Abstract
Following the rapid uplift of the Qinghai-Tibetan Plateau, the reorganization of the major river drainages in southwest China was primarily caused by river capture events. However, the impact of these past changes in drainage patterns on the current distribution and genetic structure of the endemic flora of this region remains largely unknown. Here we report a survey of amplified fragment length polymorphism (AFLP) in Terminalia franchetii, an endemic shrub or small tree of the deep and dry-hot river valleys of this region. We surveyed AFLP variation within and among 21 populations (251 individuals) of T. franchetii, distributed disjunctively between northern and southern drainage systems. Using STRUCTURE, principal coordinates analysis, and genetic distance methods, we identified two main population genetic groups (I and II) and four subgroups within the species, as follows: (I) the Upper Jinshajiang Valley (subgroup I(north)) and the Honghe drainage area (subgroup I(south)); (II) the Middle and Lower Jinshajiang and Yalongjiang Valleys (subgroup II(north)) and the Nanpanjiang drainage area (subgroup II(south)). Genetic diversity was lower in group I than in group II. According to the genetic diversity and genetic structure results, we suggest that the modern disjunctive distribution and associated patterns of genetic structure of T. franchetii result from vicariance caused by several historical drainage capture events, involving the separation of the Upper Jinshajiang, Yalongjiang and Daduhe from the Honghe or Nanpanjiang in southwest China.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The mountains of southwest China are recognized as one of the world’s biodiversity hotspots (Myers et al. 2000). This region harbours about 12,000 species of vascular plants, of which at least 20 genera and 3,500 species are endemic (Myers et al. 2000; Wang and Zhang 1994; Wilson 1992; Wu 1988; Ying et al. 1993). Previous studies of plant phylogeography in this region have focused mainly on taxa from the plateau areas of the Qinghai-Tibetan Plateau and adjacent mountain ranges (e.g., the Hengduan Mountains) (Yuan et al. 2008; Zhang et al. 2007a, 2005). However, from a phylogeographic perspective, far less attention has been devoted to the endemic flora of the valleys of the region, e.g. those of the Jinshajiang River (Upper Yangtze) and its tributaries (Jialingjiang, Daduhe, Yalongjiang), the Nanpanjiang (Upper Pearl River) and the Honghe (Red River) (Figs. 1, 2). Regional examples of such narrowly distributed and endemic species, often termed “floristic character species”, include Terminalia franchetii, Zizyphus yunnanensis, Acanthochlamys bracteata, Andropogon yunnanensis, Nouelia insignis, Trailliaedoxa gracilis, and Vitex negundo (Jin 1998, 1999, 2002; Jin et al. 1994; Shui et al. 2003). Recently, a few studies have used genetic markers to examine the biogeographic and conservation biological importance of landscape effects and riverine barriers in shaping the distribution of genetic diversity for the endemic flora of this region (Nouelia insignis: Luan et al. 2006; Peng et al. 2003; Vitex negundo: Zhang et al. 2007b). This region also provides an ideal location for studies of current genetic structure at the intra-specific level sculpted by past geological changes in paleo-drainage systems (Guo et al. 2005; He and Chen 2006; Peng et al. 2006; Rüber et al. 2004). However, no study to date has used molecular phylogeographic methodologies to explicitly address this issue in a plant species endemic to these river valleys in southwest China.
Summary of river captures in southwest China (adapted from Clark et al. 2004). a The drainage pattern prior to the major captures, when these rivers drained together into the South China Sea through the ancient Honghe River (as indicated by dashed lines). b The modern river pattern, after the putative capture and reversal events
Map of southwest China, showing the 21 populations of Terminalia franchetii surveyed for AFLP variation. Also indicated are the two major genetic groups (I and II) identified, and their subgroups: Group I, consisting of subgroups I(north) and I(south); and Group II, consisting of subgroups II(north) and II(south). The broken lines indicate past river flow routes before river capture
It has long been recognised that the current drainage patterns of the major East Asian rivers differ markedly from paleo-drainage patterns of the region (Brookfield 1998; Clark et al. 2004; Gregory 1925; Gregory and Gregory 1923; Hallet and Molnar 2001; Metivier et al. 1999; Seeber 1983; Shi et al. 2006). Clark et al. (2004) suggested that the rivers at the southeast margin of the Tibetan Plateau [the Jialingjiang, Daduhe, Yalongjiang, Jinshajiang, Mekong (Lancangjiang), Salween (Nujiang), Tsangpo-Brahmaputra (Yalu-Tsangpo) and Irrawaddy] were all once tributaries of a single southward flowing system, the paleo-Honghe (Red River), which still drains southeast into the South China Sea (Fig. 1a). Subsequent reorganization into the modern major river catchments was primarily the result of river capture and reversal events, associated with the rapid geological uplift of the Tibetan Plateau, which began about 3.4 Mya (Sun and Zheng 1998). Thus, the paleo-Honghe became a beheaded river (Clark et al. 2004).
