Abstract
The transfer of plastid DNA sequences into plant nuclear genomes plays an important role in the genomic evolution of plants. The abundance of nuclear-localized plastid DNA (nupDNA) correlates positively with nuclear genome size, but the genetic content of nupDNA remains unknown. In this mini review, we analyzed the number of nuclear-localized plastid gene fragments in known plant genomic data. Our analysis suggests that nupDNAs are abundant in plant nuclear genomes and can include multiple complete copies of protein-coding plastid genes. Mutated nuclear copies of plastid genes contained synonymous and nonsynonymous substitutions. We estimated the age of the nupDNAs based on the time when each integration occurred, which was calculated by comparing the nucleotide substitution rates of the nupDNAs and their respective plastid genes. These data suggest that there are two distinct age distribution patterns for nupDNAs in plants, and Oryza sativa and Zea mays were found to contain a very high proportion of young nupDNAs. Expressed sequence tags and predicted promoters of nupDNAs were identified, revealing that certain nuclear-localized plastid genes may be functional and that some have undergone positive natural selection pressure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The plant nuclear genomes acquired numerous DNA fragments from chloroplasts, which played an important role in the genomic of plants, and as a result, the majority of genes encoding chloroplast proteins reside in plant nuclear genomes (Baldauf and Palmer 1990; Gantt et al. 1991; Martin and Herrmann 1998; Rujan and Martin 2001; Martin et al. 2002). Although the transfer of most of these DNA fragments occurred at an early stage in organelle evolution, functional gene transfer events continue to occur in flowering plants (Martin et al. 1998; Adams et al. 1999, 2002; Millen et al. 2001).
In many eukaryotes, DNA transfer from organelles to the nuclear genome is ongoing (Ayliffe et al. 1998; Bensasson et al. 2001; Woischnik and Moraes 2002; Yuan et al. 2002; Huang et al. 2003, 2004; Stegemann et al. 2003). The transfer rate of chloroplast DNAs to the nuclear genome of tobacco has been measured using specific marker genes that were functional only when integrated into the nuclear genome (Huang et al. 2003; Stegemann et al. 2003). Gene transfer events were found to occur more often than that detected under experimental conditions (Martin 2003). The nuclear-localized plastid DNAs (nupDNAs) also tended to be located in close proximity. Once plastid DNAs become integrated into the rice nuclear genome, they are rapidly fragmented and shuffled, and newly integrated nupDNAs tend to be eliminated rapidly. Large nupDNA fragments preferentially localize at the pericentromeric regions of chromosomes, where integration and elimination frequencies are markedly higher (Matsuo et al. 2005; Noutsos et al. 2005; Sheppard and Timmis 2009). The greatest number of chloroplast DNA insertions occurs at nuclear regions characterized by sharp changes in repetitive sequence density (Guo et al. 2008). The abundance and composition of organellar DNA fragments have been investigated in model plants, such as Arabidopsis thaliana and rice (Martin et al. 2002; Shahmuradov et al. 2003). Compared with the small Arabidopsis genome, the rice nuclear genome is essentially saturated with plastid DNA sequences, and the abundance of nupDNAs correlates positively, on average, with nuclear genome size (Smith et al. 2011). The density and pattern of nupDNA integration events have been investigated in several species, and the mechanisms of integration and genomic organization have been analyzed in detail (Timmis et al. 2004; Kleine et al. 2009). The present genomic constitutions of nupDNAs could be explained by the combination of rapidly eliminated deleterious fragments and a few less deleterious, but more stable, fragments (Yoshida et al. 2013). However, the abundance, age, and predicted promoters of plastid genes in the nuclear genome of most plant species have not been thoroughly investigated.
