Abstract
Gene coexpression analyses are a powerful method to predict the function of genes and/or to identify genes that are functionally related to query genes. The basic idea of gene coexpression analyses is that genes with similar functions should have similar expression patterns under many different conditions. This approach is now widely used by many experimental researchers, especially in the field of plant biology. In this review, we will summarize recent successful examples obtained by using our gene coexpression database, ATTED-II. Specifically, the examples will describe the identification of new genes, such as the subunits of a complex protein, the enzymes in a metabolic pathway and transporters. In addition, we will discuss the discovery of a new intercellular signaling factor and new regulatory relationships between transcription factors and their target genes. In ATTED-II, we provide two basic views of gene coexpression, a gene list view and a gene network view, which can be used as guide gene approach and narrow-down approach, respectively. In addition, we will discuss the coexpression effectiveness for various types of gene sets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gene expression patterns obtained by microarray experiments provide valuable information about gene-to-gene functional relationships, and thus they have been used to cluster functionally related genes since the dawn of microarray technology (Eisen et al. 1998). The use of microarray technology has rapidly spread and produced vast amounts of gene expression data for various species, and now these expression data are available in the public databases, such as NCBI GEO (Barrett et al. 2007), ArrayExpress (Rocca-Serra et al. 2003), TAIR (Swarbreck et al. 2008) and NASCArrays (Craigon et al. 2004). By analyzing these coexpression data, we can evaluate the similarity of expression patterns, and the gene pairs with similar expression patterns are called coexpressed gene pairs. Conceptually, gene coexpression can be defined with a small number of microarray experiments in a similar manner to traditional gene clustering, but “more is different”, as proposed by Anderson (1972). A large amount of expression data will yield a dramatically different scope of gene coexpression, and this gene coexpression information can be used as a fundamental gene map, rather than a simple gene classification.
In the past several years, coexpression approaches have been intensively applied to many biological targets, such as enzymes in a metabolic pathway, subunits of protein complexes and transcription factors (see reviews Aoki et al. 2007; Saito et al. 2008; Usadel et al. 2009). The coexpression data enable us to speculate about the functions of uncharacterized genes of interest and to search for new genes that are functionally related to a phenomenon under investigation. The success of gene coexpression approaches has given rise to several gene coexpression databases in the field of plant biology (Table 1).
We started to develop our coexpression database, ATTED (Obayashi et al. 2004), in November 2003. ATTED is one of the oldest gene coexpression databases, and it is presently available on-line as ATTED-II at http://atted.jp (Obayashi et al. 2007, 2009), with remarkable extensions from the original version. Comparisons of some of the details among these coexpression databases were provided in another recent review (Usadel et al. 2009). In this review, we would like to summarize a few successful examples of coexpression analyses with ATTED-II and to describe the most effective usages of ATTED-II based on the examples.
Brief introduction of ATTED-II
ATTED-II provides two different ways to examine gene coexpression information, a gene list view and a gene network view. The coexpressed gene list and the gene network for each gene in the Arabidopsis genome with the expression data were previously constructed, and the user can easily access the information. On the other hand, the coexpressed gene lists for multiple guide genes and the coexpression networks for the query genes are provided upon request, using the CoexSearch and NetworkDrawer tools, respectively. These are the two most popular tools in ATTED-II. The former is used for the “guide gene” approach, to find related genes with one or more guide genes, while the latter is used for the “narrow-down” approach, to analyze internal relationships among a set of genes and to identify the core genes in the set.
CoexSearch tool
The CoexSearch tool provides a list of genes that are coexpressed with the guide genes. Therefore, the guide genes are expected to have strong coexpression with each other, because this tool identifies the coexpressed genes based on the average coexpression strength for the guide genes. There is no strict criterion to judge whether the guide genes are strongly coexpressed, but the average values will not be meaningful when each of the guide genes is involved in different regulatory mechanisms.
NetworkDrawer tool
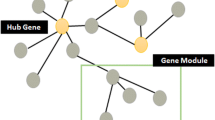
The NetworkDrawer tool accepts any set of genes and analyzes the internal relationships among the query genes. To draw the gene network from the lists of coexpressed genes, a threshold must be determined to define the coexpressed gene pairs. In ATTED-II, the three most strongly coexpressed genes for each gene are used to draw the network. This criterion was determined from the viewpoint of the user’s visibility. Greater numbers of genes can be incorporated into the network, and while the network may become more informative, it also tends to be more difficult to understand.
