Abstract
Phytophthora cinnamomi is a soil-borne plant pathogen that has caused widespread damage to vulnerable native ecosystems and agriculture systems across the world and shows no sign of abating. Management of the pathogen in the natural environment is difficult and the options are limited. In order to discover more about how resistant plants are able to defend themselves against this generalist pathogen, a microarray study of plant gene expression following root inoculation with P. cinnamomi was undertaken. Zea mays was used as a resistant model plant, and microarray analysis was conducted using the Affymetrix GeneChip Maize Genome Array on root samples collected at 6- and 24-h post-inoculation. Over 300 genes were differentially expressed in inoculated roots compared with controls across the two time points. Following Gene Ontology enrichment analysis and REVIGO visualisation of the up-regulated genes, many were implicated in plant defence responses to biotic stress. Genes that were up-regulated included those involved in phytoalexin biosynthesis and jasmonic acid/ethylene biosynthesis and other defence-related genes including those encoding glutathione S-transferases and serine-protease inhibitors. Of particular interest was the identification of the two most highly up-regulated genes, terpene synthase11 (Tps11) and kaurene synthase2 (An2), which are both involved in production of terpenoid phytoalexins. This is the first study that has investigated gene expression at a global level in roots in response to P. cinnamomi in a model plant species and provides valuable insights into the mechanisms involved in defence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The soil-borne oomycete Phytophthora cinnamomi Rands is the causal agent of disease in a wide range of hosts from diverse environments including the forests of the Mediterranean (Vettraino et al. 2005; Gómez-Aparicio et al. 2012) and the eastern USA (Balci et al. 2007; Nagle et al. 2010), and in a range of horticultural species, for example, macadamia (Mbaka et al. 2009) and avocado (Zentmyer 1984; Acosta-Muñiz et al. 2011) and the species-rich ecosystems of Australia (Cahill et al. 2008). In Australia, invasion of native vegetation communities across vast tracts of the continent has resulted in irreparable changes to their floristic and structural components (Fig. 1). The disease and its impacts are so serious that P. cinnamomi is listed by the Australian Government as a key-threatening process to Australia’s biodiversity (Environment Australia 2001). It has been estimated, for example, that for the South-West botanical province of Western Australia over 3,000 of the 5,710 described plant species are either susceptible or highly susceptible to this generalist pathogen (Shearer et al. 2004). Resistance to P. cinnamomi is rare in ecosystems where the pathogen is invasive. Currently, there are no effective, long-term methods of eradicating the pathogen from either soil or plants without subsequent vegetation damage and loss (Dunstan et al. 2010), although the use of the systemic chemical, phosphite, provides some protection against disease (Eshraghi et al. 2011).
Disease caused by P. cinnamomi in an Australian open forest ecosystem. a Healthy, uninfected vegetation at Anglesea, south-west Victoria, Australia with an intact over-storey of eucalypt species and an understorey dominated by the iconic Australian species, Xanthorrhoea australis. b Recent disease spread that has resulted in collapse and death of X. australis (arrow) and associated understorey species. c Older diseased site, with large areas of bare ground where susceptible plants have died and decomposed. Where X. australis was present depressions in the soil are all that remain (black arrows). Over time these bare sites become colonised by resistant species such as the sedges Lomandra filiformis and Lepidosperma semiteres (white arrow) resulting in a complete change in structural and floristic diversity (Cahill et al. 2008)
The pathogen invades plant root systems predominantly via a motile zoosporic stage that is released from asexual sporangia that are initiated during conducive environmental conditions. Root infection occurs also through root-to-root contact (Cahill et al. 2008). The biflagellate zoospores exhibit chemotaxis towards roots where they encyst at the zone of elongation and form a single germ tube. P. cinnamomi produces effectors that allow it to establish within host cells (Hardham and Cahill 2010; Horta et al. 2010) and in the case of the polygalacturonases enable penetration into the epidermis through degradation of anticlinal cell walls within a very short time after cyst germination (Hardham 2005). Hyphae then penetrate both inter- and intracellularly to invade successive layers of the cortex and move into the vascular tissue to colonise both phloem and xylem and at the same time move laterally along the root in all cell layers (Cahill et al. 1989). In susceptible species, colonisation and degradation of root cells leads to lesion formation, root rot and then, over time, the inhibition of water and nutrient uptake (Dawson and Weste 1984; Aberton et al. 2001) that lead to chlorosis of leaves and then plant death (Weste and Marks 1987; Wilson et al. 2000; Laidlaw and Wilson 2003; Cahill et al. 2008).
Resistance to P. cinnamomi is still quite poorly understood. For the species that do restrict colonisation of roots by the pathogen, for example, grasses and sedges, there are some responses that have been found to be associated with resistance including the involvement of reactive oxygen species (ROS), cell wall fortification, an accumulation of resistance-related proteins and antimicrobial secondary metabolites, alterations in phytohormones levels, and induction of defence-related genes (Cahill et al. 1986; Rookes et al. 2008; Sánchez-Pérez et al. 2009; Panjehkeh et al. 2010; Acosta-Muñiz et al. 2011; Eshraghi et al. 2011). In contrast, the interactions of host-specific Phytophthora species with their hosts, for example, Phytophthora infestans with potato (Fry 2008) and Phytophthora sojae with soybean (Tyler 2007), are very well characterised. The shortfall in knowledge of interactions with P. cinnamomi can now be addressed by the availability of fast throughput genomic and proteomic approaches. For example, two recent studies of the interaction of P. cinnamomi with Persea americana roots have provided new insights into the interaction through the use of protein profiling (Acosta-Muñiz et al. 2011) and EST sequencing and gene expression profiling (Mahomed and van den Berg 2011).
In this paper, we have used gene expression profiling to examine the response to P. cinnamomi induced in roots of Z. mays and show that this host rapidly responds to the pathogen. We have also found in this model system that resistance resulted from an active defence response and that it was associated with the strong up-regulation of key defence-related genes, including those related to oxidative stress, hormone biosynthesis and secondary metabolite synthesis. This new information provides insights into the fundamental molecular components that comprise a resistance response against this intractable oomycete pathogen.