Several molecular phylogenetic studies tentatively highlight the importance of river capture events in southwest China and the uplift of the Tibetan Plateau in understanding the vicariant evolution of many Asian freshwater fish species (Guo et al. 2005; He and Chen 2006; He et al. 2004; Kottelat 1989; Peng et al. 2004, 2006; Rüber et al. 2004). In general, the molecular estimates of divergence times reported in the above studies date major vicariance events to the late Tertiary or Pleistocene era, agreeing well with geological time estimates of the separation of these river capture events caused by tectonic uplifts in south-eastern Tibet.
In contrast to what has been learned from the above fish phylogenies, however, little has yet been inferred from current plant population genetic structure about historical river capture events in the region. To the best of our knowledge, there is only a single study, albeit from another region (Japan), that has used a plant species (Rhododendron ripense) to investigate the shaping of genetic structure through past (Pleistocene) geological drainage events (Kondo et al. 2009). A major complicating factor in addressing this issue is that, while fish distributions tend to change directly with changes in catchment patterns, plant distributions are likely to be influenced to a much lesser extent in this respect. Nevertheless, the above mentioned floristic character species of the river valleys of southwest China are probably highly reliant on the generally dry-hot climate conditions of these valleys (with average annual temperatures/precipitation of 18–23°C/500–800 mm), even though conditions likely vary across their 350–1,600 m above-sea-level (a.s.l.) altitudes (Jin 1998, 1999, 2002; Jin et al. 1994). Hence, it is reasonable to assume that such species tracked the climate (or other environmental conditions) of these valleys even when river capture and reversal events happened, or at least would have formed relic populations in separate valleys where similarly suitable climates developed. In this study, we used amplified fragment length polymorphism (AFLP) markers to test whether a species endemic to the dry-hot river valleys of southwest China, Terminalia franchettii Gagnep. (Combretaceae), retained genetic signatures of past hydrological landscape structures.
Terminalia franchetii is endemic to several valleys in southwest China (Fig. 2; Jin 1999, 2002; Jin et al. 1994; Shui et al. 2003). Here, the species occurs on open, stony river deposits and cliff ledges (Chen and Turland 2007). It has rather small wind-pollinated flowers, and small (<0.5 cm) three-winged fruits, and likely adapted to dispersal by water (Chen and Turland 2007; T.C. Zhang, unpublished data). As a characteristic distributional feature, T. franchetii exhibits a disjunctive distribution between the Jinshajiang/Yalongjiang and the Honghe/Nanpanjiang drainage areas, two drainage systems that were likely connected via north-to-south river-flow routes in the geological past, i.e. the Late Pliocene/Pleistocene (Clark et al. 2004; Shi et al. 2006; Figs. 1, 2). This leads to the hypothesis that if T. franchetii tracked such past river flow routes, this should have left detectable traces in the extant population genetic structure of this species, reflecting such former connections. To test this hypothesis, we used the AFLP method to explore the phylogeographic history of T. franchetii in these dry-hot river valleys, with the following specific questions addressed:
-
1.
What is the phylogeographic structure associated with the genetic diversity of T. franchetii?
-
2.
Have past geological changes in drainage systems had a major role in shaping the disjunctive range of T. franchetii and its current geographic distribution of genetic variation?
Materials and methods
Study species
Terminalia franchetii, a small deciduous shrub or tree (up to ca. 10 m high), is endemic to several valleys in the Jinshajiang, Yalongjiang, Honghe and Nanpanjiang catchments (Fig. 2; Jin 1999, 2002; Jin et al. 1994; Shui et al. 2003). It has bisexual, apetalous and rather small wind-pollinated flowers, arranged in spicate inflorescences, and small three-winged fruits (pseudodrupes) that are dry/leathery with an aerenchymatous mesocarp (Chen and Turland 2007; T.C. Zhang, unpublished data).
Population sampling
Extensive field investigations were conducted between May 2007 and August 2008. A total of 251 T. franchetii individuals were sampled from 21 populations. Sampling covered almost the entire distribution of this species in the dry-hot valleys of the region (Table 1; Fig. 2). Young, green leaves from nine to 16 individuals were collected from each population. The individual trees sampled per population were at least 20 m apart. Vegetative tissues were stored in tubes with silica gel until DNA extraction.
