Abstract
In this study, we explored the pyroptosis-related biomarkers and signatures of colorectal cancer (CRC). Gene expression profiles were downloaded from the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA)-COADREAD and were analyzed for differentially expressed genes (DEGs). DEGs in CRC‒pyroptosis-related genes (CRC‒PRGs) were obtained by intersecting DEGs associated with CRC and PRGs. The CRC‒PRGs were verified; functional enrichment analysis was performed with Gene Ontology (GO) followed by cluster analysis. Cox analyses and LASSO regression were used in TCGA dataset to construct a prognostic model for patients with CRC. A prognostic risk assessment model was constructed and efficacy was evaluated. Decision curve analysis was utilized to assess the role of the Lasso-Cox regression prognostic model for clinical utility at 1, 3, and 5 years. Twelve CRC‒PRGs were identified as prognostic pyroptosis-related DEGs. CXCL8, IL13RA2, MELK, and POP1 were selected as prognostic genes to construct features with a good prognostic performance in GEO and TCGA. Functional enrichment indicated that the 4-gene signature might be involved in CRC tumorigenesis and development through various pathways by playing a prognostic role in CRC. Furthermore, the results of the immune landscape analysis showed that the expression of CXCL8 and IL13RA2 in TCGA-COADREAD dataset was positively correlated with significant differential enrichment of most immune cells. A novel prognostic model consisting of four key genes, CXCL8, IL13RA2, MELK, and POP1, can accurately predict the survival of patients with CRC. This finding may provide a new perspective for the treatment of pyroptosis-related CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC) is a common malignancy arising from the digestive system in humans, including colon and rectal cancers, and is highly prevalent. It has a high global mortality rate and currently exhibits a rising tendency in both morbidity and mortality (Siegel et al. 2021). CRC is the third and fifth leading cause of cancer-related mortality in the USA and China, respectively (Siegel et al. 2020). The molecular mechanism of CRC is a multistage process that involves multiple genetic and polygenic variations (Fearon and Vogelstein 1990). It is therefore challenging to develop new therapeutic methods for the diagnosis, treatment, and prognosis of CRC. However, the greatest drawback of TNM classification is that it cannot fully reflect the genetic heterogeneity of individual tumors (Hegde et al. 2014). With the continuous improvement in gene sequencing, epigenetic research on tumors has attracted increasing attention. However, because of the complex molecular mechanisms affecting the prognosis of CRC, the accuracy of single gene/factor prediction models is poor (Zhuang et al. 2021). In contrast, polygenic patterns provide a better prediction of the prognosis of the different tumor types (Zhang et al. 2020a; Xue et al. 2020; Bao et al. 2020). Therefore, to personalize treatment and predict survival in patients with CRC, it is necessary to have a reliable prognostic gene profile.
Pyroptosis, also known as cellular inflammatory necrosis, is a programmed death characterized by cell swelling; until the cell membrane is broken, substances in the cell are released, resulting in a strong inflammatory response (Shi et al. 2017). A long-term chronic inflammatory response can lead to the development of local tumor tissues. In particular, when there are many bacteria in the gut, it can easily cause infection, which in turn causes cells death. We believe that pyroptosis is an important factor in the development of CRC. Several studies have suggested that apoptosis is related to CRC (Yu et al. 2019; Wu et al. 2020; Tian et al. 2020). To date, there have been few scientific and clinical studies on the relationship between CRC and pyroptosis. The prognosis of patients with CRC and the expression characteristics of the main pyroptosis-related genes (PRGs) in CRC progression remains unclear. Although great progress in the study of CRC genes has been made, the use of their associated gene characteristics to establish the prognostic properties of CRC has rarely been studied. Currently, chemo-, endocrine-, and immunotherapy, and other treatments alone cannot achieve the desired effects. Exploring the role of PRGs in CRC and its relationship with the immune microenvironment can lead to new development directions for treatment (Zhuang et al. 2021).
The purpose of this study was to explore the genes related to cell pyroptosis, explore their expression characteristics in normal and tumor tissues, and predict the prognosis and immune response of patients by analyzing the prognostic indicators. Moreover, the correlation between the pyroptosis-related pathways and CRC was analyzed using the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) databases. A total of 12 CRC‒PRGs were obtained. CRC‒PRGs were verified, and functional enrichment analysis was performed using Gene Ontology (GO) annotation analysis, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, and gene set enrichment analysis (GSEA). The receiver operating characteristic (ROC) curve was used to evaluate the diagnostic predictive value of the CRC‒PRG-related genes. The single sample GSEA (ssGSEA) algorithm was used to analyze immune infiltration in CRC‒PRGs and immune infiltration levels. By studying pyroptosis, we can further understand the mechanism of CRC, thus revealing new avenues for treatment methods.
Materials and methods
Data acquisition and procession
The expression profile dataset GSE113513 (Shen et al. 2021) of patients with CRC was downloaded from the GEO database (Barrett et al. 2007) using the R package GEOquery (Davis and Meltzer 2007). The dataset GSE113513 is from Homo sapiens. The GSE113513 dataset contains gene expression profiles of colorectal tumor tissues and matched normal colorectal tissues from patients with CRC. A total of 28 samples were analyzed, including 14 CRC tumor and 14 matched normal colorectal tissue samples. The data platform used was the GPL15207 [PrimeView] Affymetrix Human Gene Expression Array, and the data set probe name annotations all use the chip GPL platform file. All expression profiling data samples in GSE113513 were included in the subsequent analysis, including the expression profiling data of the 14 colorectal tumor tissues (group: tumor) and the corresponding 14 normal colorectal tissues (grouped: normal).
In addition, we also downloaded the CRC dataset (TCGA-COADREAD) through the TCGAbiolinks package (Colaprico et al. 2016), from TCGA as a set for subsequent validation. A total of 698 CRC samples with complete clinical information were obtained, including tumor tissues from 647 patients with CRC (cancer group, group: tumor) and 51 CRC adjacent normal tissues that were partially matched. Count sequencing data of the tissue (normal group, group: normal) were normalized to FPKM (fragments per kilobase per million) format, and the corresponding clinical data were obtained from the UCSC Xena database (Goldman et al. 2020) (http://genome.ucsc.edu). The count sequencing data and corresponding clinical data of the CRC dataset (TCGA-COADREAD) were normalized using the limma package (Ritchie et al. 2015).
In addition, we collected PRGs from the GeneCards database (Stelzer et al. 2016) (https://www.genecards.org/) and the MsigDB (Molecular Signatures Database, http://www.gsea-msigdb.org/) database (Liberzon et al. 2015). We used the term “pyroptosis” as the search key to identify 254 PRGs. We used the term “pyroptosis” as the search key from the MsigDB database to obtain 27 PRGs. In addition, we also used “pyroptosis-related genes” as the search keywords on the PubMed website and obtained the pyroptosis-related gene set from the published literature (Xu et al. 2021). After merging and deduplicating, 274 PRGs were identified (see Table S1).
CRC-related differentially expressed genes
To identify the potential mechanism of action of differential genes and related biological features and pathways in CRC, we first normalized the CRC dataset GSE113513 and dataset TCGA-COADREAD using the limma package and then used a linear model to identify the results. Differentially expressed genes (DEGs) in rectal cancer (group: tumor) and normal (group: normal) samples. We used the DESeq2 (Love et al. 2014) package to perform differential analysis on the count data of the GSE113513 and TCGA-COADREAD datasets, and the genes screened by the criteria of |logFoldChange (FC)|> 1 and adjusted P-value (P.adj) < 0.05. Genes with logFC > 1 and P.adj < 0.05 were DEGs with upregulated expression, and genes with logFC < − 1 and P.adj < 0.05 were DEGs with downregulated expression.
To determine PRGs related to CRC, we first all the DEGs with |logFC|> 1 and P.adj < 0.05, obtained by the difference analysis between the TCGA-COADREAD and GSE113513 datasets. By drawing a Venn diagram, the DEGs of the dataset were obtained. Moreover, the common DEGs of the two datasets and the PRGs were intersected and a Venn diagram was drawn. The results of the differential analysis were visualized using the R package ggplot2 to draw a volcano map, and the R package pheatmap drew a heatmap display.
Functional enrichment analysis
GO analysis is a common method utilized for large-scale functional enrichment studies, including biological processes (BP), molecular functions (MF), and cellular components (CC) (Yu 2020). We used the R package clusterProfiler (Yu et al. 2012) to perform GO annotation analysis of pyroptosis-related DEGs. The entry screening criteria were P < 0.05, FDR value (q value) < 0.05 was considered statistically significant, and P values were corrected by Benjamini‒Hochberg method.
GSEA and GSVA
Gene set enrichment analysis (GSEA) was used to evaluate the gene distribution trend in a predefined gene set in the gene table and determine its contribution to the phenotype through the correlation between phenotypes (Subramanian et al. 2005). In this study, the genes in the TCGA-COADREAD and GSE113513 datasets were first sorted into two groups according to their phenotypic correlation.
The clusterProfiler package was used to perform enrichment analysis on all differential genes in the two groups with high and low phenotype correlation. The parameters used in this GSEA enrichment analysis were as follows: the number of seeds was 2020, the number of computations was 1000, the number of genes contained in each gene set was at least 10, and the maximum number of genes contained was 500. The P value correction was performed using the Benjamini‒Hochberg method. We obtained the c2.cp.v7.2 symbols gene set from the Molecular Signatures Database (MsigDB) database (Liberzon et al. 2015), and the screening criteria for significant enrichment were P < 0.05 and FDR value < 0.05.
Gene set variation analysis (GSVA) (Hanzelmann et al. 2013) is a nonparametric, unsupervised analysis method that converts the expression matrix of different genes across samples into the expression between genes. The enrichment effect of the genetic resources was assessed using a quantitative matrix of the nuclear microarray transcription. To evaluate the enrichment of the different pathways in different samples, we obtained the “h.all.v7.4. symbols.gmt” gene set from the MsigDB database and performed GSVA analysis on the pyroptosis-related prognostic differentially expressed genes in the dataset GSE113513 to calculate the pyroptosis-related prognostic DEGs in the colorectum differences in functional enrichment between cancer tumor tissue samples (group: tumor) and corresponding normal colorectal tissue samples (group: normal).