Quality of the coexpression data
In addition to its user-friendly interfaces, one of the most important features of ATTED-II is the continuous improvement of the coexpression data with the development of new calculation methods for gene coexpression (Obayashi and Kinoshita 2009; Kinoshita and Obayashi 2009). We have quantified the quality of the gene coexpression data by using Gene Ontology Annotation (Obayashi and Kinoshita 2009), and confirmed its improvement for every update (see CoexVersion for the version history; http://atted.jp/top_search.shtml#coexversion). The user can download all of the coexpression data for further analyses. Our coexpression data are actually used in several other databases and web tools, such as PRIMe (Akiyama et al. 2008) to produce network files, CoP (Ogata et al. 2009a, b) to find coexpression network modules, KaPPA-View3 (Tokimatsu et al. 2005) to integrate transcriptomics and metabolomics analyses on pathways, PosMed (Yoshida et al. 2009) for positional cloning, and Ondex (Lysenko et al. 2009) to integrate various omics data.
Other details of ATTED-II can be found in Obayashi et al. (2007, 2009).
Examples using ATTED-II with experimental verifications
To understand the strengths and weaknesses of the gene coexpression approach, we summarized successful studies that employed ATTED-II. The reports using the guide gene approach or CoexSearch tool are summarized in Table 2, and those using the narrow-down approach or NetworkDrawer tool are shown in Table 3.
The studies by Ishihara et al. (2007), Takahashi et al. (2008) and Yamada et al. (2008) represent good examples of a single guide gene approach, where one target gene of interest for each study was already specified. However, this is not the most common scenario, and we often cannot decide which gene is the best one to use as the guide gene to identify functionally related genes. In such a case, multiple guide genes may be used to generate a single coexpressed gene list. The CoexSearch tool in ATTED-II provides a unified coexpressed gene list from multiple guide genes, by merging multiple gene lists based on the average MR (Mutual Rank) value, because our studies have shown that the MR value is more effective than the Pearson’s correlation coefficient (PCC), a popular coexpression measure (Obayashi and Kinoshita 2009). The actual MR values for each study are also shown in Table 2. In most of the successful studies, the genes with low MR values (i.e., tightly coexpressed genes) were used to design the experiments to verify the coexpression analyses, and the values ranged from MR = 1.4 in Ishihara et al. 2007 to MR = 67.7 in Takabayashi et al. 2009. However, there are two studies that used weaker coexpression (Bednarek et al. 2009; Sugano et al. 2010). We are not sure why these two cases were successful, but one possibility is that construction of a unified gene list with multiple guide genes may dramatically reduce the number of unrelated genes with relatively good average MR values. For example, in the study by Sugano et al. (2010), three guide genes (TMM, SDD, EPF1) were used to search for genes related to stomata development. If they had used a single guide gene, they would not have found STOMAGEN, the gene newly identified in their study, because this gene does not appear in the list of the top 300 coexpressed genes provided by ATTED-II. However, when they used three guide genes with the CoexSearch tool in ATTED-II, STOMAGEN appeared as the 17th ranked gene (Sugano et al. 2010). Bednarek et al. (2009) also used multiple guide genes, and they restricted their target to the cytochrome P450 gene family for the expected hydroxylation reaction in the glucosinolate biosynthetic pathway. This represents another approach to use weakly coexpressed genes as guide genes.
The narrow-down approach is useful to find the core coexpression module in a set of genes. Genes are often selected by using the pathway information in other databases, such as KaPPA-View (Tokimatsu et al. 2005), KEGG (Kanehisa et al. 2008) or Gene Ontology (GO, Ashburner et al. 2000). The NetworkDrawer tool in ATTED-II provides the internal structure of gene coexpression as a network, and the EdgeAnnotation tool provides it in a table. Since the output of the NetworkDrawer tool is a picture file and cannot be edited manually, we also provide the input format for other network drawers, such as Pajek (Batagelj and Mrvar 1998) or Cytoscape (Cline et al. 2007). Narrow-down approach was intensively used to identify genes for secondary metabolite pathways (Hirai et al. 2007; Yonekura-Sakakibara et al. 2007; Tohge et al. 2007; Sano et al. 2008; Okazaki et al. 2009; Sawada et al. 2009a, b). Okazaki et al. combined both the list approach and network approach. They first used the network approach to find the core coexpression module in a gene set for lipid metabolism, and then searched the coexpression data using the genes in the core coexpression module (Okazaki et al. 2009).