Materials and methods
Plant growth and maintenance
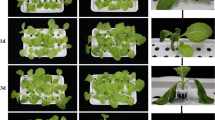
Z. mays (‘Early Leaming’, Lower Beechmont, QLD, Australia) seeds were germinated and grown in a soil-free plant growth system as described in Gunning and Cahill (2009) within a controlled-temperature cabinet under high-pressure sodium lights at a Photosynthetic Photon Flux of 250–300 μmol m−2 s−1 with a 14/10-h light/dark photoperiod at 22 °C. Each box within which plants were grown contained 1 L of one third strength hydroponic nutrient solution (THC, Excel Distributors, Doncaster East, Victoria, Australia), the plants were top misted with tap water at 2-day intervals and the nutrient solution refreshed every 5 days. The apical region of the primary root of each plant was removed, using a scalpel, 5 days after seeding to encourage lateral root growth.
Pathogen growth, maintenance and plant inoculation
P. cinnamomi isolate DU67 (A2 mating type; Deakin University culture collection) was used for inoculation of Z. mays roots (Allardyce et al. 2012). The isolate was maintained on 10 % CV8 agar plates at 25 °C in the dark and subcultured every 10 days. Pathogen virulence was maintained by passing the isolate through Lupinus angustifolius radicles every 12–16 weeks.
Zoospores of P. cinnamomi were produced using a method based on that of Byrt and Grant (1979), and the suspension diluted to a density of 1 × 105 spores mL−1 with sdH2O prior to use as inoculum. Lateral roots were inoculated when Z. mays seedlings were 10 days old. For microarray experiments, a minimum of five 20-μL droplets were applied contiguously to lateral roots, starting 0.5 cm from each root tip. Control plants were treated in an identical manner except for the use of sdH2O as the inoculum. For microscopic analysis of the interaction over time, a single 20-μL drop of zoospores was applied 0.5 cm from root tips.
Microscopic analysis of the interaction of Z. mays roots with P. cinnamomi
Using fine forceps, epidermal peels were taken from directly beneath the point of inoculation from control and inoculated Z. mays at half hourly intervals to 2-h post-inoculation (hpi) then 3 and 6 hpi and then from 24 hpi every 24 h until 120 hpi. The peels (one peel per root from a minimum of three roots per time point) were then mounted in distilled water on a microscope slide and viewed with light and epi-fluorescence microscopy (365 nm excitation and 420 nm emission) (Axioskop2, Zeiss, Clayton, Victoria, Australia). At the same time points H2O2 production was examined using 3′3′ di-aminobenzidine (DAB; 1 mg mL−1 DAB, pH 3.5) (Thordal-Christensen et al. 1997) and additional defence responses such as the production of callose and lignin were assessed as previously described (Mauch-Mani and Slusarenko 1996; Rookes et al. 2008).
Microarray analysis—RNA preparation, labelling, hybridisation and scanning
Z. mays plants were arranged in a randomised block design (Nettleton 2006) within the growth cabinet and inoculated as described above. All replicates were conducted at the same time of day across each experimental repeat to exclude variations due to differences in circadian rhythm. Root tissue was collected from the inoculated root zones and snap frozen in liquid nitrogen prior to RNA extraction at 6 and 24 hpi. Three experimental repeats were conducted; each consisted of 60 seedlings in total, with 15 control and 15 inoculated plants at each of the two time points. All harvested root tissue within each treatment group at the two time points was pooled for each RNA extraction.
Total RNA was isolated and purified from root tissue using a TRIzol® Plus RNA Purification System (which includes a DNAse 1 treatment step) according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA). A minimum of 15 μg of purified total RNA per sample was prepared for microarray analysis. The 12 samples of purified total RNA were processed at the Australian Genome Research Facility, (AGRF Melbourne Node, The Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria). Each biological replicate was then labelled, hybridised and scanned according to prescribed protocols for the GeneChip Maize Genome Array (Affymetrix, Santa Clara, CA; www.affymetrix.com).
Bioinformatics analysis
Bioinformatic analysis of raw data was conducted using Partek® Genomics Suite™ software (Partek Incorporated, St. Louis, MO). All raw data, comprising 12 separate .CEL files, were imported into Partek® Genomics Suite™ software for gene expression data, which allowed statistical interpretation and visualisation of data. All data were normalised using Robust Multi-chip Average software, to generate an expression profile from Affymetrix data (Irizarry et al. 2003). Differentially expressed genes were defined by a fold difference of >1.5 and were considered statistically significant if they had a p value cut-off of <0.05. Genes were annotated according to Affymetrix Maize Array Annotation file release 32.
Analysis of differentially regulated transcripts and promoter analysis
Gene ontology (GO) enrichment analysis of up- and down-regulated transcripts was carried out using AgriGO Singular Enrichment Analysis (Du et al. 2010) using a minimum of two mapping entries and the remainder of the parameters as specified by the programme defaults. The GO enrichment data were transferred to REVIGO (Supek et al. 2011) for reduction and visualisation. The allowed similarity value was set at 0.5 and the Z. mays genome database was used as the source for the GO term sizes. For analysis of the promoter regions of genes encoding up-regulated transcripts, the 500-bp upstream regions were downloaded from the Z. mays Genomic Database (Dong et al. 2004; Lawrence et al. 2004) and analysed using Regulatory Sequence Analysis Tools (RSAT; van Helden 2003). Background frequencies were calculated from input data using a higher order Markov model order value of 2. Significantly over-represented 6–8 mers were assembled to identify consensus motifs.
