Abstract
To identify discriminating protein patterns in serum samples between gastric cancer patients (early and advanced stages) and healthy controls. We used magnetic bead-based separation followed by matrix-assisted laser desorption ionization (MALDI) time-of-flight (TOF) mass spectrometry (MS) to identify patients with gastric cancer. In total, serum samples from 62 gastric cancer patients (32 in the training set and 30 in the test set; 19 of which had early-stage tumors and 43 of which had advanced-stage tumors) and 64 healthy controls (32 in the training set and 32 in the test set) were analyzed. The mass spectra, analyzed using ClinProTools software, distinguished between cancer patients and healthy individuals based on three different algorithm models. In the training set, patients with gastric cancer could be identified with a mean sensitivity of 94.7% and a mean specificity of 99%. Similar results were obtained with the test set, showing 79.3% sensitivity and 86.5% specificity. Our study demonstrates the high sensitivity and specificity of screening serum protein patterns using MALDI-TOF MS for the identification of patients with gastric cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer is the fourth most common cancer and the second most common cause of cancer-related death worldwide [1]. It affects approximately one million individuals, with 8,70,000 deaths annually, and it remains a global killer with a shifting burden from the developed to the developing world. Almost two-thirds of gastric cancer cases occur in developing countries and 42% of cases occur in China alone [2]. Gastric cancer mortality rates are higher in rural areas than in urban areas and are higher in men than in women, according to the data from the Center of Health Information Statistics (CHIS) under the Ministry of Public Health, China [3]. The factors that influence in incidence of gastric cancer in China have been reported to be lifestyle, nutrition and diet, education, and the health care system. Despite the remarkable decline in gastric cancer mortality during the last two decades due to dramatic improvements in socialeconomic factors, gastric cancer is still the predominant form of cancer and remains a significant cancer burden in China with a poor prognosis [3, 4].
Most gastric cancer patients are diagnosed with advanced-stage disease for which treatment options are limited contributing to the overall 5-year survival rate of less than 25% [5]. Early detection and accurate preoperative staging of an early gastric cancer offer the best prognosis and are essential for planning optimal therapy [1]. In Japan, experience has shown that the detection and treatment for gastric cancer in its early stages can significantly improve disease outcome and patient survival [6, 7]. Because of the limited sensitivity and specificity of the commonly used gastric tumor markers, which include carcinoembryonic antigen (CEA), CA 19-9, and α-fetoprotein antigen (AFP) [8], there is an urgent need to identify novel biomarkers for early diagnostic screening purposes.
An effective, clinically useful biomarker should be measurable in a readily accessible body fluid, such as serum, urine, or saliva [9]. The discovery of biomarkers in body fluids has been advanced by the recent introduction of mass spectrometry (MS)-based screening methods, such as surface-enhanced laser desorption/ionization time-of-flight (SELDI-TOF) MS, and matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) MS. The analysis of MS-based serum proteomic patterns is a well established diagnostic tool in routine clinical practice [10]. The SELDI-TOF MS analysis of various body fluids was first used to investigate a variety of different cancer types (e.g., ovarian, lung, prostate, bladder, breast, brain, and liver) [11–14], and this method has been applied to the identification of gastric cancer patients in many cases via serum protein profiling [15–18]. However, spectra generated by SELDI-TOF MS usually show limited peak and binding capacity due to the smaller surface area of the plane protein chip [19]. Moreover, SELDI-TOF MS does not allow for direct identification of the discriminatory proteins and suffers from low reproducibility and accuracy [20–23]. As for clinical diagnostic studies using MS-based proteomic approaches, MALDI-TOF MS has been the most commonly used method due to its relative ease-of-use, its high automation and throughput potential, and good sensitivity [24]. Moreover, magnetic bead-based purification approaches were developed to reduce costs and make serum profiling suitable for general MS analysis. Using magnetic beads, multiple MALDI-TOF measurements can be made using the same preparation.
Identifying differentially expressed peptides and proteins by MS, together with software-generated models that are capable of discriminating between the spectra of patients with gastric cancer and healthy controls, could lead to the identification of potential new marker molecules for gastric cancer. Here, we report an initial clinical study using magnetic bead-based purification approaches coupled with MALDI-TOF MS for the analysis of sera from patients with gastric cancer as well as from healthy controls. The identified marker molecules could aid the early diagnosis of this disease.