As described above, the MR and unified list methods are the keys for understanding the reason why the coexpression analyses worked well. In addition to these two reasons, the expression levels of the identified genes in Tables 2 and 3 were also investigated. As shown in Fig. 1, the identified genes were accumulated at the 50th to 80th percentile expression levels, and thus there may be some tendencies for highly expressed genes to be more suitable for coexpression analyses. The reason for this tendency is not straightforward, but one possibility is that the coexpression data of genes with low expression are less accurate, due to microarray noise, or that the phenotype of gene disruption appears more readily for highly expressed genes, as compared to those with lower expression.
Expression levels of the experimentally verified genes shown in Tables 2 and 3. Expression levels were determined using the average MAS5 value against all AtGenExpress developmental series experiments (Schmid et al. 2005). Following genes were used in this plot. As protein complex genes: PPL2, MCM2, MCM3, MCM4, MCM5, MCM7, NDF1, NDF2, NDF4, NDF6. As enzyme genes: UGT78D3, RHM1, CYP81F2, UGT89C1, OMT1, UGP3, MAM-IL, MAM-D. As TF genes: Myb28, Myb29, bHLHl15, LIM, C3H, HMG, hp_5g26. The other genes: PPL1, STOMAGEN, BASS5, NAI2. Two enzyme genes with low expression level are UGT78D3 and CYP81F2
Suitable types of genes as targets of gene coexpression analyses
As shown in Tables 2 and 3, coexpression approaches have been applied to several experimental targets. Since these targets are quite diverse, we wondered whether there were any tendencies of the types of genes that are suitable targets of the gene coexpression analysis. To answer this question, we examined the coexpression tendencies in functional groups of genes with the same Gene Ontology (GO) term and with the coexpression data in ATTED-II version 5.5.
First, we evaluated background distribution by randomly selecting 20–30 genes and ploting the cumulative frequency distribution of their MR values among all pairs of the selected genes. Figure 2a shows the results for 200 repetitions. Most of the MR distributions were uniform, indicating that the MR values do not have strong biases in the ATTED-II coexpression data.
Degree of coexpression strength of Gene Ontology (GO) terms. MR is the coexpression measure used in ATTED-II. MR = 1 indicates the strongest coexpression. a Coexpression strength in randomly selected genes, b that in genes in each of 64 GO Biological Process (BP) annotations, c that in genes in each of 14 GO Cellular Component (CC) annotations, d that in genes in each of 34 GO Molecular Function (MF) annotations
We then checked the MR distributions of the gene pairs in the gene groups sharing the same functional annotation and including 20–30 genes. The MR distributions for the genes for every GO Biological Process (BP) term are shown in Fig. 2b. Most of the GO BP groups are located in the upper-left area in this graph (Fig. 2b), indicating that the genes are coexpression modules in the gene set, because the preference of this region in the figure means the existence of strong gene coexpression, or a smaller MR value, among the selected genes. Figure 2c, d show the cases for GO Cellular Component (CC) and GO Molecular Function (MF), and the characteristics of these graphs were almost the same as those of GO BP.
To discuss these results quantitatively, the differences between each curve and diagonal line, or uniform distribution, in Fig. 2 were calculated, and the lists of GO terms with large deviations from the diagonal lines were generated, as potentially effective functional groups for gene coexpression analysis (Supplemental Table S1). From these lists, we chose some of the most, medium and least strongly coexpressed GO terms, as shown in Table 4. Basically almost all of the gene sets for the GO terms showed significant coexpression, as compared with the random distribution. It should be noted that statistical significance does not guarantee biological relevance, but it is possible that almost all of the gene pairs sharing the same GO annotation will have some weak but definite coexpression, as described later.
