Abstract
Plant and animal microRNAs (miRNAs) are evolutionarily ancient small RNAs, ∼19–24 nucleotides in length, that are generated by cleavage from larger highly structured precursor molecules. In both plants and animals, miRNAs posttranscriptionally regulate gene expression through interactions with their target mRNAs, and these targets are often genes involved with regulating key developmental events. Despite these similarities, plant and animal miRNAs exert their control in fundamentally different ways. Generally, animal miRNAs repress gene expression by mediating translational attenuation through (multiple) miRNA-binding sites located within the 3′ untranslated region of the target gene. In contrast, almost all plant miRNAs regulate their targets by directing mRNA cleavage at single sites in the coding regions. These and other differences suggest that the two systems may have originated independently, possibly as a prerequisite to the development of complex body plans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the last 6 years, the existence and the mechanism of double-stranded RNA-directed gene silencing have become a major area of plant and animal research. When double-stranded RNAs or self-complementary single-stranded “hairpin” RNAs are introduced into eukaryotic cells, their duplexed regions are cut into ∼21 nucleotide (nt) fragments by an enzyme called Dicer. These 21 nt short interfering RNAs (siRNAs) guide nuclease complexes to cognate single-stranded RNAs, which they cleave. It was initially thought that the sole purpose of this mechanism was to defend plants against RNA viruses and transposons. However, it has recently become apparent that the pathway also provides essential regulation of some key developmental processes in both plants and animals by producing ∼21 nt microRNAs (miRNAs). These miRNAs, excised from endogenously encoded hairpin RNAs, negatively regulate endogenous target genes by cleavage or translational inhibition of their mRNA (Fig. 1a). Mutations in the gene encoding Dicer1 in Arabidopsis can have major consequences as a result of defective miRNA production (Fig. 1b).
a Comparison of the mechanisms of miRNA biogenesis and action. The biogenesis of plant and animal miRNAs differ in that in silico stem loop predictions have yielded larger and more variable precursor miRNA molecules for plants than for animals (Reinhart et al. 2002; Voinnet 2003). The Drosha gene that processes the pri-miRNA to the pre-miRNA in animals is absent from plant genomes. In plants, the Dicer-like 1 (DCL1, a RNase-III-like protein) appears to catalyze the processing of the primary miRNA transcript to form the miRNA:miRNA* complex. The multisubunit endonuclease, shown as RISC, is the RNA-induced silecing complex. b Mutant of the Dicer-like 1 gene (DCL1) in Arabidopsis (top), showing extreme floral abnormalities, and a wild-type Arabidopsis plant (bottom). miRNAs are small RNAs that regulate a large number of genes, many of which are involved in key developmental processes. The floral abnormalities, such as distorted petals and multiple female floral organs (carpels) per flower, reflect the inability of mutant DCL1 to produce the appropriate miRNAs needed to regulate normal floral development
To date, miRNAs have been found in all plant and animal multicellular organisms examined and, among other roles, appear to regulate the development of multicellular body plans such as leaf and floral development in plants (see Fig. 1b) and early larval transitions in nematodes. In both animals and plants, miRNAs are evolutionarily ancient—at least 400 million years old (Pasquinelli et al. 2000; Floyd and Bowman 2004)—and many miRNA:target–gene interactions are broadly conserved. However, the conservation is restricted to within kingdoms; there is no report of any miRNA gene that is conserved between plants and animals. Thus, despite the apparent similarities of miRNAs from animals and plants and their critical role in development, it is possible that the miRNA system was not operating in a common ancestor, but originated independently from a more ancient system. This may not be so surprising considering that the last common ancestor of plants and animals was unicellular, and developmental comparisons have shown that the molecular mechanisms that gave rise to multicellular forms evolved independently in each lineage (Meyerowitz 2002). Small regulatory RNAs are present in prokaryotes (Altuvia 2004). For example, in Escherichia coli, more than 50 small RNA-encoding genes have been identified, some are acting in trans by hybridizing to their target gene(s). Perhaps something akin to this system was the progenitor from which both plant and animal miRNA systems evolved.
