Abstract
Mycoplasma pneumoniae is one of the main pathogens causing community-acquired respiratory tract infections in children and adults. Macrolide (ML) antibiotics are recognized generally as first-choice agents for M. pneumoniae infections, and these antibiotics were thought to have excellent effectiveness against M. pneumoniae for many years. In 2000, however, M. pneumoniae showing resistance to macrolides was isolated from clinical samples obtained from Japanese pediatric patients with community-acquired pneumonia (CAP). Since then, prevalence of ML-resistant M. pneumoniae isolates in pediatric patients has increased rapidly. In 2007, ML-resistant M. pneumoniae isolates were obtained from Japanese adults with CAP; numbers of such isolates also have gradually increased in Japan. Recently, similar antimicrobial resistance in M. pneumoniae has begun to emerge worldwide. In this review, we focus on changes of ML-resistant M. pneumoniae from year to year and consider resistance mechanisms as well as clinical features of patients with resistant M. pneumoniae infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sixteen Mycoplasma species have been isolated from human respiratory and urogenital specimens [1]. One of these species, M. pneumoniae, is a main pathogen in respiratory tract infections (RTI) acquired in the community. In school-aged children and young adults with community-acquired pneumonia (CAP), M. pneumoniae accounts for as many as 10–30% of cases [2–8]. In distinction, M. pneumoniae pneumonia in younger children and in the elderly is infrequent but not rare [9–15].
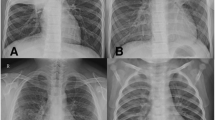
M. pneumoniae pneumonia is diagnosed based on characteristic chest radiographic abnormalities, patient symptoms, and clinical laboratory data. Diagnosis ultimately is confirmed using serologic tests performed upon paired sera obtained during both acute and convalescent phases. Conventional culture using pleuropneumonia-like organism (PPLO) broth, which requires more than 2 weeks, has not been carried out routinely. As a consequence, antibiotic choice usually is empirical.
Development of molecular methods such as polymerase chain reaction (PCR) assays to be used in combination with conventional diagnostic tests using serology and culture have contributed to improved diagnosis and characterization of M. pneumoniae infection in pediatric and adult patients [16–28]. Macrolides (ML) such as clarithromycin (CAM) and azithromycin (AZM) generally are recognized as first-choice agents against M. pneumoniae [29–33]. However, in parallel with increased use of oral ML with a 14-membered ring (14-ML) and AZM for RTI, M. pneumoniae showing resistance to ML has been isolated increasingly in clinical samples from Japanese pediatric patients with CAP [31, 34, 35]. Prevalence of ML-resistant (MLr) M. pneumoniae in pediatric patients has increased rapidly [35, 36] and MLr M. pneumoniae is beginning to be isolated worldwide [37–41].
In this review, we summarize the emergence and increase of MLr M. pneumoniae isolates, resistance mechanisms, and clinical features of MLr M. pneumoniae infection.
Diagnosis of infection
M. pneumoniae infection tends to show cyclic epidemics every 3–5 years; these outbreaks are particularly likely to occur in summer or early fall [42–46]. Although the incidence of M. pneumoniae pneumonia is highest in children 5–9 years old, such pneumonia also occurs frequently in younger children and in the elderly. Prevalence of M. pneumoniae infection also may vary according to population and diagnostic methods used.
Isolation of M. pneumoniae by culture from clinical samples such as throat or nasopharyngeal swabs was considered standard for diagnosis several years ago. However, M. pneumoniae is difficult to grow in liquid culture; cultures using PPLO broth require at least 2 weeks for completion. Compared with serologic tests or molecular techniques, including PCR, sensitivity of cultures may be <60% or 70%, even in laboratories with expertise [6, 18, 20]. Therefore, culture methods are only used rarely for routine diagnosing M. pneumoniae infection.
Serologic methods are used most frequently to diagnose M. pneumoniae infection. Frequently used, widely available serologic tests for M. pneumoniae include complement fixation (CF), enzyme immunoassay (EIA), and particle agglutination (PA) assays. However, these ordinarily require paired sera obtained during acute and convalescent phases in order to demonstrate rises in antibody titers; fourfold increase is thought to be significant. The second sample should be obtained 7–14 days after the first, provided that M. pneumoniae infection still is suspected.
Although one alternative, the ImmunoCard based on an immunoglobulin M (IgM) assay, is rapid and easy to perform, reliable diagnosis of M. pneumoniae infection still cannot be made on the basis of single acute-phase sera; confirmation using convalescent-phase samples still is necessary, as otherwise, both false-positive and false-negative results are frequent [24, 47, 48]. Additionally, sensitivity of IgM assays tends to be low, particularly for adult patients who are known to have weak IgM responses during primary infection or reinfection [49, 50]. Neither culture nor serologic testing can provide timely information for guiding choice of chemotherapeutic agents to use for early intervention. Serologic methods generally have only low sensitivity during the acute phase of disease.