Data submission to public repository
All supporting information including raw gene expression data (as .CEL files), normalised data, experimental design details, array annotation and experimental protocols were submitted to the Gene Expression Omnibus (GEO; Platform Accession Number: GPL4032) in accordance with the Minimum Information About a Microarray Experiment (MIAME) standard by the Microarray Gene Expression Data society (Brazma et al. 2001; Rayner et al. 2006). The records were assigned the accession number GSE27626.
Validation of microarray analysis using semi-quantitative reverse transcription-PCR
Semi-quantitative RT-PCR was conducted to validate microarray results using the same RNA samples. For cDNA synthesis, 1 μg of total RNA was reverse-transcribed in a reaction containing Tetro reverse transcriptase, reaction buffer, and 2 μM random hexamers as per the manufacturer’s specifications (Bioline, Alexandria, NSW, Australia). Primers were designed to examine the relative expression of three genes (An2, Gst30 and Opr1) that were identified as being up-regulated in the microarray analysis as well as Gapc2 that encodes glyceraldehyde 3-phosphate dehydrogenase for use as a control (Table S3). PCR reactions were carried out with GoTaq green master mix and reagents (Promega, Madison, WI). PCR reactions contained 1 μL of cDNA from the RT reaction and 0.5 μM of the appropriate oligonucleotides. PCR cycles consisted of an initial denaturing step of 3 min at 94 °C, followed by repetitions (18–34 cycles, depending on the primer set) of the following three steps: a 30-s denaturation step at 94 °C, 30-s annealing step ranging between 50 and 52 °C and a 1-min elongation step at 72 °C. Initial reactions were performed to determine the appropriate cycle number for analysis to be conducted within the exponential phase of the PCR reaction (data not shown). PCR products were analysed on 0.5 × TBE-agarose gels and visualised using ethidium bromide staining. Digital analysis of gel images was used to calculate fold change differences (Image J software, US National Institutes of Health, Bethesda, MD http://imagej.nih.gov/ij/, 1997–2011).
Results
Timing of penetration of root cells and response to infection
When grown under the conditions described, the roots of Z. mays were healthy and white with a clearly differentiated root cap, and the various zones of differentiation, elongation and maturation could be discerned (Fig. 2a). Following inoculation, P. cinnamomi zoospores were attracted to the root surface within 30 min and within 1 h had shed their flagella and encysted, subsequently forming germ tubes which penetrated the unsuberised primary root tissue within 3 h (Fig. 2b). At 6 hpi, cell wall thickening occurred directly below the point of inoculation and there was autofluorescence of epidermal cell walls (Fig. 2c). Larger, discrete areas of autofluorescence associated with hyphal penetration, as well as cell wall thickening, were apparent by 24 hpi (Fig. 2d). The production of hydrogen peroxide was examined using DAB staining. In control roots there was little staining but there was intense brown staining consistent with H2O2 production concentrated in the intracellular regions of epidermal cells associated with areas of germ tube penetration (Fig. 2e, f). At 24 hpi, restricted brown lesions were found beneath the point of inoculation (Fig. 2g). An overview of observed Z. mays defence responses following challenge by P. cinnamomi is provided in Table S1. Based on these observations and previous studies (Hinch et al. 1985; Rookes et al. 2008; Allardyce et al. 2012), 6 and 24 hpi were selected for subsequent microarray analysis.
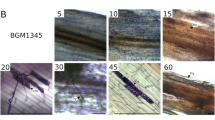
General appearance of Z. mays roots and response to infection by P. cinnamomi. a Uninoculated root tip region showing locations of the root cap (RC) the zone of differentiation (D), zone of elongation (E) and zone of maturation (M). Zoospores of P. cinnamomi are chemotactically attracted to the zone of elongation. Scale bar = 3mm, b zone of elongation observed under the light microscope 180 min post-inoculation with P. cinnamomi zoospores. Zoospores formed walled cysts (black arrow) which extended germ tubes (white arrow) towards the root followed by their penetration of the epidermis. Scale bar = 20 μm. c, d Root epidermal peels taken 6 (c) and 24 h (d) after inoculation that show autofluorescence at the site of inoculation. Hyphae can be observed in association with the autofluorescence (arrows). Scale bar = 10 μm. e, f Epidermal peel of control (e) and inoculated (f) root 6 h after inoculation stained with DAB. Intense DAB staining (arrow) was associated with cells and cell walls following inoculation and indicated the production of hydrogen peroxide. Scale bar = 10 μm. g Restricted lesion beneath the point of inoculation (arrow). Scale bar = 10 mm
Differential gene expression in Z. mays roots following inoculation with P. cinnamomi zoospores
Transcriptional profiling of Z. mays roots in response to infection with P. cinnamomi was conducted using the Affymetrix GeneChip® maize genome array, representing 13,339 genes. Levels of gene expression at 6 and 24 hpi in inoculated plants were compared with those of control plants. Over the two time points examined, a combined total of 317 genes were identified as being differentially expressed (p < 0.05, 1.5 fold cut-off) in inoculated root tissue, of which 257 were annotated (Table S2). By 6 hpi, 234 genes were differentially expressed in inoculated roots, of which 186 were annotated. Of these differentially expressed genes, 212 were up-regulated while 22 genes were down-regulated. By 24 hpi, the number of differentially expressed genes in inoculated roots was reduced to 83, 71 of which were annotated. Of the genes that showed differential expression at 24 hpi, 46 genes were up-regulated while 37 were down-regulated.