The most strongly coexpressed gene sets were the genes encoding the components of supramolecules, such as photosynthesis machinery, ribosomes and transcription machinery (Table 4). This observation is quite reasonable, because all of the components of a supramolecule should co-exist, to establish the three-dimensional structure of the molecule. In a similar manner, the genes encoding proteins that form tight complex structures will be strongly coexpressed.
The moderately coexpressed gene sets involved quite divergent genes, such as those for biosynthesis, stress responses or intercellular complexes (Table 4). The genes in this category have several strong coexpression cores and weak coexpression between the cores in the pathway.
Since almost all of the gene sets participate in coexpression modules, understanding the rarely coexpressed gene sets is important when using the coexpression data. The most rarely coexpressed gene sets were the genes for signaling pathways, such as transport and receptor binding (Table 4). Signaling pathways are mainly regulated at the protein-level, by modifications such as phosphorylation, whereas gene coexpression reflects the regulatory relationships of mRNA, and protein level regulation is not directly reflected. This limitation was clarified by Pitzschke and Hirt (2010) using the MAPK cascade as an example. The data in Table 4 agree with their statement, as in the case of the lower D values of signaling pathways. However, this result does not mean that the gene coexpression data will not be useful for studying signaling pathways, because the proteins regulating the signaling proteins, such as phosphatases and kinases, should also be regulated at the mRNA-level, since all of the participants in the signaling pathway must co-exist. When such weak coexpression information is used as in the signaling pathway, one of the important points to consider is to adopt other information about gene function at the same time, as in the case of Vicinanza et al. (2008). They discussed the coexpression network of phosphatases and kinases for the phosphoinositide system, using the human coexpression data provided in COXPRESdb, a coexpression database for animals (Obayashi et al. 2008). In their discussion, they employed a relatively low coexpression threshold (PCC > 0.4) and drew a gene network only for kinases and phosphatases (Vicinanza et al. 2008). The restriction of gene coexpression data by using the gene family is a promising approach, because it naturally combines genome information and coexpression information as in the guide gene approach. Although the identification of a signaling pathway is generally difficult, Sugano et al. (2010) recently reported a successful study about the identification of an intercellular signaling factor for stomata, STOMAGEN. In the case of transporters, which are also one of the most difficult targets, Sawada et al. (2009b) identified a transporter for glucosinolate biosynthesis.
How many genes should be examined?
Gene coexpression is defined by continuous values (MR or PCC), and there is no optimal threshold to define gene coexpression. In ATTED-II, we provide the top 300 MR genes for each guide gene. The reason why we selected 300 genes was based on a practical reason, since a gene list of this size can easily be checked visually. However, MR values around MR = 5,000 can still be meaningful to indicate the existence of gene relationships (Fig. 2), although such coexpression may also include indirect gene-to-gene relationships.
Based on the average distributions for each GO type in Fig. 2, we estimated the effective threshold to distinguish real coexpression pairs and random pairs. As a result, the most effective threshold was MR = 5,769 for BP, 3,625 for CC and 5,369 for MF. In ATTED-II, we provide the 300 most highly coexpressed genes for each gene, but it may be worthwhile to check for more weakly coexpressed genes, which can be obtained from the download page.
As one of the strategies to increase the reliability of coexpression data, ATTED-II provides a comparative view between Arabidopsis coexpression and rice coexpression, using orthologous genes. According to the determination of the significant MR threshold, we roughly set MR = 1,000, to highlight the conserved coexpression in other species.
Future directions
In this review, we have described the potential power of gene coexpression analyses by summarizing some successful examples with ATTED-II, and we also discussed a few limitations of these types of analyses. To overcome these limitations and enhance the power of ATTED-II, we are planning to integrate genome information and protein information into the coexpression information, in addition to continuously improving the coexpression data. The cis element analysis is one such approach to use genome information. ATTED-II also provides cis element prediction near the transcriptional start site, and the results were validated for some cases (Masuda and Fujita 2008). We are improving this cis element prediction function to understand gene coexpression. Although we did not discuss a comparison of coexpression among different species, the coexpression of orthologous genes in different species provides valuable information to enhance the reliability of coexpression data, and thus we will increase the number of target species in ATTED-II.