DNA extraction and AFLP protocol
Genomic DNA was extracted using a modified cetyltrimethyl ammonium bromide (CTAB) method (Doyle and Doyle 1987). The AFLP procedure was carried out according to the Beckman Coulter protocol, with only minor modifications, as described by Reisch (2007). Double digestion of genomic DNA was performed for 2 h at 37°C in a 20 μL mix, using 2 units (U) of MseI and 10 U of EcoRI. Adapters were then ligated to the DNA, for 2 h at 37°C, using 2 U of T4 DNA Ligase in a 21 μL volume. Pre-selective polymerase chain reactions (PCRs) were run in a reaction volume of 25 μL. Diluted 20× pre-selective products underwent selective PCR with the following primer combinations: E-AGG/M-AGT, E-ACC/M-CTC and E-AGG/M-CTT. These three primer combinations were chosen from 20 primer pairs screened using eight randomly selected samples. Selective amplifications were run in a 25 μL volume. Finally, the PCR products were added to a mixture of Sample Loading Solution (Beckman Coulter, Fullerton, California, USA) and CEQ Size Standard 400 (Beckman Coulter). The fluorescence-labeled selective amplification products were separated by capillary gel electrophoresis on an automated sequencer (CEQ 8000, Beckman Coulter). Raw data were collected and analyzed with the aid of CEQ Size Standard 400, using CEQ 8000 software (Beckman Coulter). Fragments were sorted into bins created using CEQ 8000 parameter options: the Y threshold (peak height) for export was 1000 relative fluorescence units (RFU) and the maximum bin width was two nucleotides (nt). Fragments were then assigned to bins according to the selection parameters and checked manually.
Data analysis
In the AFLP data matrix, the presence of a band was recorded with a score of 1, and its absence with 0. POPGENE version 1.32 (Yeh et al. 1999) was used to estimate various measures of genetic diversity, including the percentage of polymorphic bands (PPB), Shannon’s information index (I), and Nei’s (1973) gene diversity (H) assuming Hardy–Weinberg equilibrium. These parameters were calculated at the species level, and separately for various population groups identified and each single population. Genetic differentiation among populations (G st) was estimated based on Nei’s (1978) gene diversity statistics. The amount of gene flow between these populations was estimated by N m = 0.25 (1 − G st)/G st (Wright 1951, 1969).
Genetically similar groups of samples with distinctive allele frequencies were identified by a Markov chain Monte Carlo (MCMC) Bayesian clustering method in STRUCTURE version 2.2 (Pritchard et al. 2000). We used the ‘no-admixture’ model and assumed uncorrelated allele frequencies for the analysis. The number of groups was estimated on the basis of 10,000 iterations, following a burn-in of 10,000 iterations. Analyses for the predefined value of K (number of groups) were run 10 times for K ≤ 10 and five times for 10 < K ≤ 15, to ensure consistent results. The best estimate of K for the data set is usually selected by choosing the model that gave both the highest probability of the data and consistent results after multiple runs. So two criteria were applied to choose the best value of K for our data set: the estimated posterior log probability of the data, L(K), and the stability of assignment patterns across different runs. Since L(K) continued to grow slightly with increasing values of K, selecting the value of K that maximized the probability of fitting the data was not possible. We therefore calculated another ad hoc quantity, based on the rate of change in probability (ΔK) between successive K values, as proposed by Evanno et al. (2005).
The relationships between individuals were examined using principal coordinates analysis (PCoA), based on the Jaccard distance between individuals, using the software NTSYS-pc 211 (Rohlf 2000). Hierarchical analyses of molecular variance (AMOVAs) were performed to assess genetic structure within and between population genetic groups identified using ARLEQUIN, version 3.0 (Excoffier et al. 2005), and significance tests were performed on the basis of 10,000 permutations. To estimate the relationships among populations, we calculated observed and bootstrapped (1,000 permutations) data matrices of pairwise F st values between populations using program AFLP-SURV version 1.0 (Vekemans et al. 2002). These distance matrices based on pairwise F st comparisons were then used as input files for the PHYLIP 3.6 software (Felsenstein 1993). First, the NEIGHBOR program was run to cluster populations according to the neighbor-joining (NJ) method based on the observed distance matrix. Then, the CONSENSE program was used to construct a consensus tree, which was edited in TreeView, version 1.6.6 (Page 1996).
Results
With the three primer combinations used, 317 AFLP fragments were generated across the 251 Terminalia franchetii individuals surveyed. Of those fragments, 309 (97.48%) were polymorphic. The highest level of gene diversity (H), also indicated by Shannon’s Information index (I), was in population DM (H = 0.165, I = 0.255). The lowest diversity was recorded in population ES (H = 0.086, I = 0.133) (Table 2). The genetic diversity indicators revealed that at the species level H is 0.1679, and I is 0.2735 (Table 2).