Overview of functional categories and promoter analysis of up-regulated genes
The web-based AgriGO Singular Enrichment Analysis tool was used to obtain significantly over-represented GO terms from the up-regulated gene datasets. The REVIGO web server was then used to summarise identified GO terms and visualise the data. At 6 hpi, REVIGO assigned 247 GO terms to Biological Process, 48 to Cellular Component and 99 to Molecular Function. Data were displayed as scatterplots for biological process and molecular function (Fig. 3; Fig. S1). Following analysis for biological process, several clusters of GO terms with related function were identified. At 6 hpi, a tight cluster of frequently represented biological process GO terms were associated with defence responses and responses to biotic stimulus and other organisms (Fig. 3a). A further cluster was associated with glutathione metabolism and related metabolic processes. Other categories of enriched biological process GO terms included those related to carbohydrate metabolism and terpenoid metabolism. Of the GO terms related to molecular function (Fig. 3b), those for glutathione transferase activity and other transferase activities were enriched, while others related, presumably, to activities around the cell wall included pattern binding, polysaccharide binding and chitinase activity. With fewer genes up-regulated at 24 hpi, REVIGO summarised 69 GO terms to biological process, 24 to cellular component and 47 to molecular function. While a smaller set of GO terms were identified at 24 hpi in comparison with the 6 hpi dataset, the categories of terms across the two time points were similar (Fig. S1).
Genes that were up-regulated at 6 h after inoculation were subjected to GO enrichment analysis, then summarised and visualised as a scatter plot using the REVIGO Web server. a Summarised GO terms related to biological processes and b molecular function. GO terms are represented by circles and are plotted according to semantic similarities to other GO terms (adjoining circles are most closely related). Circle size is proportional to the frequency of the GO term, while colour indicates the log10 p value (red higher, blue lower). GO terms with a log10 p value of −4.3 or less for biological process and −0.96 or less for molecular function are labelled
Promoter analysis revealed three motifs that were enriched in the genes that were up-regulated at 6 h (Table 1). Interestingly, the RY repeat motif that is associated with genes that are related to ABA response was found to be prominent, followed by the GCC-Box that is related to ethylene (Et) and jasmonic acid (JA) signalling, and then the A-Box motif related to sugar (carbohydrate) signalling. REVIGO and promoter analysis of down-regulated genes was also performed but due to the small number of genes that were down-regulated at the two time points the analysis tools provided limited information (data not shown).
Principal categories of genes that were found to be up-regulated in roots in response to P. cinnamomi at 6 and 24 h after inoculation
Highly expressed genes encoding enzymes of secondary metabolite synthesis and defence-related proteins
The two most highly expressed genes at 6 hpi were both associated with the biosynthesis of terpenoid phytoalexins: Tps11 (Zm.14496.1.A1_at) that encodes terpene synthase was up-regulated 18.1-fold, and An2 (ZmAffx.12.1.S1_at) which encodes ent-copalyl diphosphate synthase and is associated with kauralexin production (Schmelz et al. 2011) was up-regulated 10-fold (Table 2; Table S2). By 24 hpi, the expression levels of both these genes decreased to the level of controls. A number of key defence-related genes also showed high up-regulation (>5-fold) at 6 hpi including Wip1 (Zm186.1.S1_at), that encodes a wound-induced proteinase inhibitor, Pr5 (Zm.6659.1.A1_at) that encodes the pathogenesis-related protein 5 and Glp2 (Zm.503.1.A1_at) which encodes the germin-like protein 2. Both Wip1 and Glp2 remained up-regulated at 24 hpi, although to lesser levels (3.3- and 3.4-fold, respectively).
Glutathione S-transferase transcripts involved in detoxifying reactive oxygen species
Of the annotated genes up-regulated at 6 hpi in P. cinnamomi inoculated Z. mays roots, seven were glutathione S-transferases (GSTs), which have previously been found to be induced under stress conditions, including pathogen attack, and serve to protect plant cells and tissues from oxidative stress caused by the generation of reactive oxygen species (Marrs 1996; Sappl et al. 2009) (Table S2). The GST genes included Gst30 (Zm864.1.S1_at) which was up-regulated 6.8-fold, Gst8 (Zm.539.1.A1_at) up-regulated 3.9-fold, Gst24 (Zm.627.1.A1_at) up-regulated 3.6-fold and Gst7 (Zm.246.1.A1_at), Gst25 (Zm.545.1.S1_at), Gst23 (Zm.561.1.A1_at) and Gst15 (Zm.545.1.S1_at), all of which were up-regulated between 1.6–2.0-fold. None of the GST-encoding genes were up-regulated at 24 hpi.
Biosynthesis of defence-related plant hormones
Several genes that encode enzymes involved in JA biosynthesis from linolenic acid (Turner et al. 2002) were up-regulated at 6 hpi, including Opr1 (Zm.10676.1.S1_at) that encodes 12-oxo-phytodienoic acid reductase 1 (induced 5.2-fold), Aos1 (Zm.6107.1.A1_at), allene oxide synthase 1 (1.6-fold) and a lipoxygenase gene, LOC542495 (Zm.445.1.S1_at; 1.5-fold) (Fig. 4). An Et biosynthesis gene, acc oxidase Aco31 (Zm.8714.1.A1_at) that catalyses the final step in the Et biosynthetic pathway, was up-regulated 4.1-fold at 6 hpi; whereas, the ethylene-responsive-factor-like protein Erf1 (Zm.11441.1.S1_at) was down-regulated by 24 hpi (−3.1-fold). In contrast, there was no change in the expression of genes associated with salicylic acid (SA) biosynthesis or SA-signalling pathways.