Assessment of the tumor microenvironment
We used the single-sample gene-set enrichment analysis (ssGSEA) algorithm to quantify the relative abundance of each immune infiltration cell. Each infiltrating immune cell was labeled, such as activated CD8 T cells, activated dendritic cells, macrophages, T cells, regulatory T cells, and various other subtypes of natural killer cells. The degree of infiltration of each immune cell in each sample was expressed as the abundance calculated using ssGSEA analysis (Charoentong et al. 2017; Barbie et al. (n.d.)). In GSE113513, we used the ggplot2 package to predict the correlation between the expression of DEGs and the invasion of immune cells in different tumor samples and predicted them on the TCGA-COADREAD dataset.
CIBERSORT (Newman et al. (n.d.)) is an immune infiltration analysis algorithm that deconvolution the transcriptome expression matrix based on the principle of linear support vector regression to estimate the composition and abundance of immune cells in the mixed cells.
We uploaded the matrix data of the TCGA-COADREAD dataset to CIBERSORT, and combined with the LM22 characteristic gene matrix, to screen out the data with the immune cell enrichment fraction greater than zero, and finally obtained and displayed the specific results of the immune cell infiltration abundance matrix.
The proportion of immunocyte infiltration abundance for samples from the TCGA-COADREAD dataset was displayed as a stacked bar graph, while the difference in infiltration abundance of immunocytes between subgroups (tumor/normal) was displayed as a boxplot. The correlation of the immune cells in the different subgroups was calculated by the spearman algorithm and visualized by R pack bag ggplot2.
Construction of protein–protein interaction network
To interact in many aspects of life processes, including biological signal transmission, gene expression regulation, energy and material metabolism, and cell cycle regulation, individual proteins interact with each other to form a protein‒protein interaction (PPI) network. Understanding the functioning of proteins in biological systems, the response mechanism of biological signals, energy metabolism in particular physiological states such as diseases, and the functional relationships between proteins all depend on the systematic analysis of the interaction of many proteins in biological systems which possesses significant meaning. The STRING database (Szklarczyk et al. 2019) is a database that searches existing proteins and predicts their role. In this study, a PPI network (confidence level 0.4) related to DEGs was established using the STRING database, and the PPI network was visualized using Cytoscape.
Construction of mRNA-RBP, mRNA-TF, mRNA-drugs interaction network
The Starbase database (Li et al. 2014) uses high-throughput experimental data of CLIP-Seq, combined with degradome experimental data, to find miRNA targets and provides a variety of visualization interfaces. The database contains abundant RNA binding proteins (RBP)-ncRNA, RBP-mRNA, RBP-RNA, and RNA-RNA data. The miRNA Target Prediction Database, miRDB, (Chen and Wang 2020) was utilized for RBP target gene prediction and functional annotation. We used the starBase database for RBPs that interact with pyroptosis-related mRNAs.
The CHIPBase database (Zhou et al. 2017) (version 2.0) (https://rna.sysu.edu.cn/chipbase/) I identifies thousands of binding motif sequences and their binding sites from the DNA-binding protein ChIP-seq data and predicted the relationship between millions of transcription factors (TFs) and genes. The hTFtarget database (Zhang et al. 2020b) (http://bioinfo.life.hust.edu.cn/hTFtarget.) is a comprehensive database of human TFs and their target regulation. We used the CHIPBase and hTFtarget database to identify TFs associated with DEGs related to pyroptosis and visualized them using Cytoscape software.
In addition, we utilized the drug-gene interaction database (DGIdb) (Freshour et al. 2021) (https://www.dgidb.org) to predict possible drugs or small molecule compounds with DEG interactions associated with pyroptosis. The mRNA-RBP, mRNA-TF, and mRNA‒drug interaction networks were visualized using the Cytoscape software.
ROC
Receiver operating characteristic curve (ROC) (Mandrekar 2010) is a graphical analysis tool that can select the best model, discard the suboptimal model, or set the best threshold within the same model. The ROC curve is a comprehensive index that reflects the sensitivity and specificity. The relationship between sensitivity and specificity was analyzed using combinatorial methods. The area under the ROC curve was typically 0.5‒1. When the area under the curve (AUC) is closer to 1, the diagnostic effect was better. The AUC has low accuracy when it is 0.5 to 0.7, the AUC has a certain accuracy when it is 0.7 to 0.9, and the AUC has high accuracy when it is above 0.9. We used the R survivalROC package to draw the ROC curve of the pyroptosis-related DEGs and patient survival time and survival status and calculated the AUC to evaluate the diagnostic effect of gene expression on the survival of patients with CRC.
Clinical correlation analysis
To study the clinical prognostic value of pyroptosis-related prognostic DEGs in CRC, we performed univariate Cox regression analysis to analyze the expression of prognostic DEGs related to pyroptosis in CRC. Factors with P < 0.1 were selected for multivariate Cox regression analysis, and a multivariate Cox regression model was established. Based on the results of univariate Cox regression analysis, we established a nomogram to predict the 1-, 3-, and 5-year survival rates of patients with CRC. A nomogram is a graph in which a cluster of disjoint line segments that is used to represent the functional relationship between multiple independent variables in a planar rectangular coordinate system. The accuracy and resolution of the calibration plots were evaluated using calibration curves. Decision curve analysis (DCA) is a convenient method to evaluate clinical predictive models, diagnostic tests, and molecular markers. We used the R package ggDCA (Tataranni and Piccoli 2019) to evaluate the predictive effect of Cox regression models on the 1-, 3-, and 5-year survival outcomes of patients with CRC.
Gene expression levels and clinical characteristics are associated with patient prognosis. We conducted differential analyses of the expression levels of pyroptosis-related prognostic DEGs in the TCGA-COADREAD dataset to further evaluate the effect of pyroptosis-related prognostic DEGs on patient prognosis. The influence of clinicopathological features and expression differences of pyroptosis-related prognostic DEGs were compared among different clinical features. We analyzed the effect of the expression levels of pyroptosis-related prognostic DEGs in CRC tissues of the tumor, including overall survival (OS), disease-specific survival (DSS), and progression-free interval (PFI).
Gene mutation analysis and single gene analysis
The cBioPortal database (Subramanian et al. 2005) (cBioPortal for Cancer Genomics) (http://cbioportal.org) provides a web resource for exploring, visualizing, and analyzing multiple tumor genetic data. This database summarizes the molecular analysis data from tumor tissues and cell lines into easy-to-understand genetic, epigenetic, gene expression, and protein groups. Using the cBioPortal database, we analyzed the gene mutation status of the final selected pyroptosis-related prognostic DEGs in the TCGA-COADREAD (CRC) dataset and displayed the final analysis results.
We also used Human Protein Atlas (HPA) database (Thul and Lindskog 2018) (www.proteinatlas.org/) to conduct single cell analysis on the expression of the differentially expressed genes pyroptosis-related prognostic DEGs in CRC. Based on the expression of genes in different tissues and cells in human body, HPA database conducted single cell analysis on the differentially expressed genes pyroptosis-related prognostic DEGs in CRC in human kidney cells in human colon tissue samples and human Rectum tissue samples, and displayed the results.
Statistical analysis
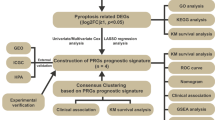
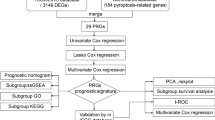
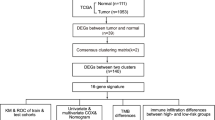
All analyses in this study were conducted in R software (Version 4.1.2) using the various mentioned packages, and continuous variables are presented as mean ± standard deviation. The Wilcoxon rank-sum method was used to compare two groups, the Kruskal‒Wallis test was used to compare more than three populations, and the Kaplan‒Meier (KM) method combined with the log-rank test was used to compare the progression-free survival between the two groups. Unless otherwise specified, P < 0.05 was the criterion for significant difference (Fig. 1).
Workflow. TCGA The cancer genome atlas, COADREAD colon and rectal cancer, DEGs differentially expressed genes, GO Gene Ontology, GSEA gene set enrichment analysis, PPI network: protein–protein interaction network, RBP RNA binding protein, TF transcription factors, LASSO least absolute shrinkage and selection operator, GSVA gene set variation analysis
Results
Metabolism-related DEGs in CRC
We normalized the data from the CRC tumor tissue samples (cancer group, group: tumor) and normal colorectal tissue samples (normal group, group: normal) in the TCGA-COADREAD dataset, GSE113513 dataset, using the limma package. To analyze the differences in gene expression values in the CRC group (tumor) relative to the normal control group (normal), we performed differential analysis on the TCGA-COADREAD dataset and the GSE113513 dataset using the DESeq2 package to obtain the DEGs of the two groups of data. A total of 18,670 DEGs were obtained from TCGA-COADREAD, of which 5470 met the thresholds of |logFC|> 1 and P.adj < 0.05. At this threshold, the number of high (low expression in normal group, logFC is positive, upregulated gene) and low expression (high expression in normal group, logFC is negative) in the cancer group was 2785 and 2685 individuals, respectively.
We drew a volcano plot of the differential analysis results of TCGA-COADREAD dataset (Fig. 2A), and a total of 17,009 DEGs were obtained from dataset GSE113513, of which 1406 met the threshold of |logFC|> 1 and P.adj < 0.05. At this threshold, the number of logFC positive (upregulated genes) was 609 and the logFC negative (downregulated genes) was 797; we plotted a volcano plot from the variance analysis of this dataset (Fig. 2B). To determine the pyroptosis-related DEGs, we first obtained the intersection of all the DEGs obtained from the TCGA-COADREAD and GSE113513 datasets with |logFC|> 1 and P.adj < 0.05 and used this to establish the CRC dataset. A Venn diagram was drawn for the 1215 common DEGs (Fig. 2C). We used the intersection of the common DEGs and PRGs in the dataset to obtain a total of 12 pyroptosis-related DEGs in CRC and drew another Venn diagram (Fig. 2D). The 12 pyroptosis-related DEGs included DPEP1, CTSG, GZMB, POP1, IL13RA2, CHI3L1, BHLHE40, CASP5, MELK, PCSK9, CXCL8, and MPEG1. Based on the results obtained from the Venn diagram, we analyzed the expression differences of 12 pyroptosis-related DEGs in the TCGA-COADREAD dataset (Fig. 2E) and the GSE113513 dataset (Fig. 2F). The R package pheatmap was used to draw a heat map showing the differential analysis results of the 12 pyroptosis-related DEGs (Fig. 2E and F).
