Abstract
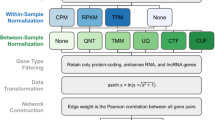
The use of systems biology methods to establish gene co-expression networks to reveal the relationship between genes at the system level has become a major area of research. Gene co-expression networks are increasingly used to explore the system-level functions of genes. They have been extensively studied and used to predict new gene functions, discover new disease biomarkers, and detect genetic variants in cancer. The gene co-expression network is conceptually very simple. The nodes represent genes, and the lines between nodes represent the interaction between genes and genes. The data set used in this article is the RNA-Seq data set of corn. The innovation of this article is to propose a new data standardization method, which overcomes the data inconsistency caused by different experimental platforms and different software analysis tools. In addition, this article also proposes a new similarity measurement method to prove the effectiveness of this method through the ROC curve and AUROC.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wang, Z., et al.: VCNet: vector-based gene co-expression network construction and its application to RNA-seq data. Bioinformatics 33(14), 2173–2181 (2017)
Hou, J., Ye, X., Li, C., et al.: K-Module algorithm: an additional step to improve the clustering results of WGCNA co-expression networks. Genes 12(1), 87 (2021)
Tang, J., et al.: Prognostic genes of breast cancer identified by gene co-expression network analysis. Front. Oncol. 8, 374 (2018)
McKenzie, A.T., et al.: Brain cell type specific gene expression and co-expression network architectures. Sci. Rep. 8(1), 1–19 (2018)
Zhang, Z., Cui, F., Wang, C., et al.: Goals and approaches for each processing step for single-cell RNA sequencing data. Brief. Bioinform. 22(4), bbaa314 (2020)
Wang, B., Liu, D., Peng, X., et al.: Data-driven anomaly detection of UAV based on multimodal regression model. In: 2019 IEEE International Instrumentation and Measurement Technology Conference (I2MTC), pp. 1–6. IEEE (2019)
Huang, J., Vendramin, S., Shi, L., et al.: Construction and optimization of a large gene coexpression network in maize using RNA-Seq data. Plant Physiol. 175(1), 568–583 (2017)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Wu, X., Song, X. (2022). Research on Gene Coexpression Network Based on RNA-Seq Data. In: Liu, Q., Liu, X., Chen, B., Zhang, Y., Peng, J. (eds) Proceedings of the 11th International Conference on Computer Engineering and Networks. Lecture Notes in Electrical Engineering, vol 808. Springer, Singapore. https://doi.org/10.1007/978-981-16-6554-7_67
Download citation
DOI: https://doi.org/10.1007/978-981-16-6554-7_67
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-6553-0
Online ISBN: 978-981-16-6554-7
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)