Schematic representation of the JA biosynthetic pathway including the genes found to be up-regulated in this study (grey boxes) at 6 hpi in inoculated Z. mays roots. Membrane derived linolenic acid is oxidised to 13-hydroperoxy-linolenic acid via the enzyme lipoxygenase (LOX). Allene oxide synthase (AOS) converts 13-hydroperoxy-linolenic acid to an unstable epoxide 12,13-epoxy-linolenic acid which rapidly becomes 12-oxo-phytodienoic acid (12-oxo-PDA) and defence responses may be initiated at this point (Turner et al. 2002). The enzyme 12-oxo-PDA reductase is involved in the final step of the conversion of 12-oxo-PDA to JA. JA may then be further catabolised to its methyl ester (MeJA). Genes up-regulated: LOX, 1.5-fold; AOS, 1.6-fold; and 12-oxo-PDA-reductase, 5.2-fold. AOC not differentially expressed (white box)
Genes encoding pathogenesis-related proteins and other defence-associated genes
A number of proteins directly involved in defence against invading pathogens were differentially expressed at both 6 and 24 hpi in inoculated root tissue (Table S2). These included protease inhibitors that contribute to plant defence by inhibiting proteases produced by invading microorganisms. At 6 hpi, several protease-inhibitor type genes were induced including genes encoding two Bowman–Birk serine proteinase inhibitors (Wip1, Zm.186.1.S1_at, Zm.17271.6.S1_x_at) that were up-regulated by 7.7- and 3.4-fold, respectively. Two thaumatin-like, pathogenesis-related protein encoding genes were also up-regulated at 6 hpi, namely Pr5 (Zm.6659.1.A1_at; 7.5-fold) and Sip1 (Zm.281.1.S1_s_at; 4.2-fold) which has 100 % amino acid sequence identity to the anti-fungal protein, zeamatin. By 24 hpi, four putative serine-protease inhibitor genes were up-regulated, including Wip1 (Zm.186.1.S1_at; 3.3-fold) and the putative Bowman–Birk serine protease inhibitor gene LOC100037814 (Zm.4270.2.A1_a_at; 2-fold) both of which were also up-regulated at 6 hpi, and two trypsin inhibitor genes, LOC541688 (Zm.375.1.S1_at) at 2.7-fold and the subtilisin-chymotrypsin inhibitor gene Sci1 (Zm.301.1.A1_a_at) at 2.2-fold. A number of genes encoding chitinase proteins were up-regulated over both time points, with a propensity towards their expression at 24 hpi.
Validation of differential gene expression using semi-quantitative RT-PCR
To verify the results of the microarray analysis, semi-quantitative RT-PCR was performed to examine expression levels of three genes (An2, Gst30 and Opr1) that were shown in the microarray analysis to display differential expression in inoculated root tissue at 6 hpi. The Gapc2 gene was used as an internal control to confirm even RNA loading and reaction efficiency across all cDNA samples. All three genes that displayed differential gene expression in the microarray analysis were shown to exhibit similar gene expression patterns in semi-quantitative RT-PCR reactions (Fig. 5a). There was some variation between the three experimental repeats but the pattern of differential gene expression was consistent. Average fold change differences in expression of the four genes were calculated through image analysis and were found to correspond with the microarray analysis results (Fig. 5b).
Semi-quantitative RT-PCR was used to validate the microarray analysis for untreated controls (C1–C3) and for the 6- h inoculated root samples (I1–I3) for each of three biological replicates. a The expression levels of An2, Gst30 and Opr1 were up-regulated following inoculation. b Comparison of fold change for each gene between the semi-quantitative RT-PCR data (grey bar) and microarray data (white bar). Data presented are the mean of the three independent replicates, and error bars represent standard error of the mean
Discussion
This study provides the first insights into global gene expression changes of a plant in response to challenge by the pathogen P. cinnamomi. By analysing the changes in gene expression in the resistant monocot species, Z. mays, our analysis provides important new information on the molecular basis of resistance to this pathogen. Changes in the Z. mays transcriptome in response to P. cinnamomi infection demonstrates that the plant both recognises and responds rapidly to the pathogen by inducing a suite of key defence-related genes by 6 hpi. By 24 hpi, several defence-related genes showed reduced expression levels, or their expression had returned to basal levels, reflecting the rapid, transient nature of this response. Our observations show the rapidity with which encystment, germination and then penetration occurs (within 3 h of inoculation) in this system and that resistance requires a rapid host response. Furthermore, the up-regulation of genes associated with the production of plant-derived pathogen-specific toxins, as well as inhibitors of pathogen effectors, including a number of serine-protease inhibitors, indicates that Z. mays responds to penetration by P. cinnamomi in an active and controlled manner.
Phytoalexins have been previously recorded as being involved in the resistance responses of Z. mays and accumulate in response to specific elicitors derived directly from pathogens or in vitro elicitation (Shen et al. 2000; Doehlemann et al. 2008). Of all the up-regulated genes identified in this study, the two most highly expressed were Tps11 and An2 which are both associated with the biosynthesis of terpenoid phytoalexins. Tps11 encodes (s)-beta-macrocarpene synthase, which is putatively involved in zealexin production (Huffaker et al. 2011), while An2 encodes ent-copalyl diphosphate synthase and is associated with kauralexin production. Both genes have recently been shown to be highly expressed in Z. mays following inoculation with several fungal pathogens including Fusarium graminearum, and it has been proposed that terpenoid phytoalexins are important components of defence (Huffaker et al. 2011; Schmelz et al. 2011). It is clear from these recent studies and our own that terpenoid phytoalexins are likely to be central to the Z. mays defence response and that these metabolites may offer alternatives for plant breeding programs and transgenic approaches towards developing disease resistant lines.
It is now well established that the SA pathway is usually associated with resistance to biotrophic pathogens, while the JA/Et pathways are important for defence against necrotrophs (Glazebrook 2005; Trusov et al. 2009; Antico et al. 2012). Our study, which showed the up-regulation of key genes in the JA and Et biosynthesis pathways at 6 hpi, including the gene encoding 12-oxo-PDA reductase, the enzyme that catalyses the final conversion of 12-oxo-PDA to JA, also the gene encoding the enzyme involved in the final step in Et biosynthesis (1-aminocyclopropane-1-carboxylate oxidase) indicates that the JA/Et pathways are important in the development of resistance of Z. mays to P. cinnamomi. Promoter analysis also revealed JA/Et-related motifs (GCC-BOX) as prominent within the up-regulated gene set and also, interestingly, an ABA-related motif (RY Repeat) that may indicate a role, as yet undetermined, for ABA in this interaction. Previous studies have suggested that the JA/Et pathways are induced in defence against P. cinnamomi (Rookes et al. 2008; Mahomed and van den Berg 2011). The results described here further substantiate JA/Et involvement in the development of resistance to P. cinnamomi and also demonstrates that the defence response exhibited by Z. mays is consistent with that of plants challenged by necrotrophic pathogens.