STRUCTURE analysis revealed that the largest increase in the posterior log probability of the data, L(K), occurred at K = 2, as seen in the graphical representation of L(K) over five or 10 runs for each K value (Fig. 3b). Considering the rate of change between successive runs, the maximum value of ΔK was even more clearly associated with K = 2 (Fig. 3c). At K = 2, all runs generally indicated the same two genetic groups of T. franchetii, but these did not correspond to the two disjunctive distribution ranges of this species from northern vs. southern drainage systems (Figs. 2, 3). Group I samples were from the northern Upper Jinshajiang Valley (TD, HTX, DJ, JA, populations 1–4) and the southern Honghe drainage area (MZ, ES, SP, YJ, pops. 18–21). By contrast, Group II samples were from the northern Middle and Lower Jinshajiang and Yalongjiang Valleys (DD, DM, RH, YY, MY, YB, LQ, JY, PDH, pops. 5–14) and the southern Nanpanjiang drainage area (JC, YS, DQ, pops. 15–17; Fig. 2). However, the STRUCTURE analysis also revealed that when K was increased to 3, the ΔK value was still considerably higher (about 20) than those ΔK values obtained for K ≥ 4 (Fig. 3c). We note that STRUCTURE with K = 3 also recovered Group I as a geographically disjunctive but genetically coherent unit, however subdivided former Group II into a cluster comprising samples from Yalongjiang and Middle Jinshajiang (pops. 5–10) vs. those from Lower Jinshajiang and Nanpanjiang/Eastern Honghe (pops. 11–17) (data not shown).
Genetic structure of Terminalia franchetii inferred by Bayesian clustering of 251 AFLP phenotypes (individuals) using the program STRUCTURE. a Assignment of individuals into K = 2 genetically distinguishable groups. Each individual is represented by a vertical bar. b Log probability of data L(K) as a function of K for 10 STRUCTURE runs at K = 1–10 or 5 runs at K = 11–15. c Rate of change in the probability between successive runs, ΔK, as a function of K (see Evanno et al. 2005)
The individual-based PCoA plot (Fig. 4) mainly subdivided all AFLP phenotypes according to Groups I and II along the first axis, which explained 11.49% of the total genetic variance. In addition, mainly along axis 2 (6.73%), each group was further subdivided into two subgroups (Fig. 4; see also Fig. 2), hereafter referred to as: subgroup I(north) (Upper Jinshajiang Valley; pops. 1–4) and subgroup I(south) (Honghe drainage area; pops.18–21), as well as subgroup II(north) (Middle and Lower Jinshajiang and Yalongjiang Valleys; pops. 5–14) and subgroup II(south) (Nanpanjiang drainage area; pops. 15–17). However, there was also some intermixing of individuals belonging to different subgroups in the centre of the PCoA plot (Fig. 4). By contrast, the population-based NJ network (Fig. 5) clearly assigned populations to their respective major groups and subgroups, and also revealed the genetic distinctness of their population constituents despite geographic proximity (e.g., populations JA vs. DD, or SP vs. DQ; see also Fig. 2).
Principal coordinates analysis (PCoA) of 251 AFLP phenotypes (individuals) of Terminalia franchetii. Subgroups delineated by ellipses include individuals from the following areas: subgroup I(north): Upper Jinshajiang Valley (pops. 1–4); subgroup I(south): Honghe drainage area (pops. 18–21), subgroup II(north): Middle and Lower Jinshajiang and Yalongjiang Valleys (pops. 5–14); and subgroup II(south): Nanpanjiang/Eastern Honghe drainage area (pops. 15–17). Open symbols and closed symbols correspond to subgroup I and subgroup II, identified by the program STRUCTURE, respectively
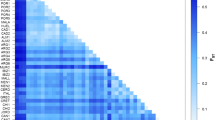
Neighbor-joining (NJ) network based on pairwise F st estimates among populations. Bootstrap values equal to or higher than 45% are shown along branches (1,000 permutations). Population codes are identified in Table 2
Total genetic diversity was lower in Group I (H = 0.130, I = 0.210) than in Group II (H = 0.173, I = 0.280) (Table 2). At the subgroup level, there was a similar result, with subgroups I(north) and I(south) having lower values (H = 0.123, 0.111; I = 0.198, 0.177, respectively) than subgroups II(north) and II(south) (H = 0.171, 0.150; I = 0.277, 0.247, respectively; Table 2). The species-wide estimate of genetic structure (G st) was 0.249, indicating that clear genetic differentiation existed among the populations. Estimates of gene flow between populations based on the G st value revealed that the number of migrants per generation (N m) was 0.754 (Table 2).