Materials and methods
Patients and sample preparation
The study protocol was approved by the ethics committee and the human research review committee of Xi’an Jiaotong University. A total of 62 serum samples (39 men and 23 women; 19 with early-stage disease and 43 with advanced-stage disease) from gastric cancer patients were collected at the First Affiliated Hospital of Xi’an Jiaotong University. Their average age was 59 (ranging from 32 to 78). All of the patients had recently been diagnosed. The histotypes of gastric carcinoma patients were as follows: 24 with poorly differentiated adenocarcinoma, 16 with moderate differentiated adenocarcinoma, 12 with tubular adenocarcinoma, and 10 with mucinous adenocarcinoma.
The 64 healthy control serum samples, consisting of 33 males and 31 females, ranging in age from 31 to 78, with an average age of 51.7, were obtained from health donors recruited for this study. All of the samples used in this study were collected between January and April, 2009. All blood samples were drawn while the patients or healthy controls were seated and non-fasting. The samples were collected in 10 cc serum separator tubes and were kept at 4°C for 1 h, then centrifuged at 3,000g for 20 min at 4°C. The serum samples were distributed into 500 μl aliquots and stored at –80°C until use.
MS analysis: WCX fractionation and MALDI-TOF MS
Magnetic bead-based weak cation-exchange chromatography (MB-WCX) (ClinProt™ purification reagent sets from Bruker Daltonics, Bremen, Germany) was used for peptidome separation of samples. MB-WCX purifications were performed according to the manufacturer’s protocol for serum, using a Bruker Magnetic Separator (8-well, #65554). The protocol was adapted with regard to sample and binding solution volumes. With the magnet lowered, 5 μl serum samples were diluted in 10 μl binding solution in a standard thin wall PCR tube, added to 10 μl of MB-WCX beads and then carefully mixed using the mixing feature of the robot. After thorough stirring, samples were incubated at room temperature for 5 min, then the tubes were placed into the magnetic separator to collect the beads on the wall of the tube until the supernatant was clear (~1 min). The supernatant was then removed and the magnet was lowered again. The magnetic beads were washed three times in MB-WCX washing solution, lifting and lowering the magnet as needed. Following the stepwise application of sample and MB-WCX separation, we eluted the peptide fraction from the magnetic beads with 5 μl of elution solution and 4 μl of stabilization buffer. To prepare the MALDI target, we spotted 1 μl of a mixture containing 10 μl of 0.3 g/l α-cyano-4-hydroxy cinnamic acid in 2:1 ethanol/acetone (v/v) and 1 μl of the eluted peptide fraction onto the MALDI AnchorChip™ (Bruker Daltonics). To evaluate the reproducibility of each serum sample, samples were spotted in triplicate.
Data processing with ClinProt software
Air-dried targets were measured immediately using a calibrated Autoflex III MALDI-TOF MS (Bruker), FlexControl software (version 3.0; Bruker) and optimized measuring protocols. For matrix suppression up to 700 Da, mass calibration was performed with a standard calibration mixture of peptides and proteins (mass range, 1,000–10,000 Da). All measurements were performed in a blinded manner, including the analysis of patient and control sera, which was performed using a mixed approach. The Flex analysis software (version 3.0; Bruker) was applied for data analysis. Clinprotools software (version 2.2; Bruker) was used for the recognition of peptide patterns. This program uses a standard data preparation workflow including spectra pretreatment, peak picking and peak calculation operation.
For statistical analysis, three different algorithms of mathematical models were used: Genetic Algorithm (GA), Supervised Neural Network (SNN), and Quick Classifier (QC). Each model selects and weights a combination of protein peaks that provide high distinguishing ability between the cancer patients and healthy controls and are briefly described. The GA algorithm derives from evolutionary survival in which the best peak clusters are combined into a new feature, and the poor clusters are discarded. This process is iteratively repeated until the optimal peak combination is found. The SNN algorithm maximizes the distance of multiple local peak clusters specific to each group. Clusters that provide greater separation are prioritized over those with low separation. Finally, the QC algorithm generates an average spectrum for each group with weighted P values for each peak. Based on the peak weights, spectra are categorized into either group along with a likeliness value. In this study, the gastric cancer and healthy control groups were subdivided into a training set (32 gastric cancer patients and 32 healthy controls) and a testing set (30 gastric cancer patients and 32 healthy controls) to test the robustness of the classification model.