References
Akiyama K, Chikayama E, Yuasa H, Shimada Y, Tohge T, Shinozaki K, Hirai MY, Sakurai T, Kikuchi J, Saito K (2008) PRIMe: a Web site that assembles tools for metabolomics and transcriptomics. In Silico Biol 8:339–345
Anderson PW (1972) More is different. Science 177:393–396
Aoki K, Ogata Y, Shibata D (2007) Approaches for extracting practical information from gene co-expression networks in plant biology. Plant Cell Physiol 48:381–390
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles—database and tools update. Nucleic Acids Res 35:D760–D765
Batagelj V, Mrvar A (1998) Pajek program for large network analysis. Connections 21:47–57
Bednarek P, Pislewska-Bednarek M, Svatos A, Schneider B, Doubsky J, Mansurova M, Humphry M, Consonni C, Panstruga R, Sanchez-Vallet A, Molina A, Schulze-Lefert P (2009) A glucosinolate metabolism pathway in living plant cells mediates broad-spectrum antifungal defense. Science 323:101–106
Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N, Workman C, Christmas R, Avila-Campilo I, Creech M, Gross B, Hanspers K, Isserlin R, Kelley R, Killcoyne S, Lotia S, Maere S, Morris J, Ono K, Pavlovic V, Pico AR, Vailaya A, Wang PL, Adler A, Conklin BR, Hood L, Kuiper M, Sander C, Schmulevich I, Schwikowski B, Warner GJ, Ideker T, Bader GD (2007) Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2:2366–2382
Craigon DJ, James N, Okyere J, Higgins J, Jotham J, May S (2004) NASCArrays: a repository for microarray data generated by NASC’s transcriptomics service. Nucleic Acids Res 32:D575–D577
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Hirai MY, Sugiyama K, Sawada Y, Tohge T, Obayashi T, Suzuki A, Araki R, Sakurai N, Suzuki H, Aoki K, Goda H, Nishizawa OI, Shibata D, Saito K (2007) Omics-based identification of Arabidopsis Myb transcription factors regulating aliphatic glucosinolate biosynthesis. Proc Natl Acad Sci USA 104:6478–6483
Ishihara S, Takabayashi A, Ido K, Endo T, Ifuku K, Sato F (2007) Distinct functions for the two PsbP-like proteins PPL1 and PPL2 in the chloroplast thylakoid lumen of Arabidopsis. Plant Physiol 145:668–679
Ishikawa N, Takabayashi A, Ishida S, Hano Y, Endo T, Sato F (2008) NDF6: a thylakoid protein specific to terrestrial plants is essential for activity of chloroplastic NAD(P)H dehydrogenase in Arabidopsis. Plant Cell Physiol 49:1066–1073
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–484
Kinoshita K, Obayashi T (2009) Multi-dimensional correlations for gene coexpression and application to the large-scale data of Arabidopsis. Bioinformatics 25:2677–2684
Lee TH, Kim YK, Pham TT, Song SI, Kim JK, Kang KY, An G, Jung KH, Galbraith DW, Kim M, Yoon UH, Nahm BH (2009) RiceArrayNet: a database for correlating gene expression from transcriptome profiling, and its application to the analysis of coexpressed genes in rice. Plant Physiol 151:16–33
Lysenko A, Hindle MM, Taubert J, Saqi M, Rawlings CJ (2009) Data integration for plant genomics—exemplars from the integration of Arabidopsis thaliana databases. Brief Bioinform 10:676–693
Manfield IW, Jen C-H, Pinney JW, Michalopoulos I, Bradford JR, Gilmartin PM, Westhead DR (2006) Arabidopsis Co-expression Tool (ACT): web server tools for microarray-based gene expression analysis. Nucleic Acids Res 34:W504–W509
Masuda T, Fujita Y (2008) Regulation and evolution of chlorophyll metabolism. Photochem Photobiol Sci 7:1131–1149
Mutwil M, Obro J, Willats WGT, Persson S (2008) GeneCAT—novel webtools that combine BLAST and co-expression analyses. Nucleic Acids Res 36:W320–W326
