (2006) Differential regulation of transcript accumulation and alternative splicing of a DREB2 homolog under abiotic stress conditions in common wheat. Genes Genet Syst 81:77–91
Gutha LR, Reddy AR (2008) Rice DREB1B promoter shows distinct stress-specific responses, and the overexpression of cDNA in tobacco confers improved abiotic and biotic stress tolerance. Plant Mol Biol 68:533–555
Hao L, Wang Y, Zhang J, Xie Y, Zhang M, Duan L, Li Z (2013) Coronatine enhances drought tolerance via improving antioxidative capacity to maintaining higher photosynthetic performance in soybean. Plant Sci 210:1–9
Hossain MA, Fujita M (2010) Evidence for a role of exogenous glycinebetaine and proline in antioxidant defense and methylglyoxal detoxification systems in mung bean seedlings under salt stress. Physiol Mol Biol 16:19–29
Huang B, Jin LG, Liu JY (2008) Identification and characterization of the novel gene GhDBP2 encoding a DRE-binding protein from cotton (Gossypium hirsutum). J Plant Physiol 165:214–223
Ke SH, Madison EL (1997) Rapid and efficient site-directed mutagenesis by single-tube ‘megaprimer’ PCR method. Nucleic Acids Res 25:3371–3372
Kim YJ, Kim JE, Lee JH, Lee MH, Jung HW, Bahk YY, Hwang BK, Hwang I, Kim WT (2004) The Vr-PLC3 gene encodes a putative plasma membrane-localized phosphoinositide-specific phospholipase C whose expression is induced by abiotic stress in mung bean (Vigna radiata L.). FEBS Lett 556:127–136
Kim JS, Mizoi J, Yoshida T, Fujita Y, Nakajima J, Ohori T, Todaka D, Nakashima K, Hirayama T, Shinozaki K, Yamaguchi-Shinozaki K (2011) An ABRE promoter sequence is involved in osmotic stress-responsive expression of the DREB2A gene, which encodes a transcription factor regulating drought-inducible genes in Arabidopsis. Plant Cell Physiol 52:2136–2146
Kizis D, Lumbreras V, Pages M (2001) Role of AP2/EREBP transcription factors in gene regulation during abiotic stress. FEBS Lett 498:187–189
Li XP, Tian AG, Luo GZ, Gong ZZ, Zhang JS, Chen SY (2005) Soybean DRE-binding transcription factors that are responsive to abiotic stresses. Theor Appl Genet 110:1355–1362
Li D, Zhang Y, Hu X, Shen X, Ma L, Su Z, Wang T, Dong J (2011) Transcriptional profiling of Medicago truncatula under salt stress identified a novel CBF transcription factor MtCBF4 that plays an important role in abiotic stress responses. BMC Plant Biol 11:109
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10:1391–1406
Liu L, Zhu K, Yang Y, Wu J, Chen F, Yu D (2008) Molecular cloning, expression profiling and trans-activation property studies of a DREB2-like gene from chrysanthemum (Dendranthema vestitum). J Plant Res 121:215–226
Mishra S, Alavilli H, Lee BH, Panda SK, Sahoo L (2014) Cloning and functional characterization of a vacuolar Na+/H+ antiporter gene from mungbean (VrNHX1) and its ectopic expression enhanced salt tolerance in Arabidopsis thaliana. PLoS One 9:e106678
Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:86–96
Mizoi J, Ohori T, Moriwaki T, Kidokoro S, Todaka D, Maruyama K, Kusakabe K, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2013) GmDREB2A;2, a canonical dehydration-responsive element-binding protein 2-type transcription factor in soybean, is posttranslationally regulated and mediates dehydration-responsive element-dependent gene expression. Plant Physiol 161:346–361
Nakabayashi R, Saito K (2015) Integrated metabolomics for abiotic stress responses in plants. Curr Opin Plant Biol 24C:10–16
Nakashima K, Shinwari ZK, Sakuma Y, Seki M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2000) Organization and expression of two Arabidopsis DREB2 genes encoding DRE-binding proteins involved in dehydration- and high-salinity-responsive gene expression. Plant Mol Biol 42:657–665
Qin F, Sakuma Y, Li J, Liu Q, Li YQ, Shinozaki K, Yamagushi-Shinozaki KY (2004) Cloning and functional analysis of a novel DREB1/CBF transcription factor involved in cold-responsive gene expression in Zea mays L. Plant Cell Physiol 45:1042–1052
Qin F, Kakimoto M, Sakuma Y, Maruyama K, Osakabe Y, Tran LS, Shinozaki K, Yamaguchi-Shinozaki K (2007a) Regulation and functional analysis of ZmDREB2A in response to drought and heat stresses in Zea mays L. Plant J 50:54–69
Qin QL, Liu JG, Zhang Z, Peng RH, Xiong AS, Yao QH, Chen JM (2007b) Isolation, optimization, and functional analysis of the cDNA encoding transcription factor OsDREB1B in Oryza Sativa L. Mol Breed 19:329–340
Roorkiwal M, Nayak SN, Thudi M, Upadhyaya HD, Brunel D, Mournet P, This D, Sharma PC, Varshney RK (2014) Allele diversity for abiotic stress responsive candidate genes in chickpea reference set using gene based SNP markers. Front Plant Sci 5:248
Sadhukhan A, Kobayashi Y, Kobayashi Y, Tokizawa M, Yamamoto YY, Iuchi S, Koyama H, Panda SK, Sahoo L (2014) VuDREB2A, a novel DREB2-type transcription factor in the drought-tolerant legume cowpea, mediates DRE-dependent expression of stress-responsive genes and confers enhanced drought resistance in transgenic Arabidopsis. Planta 240:645–664
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006a) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18:1292–1309
Sakuma Y, Maruyama K, Qin F, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2006b) Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc Natl Acad Sci USA 103:18822–18827
Schramm F, Larkindale J, Kiehlmann E, Ganguli A, Englich G, Vierling E, von Koskull-Doring P (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53:264–274
Somers DA, Samac DA, Olhoft PM (2003) Recent advances in legume transformation. Plant Physiol 131:892–899
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang QY, Guan YC, Wu YR, Chen HL, Chen F, Chu CC (2008) Overexpression of a rice OsDREB1F gene increases salt, drought, and low temperature tolerance in both Arabidopsis and rice. Plant Mol Biol 67:589–602
Yoshida T, Mogami J, Yamaguchi-Shinozaki K (2014) ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr Opin Plant Biol 21:133–139
Acknowledgments
This work was supported by the Ministry of Agriculture of China [the earmarked fund for China Agriculture Research System (CARS-09)] and the Agricultural Science and Technology Innovation Program (ASTIP) of CAAS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10265_2015_773_MOESM1_ESM.pdf
Additional Supporting Information may be found in the online version of this article: Table S1 Sequence of oligonucleotides used for cloning and sequencing analyses of VrDREB2A (CDS), Localization and transactivation ability of VrDREB2A and qRT-PCR of transgenic Arabidopsis. Table S2 Primers used for qRT-PCR expression analysis of genes downstream of VrDREB2A in Arabidopsis (PDF 117 kb)
Rights and permissions
About this article
Cite this article
Chen, H., Liu, L., Wang, L. et al. VrDREB2A, a DREB-binding transcription factor from Vigna radiata, increased drought and high-salt tolerance in transgenic Arabidopsis thaliana . J Plant Res 129, 263–273 (2016). https://doi.org/10.1007/s10265-015-0773-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-015-0773-0