In this review, we evaluated the abundance and age of nupDNAs in genomic data of 23 plant species (Arabidopsis Genome Initiative 2000; Nishiyama et al. 2003; Shrager et al. 2003; Project IRGS 2005; Tuskan et al. 2006; Jaillon et al. 2007; Ming et al. 2008; Huang et al. 2009; Paterson et al. 2009; Schnable et al. 2009; Schmutz et al. 2010; Shulaev et al. 2010; Vogel et al. 2010; Argout et al. 2011; Banks et al. 2011; Potato Genome Sequencing Consortium 2011; Young et al. 2011a; Prochnik et al. 2012; Tomato Genome Consortium 2012; Xu et al. 2013). The analysis shows that significant differences in the composition of nupDNAs exist, and that based on their age, there are two distinct distribution patterns for nupDNAs in plants. Expressed sequence tags (ESTs) indicated that certain nupDNAs may be functional. An analysis of predicted promoters of nupDNAs revealed that some were shuffled and some were eliminated. This review also reveals that the relationship between transcription output and the efficiency of nupDNA gene promoters needs to be further investigated.
Analytical approach
The plastid genome sequences of the following species were obtained from GenBank: A. thaliana (GenBank NC_000932), Brachypodium distachyon (NC_011032), Carica papaya (NC_010323), Chlamydomonas reinhardtii (NC_005353), Cucumis sativus (NC_007144), Citrus sinensis (NC_008334), Eucalyptus grandis (NC_014570), Fragaria vesca (NC_015206), Glycine max (NC_007942), Manihot esculenta (NC_010433), Medicago truncatula (NC_003119), Oryza sativa Japonica group (NC_001320), Physcomitrella patens (NC_005087), Populus trichocarpa (NC_009143), Panicum virgatum (NC_015990), Phaseolus vulgaris (NC_009259), Sorghum bicolor (NC_008602), Solanum lycopersicum (NC_007898), Selaginella moellendorffii (NC_013086), Solanum tuberosum (NC_008096), Theobroma cacao (NC_014676), Vitis vinifera (NC_007957), and Zea mays (NC_001666) (Hiratsuka et al. 1989; Maier et al. 1995; Sato et al. 1999; Maul et al. 2002; Sugiura et al. 2003; Gargano et al. 2005; Saski et al. 2005; Bausher et al. 2006; Jansen et al. 2006; Kahlau et al. 2006; Tuskan et al. 2006; Guo et al. 2007; Pląder et al. 2007; Saski et al. 2007; Bortiri et al. 2008; Daniell et al. 2008; Smith 2009; Shulaev et al. 2010; Paiva et al. 2011; Young et al. 2011a, b).
Pairwise comparisons of plastid genes and nuclear DNA sequences were performed using a BLAST program (http://www.phytozome.net; Goodstein et al. 2012). The number (K) of substitutions per nucleotide site between each of nupDNAs and chloroplast genes was calculated based on the BLAST alignment (Matsuo et al. 2005; Yoshida et al. 2013). For every plastid and nupDNA gene, 1 kb upstream of the translation start site was considered as promoter sequence. Promoters were detected using the TSSP program (http://softberry.com). We used the BLAST, the nupDNA fragments, and plastid DNA that were searched against the Expressed Sequence Tags (EST) Database (http://www.ncbi.nlm.nih.gov/) with no mismatch to identify whether ESTs are derived from nupDNA or plastid genome.
Plastid genes are abundant in plant nuclear genomes
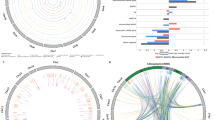
To evaluate the abundance of plastid genes in plant nuclear genomes, we used the plastid genes as the query when searching plant nuclear genome databases (http://www.phytozome.net). BLASTN identified many complete and partial gene sequences with high levels of sequence identity. Matches with E values lower than 10−10 were defined as nupDNA fragments. The sequences were related to photosynthesis, energy metabolism, fatty acid metabolism, transporters, cellular processes, and biosynthesis of cofactors. This analysis was applied to nupDNA fragments larger than 50 bp because it was difficult to confirm the origin of smaller fragments. The analysis revealed that the number of nupDNAs varies among plant species (Fig. 1 and Online Resource 1).