Analysis of metabolically related differential genes in CRC. A, B Results normalized by GSE7014 and GSE25724. Blue represents the normal group, pink represents the disease group. A Differential gene analysis volcano plot of colorectal cancer tissues (group: tumor) and adjacent normal tissues (group: normal) of the TCGA-COADREAD dataset. B Differential gene analysis volcano plot of colorectal cancer tissue (group: tumor) and normal colorectal tissue (group: Normal) of GSE113513 dataset. C Venn diagram of DEGs in TCGA-COADREAD dataset and GSE113513 dataset. D Venn diagram of common DEGs (dataset DEGs) and pyroptosis-related genes in the dataset. E, F Complex numerical heatmaps of pyroptosis-related DEGs in TCGA-COADREAD dataset (E), GSE113513 dataset (F). TCGA The cancer genome atlas, COADREAD colon and rectal cancer, DEGs differentially expressed genes
In addition, we compared the expression of 12 pyroptosis-related DEGs in the rectal cancer data set (READ) with the clinical prognosis overall survival (OS) to determine the correlation between the expression levels of the 12 pyroptosis-related DEGs and the prognostic clinical overall survival of patients with READ, as shown in Fig. S1. We also performed correlation analysis using the Spearman statistical method on the expressions of 12 pyroptosis-related DEGs in the colon cancer dataset(COAD) and READ by using the TIMER2.0 (Li et al. 2009) database (http://timer.cistrome.org/) and retained the results with the correlation coefficient greater than 0.2 and displayed the specific results. In the TIMER2.0 database, we found the correlation of eight differentially expressed genes related to cell scorch (BHLHE40, CHI3L1, CASP5, CTSG, GZMB, MPEG1, POP1, MELK) in the tumor data sets of COAD and READ. The specific results are shown in Fig. S2.
Functional enrichment analysis of pyroptosis-related DEGs
To analyze the BPs, MFs, CCs, biological pathways, and their relationship with CRC, we first performed GO function enrichment analysis of the DEGs related to pyroptosis (Table 1). It is considered statistically significant that the entry screening criteria is P value < 0.05 and the FDR value (q value) < 0.05. The results showed that the 12 DEGs related to pyroptosis were mainly enriched in neutrophil-mediated cytotoxicity, leukocyte-mediated cytotoxicity, receptor internalization, antimicrobial humoral response, and other BPs (Fig. 3A); as well as secretory granule lumen, cytoplasmic vesicle lumen, vesicle lumen, external side of plasma membrane, and other CCs (Fig. 3B); and also enriched in endopeptidase activity, serine-type endopeptidase activity, serine-type peptidase activity, serine hydrolase activity, and other MFs (Fig. 3C). We present the results of the GO functional enrichment analysis (Fig. 3A‒C), where the abscissa is − log(p.adjust), the ordinate is GO terms, and the color of the bubble chart indicates the activation or inhibition of GO terms. In addition, we also displayed the BP (Fig. 3D), CC (Fig. 3E), and MF (Fig. 3F) analysis results of GO gene function enrichment in the form of a ring network diagram (Fig. 3D‒F).
Functional enrichment analysis (GO) of pyroptosis-related DEGs. A–C GO functional enrichment of pyroptosis-related DEGs. BP analysis results in bubble chart display (A), CC analysis results in bubble chart display (B), and MF analysis results in bubble chart display (C). D–F GO function enrichment of pyroptosis-related DEGs. BP analysis results of ring network diagram display (D), CC analysis results ring network diagram display (E), MF analysis results ring network diagram display (F). The bubble color in the bubble plot (A–C) indicates the size of the Padj value for GO terms, red indicates a small Padj value, and blue indicates a large Padj value. In the circular network diagram (D–F), red dots represent specific genes, and blue circles represent specific pathways. P value < 0.05 and FDR value (q value) < 0.05 were considered to be statistically significant for the functional enrichment analysis entry screening criteria. GO Gene Ontology, BP biological process, CC cellular component, MF molecular function
GSEA
To determine the effect of the expression levels of metabolically related DEGs in CRC on the occurrence of colorectal carcinogenesis, we analyzed the relationship between all the expression of DEGs in the TCGA-COADREAD and GSE113513 datasets through GSEA enrichment analysis. The screening criteria for significant enrichment of results from GSEA enrichment analysis were P < 0.05 and FDR value (q value) < 0.05. Links between BPs, affected CCs, and MFs showed that the DEGs in TCGA-COADREAD were significantly enriched in Reactome keratinization (Fig. 4B), Reactome amyloid fiber formation (Fig. 4C), Reactome DNA methylation (Fig. 4D), Reactome deacetylate histones (Fig. 4E), and other pathways (Fig. 4A‒E, Table 2). The DEGs in dataset GSE113513 were significantly enriched in the Reactome cell cycle checkpoints (Fig. 4G), Reactome mitotic spindle checkpoints (Fig. 4H), Reactome s phase (Fig. 4I), Reactome snRNP assembly (Fig. 4J), and other pathways (Fig. 4F–J, Table 3). In addition, Reactome meiotic recombination, Reactome condensation of prophase chromosomes, and a total of 180 functional pathways, such as Reactome chromosome maintenance, were significantly enriched by both datasets simultaneously.
GSEA enrichment analysis of colorectal cancer dataset. A GSEA enrichment analysis of the TCGA-COADREAD dataset for the main 4 main biological features. B–E The DEGs in the TCGA-COADREAD dataset were significantly enriched in pathways such as Reactome keratinization (B), Reactome amyloid fiber formation (C), Reactome DNA methylation (D), Reactome HDACs deacetylate histones (E). F The GSEA analysis of the GSE113513 dataset mainly includes 4 main biological characteristics. G–J DEGs in the GSE113513 dataset were significantly enriched in Reactome cell cycle checkpoints (G), Reactome mitotic spindle checkpoints (H), Reactome S phase (I), Reactome snrnp assembly (J). The screening criteria for significant enrichment of GSEA analysis results are P < 0.05 and FDR value (q value) < 0.05. TCGA The cancer genome atlas, COADREAD colon and rectal cancer, DEGs differentially expressed genes
Construction of protein–protein interaction network, mRNA-RBP, mRNA-TF, and mRNA-drugs interaction network
Protein‒protein interaction (PPI) analysis of the 12 pyroptosis-related DEGs using the STRING database (confidence level 0.4), we constructed a PPI of DEGs related to pyroptosis, and used Cytoscape software to visualize the interaction (Fig. 5A). Only five pyroptosis-related DEGs (CTSG, CXCL8, CHI3L1, IL13RA2, and GZMB) were related to other genes in the PPI network.
Construction of protein–protein interaction network (PPI), mRNA-RBP, mRNA-TF, and mRNA-drugs interaction network. A Protein interaction network (PPI) of DEGs related to pyroptosis. B–D The mRNA-RBP (B), mRNA-TF (C), and mRNA-drugs (D) interaction networks of DEGs related to pyroptosis. In the mRNA-RBP (B) interaction network, the sky blue oval block is mRNA; the light dark green block is RBP. In the mRNA-TF (C) interaction network, the sky blue oval block is mRNA; the pink diamond block is TF; the light green diamond block is both mRNA and TF. In the mRNA-drugs (D) interaction network, the sky blue oval block is mRNA; the orange hexagonal block is drug. TF: transcription factor; RBP: RNA binding protein
We used the mRNA-RBP data in the starBase database to predict interactions with 12 pyroptosis-related DEGs (mRNAs). The acting RBP was then visualized by drawing the mRNA‒RBP interaction network using the Cytoscape software (Fig. 5B). According to the mRNA-RBP interaction network, our mRNA-RBP interaction network consists of seven mRNAs (DEGs related to pyroptosis) (BHLHE40, PCSK9, CXCL8, MELK, POP1, CHI3L1, and DPEP1), 40 RBP molecules, a total of 64 pairs of mRNA-RBP interaction relationships, and specific mRNA‒RBP interaction relationships (Table S2).
We used the CHIPBase and hTFtarget databases to search for TFs associated with the DEGs related to pyroptosis. After downloading the interaction relationships found in the two databases, we used the intersection with the 12 pyroptosis-related DEGs, and finally obtained nine pyroptosis-related DEGs (BHLHE40, CASP5, CHI3L1, CXCL8, DPEP1, MELK, MPEG1, PCSK9, and POP1) and the interaction data of 58 TFs and visualized them using Cytoscape software. The sky-blue oval block was mRNA; pink diamond block was TF; light green diamond-shaped blocks were both mRNAs and TFs (Fig. 5C). In the mRNA‒TF interaction network, the pyroptosis-related DEG BHLHE40 had the strongest interaction with TFs. There were 28 pairs of mRNA-TF interaction relationships in the BHLHE40 gene (Table S3).
The DGIdb database was used to identify potential drugs or molecular compounds of the 12 DEGs (mRNAs) associated with pyroptosis. We identified 89 potential drugs or molecular compounds corresponding to 8 mRNAs (CASP5, CTSG, CXCL8, DPEP1, GZMB, IL13RA2, MELK, and PCSK9) through the DGIdb database, as shown by the mRNA‒drug interaction network, sky blue oval blocks are mRNAs; orange hexagonal blocks are drugs (Fig. 5D). Among them, we found that 56 drugs or molecular compounds target the CXCL8 gene and the specific mRNA‒drug interaction relationship (Table S4).