In this brief review, we explore some of the similarities and differences between the miRNA systems of plants and animals and examine whether they are fundamentally different or simply variations of a theme.
Genomic organization of miRNA genes
The total number of miRNAs in any organism is unknown, but it has been estimated that Caenorhabditis elegans and Drosophila contain at least 100 miRNAs, while vertebrate genomes contain ∼250 miRNAs (Ambros 2004), thus equating to nearly 1% of the predicted genes in these organisms (Bartel 2004). In the Arabidopsis plant whose genome has been fully sequenced, over 100 miRNA-encoding loci have been identified (Llave et al. 2002a; Park et al. 2002; Reinhart et al. 2002; Jones-Rhoades and Bartel 2004; Bonnet et al. 2004; Sunkar 2004; http://cgrb.orst.edu/smallRNA/db/search_user_seq.html) although many of the miRNAs only differ from one another by single or several nucleotides and currently correspond to ∼40 “families” of miRNAs. The fact that 15 of these families were only identified recently in a study of stress-induced miRNAs implies that the upper limit of the number of miRNAs in a plant is far from known (Sunkar 2004).
Both plant and animal miRNA genes are predominantly located in what is conventionally termed the intergenic regions. The miRNA genes are mostly discrete independent transcription units that are not located near to their target genes. However, significant numbers of animal miRNAs are located in the introns of pre-mRNAs. This arrangement will give coordinate expression of the gene from its mRNA and the miRNA from the intron. Of the human miRNA genes, ∼25% is encoded within introns (Bartel 2004). In Arabidopsis, only one miRNA (miR402) gene has been identified within an intron so far (Sunkar 2004). Both the animal intron-encoded miRNAs and the plant miR402 are in the same orientation as the pre-mRNAs which carry them. This suggests that each transcript is processed to produce both a translatable spliced mRNA and a functional miRNA.
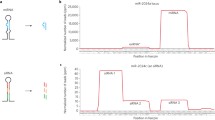
The presence of clusters of miRNA genes, being transcribed in large polycistronic primary transcripts (Fig. 2), is probably another way miRNAs are coordinately expressed. This appears to be a significant regulatory mechanism in Drosophila—∼50% of its predicted miRNAs genes are located within clusters (Aravin et al. 2003). In C. elegans and mammals, large numbers of miRNA clusters have also been found (Lagos-Quintana et al. 2001; Lau et al. 2001; Lim et al. 2003a, b), e.g., the C. elegans miR35–41 cluster (Fig. 2d) and the 14q32 domain in mouse that contains a cluster of more than 40 miRNAs (Seitz et al. 2004). miRNAs within clusters can be sequence-related. Other clusters can contain miRNAs that, although not sequence-related, appear to be involved in controlling the same functional process (Bartel 2004). In plants, most miRNAs are encoded by their own primary transcript; there have only been a few cases described of multiple miRNAs being within a polycistronic transcript such as miR395, which is present four times within a single transcript in rice (Fig. 2c). Also, three members of the miR399 family are within a 2-kb region in Arabidopsis, and a similar cluster exists in rice. This conservation suggests that this clustering is critical for the coordinate regulation of these miRNAs (Sunkar 2004). Despite these few examples of miRNA clusters in plants, in animals, miRNA clusters appear to have developed to a much greater extent.
miRNA biogenesis
In both animals and plants, miRNAs precursors seem to be encoded in capped and polyadenylated RNAs transcribed by polymerase II. These RNAs form stable stem-loop structures. Nonetheless, the biogenesis of plant and animal miRNAs differ in some aspects (see Fig. 1). In silico stem-loop predictions have yielded larger and more variable precursor miRNA molecules for plants than for animals (Reinhart et al. 2002). Most notably, the Drosha gene that processes the pri-miRNA to the pre-miRNA in animals is absent from plant genomes; this function is carried out by the plant RNase-III-like protein, Dicer-like 1 (DCL1). DCL1 appears to catalyze the processing of the primary miRNA transcript to form the miRNA:miRNA* duplex in the nucleus; the same enzyme carrying out both cleavage steps implies that the stem-loop precursor in plants are very transient molecules compared to their counterparts in animals. In contrast, this reaction occurs in the cytoplasm in animals, but is carried out by Dicer. More subtle differences include somewhat more pairing between the miRNA and the other arm of the stem loop in plants compared to animals, a tighter distribution of plant miRNA lengths that centers on 21 nt rather than the 22- to 23-nt lengths most often seen in animals and perhaps a stronger preference for a U at the 5′ terminus of the plant miRNAs (Lau et al. 2001; Reinhart et al. 2002; Bartel and Bartel 2003).