Distribution of plant nuclear-localized plastid DNA (nupDNA) and the presence or absence of complete or partial nuclear homologs (Online Resource 1). The black, gray, and white boxes indicate the nupDNA of intact coding DNA sequences (CDSs), truncated CDSs, and CDSs, respectively, with only partial nuclear homologs. a Genes related to photosynthesis; b genes related to energy metabolism; c genes related to fatty acid metabolism, transporters, cellular processes, and the biosynthesis of cofactors
Previous work determined that plants with relatively large genomes contain more nupDNAs than those with smaller genomes (Shahmuradov et al. 2003; Smith et al. 2011), and we obtained similar results in our present analysis. F. vesca, G. max, O. sativa, P. vulgaris, S. bicolor, and Z. mays were found to contain more nupDNAs than A. thaliana, C. reinhardtii, P. patens, and S. moellendorffii (Fig. 1). For example, G. max contains 1718 nupDNAs related to photosynthesis, metabolism, fatty acid metabolism, transporters, cellular processes, and the biosynthesis of cofactors, of which, 485 are involved in photosynthesis. On the other hand, A. thaliana contains only 85 nupDNAs, of which, 30 are related to photosynthesis (Online Resource 1). Compared with the genomes of lower plants, such as C. reinhardtii and P. patens, the nuclear genomes of higher plants, such as A. thaliana, F. vesca, and G. max (with the exception of S. moellendorffii), have more plastid DNA sequences (Fig. 1). The lower level of nupDNAs may be a characteristic of lower plant genomes, and sequencing additional lower plant genomes could reveal if this is a typical difference between lower and higher plants.
Interestingly, the ratio of complete coding DNA sequences (CDSs) to total nupDNAs was not constant among plant species. For O. sativa, 51.29 % of the identified nupDNA genes related to photosynthesis contained intact CDSs (Fig. 1a and Online Resource 1). In contrast, in G. max, most of the identified nupDNA genes were partial sequences, or truncated CDSs, and only 8.45 % contained intact CDSs despite having a larger number of nupDNAs than O. sativa. For O. sativa, 15 of the 21 genes related to photosynthesis had at least one nuclear intact CDS copy without mutations and 10 of the 25 genes related to energy metabolism had at least one nuclear intact CDS copy without mutations (Online Resource 1). By contrast, the only nuclear copy of plastid atpI in A. thaliana contained several single nucleotide deletions, which produced mutations. Similar results were found for C. sinensis using the plastid gene petG and in T. cacao using plastid ndhG.
The numbers of intact nuclear copies of different plastid genes varied among plants. For example, B. distachyon psbI, encoding photosystem II protein I, had one intact CDS copy (Online Resource 2), but psbF, encoding photosystem II protein VI, had eight intact CDS copies (four were nonmutated intact CDSs). Interestingly, atpF, petB, petD, ndhB, and ndhB had no intact CDS copies according to our data. By contrast, G. max plastid-derived nuclear sequences covered almost the entire plastid genome. Online Resource 1 revealed that these nupDNA fragments became integrated into the nuclear genome at different frequencies. A. thaliana had 12 nupDNAs corresponding to rbcL but had only two nupDNAs corresponding to psbC, clearly indicating that these chloroplast genes differed in their propensity to undergo integration into the nuclear genome.
Characteristics of intact nuclear copies of plastid genes
To study the features of intact plastid genes in plant nuclear genomes, we classified them as nonmutated or mutated (Fig. 2, Online Resource 1, and Online Resource 2). In A. thaliana, G. max, O. sativa, P. vulgaris, S. bicolor, and Z. mays, we subclassified mutated intact genes into those containing nonsynonymous or synonymous substitutions. The analysis clearly showed that the ratio of nonmutated intact genes to total intact genes varied among plant species. For O. sativa, 42.86 % of genes had nonmutated intact CDSs (Fig. 2a and Online Resource 1), whereas for G. max, 24.39 % of genes had nonmutated intact CDSs.