Development of molecular methods (e.g. PCR) has lessened the importance of culture as a way to detect M. pneumoniae directly. Two specific targets in the P1 adhesion gene and the 16S ribosomal RNA (rRNA) gene are the main ones used in PCR assays to detect M. pneumoniae DNA. Various PCR methods have been developed in several laboratories. M. pneumoniae detection by conventional PCR or real-time PCR has been examined as an approach to early diagnosis. Many studies have described application of multiplex PCR, hybridization assays, nucleic acid sequence-based amplification (NASBA), real-time PCR using a molecular beacon probe or TaqMan probe, or cycling probe, to the identification of M. pneumoniae DNA [16–28]. Among these methods, real-time PCR has both high sensitivity and high specificity and can detect pathogen DNA even when damaged by empirical administration of antibiotics. Sensitivity (60–100%) and specificity (96.7–100%) of real-time PCR are both higher than those of serologic assays for M. pneumoniae [18, 19, 22, 24–26]. Almost all PCR-positive cases (>90%) also were confirmed serologically [22, 24]. In addition, real-time PCR using a fluorescent probe allows continuous monitoring of in vitro DNA amplification, eliminating nonspecific amplification product using fluorescence; no gel electrophoresis is needed. This makes the method suitable for clinical laboratory settings. A molecular approach including real-time PCR is useful for confirming M. pneumoniae and providing rapid diagnosis of CAP, thus permitting rational choice of effective antimicrobial agents.
Mechanisms of ML resistance in M. pneumoniae
Intrinsically, absence of cell walls in M. pneumoniae confers resistance to β-lactams, and indeed to all antibiotics that inhibit synthesis of cell walls. In M. pneumoniae infections, 14-ML and 15-membered ring ML (15-ML) usually are considered the first-line agents. M. pneumoniae ordinarily are susceptible to all ML, including ketolides.
Main mechanisms of microorganism resistance to ML identified in previous studies include modification of the target sites in 23S rRNA by methylation or mutation, as well as on efflux pump [51–53]. ML act on the 50S ribosomal subunit to inhibit protein synthesis. The site of peptide bond formation on the large 50S ribosomal subunit is associated with the central loop in domain V of 23S rRNA. Nucleotides present in the central loop are necessary constituents of the ML binding site, which implies that all ML bind to the same site (Fig. 1a). A loop of hairpin 35 in domain II also may be part of binding site of ketolides; mutations involving ribosomal proteins L4 and L22 may contribute as well [51–54].
Binding site of macrolides on the 23S ribosomal RNA. a Secondary structure of the large subunit of ribosomal RNA (Escherichia coli; http://www.rna.ccbb.utexas.edu/). b Central loop in domain V (Mycoplasma pneumoniae numbering) and the loop of hairpin 35 in domain II (E. coli numbering) of 23S rRNA as modified according to [34, 51]. Circles indicate positions of mutations associated with macrolide resistance in clinical isolates. The square shows the position of a substitution associated with 16-membered-ring macrolide resistance in a mutant selected in vitro
ML inhibit protein synthesis mainly by binding to domain V of 23S rRNA at nucleotide positions 2063 and 2064 (M. pneumoniae numbering); these positions appear essential for binding (Fig. 1b) [55]. Mutations at A2063 or A2064 confer the highest resistance to these antimicrobials. A lower level of antibiotic resistance is caused by mutations at positions A2067 and C2617, which are near A2063 and A2064 in the secondary structure, although slightly present outside the central point of ML interaction. The A2067G mutation was reported to cause highest levels of resistance only for 16-membered-ring ML (16-ML), as it interferes with formation of the covalent bond specific to 16-ML [56].
Ketolides are a new class of antimicrobial agent characterized by the substitution at position 3 from l-cladinose to keto group. These antibiotics are active against Streptococcus pneumoniae strains resistant to ML because of interaction at both A2058/2059 (Escherichia coli numbering) in domain V and A752 in domain II [51, 54, 57]. Substitutions in ribosomal proteins L4 and L22 have been found to confer resistance in S. pneumoniae [58, 59], but mutations in domain II of 23S rRNA and in ribosomal proteins L4 and L22 are not involved in ML resistance of clinical M. pneumoniae isolates [60]. Furthermore, neither plasmids nor erm genes mediating ribosomal modification have been described with respect to M. pneumoniae.
Fluoroquinolone (FQ)-resistant M. pneumoniae mutants may be obtained in vitro by exposure to subinhibitory concentrations of FQ [61]. These mutants possess mutations in gyrA and gyrB genes, which express topoisomerase II, and in parC and parE genes, which express topoisomerase IV; both of these enzymes are essential for bacterial DNA replication. FQ often are chosen as empiric treatment of respiratory tract infection in adults, as they are active against the main pathogens, including S. pneumoniae, Haemophilus influenzae, and M. pneumoniae. In contrast to in vitro findings, no FQ-resistant strains have been observed in clinical settings.
Tetracyclines inhibit protein synthesis by preventing association of aminoacyl-tRNA to the acceptor (A) site of the ribosome [62, 63]. These antibiotics bind to the ribosomal 30S subunit involving 16S rRNA and several ribosomal proteins. Only the tetM determinant associated with conjugative transposon Tn916, which confers resistance to the tetracyclines in M. hominis, has been documented so far [64]. These agents are administered for treatment of M. pneumoniae infection in adults and in pediatric patients >8 years, but development of resistance to tetracyclines has not yet been reported.
Association of ML resistance and mutation site in 23S rRNA from MLr M. pneumoniae
Specific sites of the 23S rRNA mutation contribute to ML resistance in M. pneumoniae. In actuality, based on our results, clinical isolates with mutations at positions A2063 and A2064 show high resistance to 14-ML and 15-ML, with minimal inhibitory concentration (MIC) ≥ 32 μg/ml (Table 1). Especially, the A2063G mutation conveys the highest level of resistance to 14-ML, 15-ML, and telithromycin (TEL) but intermediate resistance to josamycin and rokitamycin belong to 16-ML. The A2063C and A2064G mutations result in a high level of resistance to all ML [36, 65]. The ketolide TEL is affected by both mutations, although A2063G was associated with MICs higher than those associated with A2064G.