Construction of a prognostic model of pyroptosis-related DEGs and GSVA analysis
To determine the prognostic value of the 12 pyroptosis-related DEGs in the TCGA-COADREAD dataset, we used LASSO regression analysis to construct a prognostic model (Fig. 6A). LASSO regression was based on linear regression. Overfitting of the model is reduced, and the generalization ability of the model is improved by increasing the penalty term (lambda × absolute value of slope). The ordinate of the LASSO regression pattern graph represents the likelihood deviation of the LASSO regression, the log (λ) value on the lower x-axis of the graph by default represents the logarithm of the LASSO regression after the lambda coefficient of the penalty term, and the values on the x-axis represent the logarithm. The numbers on the upper x-axis represent the number of variables with nonzero coefficients for each lambda. In addition, we visualized the LASSO regression results and obtained the LASSO variable trajectory plot (Fig. 6B). From the figure, we can see that there is a total of four genes in the LASSO regression prognostic model was constructed, namely CXCL8, IL13RA2, MELK, and POP1. In this regard, we visualized the risk factor grouping of the constructed LASSO regression prognostic model using a risk factor plot (Fig. 6C). The risk factor map consists of three parts: (1) Risk grouping: the risk score predicted by the LASSO regression prognostic model was grouped by the median; (2) Survival outcomes, displayed as a dot plot based on the TCGA-COADREAD dataset survival time and survival outcomes of clinical samples; (3) Heat map, visualization of the expression of pyroptosis-related prognostic DEGs in the LASSO regression prognostic model.
Construction of a prognostic model of pyroptosis-related DEGs and GSVA analysis. A LASSO regression prognostic model diagram of pyroptosis-related DEGs. B, C LASSO regression prognostic model variable trajectory plot (B), risk factor plot (C). D GSVA analysis results of pyroptosis-related prognostic DEGs. The screening standard of significant enrichment of GSEA analysis results is P < 0.05. GSVA gene set variation analysis, LASSO least absolute shrinkage and selection operator
To explore the differences in the hallmark gene set between CRC tumor tissue (group: tumor) and normal colorectal tissue (group: normal), we analyzed the pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, MELK, and POP1) in the dataset GSE113513 using GSVA. GSVA analysis of pyroptosis-related prognostic DEGs in the GSE113513 dataset showed hallmark pancreatic beta cells, adipogenesis, heme metabolism, and myc targets. Thirty hallmark gene sets showed differences between CRC tumor (group: tumor) and normal colorectal tissue (group: normal) (Fig. 6D, Table 4). Among them, hallmark pancreatic beta cells, adipogenesis, heme metabolism, myogenesis, and 23 other hallmark gene sets had significantly higher enrichment scores in CRC tumor tissue than in normal colorectal tissue, while hallmark e2f targets, Myc targets V1, Myc targets V2, and a total of seven gene sets had a significantly lower enrichment score in CRC tumor tissue than in normal colorectal tissue.
Assessment of pyroptosis-related prognostic DEGs and tumor microenvironment
To analyze the difference in immune infiltration between CRC tumor tissue (group: tumor) and normal colorectal tissue (group: normal) in the CRC dataset GSE113513, we used the ssGSEA algorithm to calculate the different groupings. Differences in the degree of infiltration of the 28 immune cells were calculated and the results showed that 17 types of immune cells were significantly enriched in the GSE113513 dataset, namely activated B cell, activated CD4 T cell, CD56bright natural killer cell, central memory CD8 T cell, effector memory CD4 T cell, effector memory CD8 T cell, eosinophil, gamma delta T cell, immature B cell, macrophage, mast cell, memory B cell, monocyte, natural killer cell, neutrophil, T follicular helper cell, type 1 T helper cell. We showed the immune infiltration results of 28 types of immune cells in the colorectal Cancer dataset GSE113513 in the form of a group comparison chart (Fig. 7A).
Assessment of the tumor microenvironment of DEGs associated with pyroptosis. A The immune infiltration results of the dataset GSE113513 are grouped and compared. B–E Pyroptosis-related prognostic DEGs CXCL8 (B), IL13RA2 (C), MELK (D), POP1 (E) in the TCGA-COADREAD data set. The correlation results show that the expression of immune cells. The symbol ns is equivalent to P > 0.05, not statistically significant; the symbol * is equivalent to P ≤ 0.05, which is statistically significant; the symbol ** is equivalent to P ≤ 0.01, which is highly statistically significant; the symbol *** is equivalent to P ≤ 0.01 ≤ 0.001, highly statistically significant. ssGSEA single-sample gene-set enrichment analysis, TCGA The cancer genome atlas, COADREAD colon and rectal cancer
Moreover, we used the ssGSEA algorithm to compare the expression of pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, MELK, and POP1) in the TCGA-COADREAD dataset with 24 immune cells (NK CD56bright cells, NK cells, Th17 cells, Tcm, B cells, NK CD56dim cells, TFH, pDC, TReg, CD8 T cells, Tem, Eosinophils, Mast cells, iDC, Tgd, T cells, Th2 cells, T helper cells, cytotoxic cells, aDC, DC, macrophages, Th1 cells, and neutrophils), and the results showed that the expression of pyroptosis-related prognostic DEGs CXCL8 (Fig. 7B) and IL13RA2 (Fig. 7C) in the TCGA-COADREAD dataset was positively correlated with the significant differential enrichment of most immune cells. The pyroptosis-related prognostic DEGs, including MELK (Fig. 7D) and POP1 (Fig. 7E), in the TCGA-COADREAD dataset were negatively correlated with significant differential enrichment of most immune cells (Fig. 7B‒E).
CIBERSORT immunoinfiltration analysis of TCGA data set of CRC
To explore the difference of immune infiltration among different groups (tumor/normal) in TCGA-COADREAD data set, we used CIBERSORT algorithm to calculate the infiltration abundance of 22 kinds of immune cells in two disease subtypes samples for the tumor group samples and the normal group samples in TCGA-COADREAD data set. Then, boxplot diagram is used to show the percentage of infiltration abundance of immune cells in TCGA-COADREAD data set samples (Fig. 8A). It can be seen from the figure that the percentage of infiltration abundance of immune cells Macrophages M0, T cells CD8, and T cells follicular helper in TCGA-COADREAD data set samples is relatively high.
CIBERSORT immunoinfiltration analysis of TCGA data set of CRC. A, B CIBERSORT immunoinfiltration analysis results of TCGA-COADREAD data set are shown by the accumulation histogram of infiltration abundance (A) and the grouping comparison chart (B). C The correlation analysis results of infiltration abundance of 16 kinds of immune cells in TCGA-COADREAD data set show. Ns symbol is equal to P ≥ 0.05, which has no statistical significance; The symbol * is equivalent to P < 0.05; The symbol ** is equivalent to P < 0.01. The symbol *** is equivalent to P < 0.001. TCGA The cancer genome atlas, COADREAD colon adenocarcinoma/rectum adenocarcinoma esophageal carcinoma
We also analyzed the infiltration difference of 22 kinds of immune cells among different groups by Mann–Whitney U test, and showed the results by grouping comparison chart (Fig. 8B). The results showed that there were statistically significant differences in the infiltration abundance of 16 kinds of immune cells between tumor group and normal group in TCGA-COADREAD data set (P < 0.05). They are B cells naive, dendritic cells resting, eosinophils, macrophages M0, macrophages M1, macrophages M2, mast cells activated, mast cells resting, monocytes, neutrophils, NK cells activated, NK cells resting, plasma cells, T cells CD4 memory activated, T cells CD8, T cells follicular helper.
Then, we calculated the correlation between the infiltration abundance of these 16 kinds of immune cells (B cells naive, dendritic cells resting, eosinophils, macrophages M0, macrophages M1, macrophages M2, mast cells activated, mast cells resting, monocytes, neutrophils, NK cells activated, NK cells resting, plasma cells, T cells CD4 memory activated, T cells CD8, T cells follicular helper) in TCGA-COADREAD data set and displayed the results (Fig. 8C). The results show that in the TCGA-COADREAD data set samples, the infiltration abundance of 17 kinds of immune cells has more negative correlation, among them, NK cells resting and mast cells activated have the highest positive correlation, while mast cells resting and Mast cells activated, macrophages M0, and plasma cells have the highest negative correlation (Fig. 8C).
Analysis of prognostic pyroptosis-related DEGs
The above results show that the expression levels of the four pyroptosis-related prognostic DEGs were closely related to the occurrence of CRC. The expression difference of related prognostic DEGs were further analyzed to reveal the correlation between the expression levels of the pyroptosis-related prognostic DEGs in TCGA-COADREAD dataset, GSE113513 dataset, and CRC grouping. Additionally, we performed a statistical analysis of the clinical information of patients with CRC obtained from the TCGA-COADREAD dataset (Table 5).
We first analyzed the four pyroptosis-related prognostic DEGs in CRC tissues (cancer group) in the TCGA-COADREAD dataset using the Wilcoxon signed-rank test. The results showed that the expression differences between the four pyroptosis-related prognostic DEGs in the TCGA-COADREAD dataset cancer tissue (group: tumor) and normal among colorectal tissues (group: normal) were statistically significant (Fig. 9A): CXCL8 (P < 0.001), IL13RA2 (P < 0.001), MELK (P < 0.001), and POP1 (P < 0.001). We drew ROC curves of the four pyroptosis-related prognostic DEGs in the TCGA-COADREAD dataset and displayed the results (Fig. 9B‒E). It can be seen from the ROC curves in the Fig. 8B and colorectal cancer that the expression of CXCL8 (AUC = 0.896, Fig. 9B) and IL13RA2 (AUC = 0.710, Fig. 9C) among the four genes selected by constructing the LASSO regression model in the TCGA-COADREAD dataset is related to colorectal cancer, and the occurrence of cancer showed a slight correlation, while the expression of MELK (AUC = 0.938, Fig. 9D) and POP1 (AUC = 0.970, Fig. 9E) showed a significant correlation with the occurrence of CRC.