Distribution of intact plant coding DNA sequences (CDSs) in nuclear-localized plastid DNA (nupDNA) and the presence or absence of mutations (Online Resource 1 and 2). The black and white boxes indicate nupDNAs containing intact CDSs, either with mutations or without, respectively. a Genes related to photosynthesis; b genes related to energy metabolism; c genes related to fatty acid metabolism, transporters, cellular processes, and biosynthesis of cofactors
The analysis also demonstrated that the nupDNAs included copies of plastid genes with synonymous substitutions in the intact CDSs. O. sativa had 19 synonymous substitutions in intact CDSs, and Z. mays had 10 such synonymous substitutions. However, A. thaliana and P. vulgaris had no synonymous substitutions in any intact CDS in their nupDNAs (Online Resource 1). This finding suggested that at least some of these genes have undergone strong positive natural selection.
Estimation of age distribution reveals variation in transfer frequency among different plants
To estimate when individual nupDNA fragments became integrated into the nuclear genome, we compared the nucleotide substitutions in nupDNAs with those present in the chloroplast genome. To estimate the rate of substitution, we estimated the age (million years, Myr) of nupDNA fragments (Matsuo et al. 2005; Yoshida et al. 2013). We excluded data from species having a low level of nupDNA, such as C. reinhardtii, P. patens, and S. moellendorffii. The age distribution profiles of the nupDNA fragments in plants suggested that nupDNAs were repeatedly integrated into the nuclear genome. Furthermore, the proportion of nupDNAs of specific ages (Myr) varied among plant species (Fig. 3 and Online Resource 1).
Age distribution of plant nuclear-localized plastid DNA (nupDNA) by millions of years (Myr) (see Online Resource 1). a Genes related to photosynthesis; b genes related to energy metabolism
There were two distinct age distribution patterns of nupDNAs in the plant species we analyzed (Fig. 3). In one pattern, a large number of nupDNAs were translocated, either within the past 1 Myr or from 1 to 10 Myr ago, and the amount of nupDNA decreased as the age increased. This was illustrated by O. sativa (Matsuo et al. 2005) and Z. mays. In the other pattern, a very low proportion of young nupDNAs existed and decreased slowly with time. This result was similar to that of Yoshida et al. (2013). This was found in A. thaliana, F. vesca, G. max, P. vulgaris, and S. bicolor.
Analysis of predicted promoters of nuclear copies of plastid genes
To study the expression of the nupDNA genes, we analyzed EST sequences corresponding to the nuclear copies of plastid genes. This was performed using only nupDNA fragments with mutated intact genes because it was difficult to confirm the origin of ESTs of nonmutated intact genes as they may have been derived from either nuclear or plastid genes. The analysis suggested that some nuclear-localized plastid genes are transcribed and functional (Online Resource 1 and Online Resource 2). To understand the promoters of nuclear-localized plastid genes, we searched the predicted promoter sequences in the plant nuclear genome database (http://www.phytozome.net) and plastid genome database (http://www.ncbi.nlm.nih.gov/). For every plastid gene and nuclear-localized plastid gene, the region up to 1 kb upstream of the translation start site was considered as the predicted promoter region, unless it was determined to be smaller.
The analysis showed that many of the predicted promoter sequences of nuclear-localized plastid genes had been shuffled or eliminated after integration into the nuclear genome (Fig. 4, Online Resource 1 and Online Resource 2). For example, in O. sativa, 55.93 and 70.67 % of the predicted promoter sequences of genes related to photosynthesis or energy metabolism, respectively, had been eliminated. Interestingly, in G. max, 70.73 % of the predicted promoter sequences of genes related to photosynthesis had been eliminated, yet only 33.33 % of those related to energy metabolism had been eliminated. By contrast, in M. truncatula, 37.5 % of the predicted promoter sequence of genes related to photosynthesis had been eliminated, but 96.08 % of those related to energy metabolism had been eliminated.