MICs for minocycline (MINO) and FQ in MLr strains were equivalent to those in susceptible strains. No strains having resistance to MINO and FQ have been observed among clinical isolates.
Several strains with the C2617 to G or A mutations show slight increases in MIC of ML compared with those with mutations at positions 2063 and 2064. A2063G in domain V of 23S rRNA is the most frequent mutation association with ML resistance, followed by A2064G; others (A2063C, C2617G, and C2617A) are rare.
Ribosomal protein L4 and L22 mutants with a few amino acids deleted or inserted have been described, but roles of these mutations in resistance to ML are uncertain [60]. No mutation has been detected in domain II of 23S rRNA.
Emergence and increase of MLr strains
MLr M. pneumoniae isolates possessing a nucleotide mutation in 23S rRNA first were isolated from pediatric patients with CAP, as reported by Okazaki and colleagues in 2001 [34].
Together with growing use of ML in Japan, resistant strains increased rapidly year by year (Fig. 2) [36]. Along with greater overall prevalence of M. pneumoniae infection in 2006, MLr strains increasingly were isolated and identified in diverse regions across Japan.
In Japanese adult CAP patients, MLr strains first were isolated in 2007; since then, occasional isolations have continued [66]. An increase in MLr strains among adult patients, las in pediatric patients, may occur in the future. However, FQ are recommended for CAP treatment in adults, and no FQ-resistant strains have been observed among isolates from adult patients with CAP.
Emergence of MLr isolates having either an A2063G or an A2064G mutation has been reported not only in Japan but also in other countries, including France, the USA, Denmark, and China [37–41]. A notably high isolation rate (92%) of MLr strains was reported in China [38, 40].
The emergence and increase of MLr M. pneumoniae has attracted the attention of microbiologists worldwide. Several recent studies demonstrated direct detection of MLr isolates using real-time PCR [37, 41, 67].
DNA profiles of MLr- and ML-susceptible (MLs) strains obtained by pulsed-field gel electrophoresis (PFGE) have suggested classification into two groups [36, 68]. Significant differences in these DNA patterns were not recognized between strains, geographic locations, or years.
PCR-based restriction fragment length polymorphism (RFLP) types of the P1 adhesin gene also were used to divide strains into two types. MLr isolates were present in both types [38, 39, 69, 70], showing no predominance.
Clinical aspects of MLr M. pneumoniae infection
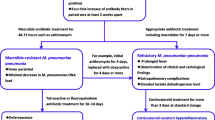
The clinical course of patients with MLr M. pneumoniae infection appears to be prolonged [36, 66, 71, 72]. Patients infected with MLr strains mostly had been treated with ML previously [36, 40]. Among CAP patients infected with MLr, treatment frequently was changed from ML to MINO or levofloxacin (LVFX) because of either persistent symptoms (i.e., fever and cough) or unresolved or worsening chest radiographic abnormalities. In our observations, treatment was changed from ML to MINO or LVFX in 8.2% of patients with MLs strains and in 39.7% of patients with MLr strains, representing a significant difference between the two patient groups (P < 0.01). ML also was more frequently discontinued in favor of MINO among MLr patients than MLs patients in a report by Suzuki et al. [71]. Neither LVFX nor MINO are recommended for pediatric patients. However, when ML are ineffective against M. pneumoniae infection, pediatricians have little choice of antimicrobials except for MINO. In adult inpatients infected with MLr strains, antibiotic agents were changed from CAM to intravenous pazufloxacin when symptoms did not improve [66]. Orally administered respiratory FQ such as moxifloxacin, sitafloxacin, and garenoxacin may be selected for adult outpatients with MLr strains.
Among clinical features of MLr M. pneumoniae infection, fever duration has been significantly longer than for MLs-strain infection. In our data, fever persisted after ML initiation for 1.6 ± 0.8 days in MLs M. pneumoniae infection and for 4.1 ± 2.3 days in MLr-strain infection, showing a significant difference (P < 0.01). In other studies, febrile days during ML administration also were significantly greater in MLr-infected patients than in MLs-infected patients (3.5–4.0 days vs. 1.0–1.5 days) [71, 72]. Furthermore, the mean duration of persistent cough after ML administration was 7.0 days in MLs-infected patients and 11.4 days in MLr-infected patients [72]. Finally, the efficacy rate of ML therapy was 91.5% and 22.7% in MLs- and MLr-strain infections, respectively [72].
Fever may have resolved spontaneously in some patients with continuing ML treatment for MLr infections; M. pneumoniae infection is associated with occasional spontaneous symptomatic recovery. According to previous reports, fever duration was 1 week in symptomatically treated M. pneumoniae infection [73, 74]. In another study, mean fever duration in hospitalized patients receiving AZM or EM was 2.1 days, whereas fever persisted for about 1 week in patients not receiving ML [75]. Similarly, mean cough duration was reported as 8.5 days [76]. Although treatment failure or serious illness has not yet been attributed to MLr-strain infection, the clinical course of patients infected with MLr strains may be prolonged.