Sensitivity and specificity, also known in statistics as classification function, are statistical measures of the performance of a binary classification test. Specificity relates to the ability of the test to identify negative results. Sensitivity measures the proportion of actual positives, which are correctly identified as such (e.g., the percentage of sick people who are correctly identified as having the condition). Specificity measures the proportion of negatives, which are correctly identified (e.g., the percentage of healthy people who are correctly identified as not having the condition). In the training set, the sensitivity and specificity were correctness of self-checking. However, in the test set, both measures were correctness of predicting.
Results
In this study, we analyzed the serum peptidome fingerprints of all 62 gastric cancer patients (19 with early-stage disease and 43 with advanced-stage disease) and 64 healthy controls (Table 1). We evaluated changes at the peptidome level in the serum samples of 32 gastric cancer patients compared with 32 healthy controls in the training set. By analyzing the spectra (screened from two groups in the training set) using the ClinProTools software 2.2, we were able to identify proteomic patterns that can clearly distinguish between gastric cancer patients and healthy controls.
MALDI spectrum generation and assay reproducibility
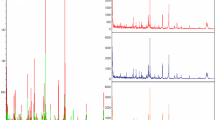
Using MB-WCX magnetic beads on pre-fractionated serum samples and MALDI-TOF MS revealed, on average, up to 119 peaks, of which 46 significant m/z peaks with a P value for the Wilcoxon rank sum test of <0.001 were detected between the gastric cancer group and the healthy controls in the training set (Table 2). Overall, the gastric cancer patients (red) and healthy controls (green) demonstrated protein profiles from 1 to 10 kDa (Fig. 1). Within this mass range, large numbers of differentially expressed proteins or peptides could be detected.
To evaluate the reproducibility and stability of the mass spectra, the Gel view of mass spectra of three different samples (three spectra for each sample), two of which were from gastric cancer patients and one was from a healthy control, showed closely reproducible peaks (Fig. 2). In addition, mass spectra were different among the different samples, even if they were both from gastric cancer patients.
Peak selection and model testing
Utilizing data from the training set, three different classification models for gastric cancer patients and healthy controls were generated using GA, SNN, and QC algorithms that could discriminate both groups with high sensitivity and specificity (Table 3). Analyzing the training set based on the GA algorithm model, gastric cancer patients could be discriminated from healthy controls with 97.1% sensitivity and 98.5% specificity. The sensitivity and specificity of the SNN model was 98.6 and 100%, respectively, and these values were 88.4 and 98.5% for the QC model. The SNN model used five peaks with following m/z values (with mean expression levels in gastric cancer patients and healthy controls): 3,316.09 (marker 1; 11.7 vs. 4.6), 6,629.59 (marker 2; 19.41 vs. 8.81), 3,217.15 (marker 3; 4.84 vs. 2.81), 3,951.98 (marker 4; 5.27 vs. 8.26), and 6,431.45 (marker 5; 7.06 vs. 3.55), among them, four peaks were up-regulated in gastric cancer patients except 3,951.98 (marker 4; 5.27 vs. 8.26).
The P values of which were all <0.000001, and the 3,316.09 peak was the most significant of the 46 discriminated peaks. These five peaks, as well as their receiver operating characteristic (ROC) curves and the area under the curve (AUC) values, are displayed in Fig. 3. Furthermore, the GA model was based on 25 peaks, which contained five SNN peaks, and the QC model was based on 13 peaks, which contained five SNN peaks. Overall, a mean sensitivity value of 94.7% and a mean specificity value of 99% could be achieved in the training set.
Depiction of the five peaks of the supervised neural network (SNN)-based model used to distinguish between serum from gastric cancer patients and controls. The chosen indicator peaks had estimated molecular masses ranging from 3,000 to 6,500 Da. This model was sufficient to discriminate gastric cancer patients (peaks in red, marked with P) from healthy controls (peaks in green, marked with C). Receiver operating characteristic (ROC) curves for each selected peak are shown together with their area under the curve (AUC) values and P values
Analysis of spectra from the completely blinded test set (30 gastric cancer patients and 32 healthy controls) accurately discriminated gastric cancer patients from healthy controls with a mean sensitivity of 79.3% and a mean specificity of 86.5%. Then, the three models mentioned above were used to compare patients with different tumor stages separately from the healthy controls. Early-stage tumors in particular could be discriminated from healthy controls with a mean sensitivity of 71.7%. The mean sensitivity for advanced-stage tumors was 87.1%. Results for different tumor stage, based on the three different algorithm models, are shown in Table 3.