Comparison of predicted promoters between plastid genes and their nuclear-localized homologs. The black, gray, and white boxes indicate high similarity, partial similarity, and dissimilarity, respectively. a Genes related to photosynthesis; b genes related to energy metabolism; c genes related to fatty acid metabolism, transporters, cellular processes, and biosynthesis of cofactors
Some genes had ESTs in both nupDNA genes and plastid genes (Table 1, Online Resource 1 and Online Resource 2), such as the O. sativa psbK (nupDNA gene EST: JK503631.1, and plastid gene EST: CI746041.1) and M. truncatula psbM (nupDNA gene EST: CO516909.1 and plastid gene EST: EX528553.1). By contrast, some ESTs were apparent in either the nupDNA gene or plastid gene, such as the O. sativa rbcL (nupDNA gene EST not found; plastid gene EST: CB672943.1) and O. sativa psbZ (nupDNA gene EST: CI741169.1; plastid gene EST not found). More data are shown in the Online Resources 1 and 2. Interestingly, the predicted rbcL promoters of the nupDNA gene and the corresponding plastid gene in O. sativa were very similar (Online Resource 3). However, although the M. truncatula nupDNA psbM and corresponding plastid gene were transcribed, the predicted promoters differed.
References
Adams KL, Song K, Roessler PG et al (1999) Intracellular gene transfer in action: dual transcription and multiple silencings of nuclear and mitochondrial cox2 genes in legumes. Proc Natl Acad Sci U S A 96:13863–13868
Adams KL, Qiu YL, Stoutemyer M et al (2002) Punctuated evolution of mitochondrial gene content: high and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc Natl Acad Sci U S A 99:9905–9912
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796
Argout X, Salse J, Aury JM et al (2011) The genome of Theobroma cacao. Nat Genet 43:101–108
Ayliffe MA, Scott NS, Timmis JN (1998) Analysis of plastid DNA-like sequences within the nuclear genomes of higher plants. Mol Biol Evol 15:738–745
Baldauf SL, Palmer JD (1990) Evolutionary transfer of the chloroplast tufA gene to the nucleus. Nature 344:262–265
Banks JA, Nishiyama T, Hasebe M et al (2011) The Selaginella genome identifies genetic changes associated with the evolution of vascular plants. Science 332:960–963
Bausher MG, Singh ND, Lee SB et al (2006) The complete chloroplast genome sequence of Citrus sinensis (L) Osbeck var’Ridge Pineapple’: organization and phylogenetic relationships to other angiosperms. BMC Plant Biol 6:21
Bensasson D, Zhang DX, Hartl DL et al (2001) Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends Ecol Evol 16:314–321
Bortiri E, Coleman-Derr D, Lazo GR et al (2008) The complete chloroplast genome sequence of Brachypodium distachyon: sequence comparison and phylogenetic analysis of eight grass plastomes. BMC Res Notes 1:61
Daniell H, Wurdack KJ, Kanagaraj A et al (2008) The complete nucleotide sequence of the cassava (Manihot esculenta) chloroplast genome and the evolution of atpF in Malpighiales: RNA editing and multiple losses of a group II intron. Theor Appl Genet 116:723–737
Gantt JS, Baldauf SL, Calie PJ et al (1991) Transfer of rpl22 to the nucleus greatly preceded its loss from the chloroplast and involved the gain of an intron. EMBO J 10:3073
Gargano D, Vezzi A, Scotti N et al (2005) The complete nucleotide sequence of potato (Solanum tuberosum cv.Desiree) chloroplast DNA. In: Abstracts Second Solanaceae Genome workshop p. 107
Goodstein DM, Shu S, Howson R et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Guo X, Castillo-Ramírez S, González V et al (2007) Rapid evolutionary change of common bean (Phaseolus vulgaris L) plastome, and the genomic diversification of legume chloroplasts. BMC Genomics 8:228
Guo X, Ruan S, Hu W et al (2008) Chloroplast DNA insertions into the nuclear genome of rice: the genes, sites and ages of insertion involved. Funct Integr Genom 8:101–108
Hiratsuka J, Shimada H, Whittier R et al (1989) The complete sequence of the rice (Oryza sativa) chloroplast genome: intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol Gen Genet 217:185–194