When applied to the whole dataset, hierarchical AMOVA (Table 3) confirmed the existence of significant differentiation between Groups I and II, with 14.98% of the total variance occurring among groups and 15.02% among populations within groups. So, the highest percentage of the genetic variance resided within populations (69.99%; Table 3). Within Group I, taken separately, 19.05% of the variance occurred among subgroups I(north) and I(south), and 8.74% among populations within subgroups. Within Group II, the variance was 8.03% among subgroups II(north) and II(south), and 11.87% among populations within subgroups (Table 3). Overall, the highest percentage of regional differentiation occurred between subgroups I(north) and I(south), followed by Groups I and II, and subgroups II(north) and II(south).
Discussion
Broad-scale phylogeographic structure
Collectively, the STRUCTURE, PCoA and NJ analyses of the present AFLP data clearly distinguish two main groups and four subgroups of Terminalia franchetii within the study area (Figs. 2, 3, 4, 5). Importantly, the two main groups (I and II), as most clearly identified by STRUCTURE (Fig. 3a), did not match the species’ present-day disjunctive distribution between northern and southern drainage systems (Fig. 2). Instead, subgroup I(north) was genetically closer to subgroup I(south), rather than to the geographically adjacent subgroup II(north) immediately to the east. In turn, the latter subgroup was found to be genetically closer to subgroup II(south). These results suggest that subgroups I(north) and I(south) probably once formed a continuous range, as did II(north) and II(south), but with each pair occupying different paleo-drainages. This broad-scale phylogeographic structure of T. franchetii is consistent with the paleo-drainage geographical pattern in this region. Consequently, we conclude that the currently disjunctive distribution of subgroups I(north) and I(south), similarly, of subgroups II(north) and II(south), resulted from past vicariant events due to shifts in paleo-drainage systems (see below).
Vicariance pattern between Jinshajiang and Honghe
The results of AFLP marker analyses show that the Upper Jinshajiang lineages are strongly clustered with the Honghe lineages (Group I; subgroups I(north) and I(south)). This result has two possible explanations: that the populations spread between the Jinshajiang Valley and the Honghe area via long-distance dispersal; or that a vicariance event occurred sometime in the past. Presently, T. franchetii mainly grows in deep river valleys surrounded by very high mountains. For example, at the local scale, the population sampled at the highest altitude, TD (1,916 m a.s.l.; Table 1) is overtopped by the adjacent Mt. Yulongxueshan (5,596 m a.s.l.) by 3,680 m. Moreover, several mountains located between the drainage areas of the Jinshajiang and Honghe reach elevations of >4,000 m (a.s.l.) and probably act as significant barriers to any potential wind-mediated long-distance dispersal of the species’ small and three-winged fruits. In addition, we have observed in field and laboratory experiments that the seed viability of T. franchetii declines rapidly (T.C. Zhang, unpublished data). In sum, long-distance seed dispersal of this species between the Upper Jinshajiang Valley and the Honghe area seems unlikely. Hence, the presently disjunctive range of Group I (with subgroups I(north) and I(south)) likely results from a past vicariant event. The most likely scenario is that T. franchetii was once more widely distributed between the Upper Jinshajiang Valley and the Honghe drainage areas, which shared the same river flowing-route into the South China Sea (Fig. 1a). It is likely that the rapid geological uplift of the eastern Tibetan Plateau, which began about 3.4 Mya (Sun and Zheng 1998), caused a massive reorganization of drainage patterns among the eastern plateau rivers through capture and reversal events. This would have included the river capture of the Upper Jinshajiang from the paleo-Honghe River, followed by an eastward re-direction (reversal) of the Upper Jinshajiang into the Middle and Lower Jinshajiang River, which subsequently formed a single river (Clark et al. 2004; Figs. 1a, b, 2). We therefore propose that prior to this river capture event, high levels of pollen and/or seed flow occurred among populations of T. tranchetii along the continuous valleys of the Upper Jinshajiang and paleo-Honghe. However, since this species is likely adapted to the dry-hot habitat conditions of these river valleys, the above river capture of the Upper Jinshajiang may help to explain its modern disjunctive distribution between the Upper Jinshajiang and the Honghe Valley, and also the genetic similarity between subgroups I(north) and I(south). Several molecular phylogenetic studies in freshwater fish have invoked the same (or similar) river capture events in southwest China to explain the biogeographic patterns observed (Guo et al. 2005; He and Chen 2006; He et al. 2004; Kottelat 1989; Peng et al. 2004, 2006; Rüber et al. 2004).