Differential expression analysis of pyroptosis-related prognostic DEGs in TCGA-COADREAD dataset and GSE113513 dataset. A Differential expression analysis of pyroptosis-related prognostic DEGs in the TCGA-COADREAD dataset. Results of grouping comparison chart show. B–E ROC curves of pyroptosis-related prognostic DEGs CXCL8 (B), IL13RA2 (C), MELK (D), POP1 (E) in the TCGA-COADREAD dataset. results of grouping comparison chart show. F Differential expression analysis of pyroptosis-related prognostic DEGs in GSE113513 dataset. G–J ROC curves of pyroptosis-related prognostic DEGs CXCL8 (G), IL13RA2 (H), MELK (I), POP1 (J) in the GSE113513 dataset. P > 0.05, no statistical significance; P < 0.05, statistically significant; P < 0.01, highly statistically significant; P < 0.001, extremely statistically significant. The closer the AUC in the ROC curve was to 1, the better the diagnosis would be. AUC ranged from 0.5 to 0.7 with low accuracy; AUC ranged from 0.7 to 0.9 with some accuracy; High accuracy above 0.9 AUC. TCGA The cancer genome atlas, COADREAD colon and rectal cancer, ROC receiver operating characteristic curve
We performed the same analysis on the expression differences of the four pyroptosis-related prognostic DEGs in the GSE113513 dataset. We first analyzed the four pyroptosis-related prognostic differential expression levels using the Wilcoxon signed-rank test. The expression levels of genes in the CRC tissue samples (cancer group, group: tumor) of the GSE113513 dataset and the corresponding matched normal colorectal tissue samples (normal group, group: normal) were analyzed. The differential analysis results showed that the expression differences of the four pyroptosis-related prognostic DEGs were statistically significant between colorectal cancer tissues (group: tumor) and normal colorectal tissues (group: normal) in the GSE113513 dataset (Fig. 9F): CXCL8 (P < 0.001), IL13RA2 (P = 0.002), MELK (P < 0.001), and POP1 (P < 0.001). We drew the ROC curves of the four pyroptosis-related prognostic DEGs in the GSE113513 dataset and displayed the results (Fig. 9G‒J). The ROC curve results were as follows: the expression of IL13RA2 (AUC = 0.827, Fig. 9H) in the GSE113513 dataset showed a slight correlation with the occurrence of CRC, while CXCL8 (AUC = 0.954, Fig. 9G), MELK (AUC = 0.944, Fig. 9I), and POP1 (AUC = 0.995, Fig. 9J) were significantly correlated with the occurrence of CRC. This indicated that the expression of the four pyroptosis-related prognostic DEGs selected by constructing the LASSO regression model was correlated with the occurrence of CRC.
Prognostic analysis and prognostic performance of pyroptosis-related prognostic DEGs
We performed prognostic analysis on the LASSO regression model constructed with the four pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, MELK, and POP1), with P < 0.05, as the standard; the related molecules were considered statistically significant, and we performed LASSO regression. The prognostic survival KM curve of the model in the TCGA-COADREAD dataset show that the constructed LASSO regression model has significant statistical significance in the prognosis and survival prediction of patients with CRC (P = 0.026, Fig. 10A).
Prognostic analysis of DEGs related to pyroptosis. A KM curve for prognostic analysis of LASSO regression model of pyroptosis-related prognostic DEGs in the TCGA-COADREAD data set. B–E Prognostic analysis KM curve of pyroptosis-related prognostic DEGs CXCL8 (B), IL13RA2 (C), MELK (D), POP1 (E) in the TCGA-COADREAD data set. P > 0.05, no statistical significance; P < 0.05, statistically significant; P < 0.01, highly statistically significant; P < 0.001, extremely statistically significant. TCGA The cancer genome atlas, COADREAD colon and rectal cancer, LASSO least absolute shrinkage and selection operator, KM curve Kaplan–Meier curve
We then drew the KM curve of prognosis and survival for the four pyroptosis-related prognostic DEGs, and took P < 0.05, as the standard to consider the related molecules to be statistically significant, and obtained three that met the requirements. Of the prognostic DEGs, three were associated with pyroptosis (Fig. 10B‒E), these genes were CXCL8 (P = 0.011, Fig. 10B), IL13RA2 (P = 0.018, Fig. 10C), and POP1 (P = 0.026, Fig. 10E). However, the results of the prognostic survival KM curve analysis of MELK (P = 0.059, Fig. 10D) showed that the expression of MELK did not significantly affect the occurrence of CRC in the TCGA-COADREAD dataset.
To further confirm our established LASSO regression prediction model, we used single and multivariate COX regression analysis methods on the TCGA-COADREAD data. Three prognostic DEGs (CXCL8, IL13RA2, and POP1) met the requirements for association with pyroptosis. In addition to the correlation between different clinical stages and prognosis of the tumor, the results showed the expression of CXCL8, IL13RA2, POP1, tumor clinical stage T, clinical N, clinical M, age, and pathologic stage showed clinically significant correlation with the prognosis (Table 6). We organized the results of the univariate and multivariate COX regressions and displayed them in the form of a forest plot (Fig. 11A). We then performed nomogram analysis to determine the prognostic power of the LASSO‒Cox regression model and drew a nomogram (Fig. 11B). The nomogram is based on multi-factor regression analysis by setting a certain scale to characterize the situation of each variable in the multi-factor regression model and finally calculating the total score to predict the probability of the event.
Prognostic performance of pyroptosis-related prognostic DEGs. A–C Forest plot (A), nomogram (B), 1, 3, and 5-year calibration curve plot (C) of univariate and multivariate COX regression analysis of pyroptosis-related prognostic DEGs, in the TCGA-COADREAD data set. D–F: 1-year (D), 3-year (E), 5-year (F) DCA plots of the LASSO-Cox regression prognostic model. TCGA The cancer genome atlas, COADREAD colon and rectal cancer, LASSO Least absolute shrinkage and selection operator, DCA decision curve analysis
In addition, we performed 1-, 3-, and 5-year prognostic calibration analyses on the nomograms in univariate and multivariate COX regression and plotted calibration curves (Fig. 11C, calibration graphs). This was used to evaluate the prediction effect of the model on the actual result by drawing the fitting situation between the actual probability and the probability predicted by the model under different conditions in the figure and is mainly used for the fitting analysis of the model established by the COX regression method and the actual situation. The horizontal axis of the calibration curve represents the survival probability predicted by the model, and the vertical axis represents the survival probability displayed by the actual data. The lines and dots in different colors represent the predictions of the model at different time points. The lines with different colors are closer to the ideal grey line, indicating that the prediction effect is better at this time point.
We then used DCA to assess the role of the constructed LASSO-Cox regression prognostic model in terms of clinical utility at 1- (Fig. 11D), 3- (Fig. 11E), and 5-years (Fig. 11F). The results were displayed (Fig. 11D‒F), and the x-axis in the DCA diagram represents the probability threshold or threshold probability, and the y-axis represents the net benefit. The results can be judged by observing that the line of the model can be stably higher than the x value range of the all-positive and all-negative lines. The larger the x value range, the better the model effect.
Clinical analysis of prognostic pyroptosis-related DEGs
To further determine whether there was a correlation between the expression levels of pyroptosis-related prognostic DEGs and prognostic clinical characteristics of patients in the TCGA-COADREAD dataset, we analyzed pyroptosis-related prognostic factors in CRC tissues. The effect of prognostic DEGs (CXCL8, IL13RA2, and POP1) expression levels on tumor OS, DSS, and PFI (Fig. 12A‒I) was assessed.
Clinical analysis of prognostic DEGs related to pyroptosis. A–C Correlation analysis of pyroptosis-related prognostic differentially expressed gene CXCL8 with clinical OS (A), DSS (B), and PFI (C) in the TCGA-COADREAD data set. D–F Correlation analysis of pyroptosis-related prognostic differentially expressed gene IL13RA2 with clinical OS (D), DSS (E), and PFI (F) in the TCGA-COADREAD data set. G–I Correlation analysis of pyroptosis-related prognostic differentially expressed gene POP1 with clinical OS (G), DSS (H), and PFI (I) in the TCGA-COADREAD data set. P > 0.05, no statistical significance; P < 0.05, statistically significant; P < 0.01, highly statistically significant; P < 0.001, extremely statistically significant. OS overall survival, DSS disease-specific survival, PFI progression-free interval
It could be seen from the results that the expression level of the DEGs CXCL8 in relation to pyroptosis has a statistically significant difference in the expression of the overall survival (OS) of the tumor, with a (P = 0.01, Fig. 12A), while the expression level was not statistically significant for DSS (P = 0.603, Fig. 12B) or PFI (P = 0.78, Fig. 12C) in the tumor group.
The expression level of pyroptosis-related prognostic DEG IL13RA2 was related to the OS of the tumor (P < 0.001, Fig. 12D), and the DSS (P = 0.026, Fig. 12E), which was statistically significant. The P value of the PFI (P = 0.06, Fig. 12F) was greater than 0.05, indicating that the expression level of the gene BHLHE40 had no statistical significance on the tumor PFI.
The expression level of the differentially expressed prognostic gene POP1 related to pyroptosis affected the OS of the tumor (P = 0.028, Fig. 12G) and PFI (P = 0.04, Fig. 12I). Differences in expression were statistically significant. However, the expression of POP1 in the TCGA-COADREAD dataset had no significant effect on the DSS (P = 0.214, Fig. 12H).
Gene mutation analysis of pyroptosis-related prognostic DEGs
The cBioPortal database converts complex genetic, epigenetic, gene expression, and proteomic events in cancer tissues and cell lines into simple genetic and epigenetic events. For the final determined pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, and POP1), we queried the gene mutation sites of the three genes in the TCGA-COADREADcolorectal Cancerdataset through the cBioPortal database and analyzed the results (Fig. 13).