A brief consideration of the host response is helpful here. M. pneumoniae resides on the surface of ciliated human respiratory epithelial cell. Several membrane proteins in M. pneumoniae have high affinity for various surface receptors on host cells [77]. Through interaction with recognition receptors, including toll-like receptors 2 and 6, mycoplasma membrane lipoproteins are able to induce host immune responses [77, 78]. Adherence of M. pneumoniae to host cells in the respiratory tract is mediated by the P1 adhesin protein and accessory proteins [79–82]. Following adherence, macrophages become activated and release cytokines, and a mononuclear cell inflammatory response is established. Human lung epithelial cells infected with M. pneumoniae induce expression of interleukin (IL)-8, which is both a potent chemoattractant and an activator of neutrophils, monocytes, and T lymphocytes [83–85]. In particular, significant production of IL-8 and IL-18, a cytokine that induces interferon gamma (IFN-γ) production and promotes a type 1 cytokine response, is associated with the severity of M. pneumoniae pneumonia in children [86–88] and adults [89].
As previously described, the inflammatory response to M. pneumoniae infection is considered to play a crucial role in the pathogenesis of the ensuing clinical disease. ML possess antimicrobial activity against M. pneumoniae, and also have anti-inflammatory effects. For example, CAM is able to suppress IL-8 induction, even in MLr M. pneumoniae infection [78, 86, 90]. Thus, clinical efficacy of ML in treating M. pneumoniae infection may reflect not only direct antimicrobial activity but also anti-inflammatory effects of inhibition of production of cytokines, including IL-8.
Symptoms and severity of illness due to M. pneumoniae were similar in younger and older patients, and the mortality rate was low, even in the elderly [13, 91, 92]. This disease ordinarily is mild, and reinfection may occur in adults who already experienced M. pneumoniae infection, as protective immunity usually is not established following initial infection [93]. However, symptoms appear to be prolonged in certain clinical situations regardless of antimicrobial resistance [15, 94]. Takahashi and colleagues reported M. pneumoniae DNA in sputum after 2 weeks of MINO therapy. Elderly persons with M. pneumoniae pneumonia may have a longer clinical course than younger persons. Further observations from various patients infected with M. pneumoniae are necessary.
In the future, alternative treatment strategies, such as promptly initiated steroid therapy, might be considered for patients with MLr strains as a symptomatic measure [95–97].
Conclusions
In pediatric patients with CAP, MLr M. pneumoniae possessing a 23S-rRNA mutation was first isolated in 2000; numbers of these isolates have increased rapidly year by year in Japan. In other countries, MLr M. pneumoniae also is emerging. Symptoms often are relatively prolonged in patients with MLr M. pneumoniae infection. MLr M. pneumoniae should be considered an emerging issue for not only pediatric patients but also adult patients with CAP.
No FQ-resistant M. pneumoniae strain has been observed thus far among specimens from adult patients with CAP. However, emergence of infections with FQ-resistant M. pneumoniae might occur, considering the increasing FQ prescription rate among adult patients. Further studies are needed to clarify the prevalence of MLr M. pneumoniae in pediatric and adult patients with RTI and to establish clinical guidelines regarding the most appropriate antimicrobial agents to use against these strains. Interventions to prevent emergence and increases of novel antibiotic-resistant strains are needed, including rapid identification of pathogens by methods such as real-time PCR, as well as antimicrobial treatments capable of eliminating the organisms.
References
Waites KB, Rikihisa Y, Taylor-Robinson D. Mycoplasma, Ureaplasma. In: Murray PR, Baron EJ, Jorgensen JH, Pfaller MA, Yolken RH, eds. Manual of clinical microbiology. 8th edn. Washington: American Society for Microbiology; 2003. p. 972–90.
Hammerschlag MR. Mycoplasma pneumoniae infections. Curr Opin Infect Dis. 2001;14:181–6.
Heiskanen-Kosma T, Korppi M, Jokinen C, Kurki S, Heiskanen L, Juvonen H, et al. Etiology of childhood pneumonia: serologic results of a prospective, population-based study. Pediatr Infect Dis J. 1998;17:986–91.
Kashyap B, Kumar S, Sethi GR, Das BC, Saigal SR. Comparison of PCR, culture & serological tests for the diagnosis of Mycoplasma pneumoniae in community-acquired lower respiratory tract infections in children. Indian J Med Res. 2008;128:134–9.
McIntosh K. Community-acquired pneumonia in children. N Engl J Med. 2002;346:429–37.
Morozumi M, Hasegawa K, Chiba N, Iwata S, Kawamura N, Kuroki H, et al. Application of PCR for Mycoplasma pneumoniae detection in children with community-acquired pneumonia. J Infect Chemother. 2004;10:274–9.
Nakayama E, Hasegawa K, Morozumi M, Kobayashi R, Chiba N, Iitsuka T, et al. Rapid optimization of antimicrobial chemotherapy given to pediatric patients with community-acquired pneumonia using PCR techniques with serology and standard culture. J Infect Chemother. 2007;13:305–13.
Oguz F, Unuvar E, Aydin D, Yilmaz K, Sidal M. Frequency of Mycoplasma pneumoniae among atypical pneumonia of childhood. Turk J Pediatr. 2002;44:283–8.
Defilippi A, Silvestri M, Tacchella A, Giacchino R, Melioli G, Di Marco E, et al. Epidemiology and clinical features of Mycoplasma pneumoniae infection in children. Respir Med. 2008;102:1762–8.
Howard LS, Sillis M, Pasteur MC, Kamath AV, Harrison BD. Microbiological profile of community-acquired pneumonia in adults over the last 20 years. J Infect. 2005;50:107–13.
Ishida T, Hashimoto T, Arita M, Ito I, Osawa M. Etiology of community-acquired pneumonia in hospitalized patients: a 3-year prospective study in Japan. Chest. 1998;114:1588–93.