Huang CY, Ayliffe MA, Timmis JN (2003) Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature 422:72–76
Huang CY, Ayliffe MA, Timmis JN (2004) Simple and complex nuclear loci created by newly transferred chloroplast DNA in tobacco. Proc Natl Acad Sci U S A 101:9710–9715
Huang S, Li R, Zhang Z et al (2009) The genome of the cucumber, Cucumis sativus L. Nat Genet 41:1275–1281
Jaillon O, Aury JM, Noel B et al (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Jansen RK, Kaittanis C, Saski C et al (2006) Phylogenetic analyses of Vitis (Vitaceae) based on complete chloroplast genome sequences: effects of taxon sampling and phylogenetic methods on resolving relationships among rosids. BMC Evol Biol 6:32
Kahlau S, Aspinall S, Gray JC et al (2006) Sequence of the tomato chloroplast DNA and evolutionary comparison of solanaceous plastid genomes. J Mol Evol 63:194–207
Kleine T, Maier UG, Leister D (2009) DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant Biol 60:115–138
Maier RM, Neckermann K, Igloi GL et al (1995) Complete sequence of the maize chloroplast genome: gene content, hotspots of divergence and fine tuning of genetic information by transcript editing. J Mol Biol 251:614–628
Martin W (2003) Gene transfer from organelles to the nucleus: frequent and in big chunks. Proc Natl Acad Sci U S A 100:8612–8614
Martin W, Herrmann RG (1998) Gene transfer from organelles to the nucleus: how much, what happens, and why? Plant Physiol 118:9–17
Martin W, Stoebe B, Goremykin V et al (1998) Gene transfer to the nucleus and the evolution of chloroplasts. Nature 393:162–165
Martin W, Rujan T, Richly E et al (2002) Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc Natl Acad Sci U S A 99:12246–12251
Matsuo M, Ito Y, Yamauchi R et al (2005) The rice nuclear genome continuously integrates, shuffles, and eliminates the chloroplast genome to cause chloroplast–nuclear DNA flux. Plant Cell 17:665–675
Maul JE, Lilly JW, Cui L et al (2002) The Chlamydomonas reinhardtii Plastid Chromosome Islands of Genes in a Sea of Repeats. Plant Cell 14:2659–2679
Millen RS, Olmstead RG, Adams KL et al (2001) Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plant Cell 13:645–658
Ming R, Hou S, Feng Y et al (2008) The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452:991–996
Nishiyama T, Fujita T, Shin T et al (2003) Comparative genomics of Physcomitrella patens gametophytic transcriptome and Arabidopsis thaliana: implication for land plant evolution. Proc Natl Acad Sci U S A 100:8007–8012
Noutsos C, Richly E, Leister D (2005) Generation and evolutionary fate of insertions of organelle DNA in the nuclear genomes of flowering plants. Genome Res 15:616–628
Paiva JAP, Prat E, Vautrin S et al (2011) Advancing Eucalyptus genomics: identification and sequencing of lignin biosynthesis genes from deep-coverage BAC libraries. BMC Genomics 12:137
Paterson AH, Bowers JE, Bruggmann R et al (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Pląder W, Yukawa Y, Sugiura M et al (2007) The complete structure of the cucumber (Cucumis sativus L.) chloroplast genome: its composition and comparative analysis. Cell Mol Biol Lett 12:584–594
Potato Genome Sequencing Consortium (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Prochnik S, Marri PR, Desany B et al (2012) The cassava genome: current progress, future directions. Trop Plant Biol 5:88–94
Project IRGS (2005) The map-based sequence of the rice genome. Nature 436:793–800
Rujan T, Martin W (2001) How many genes in Arabidopsis come from cyanobacteria? An estimate from 386 protein phylogenies. Trends Genet 17:113–120
Saski C, Lee SB, Daniell H et al (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Saski C, Lee SB, Fjellheim S et al (2007) Complete chloroplast genome sequences of Hordeum vulgare, Sorghum bicolor and Agrostis stolonifera, and comparative analyses with other grass genomes. Theor Appl Genet 115:571–590