Mutation analysis of DEGs associated with pyroptosis. A Mutation analysis results of pyroptosis-related prognostic DEGs CXCL8, IL13RA2, and POP1 in the TCGA-COADREAD dataset. B–F Pyroptosis-related prognostic DEGs CXCL8 (B), IL13RA2 (C), POP1 (D) gene mutation site analysis results in the TCGA-COADREAD dataset. All data are from the cBioPortal database. TCGA The cancer genome atlas, COADREAD colon and rectal cancer
The results showed (Fig. 13A) that the genetic mutations of the three pyroptosis-related prognostic DEGs in the TCGA-COADREAD dataset samples were mainly divided into six types: (1) Missense mutation (unknown significance), (2) splice mutation (unknown significance), (3) truncating mutations (unknown significance), (4) structural variant (unknown significance), (5) significant amplification (amplification), and (6) Deep Deletion (deep deletion).
There are three main types of mutations in the differentially expressed gene CXCL8 related to pyroptosis: missense mutations (unknown significance), significant amplification, and deep deletion. The total number of mutations in CXCL8 accounts for the total number of samples in the TCGA-COADREAD dataset. 1%, while the mutation types of the pyroptosis-related prognostic differentially expressed gene IL13RA2 mainly include missense mutations (unknown significance), truncating mutations (unknown significance), and significant amplification. The total number of mutations accounted for 1.9% of the total samples in the TCGA-COADREAD dataset; there are 5 types of mutations in the differentially expressed gene POP1 associated with pyroptosis, including missense mutations (unknown significance), splicing mutations (unknown significance), and truncation mutations (unknown significance), structural variants (unknown significance), and significant amplification. These account for 6% of the total samples in the TCGA-COADREAD dataset.
In addition, we analyzed the specific mutation sites of three pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, and POP1) in the TCGA-COADREAD dataset (Fig. 12B‒D). The results showed that the type of post translational modification (PTM) of CXCL8 was citrullination, with a total of two main mutation sites (variant of undetermined significance, VUS: E97D, R87M). In this type of missense mutation, the main function of the mutation site is to cause protein change (protein change), which is distributed in exons 3 and 4 (Fig. 13B).
The PTM type of gene IL13RA2 in the TCGA-COADREAD dataset is N-linked glycosylation, with a total of 10 major mutation sites (VUS: *381Rext*23, F376L, G360C, F344L, R343H, D209Y, H106R, A103V, D93Y, R74Q). The mutation type of *381Rext*23 mutation site is a truncating mutation, and the mutation types of other mutation sites are missense mutations. The main function of the site is to cause protein changes that are distributed in exons 3, 4, 6, 9, and 10. The topological regions involved corresponded to three parts: cytoplasmic, transmembrane, and extracellular (Fig. 13C).
The types of PTMs of POP1 include phosphorylation, acetylation, ubiquitination, methylation, and glutathionylation. A total of 23 major mutation sites (VUS: T752Lfs*12, S371Kfs*4, POP1-ERICH5, E92K, L580R, R241W, R513Q, S801N, E378K, R241Q, R954H, L276P, G820E, R55Q, K313E, H138R, R465H, S865I, D902G, G401S, R141*, W818*, X807_splice), where the mutation type of the X807 splice mutation site is splice mutation, the mutation type of the POP1-ERICH5 mutation site is fusion mutation, as well as for the mutation site R141. The mutation type of *, W818*, T752Lfs*12, S371Kfs*4 is truncating mutation, and the mutation type of the other 17 mutation sites are missense mutations. The main function of the mutation site is to cause protein changes were distributed in the region of exon 3–16 (Fig. 13D).
Analysis of expression distribution of pyroptosis-related prognostic DEGs and single cell analysis
In addition, we analyzed the distribution of RNA and protein expression of differentially expressed genes related prognostic DEGs (CXCL8, IL13RA2, POP1) in HPA database in human as well as the expression in colonic and rectal tissues. The results showed that the significant upregulation of CXCL8 was found to be characteristic of bone marrow and lymphoid tissue. In addition, the protein encoded by CXCL8 was expressed in multiple human tissues such as stomach, kidney, and male tissue with high content distribution (Fig. 14A). We also analyzed the correlation between the expression of CXCL8 and tissue cell type in human colon tissue and rectal tissue (Fig. 14B, C). The results showed that the expression of CXCL8 in human colon tissue was most significantly correlated with c-12 B-cells (Fig. 14B). The expression of CXCL8 in human rectal tissue was also correlated with many cells, but it was most significantly correlated with C-11 Entero Endocrine cells (Fig. 14C).
Analysis of expression distribution of pyroptosis-related prognostic DEGs CXCL8 and IL13RA2 and single cell analysis. A mRNA and protein expression of CXCL8, a differentially expressed gene related to pyroptosis, in normal human body tissues. B, C Display of results from single-gene analysis of CXCL8, a differentially expressed gene related to apoptosis prognosis in the HPA database, in colon (B) and rectum (C) tissues. D mRNA and protein expression of IL13RA2, a differentially expressed gene related to apoptosis, in normal human body tissues. E, F Single gene analysis of IL13RA2, a differentially expressed gene related to cell scorch in HPA database, in colon (E) and rectum (F) tissues. All data are from The Human Protein Altas database
The significant upregulation of the IL13RA2 was found to be characteristic of Male tissues testis. In addition, there is currently no clear information on the distribution of the protein expression encoded by the IL13RA2 in human tissues (Fig. 13D). We also analyzed the correlation between the expression of IL13RA2 and tissue cell type in human colon tissue and rectal tissue (Fig. 13E, F). The results showed that the expression of IL13RA2 in human colon tissue was significantly related to several cells such as C-14 Entero Endocrine cells (Fig. 13E), while the expression of IL13RA2 in human rectal tissue was only related to C-11 Entero Endocrine cells (Fig. 14F).
The POP1 is expressed in many tissues of the human body, such as the gastrointestinal tract. In addition, there is no clear information about the distribution of the expression of the protein encoded by the POP1 in human tissues (Fig. 15A). We also analyzed the correlation between the expression of POP1 in human colon tissue and rectal tissue and tissue cell type (Fig. 15B, C), and the results showed that the expression of POP1 in human colon tissue was correlated with multiple cells. In addition, the correlation with C-5 distinct enterocytes and C-8 undivided cells was more significant (Fig. 14B), while the expression of POP1 in human rectal tissue had a certain correlation with many cells, but none of them was particularly significant (Fig. 15C).
Analysis of expression distribution of pyroptosis-related prognostic DEGs POP1 and single cell analysis. A mRNA and protein expression of POP1, a differentially expressed gene related to pyroptosis, in normal human body tissues. B, C Display of results from single-gene analysis of POP1, a differentially expressed gene related to apoptosis prognosis in the HPA database, in colon (B) and rectum (C) tissues. All data are from The Human Protein Altas database. D–F Cell line analysis of differentially expressed of pyroptosis-related prognostic DEGs genes CXCL8, IL13RA2, and POP1 in homotissues and organs of normal human body
Finally, we also analyzed differential expression genes pyroptosis-related prognostic DEGs (CXCL8, IL13RA2, and POP1) in different tissues and organs of the human body and their corresponding cell lines (Fig. 15D–F). The results showed that CXCL8, one of the pyroptosis-related prognostic DEGs, was significantly expressed in BJ hTERT + cell line in mesenchymal. Second, there is low expression in GAMG cell lines in Brain (Fig. 15D). IL13RA2 was significantly expressed in both Mesenchymal cell lines and Brain cell lines (Fig. 15E). POP1 was significantly expressed in various cell lines of Brain, Mesenchymal, Lymphoid and bone marrow (Fig. 15F).
Discussion
CRC is one of the most common malignant tumors that is characterized by a high recurrence rate and poor prognosis, particularly in developed countries. It is the third most common cancer among males and ranks second among females (Kraus et al. 2014; Ferlay et al. 2010). Although great progress has been made in terms of treatment and diagnosis of CRC, its mortality and morbidity remain high. In particular, the age of patients with CRC are prominently becoming younger, and the early diagnosis and prognosis of CRC should be improved (Zhang et al. 2020c). In recent years, several indicators, such as age, sex, and pathological stage, have appeared and at present, the imaging and serum markers of CRC are the main basis for judging its prognosis. However, owing to individual differences, the prediction of improved treatment effect and prognosis by the above factors alone is often limited. With the rapid development of gene sequencing technology, multiple gene models can be constructed based on the expression characteristics of key regulators of the same signaling pathway, which can improve the prediction accuracy and explore new targeted therapies. Currently, multiple biomarkers are used to predict the prognosis of CRC (Akagi et al. 2013). Targeting pyroptosis can be used as an effective antitumor drug and is expected to become a new treatment method (Wu et al. 2020). However, current research on tumor markers cannot be fully adapted to the diagnosis and prognosis of CRC. Therefore, identifying new biomarkers for CRC is crucial (Yang et al. 2022). Studies have shown that, in a variety of tumors, an increasing number of genes are associated with pyroptosis (Du et al. 2022), but it is unknown if there is a link between genes related to pyroptosis and the prognosis of patients with CRC. This study aimed to understand the effects of PRGs on the prognosis of CRC. Patients with CRC were successfully stratified and predicted based on GEO and TCGA databases. In addition, we confirmed that many immune cells and pathways are significantly different in patients with different risk levels; this can be used as a new method for predicting CRC immunotherapy. CXCL8, IL13RA2, and MELK were selected as prognostic genes. POP1 is a prognostic gene that showed a better prognosis in GEO and TCGA. Genetic characteristics have been shown to be independent prognostic factors of CRC.