Martínez MA, Ruiz M, Zunino E, Luchsinger V, Avendaño LF. Detection of Mycoplasma pneumoniae in adult community-acquired pneumonia by PCR and serology. J Med Microbiol. 2008;57:1491–5.
Miyashita N, Ouchi K, Kawasaki K, Oda K, Kawai Y, Shimizu H, et al. Mycoplasma pneumoniae pneumonia in the elderly. Med Sci Monit. 2008;14:387–91.
Porath A, Schlaeffer F, Lieberman D. The epidemiology of community-acquired pneumonia among hospitalized adults. J Infect. 1997;34:41–8.
Takahashi T, Morozumi M, Chiba N, Asami R, Okada T, Murayama SY, et al. Prolonged Mycoplasma pneumoniae infection in an elderly patient with community-acquired pneumonia. J Infect Chemother. 2009;15:243–7.
Abele-Horn M, Busch U, Nitschko H, Jacobs E, Bax R, Pfaff F, et al. Molecular approaches to diagnosis of pulmonary diseases due to Mycoplasma pneumoniae. J Clin Microbiol. 1998;36:548–51.
Dorjgo-Zetsma JW, Verkooyen RP, Van Helden HP, Van der Nat H, Van den Bosch JM. Molecular detection of Mycoplasma pneumoniae in adults with community-acquired pneumonia requiring Hospitalization. J Clin Microbiol. 2001;39:1184–6.
Dorigo-Zetsma JW, Zaat SA, Wertheim-van Dillen PM, Spanjaard L, Rijntjes J, van Waveren G, et al. Comparison of PCR, culture, and serological tests for diagnosis of Mycoplasma pneumoniae respiratory tract infection in children. J Clin Microbiol. 1999;37:14–7.
Hardegger D, Nadal D, Bossart W, Altwegg M, Dutly F. Rapid detection of Mycoplasma pneumoniae in clinical samples by real-time PCR. J Microbiol Methods. 2000;41:45–51.
Ieven M, Ursi D, Van Bever H, Quint W, Niesters HG, Goossens H. Detection of Mycoplasma pneumoniae by two polymerase chain reactions and role of M. pneumoniae in acute respiratory tract infections in pediatric patients. J Infect Dis. 1996;173:1445–52.
Khanna M, Fan J, Pehler-Harrington K, Waters C, Douglass P, Stallock J, et al. The pneumoplex assays, a multiplex PCR-enzyme hybridization assay that allows simultaneous detection of five organisms, Mycoplasma pneumoniae, Chlamydia (Chlamydophila) pneumoniae, Legionella pneumophila, Legionella micdadei, and Bordetella pertussis, and its real-time counterpart. J Clin Microbiol. 2005;43:565–71.
Morozumi M, Nakayama E, Iwata S, Aoki Y, Hasegawa K, Kobayashi R, et al. Simultaneous detection of pathogens in clinical samples from patients with community-acquired pneumonia by real-time PCR with pathogen-specific molecular beacon probes. J Clin Microbiol. 2006;44:1440–6.
Nadala D, Bossart W, Zucol F, Steiner F, Berger C, Lips U, et al. Community-acquired pneumonia in children due to Mycoplasma pneumoniae: diagnostic performance of a seminested 16S rDNA-PCR. Diagn Microbiol Infect Dis. 2001;39:15–9.
Otomo S, Yamamura J, Hayashi E, Nakamura T, Kakinuma H, Nakamoto Y, et al. Analysis of children with Chlamydophila (Chlamydia) pneumoniae and Mycoplasma pneumoniae respiratoryinfections by real-time PCR assay and serological tests. APMIS. 2008;116:477–83.
Pitcher D, Chalker VJ, Sheppard C, George RC, Harrison TG. Real-time detection of Mycoplasma pneumoniae in respiratory samples with an internal processing control. J Med Microbiol. 2006;55:149–55.
Templeton KE, Scheltinga SA, Graffelman AW, van Schie JM, Crielaard JW, Sillekens P, et al. Comparison and evaluation of real-time PCR, real-time nucleic acid sequence-based amplification, conventional PCR, and serology for diagnosis of Mycoplasma pneumoniae. J Clin Microbiol. 2003;41:4366–71.
Tjhie JH, van Kuppeveld FJ, Roosendaal R, Melchers WJ, Gordijn R, MacLaren DM, et al. Direct PCR enables detection of Mycoplasma pneumoniae in patients with respiratory tract infections. J Clin Microbiol. 1994;32:11–6.
Welti M, Jaton K, Altwegg M, Sahli R, Wenger A, Bille J. Development of a multiplex real-time quantitative PCR assay to detect Chlamydia pneumoniae, Legionella pneumophila and Mycoplasma pneumoniae in respiratory tract secretions. Diagn Microbiol Infect Dis. 2003;45:85–95.
Felmingham D, Robbins MJ, Sanghrajka M, Leakey A, Ridgway GL. The in vitro activity of some 14-, 15- and 16-membered macrolides against Staphylococcus spp., Legionella spp., Mycoplasma spp. and Ureaplasma urealyticum. Drugs Exp Clin Res. 1991;17:91–9.
Ishida K, Kaku M, Irifune K, Mizukane R, Takemura H, Yoshida R, et al. In vitro and in vivo activities of macrolides against Mycoplasma pneumoniae. Antimicrob Agents Chemother. 1994;38:790–8.