Sato S, Nakamura Y, Kaneko T et al (1999) Complete structure of the chloroplast genome of Arabidopsis thaliana. DNA Res 6:283–290
Schmutz J, Cannon SB, Schlueter J et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Schnable PS, Ware D, Fulton RS et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115
Shahmuradov IA, Akbarova YY, Solovyev VV et al (2003) Abundance of plastid DNA insertions in nuclear genomes of rice and Arabidopsis. Plant Mol Biol 52:923–934
Sheppard AE, Timmis JN (2009) Instability of plastid DNA in the nuclear genome. PLoS Genet 5:e1000323
Shrager J, Hauser C, Chang CW et al (2003) Chlamydomonas reinhardtii genome project. A guide to the generation and use of the cDNA information. Plant Physiol 131:401–408
Shulaev V, Sargent DJ, Crowhurst RN et al (2010) The genome of woodland strawberry (Fragaria vesca). Nat Genet 43:109–116
Smith DR (2009) Unparalleled GC content in the plastid DNA of Selaginella. Plant Mol Biol 71:627–639
Smith DR, Crosby K, Lee RW (2011) Plastids and gene transfer: correlation between nuclear plastid DNA abundance and plastid number supports the limited transfer window hypothesis. Genome Biol Evol 3:365
Stegemann S, Hartmann S, Ruf S et al (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci U S A 100:8828–8833
Sugiura C, Kobayashi Y, Aoki S et al (2003) Complete chloroplast DNA sequence of the moss Physcomitrella patens: evidence for the loss and relocation of rpoA from the chloroplast to the nucleus. Nucleic Acids Res 31:5324–5331
Timmis JN, Ayliffe MA, Huang CY et al (2004) Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5:123–135
Tomato Genome Consortium (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Tuskan GA, Difazio S, Jansson S et al (2006) The genome of black cottonwood, Populus trichocarpa (Torr & Gray). Science 313:1596–1604
Vogel JP, Garvin DF, Mockler TC et al (2010) Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 463:763–768
Woischnik M, Moraes CT (2002) Pattern of organization of human mitochondrial pseudogenes in the nuclear genome. Genome Res 12:885–893
Xu Q, Chen LL, Ruan X et al (2013) The draft genome of sweet orange (Citrus sinensis). Nat Genet 45:59–66
Yoshida T, Furihata HY, Kawabe A (2013) Patterns of genomic integration of nuclear chloroplast DNA fragments in plant species. DNA Res 21:127–140
Young HA, Lanzatella CL, Sarath G et al (2011a) Chloroplast genome variation in upland and lowland switchgrass. PLoS One 6:e23980
Young ND, Debellé F, Oldroyd GED et al (2011b) The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480:520–524
Yuan Q, Hill J, Hsiao J et al (2002) Genome sequencing of a 239-kb region of rice chromosome 10 L reveals a high frequency of gene duplication and a large chloroplast DNA insertion. Mol Genet Genomics 267:713–720
Acknowledgments
This work was supported by grants from the Trans-Century Training Program’s Foundation for the Talents by Heilongjiang Provincial Education Department (1251–NCET—004) and the Innovation Team Project by Heilongjiang Provincial Education Department to A. X. Wang, the Returned Oversea Scholar Foundation by Heilongjiang Provincial Education Department (1252HQ011), and the National Science Foundation of China (31301780) to X. L. Chen.
Author information
Authors and Affiliations
Corresponding author
Supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Homologs of plastid genes in plant nuclear DNA. (PDF 728 kb)
Online Resource 2
Homologs of intact coding DNA sequences (CDSs) of plastid genes in plant nuclear DNA. (PDF 257 kb)
Online Resource 3
Sequence alignment of predicted plant gene promoter regions. (PDF 384 kb)
Rights and permissions
About this article
Cite this article
Chen, H., Yu, Y., Chen, X. et al. Plastid DNA insertions in plant nuclear genomes: the sites, abundance and ages, and a predicted promoter analysis. Funct Integr Genomics 15, 131–139 (2015). https://doi.org/10.1007/s10142-014-0422-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-014-0422-z