GSTs are a family of enzymes implicated in stress responses from both exogenous and endogenous origins, including pathogen defence (Marrs 1996). During defence, GSTs neutralise toxic components released by both pathogens and the plants themselves by facilitating conjugation of the toxicant with glutathione. Following conjugation, the compounds are then transported to the apoplast or sequestered within vacuoles (Marrs 1996; Dixon et al. 2002). A suite of seven GST genes were up-regulated at the early time-point of 6 hpi in this study. GST gene expression has been shown to be transcriptionally regulated by a broad range of stress conditions, including the production of reactive oxygen species, such as H2O2 during pathogen attack (Tenhaken et al. 1995; Marrs 1996; Lamb and Dixon 1997; Sappl et al. 2009).
A number of other genes also associated with oxidative stress were observed to be up-regulated in the Z. mays–P. cinnamomi interaction. For example, a gene (Glp2) encoding a germin-like protein was up-regulated at 6 hpi and to a lesser degree at 24 hpi. GLPs are purported to have strong anti-oxidant activity and are extremely resistant to degradation by both pathogen-derived proteases and H2O2 (Kukavika et al. 2005; Łaźniewska et al. 2010). Several GLPs have superoxide dismutase activity but a clear link between GLPs and H2O2 production is yet to be established (Christensen et al. 2004; Łaźniewska et al. 2010). While there is little information on the role of GLPs in defence to necrotrophs in Z. mays, they have been reported to be induced following pathogen invasion in other monocots (Membré et al. 2000; Zimmermann et al. 2006).
Inducible defence-related proteins that are present within healthy tissue at low levels have been identified in many plant species following infection by necrotrophic fungi and oomycetes (van Loon et al. 2006; Meier and Gehring 2008; Aswati-Nair et al. 2010). In our study two thaumatin-like protein (TLP) genes were up-regulated at 6 hpi, namely those encoding the pathogenesis-related protein PR5, which was highly up-regulated (7.5-fold), and the stress-induced protein 1 (4.2-fold) with 100 % sequence identity to zeamatin, a 22-kDa TLP which has been reported to have potent antifungal activity in vitro (Schimoler-O’Rourke et al. 2001). TLPs exhibit multiple enzymatic activities, including β-1,3 glucanase activity, which specifically targets the cell walls of pathogenic organisms, including fungi and oomycetes (Liu et al. 2010). Early up-regulation of TLPs in Z. mays roots may therefore form an integral part of an arsenal of defence genes and their products that work synergistically to limit P. cinnamomi invasion.
Our study has identified a range of key defence-related genes that are up-regulated in Z. mays roots in response to infection by P. cinnamomi. While some of these genes encode products that have been identified during resistance of plants to other root pathogens (Okubara and Paulitz 2005), our analysis has revealed new components of defence against a broad host range oomycete that include GSTs, the PR5 protein and enzymes involved in phytoalexin synthesis as well as GLPs and serine-protease inhibitors whose role in resistance is less well characterised. Furthermore, our results demonstrate that resistance to P. cinnamomi in Z. mays requires the simultaneous involvement of JA/Et pathways, terpenoid biosynthesis, ROS-related genes and the production of other potentially fungitoxic molecules. This new knowledge has opened up the potential for manipulation of molecular pathways and defence responses in other species and provides a sound molecular platform for further research on plant resistance to P. cinnamomi.
References
Aberton MJ, Wilson BA, Cahill DM (2001) Development of disease caused by Phytophthora cinnamomi in mature Xanthorrhoea australis. Aust J Bot 49:209–219
Acosta-Muñiz CH, Escobar-Tovar L, Valdes-Rodríguez S, Fernández-Pavia S, Arias-Saucedo LJ, de las Cruz Espindola Barquera M, Lim MÁG (2011) Identification of avocado (Persea americana) root proteins induced by infection with the oomycete Phytophthora cinnamomi using a proteomic approach. Physiol Plant 144:59–72
Allardyce JA, Rookes JE, Cahill DM (2012) Defining plant resistance to Phytophthora cinnamomi: a standardised approach to assessment. J Phytopathol 160:269–276
Antico CJ, Colon C, Banks T, Ramonell KM (2012) Insights into the role of jasmonic acid-mediated defences against necrotrophic and biotrophic fungal pathogens. Front Biol 7:48–56
Aswati-Nair R, Kiran AG, Sivakumar KC, Thomas G (2010) Molecular characterisation of an oomycete-responsive PR-5 protein gene from Zingiber zurumbet. Plant Mol Biol Rep 28:128–135
Balci Y, Balci S, Eggers J, MacDonald WL, Juzwik J, Long RP, Gottschalk KW (2007) Phytophthora spp. associated with forest soils in eastern and north central U.S. oak ecosystems. Plant Dis 91:705–710
Brazma A, Hingamp P, Quackenbush J, Sherlock G, Spellman P, Stoeckhart C, Aach J, Ansorge W, Ball CA, Causton HC et al (2001) Minimum information about a microarray experiment (MIAME)-toward standards for microarray data. Nature 29:365–371
Byrt P, Grant BR (1979) Some conditions governing zoospore production in axenic cultures of Phytophthora cinnamomi Rands. Aust J Bot 27:103–115
Cahill DM, Grant B, Weste G (1986) Changes in cytokinin concentrations in xylem extrudate following infection of Eucalyptus marginata Donn ex Sm with Phytophthora cinnamomi Rands. Plant Physiol 81:1103–1109