Morozumi M, Hasegawa K, Kobayashi R, Inoue N, Iwata S, Kuroki H, et al. Emergence of macrolide-resistant Mycoplasma pneumoniae with a 23S rRNA gene mutation. Antimicrob Agents Chemother. 2005;49:2302–6.
Rennie KA, Prasad ES, Wenman WM. In vitro activity of dirithromycin, a new macrolide antibiotic, against Mycoplasma species. Diagn Microbiol Infect Dis. 1994;20:57–9.
Waites KB, Cassell GH, Canupp KC, Fernandes PB. In vitro susceptibilities of mycoplasmas and ureaplasmas to new macrolides and aryl-fluoroquinolones. Antimicrob Agents Chemother. 1988;32:1500–2.
Okazaki N, Narita M, Yamada S, Izumikawa K, Umetsu M, Kenri T, et al. Characteristics of macrolide-resistant Mycoplasma pneumoniae strains isolated from patients and induced with erythromycin in vitro. Microbiol Immunol. 2001;45:617–20.
Okazaki N, Ohya H, Sasaki T. Mycoplasma pneumoniae isolated from patients with respiratory infection in Kanagawa Prefecture in 1976–2006: emergence of macrolide-resistant strains. Jpn J Infect Dis. 2007;60:325–6.
Morozumi M, Iwata S, Hasegawa K, Chiba N, Takayanagi R, Matsubara K, et al. Increased macrolide resistance of Mycoplasma pneumoniae in pediatric patients with community-acquired pneumonia. Antimicrob Agents Chemother. 2008;52:348–50.
Li X, Atkinson TP, Hagood J, Makris C, Duffy LB, Waites KB. Emerging macrolide resistance in Mycoplasma pneumoniae in children: detection and characterization of resistant isolates. Pediatr Infect Dis J. 2009;28:693–6.
Liu Y, Ye X, Zhang H, Xu X, Li W, Zhu D, et al. Antimicrobial susceptibility of Mycoplasma pneumoniae isolates and molecular analysis of macrolide-resistant strains from Shanghai, China. Antimicrob Agents Chemother. 2009;53:2160–2.
Pereyre S, Charron A, Renaudin H, Bébéar C, Bébéar CM. First report of macrolide-resistant strains and description of a novel nucleotide sequence variation in the P1 adhesin gene in Mycoplasma pneumoniae clinical strains isolated in France over 12 years. J Clin Microbiol. 2007;45:3534–9.
Xin D, Mi Z, Han X, Qin L, Li J, Wei T, et al. Molecular mechanisms of macrolide resistance in clinical isolates of Mycoplasma pneumoniae from China. Antimicrob Agents Chemother. 2009;53:2158–9.
Wolff BJ, Thacker WL, Schwartz SB, Winchell JM. Detection of macrolide resistance in Mycoplasma pneumoniae by real-time PCR and high-resolution melt analysis. Antimicrob Agents Chemother. 2008;52:3542–9.
Alexander ER, Foy HM, Kenny GE, Kronmal RA, McMahan R, Clarke ER, et al. Pneumonia due to Mycoplasma pneumoniae. Its incidence in the membership of a co-operative medical group. N Engl J Med. 1966;275:131–6.
Eun BW, Kim NH, Choi EH, Lee HJ. Mycoplasma pneumoniae in Korean children: the epidemiology of pneumonia over an 18-year period. J Infect. 2008;56:326–31.
Feikin DR, Moroney JF, Talkington DF, Thacker WL, Code JE, Schwartz LA, et al. An outbreak of acute respiratory disease caused by Mycoplasma pneumoniae and adenovirus at a federal service training academy: new implications from an old scenario. Clin Infect Dis. 1999;29:1545–50.
Hauksdóttir GS, Jónsson T, Sigurdardóttir V, Löve A. Seroepidemiology of Mycoplasma pneumoniae infections in Iceland 1987–96. Scand J Infect Dis. 1998;30:177–80.
Lind K, Benzon MW, Jensen JS, Clyde WA Jr. A seroepidemiological study of Mycoplasma pneumoniae infections in Denmark over the 50-year period 1946–1995. Eur J Epidemiol. 1997;13:581–6.
Beersma MF, Dirven K, van Dam AP, Templeton KE, Claas EC, Goossens H. Evaluation of 12 commercial tests and the complement fixation test for Mycoplasma pneumoniae-specific immunoglobulin G (IgG) and IgM antibodies, with PCR used as the “gold standard”. J Clin Microbiol. 2005;43:2277–85.
Narita M, Togashi T. Evaluation of a rapid IgM antibody detection kit for diagnosis of Mycoplasma pneumoniae infection during childhood. Kansenshogaku Zasshi. 2003;77:310–5.
Daxboeck F, Kircher K, Krause R, Heinzl H, Wenisch C, Stanek G. Effect of age on antibody titer to Mycoplasma pneumoniae. Scand J Infect Dis. 2002;34:577–9.
Sillis M. The limitations of IgM assays in the serological diagnosis of Mycoplasma pneumoniae infections. J Med Microbiol. 1990;33:253–8.
Hansen LH, Mauvais P, Douthwaite S. The macrolide-ketolide antibiotic binding site is formed by structures in domain II and V of 23S ribosomal RNA. Mol Microbiol. 1999;31:623–31.
Poehlsgaard J, Douthwaite S. Macrolide antibiotic interaction and resistance on the bacterial ribosome. Curr Opin Investig Drugs. 2003;4:140–8.