Discussion
Despite the great urgency to identify clinically useful biomarkers for early gastric cancer detection, to date no specific biomarkers have been identified facilitating the early detection and large-scale screening of gastric cancer [25]. An alternative approach to address this problem would be to use proteomic technologies. Recent advances in proteomic technologies, including both MS and protein microarrays, have enabled large-scale screening of proteins in tissues and serum from patients that are applicable for biomarker discovery [26]. Human serum contains a complex array of peptides. Some of these peptides may function as biomarkers, with their presence/absence or relative abundances being correlated with health status and thus useful for prognosis or diagnosis [27–29]. The low-molecular-weight (LMW) serum proteome contains an enormous wealth of biomarker information that has not yet been investigated. Moreover, MS exhibits optimal performance in the LMW range [10]. The SELDI MS approach has predominantly been reported for the profiling of high-molecular weight proteins (>10–20 kDa), whereas MALDI-TOF MS approaches, not using on-target peptide/protein purification, have also been applied for low-mass proteins and peptides (1–15 kDa) [24]. Thus, MALDI-TOF MS has been applied to serum proteomic profiling studies of a variety of different diseases [5].
MB-MALDI-TOF MS (magnetic bead-based fractionation followed by MALDI-TOF MS) could greatly increase the sensitivity of the mass spectra [30]. The profiling kit involving MB-WCX was developed for the enrichment and purification of peptides and proteins from complex biological samples (e.g., human serum) prior to MALDI-TOF MS analysis and is thereby suitable for sample preparation in the context of protein and peptide profiling studies. Moreover, the MB-WCX method is confirmed suitable for the fractionation of low-molecular-mass range peptides (1–10 kDa). In fact, many studies confirmed that the MB-WCX method in combination with MALDI-TOF MS was a highly sensitive and reproducible approach for serum profiling of different cancers [30–32]. In addition, MALDI protein profiling combined with advanced bioinformatics (using software such as the ClinProTools software used in this study) can be employed to identify biomarker patterns for human diseases, as well as being used to improve the reproducibility of mass spectra, offering the potential for clinical proteomic studies. In this study, we used the MB-MALDI-TOF MS technique combined with ClinProTools software (version 2.2) to analyze the serum proteomic profiles of patients with gastric cancer and generated numerous discriminating m/z peaks, which could accurately distinguish cancer patients from healthy individuals. Our findings confirmed that this method is suitable for serum proteomic analysis.
To date, most studies to identify serum biomarkers for the detection of gastric cancer have employed SELDI-TOF MS [15–18], there are only a few reports in which the MALDI-TOF MS approach was used. Ebert et al. [5] generated mass spectra from the sera of 14 patients with gastric cancer and 14 healthy controls by MALDI-TOF MS and identified one peak with a m/z value of 1,466 Da that was significantly up-regulated in the tumor sera compared with the control sera. They subsequently identified a MALDI-MS/MS spectrum of 1,466 Da corresponding to the partial sequence of fibrinopeptide A [5]. However, increased fibrinopeptide A serum levels are not specific to gastric cancer and can also been found in hepatocellular, ovarian, and urothelial cancers [33–35]. In 2008, Chang et al. identified five mass peaks that correlated with stomach cancer, with peaks corresponding to m/z values of 2187, 2387, and 3572 showing down-regulation (with sensitivities of 85, 88, and 79%) and peaks corresponding to m/z values of 2,753 and 4,132 showing up-regulation (with sensitivities of 47 and 68%), by comparing spectra generated from gastric juice samples between 34 gastric cancer patients and 106 healthy controls [26]. In the study by Chang et al. [26], samples were taken from gastric juice rather than sera and some of the identified biomarkers showed low sensitivity.