Cahill DM, Legge N, Grant B, Weste G (1989) Cellular and histological changes induced by Phytophthora cinnamomi in a group of plant species ranging from fully susceptible to fully resistant. Phytopathology 79:417–424
Cahill DM, Rookes JE, Wilson BA, Gibson L, McDougall KL (2008) Phytophthora cinnamomi and Australia’s biodiversity: impacts, predictions and progress towards control. Aust J Bot 56:279–310
Christensen A, Thordal-Christensen H, Zimmermann G, Gjetting T, Lyngkjær MF, Dudler R, Schweizer P (2004) The germin-like protein GLP4 exhibits superoxide dismutase activity and is an important component of quantitative resistance in wheat and barley. Mol Plant-Microbe Interact 17:109–117
Dawson P, Weste G (1984) Impact of root infection by Phytophthora cinnamomi on the water relations of two Eucalyptus species that differ in susceptibility. Phytopathology 74:486–490
Dixon DP, Lapthorn A, Edwards R (2002) Plant glutathione transferases. Genome Biol 3:3004.1–3004.10
Doehlemann G, Wahl R, Horst RJ, Voll LM, Usadel B, Poree F, Stitt M, Kühnemann JP, Sonnewald U, Kahmann R, Kämper J (2008) Reprogramming a maize plant: transcriptional and metabolic changes induced by the fungal biotroph Ustilago maydis. Plant J 56:181–195
Dong Q, Schlueter SD, Brendel V (2004) PlantGDB, plant genome database and analysis tools. Nucleic Acids Res 32:D354–D359
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) AgriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70
Dunstan WA, Rudman T, Shearer BL, Moore NA, Paap T, Calver MC, Dell B, Hardy GESJ (2010) Containment and spot eradication of a highly destructive, invasive plant pathogen (Phytophthora cinnamomi) in natural ecosystems. Biol Invasions 4:913–925
Environment Australia (2001) Threat abatement plan for dieback caused by the root-rot fungus (Phytophthora cinnamomi). Environment Australia, Canberra
Eshraghi L, Anderson J, Aryamanesh N, Shearer B, McComb J, Hardy GESJ, O’Brien PA (2011) Phosphite primed defence responses and enhanced expression of defence genes in Arabidopsis thaliana infected with Phytophthora cinnamomi. Plant Pathol 60:1086–1095
Fry W (2008) Phytophthora infestans: the plant (and R gene) destroyer. Mol Plant Path 9:385–402
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227
Gómez-Aparicio LG, Ibáňez B, Serrano MS, De Vita P, Ávila JM, Pérez-Ramos IM, Garcia LV, Sánchez ME, Maraňón T (2012) Spatial patterns of soil pathogens in declining Mediterranean forests: implications for tree species regeneration. New Phytol 194:1014–1024
Gunning TK, Cahill DM (2009) A soil-free plant growth system to facilitate analysis of plant–pathogen interactions in roots. J Phytopathol 157:497–501
Hardham AR (2005) Phytophthora cinnamomi. Mol Plant Path 5:589–604
Hardham AR, Cahill DM (2010) The role of oomycete effectors in plant pathogen interactions. Funct Plant Biol 37:919–925
Higo K, Ugawa IM, Higo H (1998) PLACE: a database of plant cis-acting regulatory DNA elements. Nucleic Acids Res 26:358–359
Hinch JM, Wetherbee R, Mallett JE, Clarke AE (1985) Response of Zea mays roots to infection with Phytophthora cinnamomi. Protoplasma 126:178–187
Horta M, Caetano P, Medeira C, Maia I, Cravador A (2010) Involvement of the β-cinnamomin elicitin in infection and colonisation of cork oak roots by Phytophthora cinnamomi. Eur J Plant Path 127:427–436
Huffaker A, Kaplan F, Vaughan MM, Dafoe NJ, Ni X, Rocca JR, Alborn HT, Teal PEA, Schmelz EA (2011) Novel acidic sesquiterpenoids constitute a dominant class of pathogen-induced phytoalexins in maize. Plant Physiol 156:2082–2097
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalisation, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Kukavika B, Vučinić Ž, Vuletić M (2005) Superoxide dismutase, peroxidise, and germin-like protein activity in plasma membranes and apoplast of maize roots. Protoplasma 226:191–197
Laidlaw WS, Wilson BA (2003) Floristic and structural characteristics of a coastal heathland exhibiting symptoms of Phytophthora cinnamomi infestation in the eastern Otway Ranges, Victoria. Aust J Bot 51:283–293
Lamb C, Dixon RA (1997) The oxidative burst in plant disease resistance. Annu Rev Plant Physiol Plant Mol Biol 48:251–275
Lawrence CJ, Dong Q, Polacco ML, Seigfried TE, Brendel V (2004) MaizeGDB, the community database for maize genetics and genomics. Nucleic Acids Res 32:D393–D397
Łaźniewska J, Macioszek VK, Lawrence CB, Kononowicz AK (2010) Fight to the death: Arabidopsis thaliana defence response to fungal necrotrophic pathogens. Acta Physiol Plant 32:1–10
Liu J-J, Sturrock R, Ekramoddoullah AKM (2010) The superfamily of thaumatin-like proteins: its origin, evolution, and expression towards biological function. Plant Cell Rep 29:419–436
Mahomed W, van den Berg N (2011) EST sequencing and gene expression profiling of defence-related genes from Persea americana infected with Phytophthora cinnamomi. BMC Plant Biol 11:167
Marrs KA (1996) The functions and regulation of glutathione S-tranferases in plants. Annu Rev Plant Physiol Plant Mol Biol 47:127–158
Mauch-Mani B, Slusarenko AJ (1996) Production of salicylic acid precursors is a major function of phenylalanine ammonia-lyase in the resistant response of Arabidopsis to Peronospora parasitica. Plant Cell 8:203–212
Mbaka JN, Wamocho LS, Turoop L, Waiganjo MM (2009) The incidence and distribution of Phytophthora cinnamomi Rands on macadamia in Kenya. J Anim Plant Sci 4:289–297