Vicariance pattern between Yalongjiang and Honghe
Subgroup II(north) (Yalongjiang with Middle and Lower Jinshajiang) did not form a group with subgroup I(north) (Upper Jinshajiang), although populations of these two subgroups presently occur along the same river system. Rather, subgroup II(north) formed a cluster with subgroup II(south) (Nanpanjiang). This genetic distribution pattern may reflect the geological history of the Jinshajiang and Yalongjiang rivers, which originally flowed southwards as independent tributaries to the paleo-Honghe (Clark et al. 2004; Fig. 1a). In this paleo-drainage system, the Jinshajiang and Yalongjiang were not linked as they are now in the Middle Jinshajiang area. Thus, the genetic differentiation between subgroups I(north) and II(north) (see Figs. 4, 5) is consistent with this paleo-drainage disjunction. Furthermore, following the paleo-Jinshajiang river capture, the Upper Jinshajiang then flowed eastwards and joined with the modern Yalongjiang. Together with this palaeo-evidence, our results suggest that subgroups I(north) and II(north) had geographically separate origins, but then connected after river capture and reversal. Under this scenario, the unexpectedly high gene diversity (H) in populations DD (0.160) and DM (0.165; Table 2), located at the boundary between these subgroups, may be explained by modern gene migration.
The question arises of why the Middle and Lower Jinshajiang and Yalongjiang Valley populations (subgroup II(north)) are not grouped with the Honghe area populations (subgroup I(south)), since historically the paleo-Yalongjiang was also a tributary of the Honghe (see Fig. 1a). The solution to this conundrum possibly relates to the spatio-temporal order of river capture events. Based on paleo-geological data (Clark et al. 2004) and molecular biogeographic evidence (see below), the order of river captures in the north progressed from east to west along the Jinshajiang. Thus, the Yalongjiang would have been diverted earlier from the Honghe and into the lower Yangtze River. Then the Upper Jinshajiang would have remained connected to the Honghe until its own capture occurred in the late Pleistocene (Clark et al. 2004; Guo et al. 2005; He and Chen 2006; Peng et al. 2004, 2006). Therefore, subgroup II(north) would have separated from subgroup I(south) earlier than that of subgroup I(north). This scenario could explain why the genetic difference between subgroups II(north) and I(south) is higher than that between subgroups I(north) and I(south) (see Figs. 4, 5).
Vicariance pattern between Lower Jinshajiang and Nanpanjiang/Eastern Honghe
With K = 3, STRUCTURE suggested a genetic link between populations from the Lower Jinshajiang (pops. 11–14) and those from the Nanpanjiang/Eastern Honghe area (pops. 15–17). This genetic group is also supported by chloroplast DNA sequencing data of T. franchetii (T.C. Zhang, unpublished data), showing that the populations in these two areas share one main haplotype that was only rarely found in other populations.
Similar to the above scenario for Upper Jinshajiang and Yalongjiang, this genetic link between Lower Jinshajiang and Nanpanjiang/Eastern Honghe (hereafter termed the “Z” group) is suggestive of another continuous distribution of T. franchetii in the past, most likely along the paleo-Daduhe, a more easterly located tributary to the paleo-Honghe (Figs. 1a). Today, there are several large mountain ranges, such as the Wumengshan Mountains (4,247 m a.s.l.), which act as barriers to modern gene flow between these two areas. Hence, the presently disjunctive distribution of the “Z” group likely results from an ancient vicariance event, such as the separation of the paleo-Daduhe from the Nanpanjiang/Eastern Honghe by the Lower Jinshajiang capture (Clark et al. 2004; Fig. 1). Similar to the boundary populations between subgroups I(north) and II(north), population DQ located at the border of the Nanpanjiang and Honghe areas has spuriously high genetic diversity (Table 2), which may result from ongoing gene flow between subgroups I(south) and II(south).
Conclusion
Following the rapid and extreme uplift of the Tibetan Plateau, the reorganization of the major river drainages was primarily caused by river capture and reversal events (Clark et al. 2004). The significant increase in geological and environmental diversity that accompanied this uplift, including modified terrain and isolation, promoted rapid divergence and speciation. Drawing inferences from AFLP data, we propose that a number of historical vicariance events account for the distribution and genetic structure of T. franchetii in dry-hot valleys of southwest China. First, the capture of the Upper Jinshajiang from the Honghe (by the Middle Jinshajiang) may explain the genetic link between subgroups I(north) and I(south). Second, the capture of the Yalongjiang by the Lower Jinshajiang likely explains the genetic differentiation between subgroups I(north) and II(north). Finally, we postulate that a further ancient vicariant event, sundering the former flow-routes between the Daduhe/Lower Jinshajiang and Nanpanjiang/East Honghe, could explain the genetic (AFLP and cpDNA) link between populations associated with these presently disjunct drainage systems (pops. 11–14 vs. 15–17). Against the backdrop of the known history of drainage rearrangements in these areas, these scenarios can explain the current disjunct distribution of T. franchetii and its modern, broad-scale phylogeographic structure. Moreover, our data revealed how paleo-drainage rearrangements in southwest China can change a plant species’ distribution from continuous to fragmented (Upper Jinshajiang vs. Honghe; Lower Jinshajiang vs. Nanpanjiang/East Honghe), and a disjunctive distribution to a continuous one (Yalongjiang plus Jinshajiang; Upper Jinshajiang plus Lower Jinshajiang). Overall, these results provide new insights into the phylogeography of plant species endemic to the deep river valleys of southwest China, and stress the importance of taking into account paleo-hydrological evidence, in order to gain a more integrated understanding of how present-day plant distributions and genetic structures developed in this region.