Using MALDI-TOF–MS, several previous studies also dealt with the identification of peptide peaks and the determination of their expression levels [36–38]. Using 28 gastric cancer patients and 16 healthy controls, Gao et al. reported that seven peaks (viz. 4268.05, 5636.53, 5248.49, 5754.25, 2933.15, 1450.13, and 1349.4 m/z) could be used as candidate biomarkers for gastric cancer [37]; among these peaks, four peaks (viz. 2933.15, 5248.49, 5754.25, 1349.4) were also identified in our study, despite all being non-significant with the P values of 0.34, 0.46, 0.056 and 0.38, respectively. Based on 20 gastric cancer patients and 20 healthy controls, Li et al. asserted that the two most significant peaks be 2,863.71 and 4,965.08 m/z [38]; the 2,863.71 m/z was similar to 2,862.83 m/z (Table 2) in our study. The discrepancy among different studies can be attributed to multiple factors, such as the varied sample sizes, the inherent complexity of gastric cancer, and the difference in disease stages and histotypes of gastric cancer patients used in these studies. To verify the varied results, identification of candidate biomarkers may be the best way.
In our study, we identified 46 potential biomarkers for distinguishing gastric cancer patients from healthy controls. Some discriminating m/z peaks were up-regulated in gastric cancer patients (e.g., 3316.09, 6629.59, 3217.15, 6431.45), while others were down-regulated (e.g., 3,951.98). Three different algorithms all showed high sensitivity and specificity, not only in the training set but also in the test set. A set of five discriminating m/z peaks (m/z: 3316.09, 6629.59, 3217.15, 3951.98, 6431.45) can be used as the diagnostic biomarkers to distinguish gastric cancer patients from healthy controls. On average, values of 94.7% sensitivity and 99% specificity were obtained in the training set, and values of 79.3% sensitivity and 86.5% specificity were obtained in the test set, both of which showed high predictive accuracy. However, since the number of samples in the early-stage tumor group was low, further studies using greater numbers of samples are required to confirm these results.
In conclusion, our findings demonstrate that screening for serum protein patterns using MB-MALDI-TOF MS shows high sensitivity and specificity in identifying patients with gastric cancer. Because of the high-throughput capability of this approach, the identified differentially expressed protein panel may improve the early detection of gastric cancer. However, expanding the data set of gastric cancer patients with different tumor stages, especially early-stage tumors, will aid the identification of clinically useful markers for early gastric cancer detection.
References
Lam KWK, Lo SCL (2008) Discovery of diagnostic serum biomarkers of gastric cancer using proteomics. Proteomics Clin Appl 2:219–228 (PMID: 21136826)
Li JJ, Qi RZ, Ng GKH, Xie D (2009) Proteomics in gastric cancer research: benefits and challenges. Proteomics Clin Appl 3:185–196
Yang L (2006) Incidence and mortality of gastric cancer in China. World J Gastroenterol 12(1):17–20 (PMID: 16440411)
Hohenberger P, Gretschel S (2003) Gastric cancer. Lancet 362:305–315 (PMID: 12892963)
Ebert MPA, Niemeyer D, Deininger SO, Wex T, Knippig C, Hoffmann J, Sauer J, Albrecht W, Knippig P, Rolcken C (2006) Identification and confirmation of increased fibrinopeptide a serum protein levels in gastric cancer sera by magnet bead assisted MALDI-TOF mass spectrometry. J Proteome Res 5:2152–2158 (PMID: 16944926)
Otsuka A, Yohmiya Y, Noha N (2004) Usefulness of magnifying endoscopy in the diagnosis of early gastric cancer. Endoscopy 36(2):165–169 (PMID: 14765314)