Meier S, Gehring C (2008) A guide to the integrated application of on-line data mining tools for the inference of gene functions at the systems level. J Biotechnol 3:1375–1387
Membré N, Bernier F, Staiger D, Berner A (2000) Arabidopsis thaliana germin-like proteins: common and specific features point to a variety of functions. Planta 211:345–354
Nagle AM, Long RP, Madden LV, Bonello P (2010) Association of Phytophthora cinnamomi with White Oak Decline in Southern Ohio. Plant Dis 94:1026–1034
Nettleton D (2006) A discussion of statistical methods for design and analysis of microarray experiments for plant scientists. Plant Cell 18:2112–2121
Okubara PA, Paulitz TC (2005) Root defense responses to fungal pathogens: a molecular perspective. Plant Soil 274:215–226
Panjehkeh N, Backhouse D, Taji A (2010) Role of proanthocyanidins in resistance of the legume Swainsona formosa to Phytophthora cinnamomi. J Phytopathol 158:365–371
Rayner TF, Rocca-Serra P, Spellman PT, Causton HC, Farne A, Holloway E, Irezarry RA, Liu J, Maier DS, Miller M et al (2006) A simple spreadsheet-based MIAME-supportive format for microarray data: MAGE-TAB. BMC Bioinforma 7:489
Rookes JE, Wright ML, Cahill DM (2008) Elucidation of defence responses and signalling pathways induced in Arabidopsis thaliana following challenge with Phytophthora cinnamomi. Physiol Mol Plant P 72:151–161
Sánchez-Pérez JDL, Jaimes-Lara MG, Salgado-Garciglia R, López-Meza JE (2009) Root extracts from Mexican avocado (Persea americana var. drymifolia) inhibit the mycelial growth of the oomycete Phytophthora cinnamomi. Eur J Plant Pathol 124:595–601
Sappl PG, Carroll AJ, Clifton R, Lister R, Whelan J, Harvey-Millar A, Singh KB (2009) The Arabidopsis glutathione transferase gene family displays complex stress regulation and co-silencing multiple genes results in altered metabolic sensitivity to oxidative stress. Plant J 58:53–68
Schimoler-O’Rourke R, Richardson M, Selitrennikoff CP (2001) Zeamatin inhibits trypsin and α-amylase activities. Appl Environ Microb 67:2365–2366
Schmelz EA, Kaplan F, Huffaker A, Dafoe NJ, Vaughan MM, Ni X, Rocca JR, Alborn HT, Teal PE (2011) Identity, regulation, and activity of inducible diterpenoid phytoalexins in maize. Proc Natl Acad Sci USA 108:5455–5460
Shearer BL, Crane CE, Cochrane A (2004) Quantification of the susceptibility of the native flora of the South-West Botanical Province, Western Australia, to Phytophthora cinnamomi. Aust J Bot 52:435–443
Shen B, Zheng Z, Dooner HK (2000) A maize sesquiterpene cyclase gene induced by insect herbivory and volicitin: characterisation of wild-type and mutant alleles. Proc Natl Acad Sci USA 97:14807–14812
Supek F, Bošnjak M, Škunca N, Šmuc T (2011) REVIGO summarizes and visualizes long lists of Gene Ontology terms. PLoS One 6:e21800
Tenhaken R, Levine A, Brisson LF, Dixon RA, Lamb C (1995) Function of the oxidative burst in hypersensitive disease resistance. Proc Natl Acad Sci USA 92:4158–4163
Thordal-Christensen H, Zhang Z, Wei Y, Collinge DB (1997) Subcellular localisation of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J 11:1187–1194
Trusov Y, Sewelam N, Rookes JE, Kunkel M, Nowak E, Schenk PM, Botella JR (2009) Heterotrimeric G proteins-mediated resistance to necrotrophic pathogens includes mechanisms independent of salicylic acid-, jasmonic acid/ethylene-, and abscisic acid-mediated defense signalling. Plant J 58:69–81
Turner JG, Ellis C, Devoto A (2002) The jasmonate signaling pathway. Plant Cell 14:S153–S164
Tyler BM (2007) Phytophthora sojae: root rot pathogen of soybean and model oomycete. Mol Plant Path 8:1–8
Van Helden J (2003) Regulatory sequence analysis tools. Nucleic Acids Res 31:3593–3596
Van Loon LC, Rep M, Pieterse CMJ (2006) Significance of inducible defence-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Vettraino AM, Morel O, Perlerou C, Robin C, Diamandis S, Vannini A (2005) Occurrence and distribution of Phytophthora species in European chestnut stands, and their association with Ink Disease and crown decline. Eur J Plant Pathol 111:169–180
Weste G, Marks GC (1987) The biology of Phytophthora cinnamomi in Australasian forests. Annu Rev Phytopathol 25:207–229
Wilson BA, Aberton J, Cahill DM (2000) Relationships between site factors and distribution of Phytophthora cinnamomi in the Eastern Otway Ranges, Victoria. Aust J Bot 48:247–260
Zentmyer G (1984) Avocado diseases. Trop Pest Manag 30:388–400
Zimmermann G, Bäumlein H, Mock H-P, Himmelbach H, Schweizer P (2006) The multigene family encoding germin-like proteins of barley: regulation and function in basal host resistance. Plant Physiol 142:181–192
Acknowledgments
The authors thank the Australian Commonwealth Department of Sustainability, Environment, Water, Population and Communities for financial support and the Melbourne Node of the Australian Genome Research Facility for processing microarray data. We thank Dr. Michael Gardner and Ms Sharareh Kavkani, Deakin University, for assistance with bioinformatic analyses. MIAME-compliant data are deposited in GEO (www.ncbi.nlm.nih.gov/geo), accession number GSE27626.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Allardyce, J.A., Rookes, J.E., Hussain, H.I. et al. Transcriptional profiling of Zea mays roots reveals roles for jasmonic acid and terpenoids in resistance against Phytophthora cinnamomi . Funct Integr Genomics 13, 217–228 (2013). https://doi.org/10.1007/s10142-013-0314-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-013-0314-7