References
Brookfield ME (1998) The evolution of the great river systems of southern Asia during the Cenozoic India-Asia collision: rivers draining southwards. Geomorphology 22:285–312
Chen J, Turland NJ (2007) Terminalia. In: Wu ZY, Raven PH (eds) Flora of China. Science Press/Missouri Botany Garden Press, Beijing/St. Louis, pp 310–314
Clark MK, Schoenbohm LM, Royden LH, Whipple KX, Burchfiel BC, Zhang X, Tang W, Wang E, Chen L (2004) Surface uplift, tectonics, and erosion of eastern Tibet from large-scale drainage patterns. Tectonics 23:1006–1029
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small amounts of fresh leaf tissue. Phytochem Bull 19:11–15
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol bioinform online 1:47–50
Felsenstein J (1993) PHYLIP (phylogenetic inference package) version 3.6. Department of Genetics, University of Washington. Seattle 63:188–192
Gregory JW (1925) The evolution of the river system of south-eastern Asia. Scott Geogr J 41:129–141
Gregory JW, Gregory CJ (1923) The Alps of Chinese Tibet and their geographical relations. Geogr J 61:153–174
Guo X, He S, Zhang Y (2005) Phylogeny and biogeography of Chinese sisorid catfishes re-examined using mitochondrial cytochrome b and 16S rRNA gene sequences. Mol Phylogenet Evol 35:344–362
Hallet B, Molnar P (2001) Distorted drainage basins as markers of crustal strain east of the Himalaya. J Geophys Res 106:13697–13709
He D, Chen Y (2006) Biogeography and molecular phylogeny of the genus Schizothorax (Teleostei: Cyprinidae) in China inferred from cytochrome b sequences. J Biogeogr 33:1448–1460
He D, Chen Y, Chen Z (2004) Molecular phylogeny of the specialized schizothoracine fishes (Teleostei: Cyprinidae), with their implications for the uplift of the Qinghai-Tibetan Plateau. Chin Sci Bull 49:39–48
Jin Z (1998) Study on the floristic elements of seed plant in the dry-warm valleys of Yunnan and Sichuan. Guihaia 18:313–321
Jin Z (1999) The floristic study on seed plants in the dry-hot valleys in Yunnan and Sichuan. Guihaia 19:1–14
Jin Z (2002) Floristic features of dry-hot and dry-warm valleys in Yunnan and Sichuan. Yunnan Science and Technology Press, Kunming
Jin Z, Ou X, Ou P, Chen J (1994) A preliminary study on the floristic characteristics of seed plants in the dry-hot river valley of Jinshajiang. Acta Botanica Yunnanica 16:1–16
Kondo T, Nakagoshi N, Isagi Y (2009) Shaping of genetic structure along Pleistocene and modern river systems in the hydrochorous riparian azalea, Rhododendron ripense (Ericaceae). Am J Bot 96:1532–1543
Kottelat M (1989) Zoogeography of the fishes from Indochinese inland waters with an annotated check-list. Bull Zool Mus Univ Amsterdam 12:1–56
Luan S, Chiang TY, Gong X (2006) High genetic diversity vs. low genetic differentiation in Nouelia insignis (Asteraceae), a narrowly distributed and endemic species in China, revealed by ISSR fingerprinting. Ann Bot 98:583
Metivier F, Gaudemer Y, Tapponnier P, Klein M (1999) Mass accumulation rates in Asia during the Cenozoic. Geophys J Int 137:280–318
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Page RDM (1996) Tree view: an application to display phylogenetic trees on personal computers. Bioinformatics 12:357–358
Peng Y, Hu Y, Sun H (2003) Allozyme analysis of Nouelia insignis and its meanings of biogeography and conservation biology. Acta Botanica Yunnanica 25:563–571
Peng Z, He S, Zhang Y (2004) Phylogenetic relationships of glyptosternoid fishes (Siluriformes: Sisoridae) inferred from mitochondrial cytochrome b gene sequences. Mol Phylogenet Evol 31:979–987
Peng Z, Ho SYW, Zhang Y, He S (2006) Uplift of the Tibetan plateau: evidence from divergence times of glyptosternoid catfishes. Mol Phylogenet Evol 39:568–572