Obayashi T, Kinoshita K (2009) Rank of correlation coefficient as a comparable measure for biological significance of gene coexpression. DNA Res 16:249–260
Obayashi T, Okegawa T, Sasaki-Sekimoto Y, Shimada H, Masuda T, Asamizu E, Nakamura Y, Shibata D, Tabata S, Takamiya K, Ohta H (2004) Distinctive features of plant organs characterized by global analysis of gene expression in Arabidopsis. DNA Res 11:11–25
Obayashi T, Kinoshita K, Nakai K, Shibaoka M, Hayashi S, Saeki M, Shibata D, Saito K, Ohta H (2007) ATTED-II: a database of co-expressed genes and cis elements for identifying co-regulated gene groups in Arabidopsis. Nucleic Acids Res 35:D863–D869
Obayashi T, Hayashi S, Shibaoka M, Saeki M, Ohta H, Kinoshita K (2008) COXPRESdb: a database of coexpressed gene networks in mammals. Nucleic Acids Res 36:D77–D82
Obayashi T, Hayashi S, Saeki M, Ohta H, Kinoshita K (2009) ATTED-II provides coexpressed gene networks for Arabidopsis. Nucleic Acids Res 37:D987–D991
Ogata Y, Suzuki H, Shibata D (2009a) A gene co-expression database for understanding biological process in soybean. Plant Biotechnol 26:503–507
Ogata Y, Suzuki H, Shibata D (2009b) A database for poplar gene co-expression analysis for systematic understanding of biological processes, including stress responses. J Wood Sci 55:395–400
Okazaki Y, Shimojima M, Sawada Y, Toyooka K, Narisawa T, Mochida K, Tanaka H, Matsuda F, Hirai A, Hirai MY, Ohta H, Saito K (2009) A chloroplastic UDP-glucose pyrophosphorylase from Arabidopsis is the committed enzyme for the first step of sulfolipid biosynthesis. Plant Cell 21:892–909
Pitzschke A, Hirt H (2010) Bioinformatic and systems biology tools to generate testable models of signaling pathways and their targets. Plant Physiol 152:460–469
Rocca-Serra P, Brazma A, Parkinson H, Sarkans U, Shojatalab M, Contrino S, Vilo J, Abeygunawardena N, Mukherjee G, Holloway E, Kapushesky M, Kemmeren P, Lara GG, Oezcimen A, Sansone SA (2003) ArrayExpress: a public database of gene expression data at EBI. C R Biol 326:1075–1078
Saito K, Hirai MY, Yonekura-Sakakibara K (2008) Decoding genes with coexpression networks and metabolomics—‘majority report by precogs’. Trends Plant Sci 13:36–43
Sano R, Ogata Y, Suzuki H, Ogawa Y, Dansako T, Sakurai N, Okazaki K, Aoki K, Saito K, Shibata D (2008) Over-expression of transcription associated factor genes coexpressed with genes of the mevalonate pathway, upstream of isoprenoid biosynthesis, in Arabidopsis cultured cells. Plant Biotechnol 25:583–587
Sawada Y, Kuwahara A, Nagano M, Narisawa T, Sakata A, Saito K, Hirai MY (2009a) Omics-based approaches to methionine side chain elongation in Arabidopsis: characterization of the genes encoding methylthioalkylmalate isomerase and methylthioalkylmalate dehydrogenase. Plant Cell Physiol 50:1181–1190
Sawada Y, Toyooka K, Kuwahara A, Sakata A, Nagano M, Saito K, Hirai MY (2009b) Arabidopsis bile acid: sodium symporter family protein 5 is involved in methionine-derived glucosinolate biosynthesis. Plant Cell Physiol 50:1579–1586
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Srinivasasainagendra V, Page GP, Mehta T, Coulibaly I, Loraine AE (2008) CressExpress: a tool for large-scale mining of expression data from Arabidopsis. Plant Physiol 147:1004–1016
Steinhauser D, Usadel B, Luedemann A, Thimm O, Kopka J (2004) CSB.DB: a comprehensive systems-biology database. Bioinformatics 20:3647–3651
Sugano SS, Shimada T, Imai Y, Okawa K, Tamai A, Mori M, Hara-Nishimura I (2010) Stomagen positively regulates stomatal density in Arabidopsis. Nature 463:241–244