There were five pyroptosis-related DEGs in the PPI network: CTSG, CXCL8, CHI3L1, IL13RA2, and GZMB which are related to other genes. The mRNA‒RBP interaction network consisted of 7 mRNAs. DEGs related to pyroptosis: BHLHE40, PCSK9, CXCL8, MELK, POP1, CHI3L1, and DPEP1. The pyroptosis-related DEG BHLHE40 had the most interaction relationship with TFs in the mRNA-TF interaction network. C‒X‒C motif chemokine 8 (CXCL8), also known as interleukin 8 (IL-8), is primarily derived from macrophages. In addition, it plays an important role in the inflammatory response and chemotaxis of neutrophils (Ha et al. 2017). At present, the relationship between the CXCL8 gene and tumor biology is still debated, and a study by Do HTT et al. (Do et al. 2020) showed that overexpression of CXCL8 can promote the proliferation, migration, and invasion of CRC cells. It is also associated with CRC angiogenesis, metastasis, poor prognosis, and asymptomatic survival, among other factors. The other studies (Wang et al. 2017; Li et al. 2021) showed that high expression of CXCL8 can prevent CRC liver metastasis, thereby contributing to better survival in patients with CRC, and provide a better prognosis. GO results showed that 12 pyroptosis-related DEGs were mainly enriched in neutrophil-mediated cytotoxicity, leukocyte-mediated cytotoxicity, receptor internalization, antimicrobial humoral response, and other BPs in CRC; as well as secretory granule lumen, cytoplasmic vesicle lumen, vesicle lumen, external side of plasma membrane, and other CCs; and were enriched in endopeptidase activity, serine-type endopeptidase activity, serine-type peptidase activity, serine hydrolase activity, and other MFs. The GSEA results indicated that 180 functional pathways, including Reactome chromosome maintenance, Reactome meiotic recombination, and Reactome condensation of prophase chromosomes, were significantly enriched by both datasets simultaneously. Interestingly, this finding has not been previously reported. The DEGs in the dataset GSE113513 were significantly enriched in the Reactome cell cycle checkpoints and Reactome mitotic spindle checkpoints. A previous study (Grady 2004) showed that the chromosomal region (CIN) was acquired and lost in most patients with CRC and caused different types of gene changes, thus causing tumorigenesis. CIN is mainly caused by abnormalities in DNA replication and spindle checkpoints. The DEGs were significantly enriched in TCGA-COADREAD during DNA methylation. A study by Rui Yang et al. on eight patients (Yang et al. 2019) showed that DNA methylation plays an important role in the formation of tumor responses and the observation of CD8+ tumor infiltrating lymphocytes.
The expression levels of the four pyroptosis-related prognostic DEGs were closely associated with the occurrence of CRC. Meanwhile, the ROC curve showed that the expression of MELK and POP1 was significantly correlated with the occurrence of colorectal cancer in both the TCGA-COADREAD and GSE113513 datasets. In addition, the results of analyzing the differences in immune infiltration showed that the expression of CXCL8 and IL13RA2 in the TCGA-COADREAD dataset was positively correlated with the significant differential enrichment of most immune cells, while the expression of MELK and POP1 in the TCGA-COADREAD dataset was positively correlated with the significant differential enrichment of most immune cells was negatively correlated. Liu et al. (Liu et al. 2020) believed that MELK accelerates the progression of CRC by activating the FAK/Src pathway, and Fan et al. (Fan et al. 2020) considered POP1 to play an important role in the pathogenesis of CRC and had prognostic value. After drawing the prognosis survival KM curve individually, it was found that CXCL8, IL13RA2, and POP1 were the DEGs related to pyroptosis that met the threshold requirements.
Their expression levels, as well as tumor clinical T, N, and M stage, as well as age and pathological stage, are significantly correlated with prognosis. Our analysis showed that high expression of CXCL8 or POP1 can contribute to better survival in patients with CRC and provide a better prognosis. The predicted and actual results were in agreement. The level of CXCL8 expression showed a statistically significant difference in tumor OS, and the level of POP1 expression showed a statistically significant difference in tumor OS and PFI. “Pyrin-only” 1 (POP1, POPDC1, and BVES) is a protein that can regulate the formation of tight junctions between cells and prevent the occurrence of epithelial-mesenchymal transition (EMT), and through its epigenetic silencing, can promote the occurrence of EMT. Liu et al. (Liu et al. 2021) considered POP1 to be an oncogene in breast cancer. C‒X‒C motif ligand 8 (CXCL8) is a cytokine with multiple functions that can regulate tumor proliferation, invasion, and migration in a paracrine manner. The interaction between CXCL8 and CXCR1/2 in the tumor microenvironment is key to tumor development and metastasis. The regulatory role of the CXCL8‒CXCR1/2 axis is involved in tumorigenesis and metastasis (Ha et al. 2017).
SsGESA is an extension of the GSEA method, which calculates the enrichment score for each sample and gene set pair. Each ssGSEA enrichment score represented the degree to which members of a particular gene set in the sample were coordinated upregulated or downregulated. SsGSEA transformed the gene expression profiles of a single sample into a gene set enrichment profile. This transformation enables researchers to describe the cell state based on the level of activity of biological processes and pathways rather than by the expression level of individual genes. Therefore, ssGESA can calculate the immune cell infiltration score if it uses the gene set related to the immune cell marker. The results of ssGSEA analysis showed that the expression levels of CXCL8 and IL13RA2 in the TCGA-COADREAD data set were positively correlated with significant differential enrichment in most immune cells. The expression of MELK and POP1 in the TCGA-COADREAD data set was negatively correlated with significant differential enrichment in most immune cells.
TIMER2.0 (Barbie et al. (n.d.)) is an immune infiltrate used for the systematic analysis of different types of cancer. Various immune deconvolution methods are provided to estimate the abundance of immune infiltration and to fully explore the immunological, clinical and genomic features of the tumor. The characteristic genes were identified separately for each cancer type by selecting genes negatively correlated with tumor purity from immune cell markers. It could not be directly interpreted as a cellular component or compared between different immune cell types and data sets. Due to the upgrade and revision of TIMER database, immune infiltration analysis related to immune cells cannot be performed at present. However, in TIMER2.0 database, we found eight differentially expressed genes related to cell apoptosis (BHLHE40, CHI3L1, CASP5, CTSG, GZMB, MPEG1, POP1, MELK) analyzed their correlations in the COAD and READ tumor data sets: CTSG and MPEG1 were moderately strongly correlated, BHLHE40 and MPEG1, CHI3L1 and GZMB, GZMB and MPEG1 were not correlated.
Our study had certain limitations. First, the number of CRC samples and clinical data are limited. A single microarray analysis results in a high false-positive rate and has a one-sided bias effect. Therefore, multisample data will be further integrated to improve the detection capability of the detector ii the model. Second, to confirm the predictive model, a large body of evidence must be collected from multiple research institutions. Further clinical and population data of patients with CRC await further analysis. Third, clinical, cellular, and animal functional tests are lacking; therefore, the reliability of the data analysis needs to be further tested. PCR, Western blotting, and immunohistochemistry are necessary to fully understand the function and possible mechanism of CRC.
Conclusion
In summary, from the GEO (GSE113513) and TCGA-COADREAD CRC datasets, a total of 12 CRC‒PRG‒DEGs were found, that is, in relation with CRC. The PRGs signature proposed in this paper has excellent characteristics and deserves further in-depth research and long-term use. However, large prospective studies are needed to determine the prognostic value of CRC‒PRG-DEGs, and further experimental validation should be performed to demonstrate the biological role of CRC‒PRG‒DEGs in CRC.
Data availability
Publicly available datasets were analyzed in this study. This data can be found here: GSE113513 from the GEO and TCGA-COADREAD.