Vester B, Douthwaite S. Macrolide resistance conferred by base substitutions in 23S rRNA. Antimicrob Agents Chemother. 2001;45:1–12.
Bryskier A, Agouridas C, Chantot JF. Ketolides: new semisynthetic 14-membered-ring macrolides. In: Zinner SH, Young LS, Acar JF, Neu HC, editors. Expanding indications for the new macrolides, azalides, and streptogramins. New York: Marcel Dekker Inc; 1997. p. 39–50.
Lucier TS, Heitzman K, Liu SK, Hu PC. Trasition mutations in the 23S rRNA of erythromycin-resistant isolates of Mycoplasma pneumoniae. Antimicrob Agents Chemother. 1995;39:2770–3.
Pereyre S, Guyot C, Renaudin H, Charron A, Bébéar C, Bébéar CM. In vitro selection and characterization of resistance to macrolides and related antibiotics in Mycoplasma pneumoniae. Antimicrob Agents Chemother. 2004;48:460–5.
Ubukata K, Iwata S, Sunakawa K. In vitro activities of new ketolide, telithromycin, and eight other macrolide antibiotics against Strptococcus pneumoniae having mefA and ermB genes that mediate macrolide resistance. J Infect Chemother. 2003;9:221–6.
Cattoir V, Merabet L, Legrand P, Soussy CJ, Leclercq R. Emergence of a Streptococcus pneumoniae isolate resistant to streptogramins by mutation in ribosomal protein L22 during pristinamycin therapy of pneumococcal pneumonia. J Antimicrob Chemother. 2007;59:1010–2.
Reinert RR, Al-Lahham A. Time-kill study of the activity of telithromycin against macrolide-resistant Streptococcus pneumoniae isolates with 23S rRNA mutations and changes in ribosomal proteins L4 and L22. Antimicrob Agents Chemother. 2005;49:3011–3.
Bébéar CM, Pereyre S. Mechanisms of drug resistance in Mycoplasma pneumoniae. Curr Drug Targets Infect Disord. 2005;5:263–71.
Gruson D, Pereyre S, Renaudin H, Charron A, Bébéar C, Bébéar CM. In vitro development of resistance to six and four FQ in Mycoplasma pneumoniae and, respectively. Antimicrob Agents Chemother. 2005;49:1190–3.
Anokhina MM, Barta A, Nierhaus KH, Spiridonova VA, Kopylov AM. Mapping of the second tetracycline binding site on the ribosomal small subunit of E. coli. Nucleic Acids Res. 2004;32:2594–7.
Brodersen DE, Clemons WM Jr, Carter AP, Morgan-Warren RJ, Wimberly BT, Ramakrishnan V. The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell. 2000;103:1143–54.
Roberts MC, Koutsky LA, Holmes KK, LeBlanc DJ, Kenny GE. Tetracycline-resistant Mycoplasma hominis strains contain streptococcal tetM sequences. Antimicrob Agents Chemother. 1985;28:141–3.
Matsuoka M, Narita M, Okazaki N, Ohya H, Yamazaki T, Ouchi K, et al. Characterization and molecular analysis of macrolide-resistant Mycoplasma pneumoniae clinical isolates obtained in Japan. Antimicrob Agents Chemother. 2004;48:4624–30.
Isozumi R, Yoshimine H, Morozumi M, Ubukata K, Ariyoshi K. Adult case of community-acquired pneumonia caused by macrolide-resistant Mycoplasma pneumoniae. Respirology. 2009; 14(8):1206–8.
Peuchant O, Ménard A, Renaudin H, Morozumi M, Ubukata K, Bébéar CM, et al. Increased macrolide resistance of Mycoplasma pneumoniae in France directly detected in clinical specimens by real-time PCR and melting curve analysis. J Antimicrob Chemother. 2009;64:52–8.
Cousin-Allery A, Charron A, Barbeyrac BD, Fremy G, Jensen JS, Renaudin H, et al. Molecular typing of Mycoplasma pneumoniae strains by PCR-based methods and pulsed-field gel electrophoresis. Application to French and Danish isolates. Epidemiol Infect. 2000;124:103–11.
Numazaki K, Umetsu M, Adachi N. Mycoplasma pneumoniae infection and its genotypical characterization in children of Hokkaido, Japan. In Vivo. 2003;17:421–4.
Schwartz SB, Thurman KA, Mitchell SL, Wolff BJ, Winchell JM. Genotyping of Mycoplasma pneumoniae isolates using real-time PCR and high-resolution melt analysis. Clin Microbiol Infect. 2009;15:756–62.
Suzuki S, Yamazaki T, Narita M, Okazaki N, Suzuki I, Andoh T, et al. Clinical evaluation of macrolide-resistant Mycoplasma pneumoniae. Antimicrob Agents Chemother. 2006;50:709–12.
Matsubara K, Morozumi M, Okada T, Matsushima T, Komiyama O, Shoji M, et al. A comparative clinical study of macrolide-sensitive and macrolide-resistant Mycoplasma pneumoniae infections in pediatric patients. J Infect Chemother. 2009; 15(6):380–3.
Denny FW, Clyde WA Jr, Glezen WP. Mycoplasma pneumoniae disease: clinical spectrum, pathophysiology, epidemiology, and control. J Infect Dis. 1971;123:74–92.
Smith CB, Friedewald WT, Chanock RM. Shedding of Mycoplasma pneumoniae after tetracycline and erythromycin therapy. N Engl J Med. 1967;276:1172–5.