Swarbreck D, Wilks C, Lamesch P, Berardini TZ, Garcia-Hernandez M, Foerster H, Li D, Meyer T, Muller R, Ploetz L, Radenbaugh A, Singh S, Swing V, Tissier C, Zhang P, Huala E (2008) The Arabidopsis Information Resource (TAIR): gene structure and function annotation. Nucleic Acids Res 36:D1009–D1014
Takabayashi A, Ishikawa N, Obayashi T, Ishida S, Obokata J, Endo T, Sato F (2009) Three novel subunits of Arabidopsis chloroplastic NAD(P)H dehydrogenase identified by bioinformatic and reverse genetic approaches. Plant J 57:207–219
Takahashi N, Lammens T, Boudolf V, Maes S, Yoshizumi T, De Jaeger G, Witters E, Inze D, De Veylder L (2008) The DNA replication checkpoint aids survival of plants deficient in the novel replisome factor ETG1. EMBO J 27:1840–1851
Tohge T, Yonekura-Sakakibara K, Niida R, Watanabe-Takahashi A, Saito K (2007) Phytochemical genomics in Arabidopsis thaliana: a case study for functional identification of flavonoid biosynthesis genes. Pure Apple Chem 79:811–823
Tokimatsu T, Sakurai N, Suzuki H, Ohta H, Nishitani K, Koyama T, Umezawa T, Misawa N, Saito K, Shibata D (2005) KaPPA-view: a web-based analysis tool for integration of transcript and metabolite data on plant metabolic pathway maps. Plant Physiol 138:1289–1300
Toufighi K, Brady M, Austin R, Ly E, Provart N (2005) The botany array resource: e-northerns, expression angling, and promoter analyses. Plant J 43:153–163
Usadel B, Obayashi T, Mutwil M, Giorgi FM, Bassel GW, Tanimoto M, Chow A, Steinhauser D, Persson S, Provart NJ (2009) Co-expression tools for plant biology: opportunities for hypothesis generation and caveats. Plant Cell Environ 32:1633–1651
Vicinanza M, D’Angelo G, Di Campli A, De Matteis MA (2008) Function and dysfunction of the PI system in membrane trafficking. EMBO J 27:2457–2470
Yamada K, Nagano AJ, Nishina M, Hara-Nishimura I, Nishimura M (2008) NAI2 is an endoplasmic reticulum body component that enables ER body formation in Arabidopsis thaliana. Plant Cell 20:2529–2540
Yonekura-Sakakibara K, Tohge T, Niida R, Saito K (2007) Identification of a flavonol 7-O-rhamnosyltransferase gene determining flavonoid pattern in Arabidopsis by transcriptome coexpression analysis and reverse genetics. J Biol Chem 282:14932–14941
Yonekura-Sakakibara K, Tohge T, Matsuda F, Nakabayashi R, Takayama H, Niida R, Watanabe-Takahashi A, Inoue E, Saito K (2008) Comprehensive flavonol profiling and transcriptome coexpression analysis leading to decoding gene-metabolite correlations in Arabidopsis. Plant Cell 20:2160–2176
Yoshida Y, Makita Y, Heida N, Asano S, Matsushima A, Ishii M, Mochizuki Y, Masuya H, Wakana S, Kobayashi N, Toyoda T (2009) PosMed (Positional Medline): prioritizing genes with an artificial neural network comprising medical documents to accelerate positional cloning. Nucleic Acids Res 37:W147–W152
Acknowledgments
This work was supported by a Grant-in-Aid for Scientific Research (No. 50397048) to TO and a Grant-in-Aid for Scientific Research on Priority Areas “Transportsome” to KK. Computation time was provided by the Super Computer System, Human Genome Center, Institute of Medical Science, The University of Tokyo.
Author information
Authors and Affiliations
Corresponding author
Additional information
Takeshi Obayashi is the recipient of the BSJ Special Award, 2009.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Obayashi, T., Kinoshita, K. Coexpression landscape in ATTED-II: usage of gene list and gene network for various types of pathways. J Plant Res 123, 311–319 (2010). https://doi.org/10.1007/s10265-010-0333-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-010-0333-6