Abbreviations
- CRC:
-
Colorectal cancer
- PRGs:
-
Pyroptosis-related genes
- GEO:
-
Gene Expression Omnibus
- DEGs:
-
Differentially expressed genes
- CRC-PRGs:
-
CRC-pyroptosis-related genes
- GO:
-
Gene Ontology
- BP:
-
Biological process
- MF:
-
Molecular function
- CC:
-
Cellular component
- GSEA:
-
Gene set enrichment analysis
- BH:
-
Benjamini-Hochberg
- MSigDB:
-
Molecular Signatures Database
- GSVA:
-
Gene set variation analysis
- ssGSEA:
-
Single-sample gene-set enrichment analysis
- PPI:
-
Protein–protein interaction
- RBPs:
-
RNA binding proteins
- TFs:
-
Transcription factors
- DGIdb:
-
Drug-gene interaction database
- ROC:
-
Receiver operating characteristic curve
- AUC:
-
Area under the curve
- DCA:
-
Decision curve analysis
- TNM:
-
Tumor-lymph node-metastasis
- OS:
-
Including overall survival
- DSS:
-
Disease-specific survival
- PFI:
-
Progression-free Interval progress-free interval
- HPA:
-
Human Protein Atlas
- READ:
-
Rectal cancer data set
- COAD:
-
Colon cancer dataset
- KM:
-
Kaplan‒Meier
- Tfh:
-
T follicular helper
- CXCL8:
-
C-X-C motif chemokine 8
- PTM:
-
Post-translational modification
- IL-8:
-
Interleukin 8
- CIN:
-
Chromosomal regions
- CXCR1/2:
-
C-X-C chemokine receptor 1/2
References
Akagi Y, Kinugasa T, Adachi Y, Shirouzu K (2013) Prognostic significance of isolated tumor cells in patients with colorectal cancer in recent 10-year studies. Mol Clin Oncol 1(4):582–592. https://doi.org/10.3892/mco.2013.116
Bao M, Zhang L, Hu Y (2020) Novel gene signatures for prognosis prediction in ovarian cancer. J Cell Mol Med 24(17):9972–9984. https://doi.org/10.1111/jcmm.15601
Barbie DA, Tamayo P Fau - Boehm JS, Boehm Js Fau - Kim SY, et al. (n.d.) Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. (1476–4687 (Electronic)). https://doi.org/10.1038/nature08460
Barrett T, Troup DB, Wilhite SE et al (2007) NCBI GEO: mining tens of millions of expression profiles–database and tools update. Nucleic Acids Res 35(Database issue):D760-765. https://doi.org/10.1093/nar/gkl887
Charoentong P, Finotello F, Angelova M et al (2017) Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18(1):248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Chen Y, Wang X (2020) miRDB: an online database for prediction of functional microRNA targets. Nucleic Acids Res 48(D1):D127–D131. https://doi.org/10.1093/nar/gkz757
Colaprico A, Silva TC, Olsen C et al (2016) TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res 44(8):e71. https://doi.org/10.1093/nar/gkv1507
Davis S, Meltzer PS (2007) GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23(14):1846–1847. https://doi.org/10.1093/bioinformatics/btm254
Do HTT, Lee CH, Cho J (2020) Chemokines and their receptors: multifaceted roles in cancer progression and potential value as cancer prognostic markers. Cancers (Basel) 12(2)·https://doi.org/10.3390/cancers12020287
Du W, Miao Y, Zhang G et al (2022) The regulatory role of neuropeptide gene glucagon in colorectal cancer: a comprehensive bioinformatic analysis. Dis Markers 2022(1875–8630 (Electronic)):4262600. https://doi.org/10.1155/2022/4262600
Fan X, Liu L, Shi Y et al (2020) Integrated analysis of RNA-binding proteins in human colorectal cancer. World J Surg Oncol 18(1):222. https://doi.org/10.1186/s12957-020-01995-5
Fearon ER, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. Cell 61(5):759–767. https://doi.org/10.1016/0092-8674(90)90186-i
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM (2010) Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 127(12):2893–2917. https://doi.org/10.1002/ijc.25516
Freshour SL, Kiwala S, Cotto KC et al (2021) Integration of the drug-gene interaction database (DGIdb 4.0) with open crowdsource efforts. Nucleic Acids Res 49(D1):D1144–D1151. https://doi.org/10.1093/nar/gkaa1084
Goldman MJ, Craft B, Hastie M et al (2020) Visualizing and interpreting cancer genomics data via the Xena platform. Nat Biotechnol 38(6):675–678. https://doi.org/10.1038/s41587-020-0546-8
Grady WM (2004) Genomic instability and colon cancer. Cancer Metastasis Rev 23(1–2):11–27. https://doi.org/10.1023/a:1025861527711
Ha H, Debnath B, Neamati N (2017) Role of the CXCL8-CXCR1/2 axis in cancer and inflammatory diseases. Theranostics 7(6):1543–1588. https://doi.org/10.7150/thno.15625
Hanzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14(1471–2105 (Electronic)):7. https://doi.org/10.1186/1471-2105-14-7
Hegde M, Ferber M, Mao R et al (2014) ACMG technical standards and guidelines for genetic testing for inherited colorectal cancer (Lynch syndrome, familial adenomatous polyposis, and MYH-associated polyposis). Genet Med 16(1):101–116. https://doi.org/10.1038/gim.2013.166
Kraus S, Nabiochtchikov I, Shapira S, Arber N (2014) Recent advances in personalized colorectal cancer research. Cancer Lett 347(1):15–21. https://doi.org/10.1016/j.canlet.2014.01.025
Li JH, Liu S, Zhou H, Qu LH, Yang JH (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42(Database issue):D92-97. https://doi.org/10.1093/nar/gkt1248
Li E, Yang X, Du Y et al (2021) CXCL8 associated dendritic cell activation marker expression and recruitment as indicators of favorable outcomes in colorectal cancer. Front Immunol 12(1664–3224(Electronic)):667177. https://doi.org/10.3389/fimmu.2021.667177
Li T, Fu J, Zeng Z, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells.(1362–4962 (Electronic))·https://doi.org/10.1038/nature08460. Epub 2009 Oct 21.
Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P (2015) The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst 1(6):417–425. https://doi.org/10.1016/j.cels.2015.12.004
Liu G, Zhan W, Guo W et al (2020) MELK accelerates the progression of colorectal cancer via activating the FAK/Src pathway. Biochem Genet 58(5):771–782. https://doi.org/10.1007/s10528-020-09974-x
Liu Y, Sun H, Li X et al (2021) Identification of a three-RNA binding proteins (RBPs) signature predicting prognosis for breast cancer. Front Oncol 11(2234–943 X (Print)):663556. https://doi.org/10.3389/fonc.2021.663556
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Mandrekar JN (2010) Receiver operating characteristic curve in diagnostic test assessment. J Thorac Oncol 5(9):1315–1316. https://doi.org/10.1097/JTO.0b013e3181ec173d
Newman AM, Liu CL, Green MA-O, et al. Robust enumeration of cell subsets from tissue expression profiles.(1548–7105 (Electronic))·https://doi.org/10.1038/nmeth.3337
Ritchie ME, Phipson B, Wu D et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47. https://doi.org/10.1093/nar/gkv007
Shen A, Liu L, Huang Y et al (2021) Down-regulating HAUS6 suppresses cell proliferation by activating the p53/p21 pathway in colorectal cancer. Front Cell Dev Biol 9(2296–634 X (Print)):772077. https://doi.org/10.3389/fcell.2021.772077
Shi J, Gao W, Shao F (2017) Pyroptosis: gasdermin-mediated programmed necrotic cell death. Trends Biochem Sci 42(4):245–254. https://doi.org/10.1016/j.tibs.2016.10.004
Siegel RL, Miller KD, Goding Sauer A et al (2020) Colorectal cancer statistics 2020. CA Cancer J Clin. 70(3):145–164. https://doi.org/10.3322/caac.21601
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer Statistics 2021. CA Cancer J Clin 71(1):7–33. https://doi.org/10.3322/caac.21654
Stelzer G, Rosen N, Plaschkes I, et al. (2016) The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 54(1934–340X (Electronic)): 1 30 31–31 30 33·https://doi.org/10.1002/cpbi.5
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Tataranni T, Piccoli C (2019) Dichloroacetate (DCA) and cancer: an overview towards clinical applications. Oxid Med Cell Longev 2019(1942–0994 (Electronic)):8201079. https://doi.org/10.1155/2019/8201079
Thul PJ, Lindskog C (2018) The human protein atlas: a spatial map of the human proteome. Protein Sci 27(1):233–244. https://doi.org/10.1002/pro.3307
Tian W, Wang Z, Tang NN et al (2020) Ascorbic acid sensitizes colorectal carcinoma to the cytotoxicity of arsenic trioxide via promoting reactive oxygen species-dependent apoptosis and pyroptosis. Front Pharmacol 11(1663–9812 (Print)):123. https://doi.org/10.3389/fphar.2020.00123
Wang S, Zhang C, Zhang Z et al (2017) Transcriptome analysis in primary colorectal cancer tissues from patients with and without liver metastases using next-generation sequencing. Cancer Med 6(8):1976–1987. https://doi.org/10.1002/cam4.1147
Wu LS, Liu Y, Wang XW et al (2020) LPS enhances the chemosensitivity of oxaliplatin in HT29 cells via GSDMD-mediated pyroptosis. Cancer Manag Res 12(1179–1322 (Print)):10397–10409. https://doi.org/10.2147/CMAR.S244374
Xu D, Ji Z, Qiang L (2021) Molecular characteristics, clinical implication, and cancer immunity interactions of pyroptosis-related genes in breast cancer. Front Med (Lausanne) 8(2296–858 X (Print)):702638. https://doi.org/10.3389/fmed.2021.702638
Xue Y, Li J, Lu X (2020) A novel immune-related prognostic signature for thyroid carcinoma. Technol Cancer Res Treat 19(1533–0338 (Electronic)): 1533033820935860·https://doi.org/10.1177/1533033820935860
Yang R, Cheng S, Luo N et al (2019) Distinct epigenetic features of tumor-reactive CD8+ T cells in colorectal cancer patients revealed by genome-wide DNA methylation analysis. Genome Biol 21(1):2. https://doi.org/10.1186/s13059-019-1921-y
Yang Y, Yu J, Hu J et al (2022) A systematic and comprehensive analysis of colorectal squamous cell carcinoma: Implication for diagnosis and treatment. Cancer Med 11(12):2492–2502. https://doi.org/10.1002/cam4.4616
Yu G (2020) Gene ontology semantic similarity analysis using GOSemSim. Methods Mol Biol 2117(1940–6029 (Electronic)):207–215. https://doi.org/10.1007/978-1-0716-0301-7_11
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Yu J, Li S, Qi J et al (2019) Cleavage of GSDME by caspase-3 determines lobaplatin-induced pyroptosis in colon cancer cells. Cell Death Dis 10(3):193. https://doi.org/10.1038/s41419-019-1441-4
Zhang Q, Wang J, Liu M et al (2020) Weighted correlation gene network analysis reveals a new stemness index-related survival model for prognostic prediction in hepatocellular carcinoma. Aging (Albany NY) 12(13):13502–13517. https://doi.org/10.18632/aging.103454
Zhang Q, Liu W, Zhang HM et al (2020) hTFtarget: a comprehensive database for regulations of Human Transcription Factors and Their Targets. Genomics Proteomics Bioinformatics 18(2):120–128. https://doi.org/10.1016/j.gpb.2019.09.006
Zhang Y, Liu X, Xu M, Chen K, Li S, Guan G (2020) Prognostic value of pretreatment systemic inflammatory markers in patients with locally advanced rectal cancer following neoadjuvant chemoradiotherapy. Sci Rep 10(1):8017. https://doi.org/10.1038/s41598-020-64684-z
Zhou KR, Liu S, Sun WJ et al (2017) ChIPBase v2.0: decoding transcriptional regulatory networks of non-coding RNAs and protein-coding genes from ChIP-seq data. Nucleic Acids Res 45(D1):D43–D50. https://doi.org/10.1093/nar/gkw965
Zhuang Z, Cai H, Lin H et al (2021) Development and validation of a robust pyroptosis-related signature for predicting prognosis and immune status in patients with colon cancer. J Oncol 2021:5818512. https://doi.org/10.1155/2021/5818512
Funding
The authors declare that no grants were involved in supporting this work.
Author information
Authors and Affiliations
Contributions
RBL performed the literature search, and GL conceived and designed the project. RBL and SYZ performed the data analysis. RBL wrote the paper. GL reviewed and amended the manuscript. The manuscript has been read and approved by all authors.
Corresponding author
Ethics declarations
Ethics approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, R., Zhang, S. & Liu, G. Identification and validation of a pyroptosis-related prognostic model for colorectal cancer. Funct Integr Genomics 23, 21 (2023). https://doi.org/10.1007/s10142-022-00935-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-022-00935-8