Lu YJ, Chen TH, Lin LH, Shen CM, Huang CH. Macrolide use shortens fever duration in Mycoplasma pneumoniae infection in children: a 2-year experience. J Microbiol Immunol Infect. 2008;41:307–10.
John SD, Ramanathan J, Swischuk LE. Spectrum of clinical and radiographic findings in pediatric mycoplasma pneumonia. Radiographics. 2001;21:121–31.
Wieslander A, Boyer MJ, Wroblewski H. Membrane protein structure. In: Maniloff J, McElhaney RN, Finch LR, Baseman JB, editors. Mycoplasmas: molecular biology, pathogenesis. Washington: American Society for Microbiology; 1992. p. 93–112.
Chmura K, Bai X, Nakamura M, Kandasamy P, McGibney M, Kuronuma K, et al. Induction of IL-8 by Mycoplasma pneumoniae membrane in BEAS-2B cells. Am J Physiol Lung Cell Mol Physiol. 2008;295:220–30.
Broaders SA, Hooper WC, Phillips DJ, Talkington DF. Mycoplasma pneumoniae subtype-independent induction of proinflammatory cytokines in THP-1 cells. Microb Pathog. 2006;40:286–92.
Chaudhry R, Varshney AK, Malhotra P. Adhesion proteins of Mycoplasma pneumoniae. Front Biosci. 2007;12:690–9.
Layh-Schmitt G, Podtelejnikov A, Mann M. Proteins complexed to the P1 adhesin of Mycoplasma pneumoniae. Microbiology. 2000;146:741–7.
Seto S, Miyata M. Attachment organelle formation represented by localization of cytadherence proteins and formation of the electron-dense core in wild-type and mutant strains of Mycoplasma pneumoniae. J Bacteriol. 2003;185:1082–91.
Waites KB, Talkington DF. Mycoplasma pneumoniae and its role as a human pathogen. Clin Microbiol Rev. 2004;17:697–728.
Yang J, Hooper WC, Phillips DJ, Talkington DF. Regulation of proinflammatory cytokines in human lung epithelial cells infected with Mycoplasma pneumoniae. Infect Immun. 2002;70:3649–55.
Yang J, Hooper WC, Phillips DJ, Talkington DF. Cytokines in Mycoplasma pneumoniae infections. Cytokine Growth Factor Rev. 2004;15:157–68.
Narita M, Tanaka H, Yamada S, Abe S, Ariga T, Sakiyama Y. Significant role of interleukin-8 in pathogenesis of pulmonary disease due to Mycoplasma pneumoniae infection. Clin Diagn Lab Immunol. 2001;8:1028–30.
Narita M, Tanaka H, Abe S, Yamada S, Kubota M, Togashi T. Close association between pulmonary disease manifestation in Mycoplasma pneumoniae infection and enhanced local production of interleukin-18 in the lung, independent of gamma interferon. Clin Diagn Lab Immunol. 2000;7:909–14.
Narita M, Tanaka H. Cytokines involved in the severe manifestations of pulmonary diseases caused by Mycoplasma pneumoniae. Pediatr Pulmonol. 2007;42:397.
Tanaka H, Narita M, Teramoto S, Saikai T, Oashi K, Igarashi T, et al. Role of interleukin-18 and T-helper type 1 cytokines in the development of Mycoplasma pneumoniae pneumonia in adults. Chest. 2002;121:1493–7.
Abe S, Nakamura H, Inoue S, Takeda H, Saito H, Kato S, et al. Interleukin-8 gene repression by clarithromycin is mediated by the activator protein-1 binding site in human bronchial epithelial cells. Am J Respir Cell Mol Biol. 2000;22:51–60.
Beović B, Bonac B, Kese D, Avsic-Zupanc T, Kreft S, Lesnicar G, et al. Aetiology and clinical presentation of mild community-acquired bacterial pneumonia. Eur J Clin Microbiol Infect Dis. 2003;22:584–91.
Marrie TJ. Epidemiology of mild pneumonia. Semin Respir Infect. 1998;13:3–7.
Waites KB. New concepts of Mycoplasma pneumoniae infections in children. Pediatr Pulmonol. 2003;36:267–78.
Miyashita N, Obase Y, Ouchi K, Kawasaki K, Kawai Y, Kobashi Y, et al. Clinical features of severe Mycoplasma pneumoniae pneumonia in adults admitted to an intensive care unit. J Med Microbiol. 2007;56:1625–9.
Kim DH, Lee KY, Kim MS, Youn YS, Hwang JY, Rhim JW, et al. Corticosteroid treatment in siblings affected with severe Mycoplasma pneumoniae pneumonia. Infect Chemother. 2009;41(3):190–5.
Tagliabue C, Salvatore CM, Techasaensiri C, Mejias A, Torres JP, Katz K, et al. The impact of steroids given with macrolide therapy on experimental Mycoplasma pneumoniae respiratory infection. J Infect Dis. 2008;198:1180–8.
Radisic M, Torn A, Gutierrez P, Defranchi HA, Pardo P. Severe acute lung injury caused by Mycoplasma pneumoniae: potential role for steroid pulses in treatment. Clin Infect Dis. 2000;31:1507–11.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Morozumi, M., Takahashi, T. & Ubukata, K. Macrolide-resistant Mycoplasma pneumoniae: characteristics of isolates and clinical aspects of community-acquired pneumonia. J Infect Chemother 16, 78–86 (2010). https://doi.org/10.1007/s10156-009-0021-4
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10156-009-0021-4