Ebert MPA, Lamer S, Meuer J, Malfertheiner P, Reymond M, Buschmann T, Rolcken C, Seibert V (2005) Identification of the thrombin light chain A as the single best mass for differentiation of gastric cancer patients from individuals with dyspepsia by proteome analysis. J Proteome Res 4:586–590 (PMID:15822938)
Oue N, Sentanil K, Noguchi T, Ohara S, Sakamoto N, Hayashi T, Anami K, Motoshital J, Ito M, Tanaka S, Yoshida K, Yasui W (2009) Serum olfactomedin 4 (GW112, hGC-1) in combination with Reg IV is a sensitive biomarker for gastric cancer patients. Int J Cancer 125:2383–2392 (PMID:19670418)
Petricoin EF, Zoon K, Kohn ES, Barrett JC, Liotta LA (2002) Clinical proteomics: translating benchside promise into bedside reality. Nat Rev 1:683–695 (PMID:12209149)
Petricoin E III, Liotta L (2003) Mass spectrometry—based diagnostics: the upcoming revolution in disease detection. Clin Chem 49:533 (PMID:12651801)
Wang CL, Liao PC, Chen CD, Liang Y, Chuang WY, Tsai YH, Chen HC, Chang YS, Yu JS, Wu CC, Yu CJ (2009) Discovery of retinoblastoma—associated binding protein 46 as a novel prognosis marker for distant metastasis in nonsmall cell lung cancer by combined analysis of cancer cell secretome and pleural effusion proteome. J Proteome Res 8:4428–4440 (PMID:19655816)
Petricoin EF, Ardekani AM, Hitt BA, Levine PJ, Fusaro VA, Steinberg SM, Mills GB, Simone C, Fishman DA, Kohn EC, Liotta LA (2002) Use of proteomic patterns in serum to identify ovarian cancer. Lancet 359:572–577 (PMID:11867112)
Pusch W, Flocco MT, Leung SM, Thiele H (2003) Mass spectrometry-based clinical proteomics. Pharmacogenomics 4:463–476 (PMID:12831324)
Carrette O, Demalte I, Scherl A, Yalkinoglu O, Corthals G, Burkhard P, Hochstrasser DF, Sanchez JC (2003) A panel of cerebrospinal fluid potential biomarkers for the diagnosis of Alzheimer’s disease. Proteomics 3:1486–1494 (PMID:12923774)
Liang Y, Fang M, Li J, Liu CB, Rodd JA, Kung HF, Yew DTW (2006) Serum proteomic patterns for gastric lesions as revealed by SELDI mass spectrometry. Exp Mol Pathol 81:176–180 (PMID:16828742)
Ren H, Du N, Liu G, Hu HT, Tian W, Deng ZP, Shi JS (2006) Analysis of variabilities of serum proteomic spectra in patients with gastric cancer before and after operation. World J Gastroenterol 12:2789–2792 (PMID:16718772)
Poon TCW, Sung JJY, Chow SM, Ng EKW, Yu ACW, Chu ESH, Hui AMY (2006) Diagnosis of gastric cancer by serum proteomic fingerprinting. Gastroenterology 130:1858–1864 (PMID:16697748)
Su Y, Shen J, Qian H, Ma H, Ji J, Ma L, Zhang W, Meng L, Li Z, Wu J, Jin G, Zhang J, Shou C (2007) Diagnosis of gastric cancer using decision tree classification of mass spectral data. Cancer Sci 98:37–43 (PMID:17052262)
Villanueva J, Philip J, Entenberg D, Chaparro CA, Tanwar MK, Holland EC, Tempst P (2003) Serum peptide profiling by magnetic particle-assisted, automated sample processing and MALDI-TOF mass spectrometry. Anal Chem 76:1560–1570 (PMID:15018552)
Diamandis EP (2003) Point: proteomic patterns in biological fluids: do they represent the future of cancer diagnostics? Clin Chem 49:1272–1275 (PMID:12881441)
Diamandis EP (2004) Analysis of serum proteomic patterns for early cancer diagnosis: drawing attention to potential problems. J Natl Cancer Inst 96:353–356 (PMID:14996856)
Diamandis EP (2004) Mass spectrometry as a diagnostic and a cancer biomarker discovery tool: opportunities and potential limitations. Mol Cell Proteomics 3:367–378 (PMID:14990683)
Diamandis EP, van der Merwe DE (2005) Plasma protein profiling by mass spectrometry for cancer diagnosis: opportunities and limitations. Clin Cancer Res 11:963–965 (PMID:15709159)
Palmblad M, Tiss A, Cramer R (2009) Mass spectrometry in clinical proteomics—from the present to the future. Proteomics Clin Appl 3:6–17 (PMID:21136932)
Ramachandran N, Srivastava S, LaBaer J (2008) Applications of protein microarrays for biomarker discovery. Proteomics Clin Appl 2:1444–1459 (PMID:21136793)