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reisch C (2007) Genetic structure of Saxifraga tridactylites (Saxifragaceae) from natural and man-made habitats. Conserv Genet 8:893–902
Rohlf F (2000) NTSYSpc: numerical taxonomy and multivariate analysis system, Version 2.1 user guide. Applied Biostatistics Inc, New York
Rüber L, Britz R, Kullander SO, Zardoya R (2004) Evolutionary and biogeographic patterns of the Badidae (Teleostei: Perciformes) inferred from mitochondrial and nuclear DNA sequence data. Mol Phylogenet Evol 32:1010–1022
Seeber L (1983) River profiles along the Himalayan arc as indicators of active tectonics. Tectonophysics 92:335–367
Shi Z, Ming Q, Dong M (2006) A new discussion on the origin of the east turn of Yangtze River. Yunnan Geogr Environ Res 18:1–6
Shui Y, Chen W, Li Z, Huang S, Zhang K (2003) Seed plants of Honghe region in SE Yunnan, China. Yunnan Science and Technology Press, Kunming
Sun HL, Zheng D (1998) Formation, evolution and development of Qinghai-Xizang (Tibetan) Plateau. Guangdong Science and Technology Press, Guangzhou
Vekemans X, Beauwens T, Lemaire M, Roldan-Ruiz I (2002) Data from amplified fragment length polymorphism (AFLP) markers show indication of size homoplasy and of a relationship between degree of homoplasy and fragment size. Mol Ecol 11:139–151
Wang HS, Zhang YL (1994) The biodiversity and characters of spermatophytic genera endemic to China. Acta Botanica Yunnanica 16:209–220
Wilson EO (1992) Tht diversity of life. The Belknap Press of Harvard University Press, Cambridge
Wright S (1951) The genetical structure of populations. Ann Eugen 15:323–354
Wright S (1969) Theory of gene frequencies: evolution and the genetics of populations, vol 2. University of Chicago Press, Chicago
Wu ZY (1988) Hengduan mountain flora and her significance. J Jpn Bot 63:297–311
Yeh FC, Yang RC, Boyle T (1999) POPGENE: Microsoft Window-Based Freeware for Population Genetic Analysis, version 1.31. Distributed by the authors University of Alberta and Centre for International Forestry Research. Edmonton, Alberta
Ying T, Zhang Y, Boufford DE (1993) The endemic genera of seed plants of China. Science Press, Beijing
Yuan QJ, Zhang ZY, Peng H, Ge S (2008) Chloroplast phylogeography of Dipentodon (Dipentodontaceae) in southwest China and northern Vietnam. Mol Ecol 17:1054–1065
Zhang Q, Chiang TY, George M, Liu JQ, Abbott RJ (2005) Phylogeography of the Qinghai-Tibetan Plateau endemic Juniperus przewalskii (Cupressaceae) inferred from chloroplast DNA sequence variation. Mol Ecol 14:3513–3524
Zhang D, Chen S, Gao Q (2007a) Patterns of genetic variation in Swertia przewalskii, an endangered endemic species of the Qinghai-Tibet Plateau. Biochem Genet 45:33–50
Zhang ZY, Zheng XM, Ge S (2007b) Population genetic structure of Vitex negundo (Verbenaceae) in Three-Gorge Area of the Yangtze River: the riverine barrier to seed dispersal in plants. Biochem Syst Ecol 35:506–516
Acknowledgments
We thank Zhimin Li, Gao Chen, Xuekun Yue, Hua Zhu and Yunhui Zhu for help with the field survey and leaf collection. We are grateful to Qin Qiao, Zelong Nie, Jipei Yue, Yonghong Zhang, Liangliang Yue, Hongguang Zha and Linyang Chen for assistance with the molecular techniques and data analysis. We also appreciate the help of Daniel Kister and Prof. Hans Peter Comes in language improvement. This work was supported by the National 973 Project (2007CB411601); the Natural Science Foundation of China (30625004, 40771073); and Yunnan Natural Science Foundation (2008CC013).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, T., Sun, H. Phylogeographic structure of Terminalia franchetii (Combretaceae) in southwest China and its implications for drainage geological history. J Plant Res 124, 63–73 (2011). https://doi.org/10.1007/s10265-010-0360-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-010-0360-3