Chang WC, Hsu PI, Chen YY, Hsiao M, Lu PJ, Chen CH (2008) Observation of peptide differences between cancer and control in gastric juice. Proteomics Clin Appl 2:55–62 (PMID:21136779)
Diamandis EP (2006) Peptidomics for cancer diagnosis: present and future. J Proteome Res 5:2079–2082 (PMID:16944917)
Liotta LA, Petricoin EF (2006) Serum peptidome for cancer detection: spinning biological trash into diagnostic gold. J Clin Invest 116:26–30 (PMID:16395400)
Villanueva J, Shaffer DR, Philip J, Chaparro CA, Erdjument-Bromage H, Olshen AB, Fleisher M, Lilja H, Brogi E, Boyd J, Sanchez-Carbayo M, Holland EC, Cordon-Cardo C, Scher HI, Tempst P (2006) Differential exoprotease activities confer tumor-specific serum peptidome patterns. J Clin Invest 116:271–284 (PMID:16395409)
Fiedler GM, Baumann S, Leichtle A, Oltmann A, Kase J, Thiery J, Ceglarek U (2007) Standardized peptidome profiling of human urine by magnetic bead separation and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Clin Chem 53:421–428 (PMID:17272489)
Schaub NP, Jones KJ, Nyalwidhe JO, Cazares LH, Karbassi ID, Semmes OJ, Feliberti EC, Perry RR, Drake RR (2009) Serum proteomic biomarker discovery reflective of stage and obesity in breast cancer patients. J Am Coll Surg 208:970–978 (PMID:19476873)
Schwamborn K, Krieg RC, Grosse J, Reulen N, Weiskirchen R, Knuechel R, Jakse G, Henkel C (2009) Serum proteomic profiling in patients with bladder cancer. Eur Urol 56:989–999 (PMID:19282097)
Theodorescu D, Wittke S, Ross MM, Walden M, Conaway M, Just I, Mischak H, Frierson HF (2006) Discovery and validation of new protein biomarkers for urothelial cancer: a prospective analysis. Lancet Oncol 7:230–240 (PMID:16510332)
Bergen HR, Vasmatzis G, Cliby WA, Johnson KL, Oberg AL, Muddiman DC (2003) Discovery of ovarian cancer biomarkers in serum using NanoLC electrospray ionization TOF and FT-ICR mass spectrometry. Dis Markers 19:239–249 (PMID:15266667)
Orvisky E, Drake SK, Martin BM, Abdel-Hamid M, Ressom H, Varghese RS, An Y, Saha D, Hortin GL, Loffredo CA, Goldman R (2006) Enrichment of low molecular weight fraction of serum for mass spectrometric analysis of peptides associated with hepatocellular carcinoma. Proteomics 6:2895–2902 (PMID: 16586431)
Liu WT, Gao X, Cai Q, Li JF, Zhu ZL, Li C, Yao XX, Yang QM, Xiang M, Yan M, Zhu ZG (2010) Identification of novel serum biomarkers for gastric cancer by magnetic bead. Front Biosci 2:961–971 (PMID: 20515767)
Gao X, Liu WT, Yang QM, Liu BY, Cai Li, Li JF, Zhu ZL, Xiang M, Yan M, Zhu ZG (2010) Proteomic profiling of gastric cancer in serum using magnetic bead based sample fractionation and MALDI-TOF-MS. Chin J Clin Oncol 16:933–936
Li T, Shen HM (2009) Screen tumor specific biomarkers of gastric cancer in serum using magnetic bead based sample fractionation and MALDI-TOFMS. Modern Oncol 17:1513–1515
Acknowledgments
This work was supported by the National Science Foundation for Postdoctoral Scientists of China (Grant number: 20090461301); the Young Scientist Foundation from the Medical School in Xi’an Jiaotong University (Grant number: YQN0809); the Scientific Research Support Program for New Teachers (Grant number: 0116-081410-05); the Postdoctoral Science Foundation of Xi’an Jiaotong University (Grant number: 2116-04212217); Guang Hua Medical Innovation Research Foundation (Grant number: 0203407); and the International Sci-tech Cooperation Program (Grant number: 2009DFA31420).
Conflicts of interest
The authors confirm that they have no financial or personal relationships that cause a conflict of interest regarding the work in the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, J., Song, YC., Dang, CX. et al. Serum peptidome profiling in patients with gastric cancer. Clin Exp Med 12, 79–87 (2012). https://doi.org/10.1007/s10238-011-0149-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-011-0149-2