miRNA target genes
To date, plant miRNAs share much higher complementarity to their target genes (zero to three mismatches) than animal miRNAs to their target genes, although in both cases, high to perfect complementarity between the target mRNA and the 5′-half of the miRNA is required (Parizotto et al. 2004; Doench and Sharp 2004). In plants, the high complementarity, together with evolutionary conservation between Arabidopsis and rice, as well as the presence of the miRNA-binding motifs in multiple members of a gene family, have enabled accurate prediction of miRNA targets. Curiously, in plants, there are families of sequence-related miRNA genes that are predicted to target multiple members of a gene family (Table 1). Thus, multiple miRNA genes could be targeting a single member, with tissues and stage specificity, and/or a single miRNA gene could be regulating multiple family members. Either way, it appears that there could be many examples of gene duplication and divergence in broadening the role of miRNAs. The putative targets in plants are predominantly regulatory genes, such as transcription factors (Table 2), F-box proteins, and ubiquitin-conjugating enzymes (Rhoades et al. 2002; Jones-Rhoades and Bartel 2004; Sunkar 2004), many of which have been implicated in pivotal developmental roles. However, targets that fall outside of this broad classification have now been discovered such as ATP sulfurylases, laccases, cytochrome c oxidase, and superoxide dismutases (Jones-Rhoades and Bartel 2004; Sunkar 2004). The recent identification of these latter targets implies that miRNA-gene regulation may be involved in many facets of plant biology, not just development.
In contrast to the plant situation, simple homology-based searches have failed to uncover targets for miRNAs in animals (Ambros et al. 2003; Bartel and Bartel 2003). It has been necessary to create complex programs, relying on finding short segments of conserved complementarity to predict miRNA targets in animals (Enright et al. 2003; Lewis et al. 2003; Stark et al. 2003). Thousands of target genes have been predicted for mammals, and like plants, there is a strong bias toward genes involved in gene regulation, such as mRNA-encoding transcription factors, components of miRNA, and ubiquitin machinery, and proteins involved in translational repression. But again, there are also other classes of genes, including many structural proteins and enzymes, implying that miRNAs have regulatory roles in a diverse range of physiological processes (Lewis et al. 2003; John et al. 2004). It has been estimated that human miRNAs have the potential to regulate between 10 and 30% of all human genes (John et al. 2004; Lewis et al. 2005). However, of all the putative animal miRNA targets predicted to date, only several dozens have been validated experimently (Lewis et al. 2003; Stark et al. 2003).
One of the most notable differences between animal and plant miRNA systems is the location of the miRNA-binding sites within the target genes. These binding sites in known animal miRNA-target genes usually occur in multiples and always within the in the 3′ untranslated region (3′-UTR) of the mRNA (Lewis et al. 2003; Enright et al. 2003; Stark et al. 2003). For example, the lin-14 gene has seven lin-4 target sites (Lee et al. 1993). However, there may have been a bias in the discovery of animal miRNA-binding sites, because this has been primarily based upon computer predictions using databases composed of only 3′-UTR sequences (Lewis et al. 2003; Enright et al. 2003). Animal miRNA-mediated control seems likely to occur in a combinatorial way, with the presence and number of multiple binding sites in an mRNA likely to reflect the degree of potential repression.
Plant miRNA-binding sites are found almost exclusively within the open-reading frames of the target genes and with one site per mRNA. In some cases, the binding site can span intron/exon splice junctions and is only generated after the excision of the intron (Xie et al. 2003). However, miRNA-binding sites have recently been predicted to occur in the 3′-UTRs of a few plant mRNAs and, in one case, in the 5′-UTR of putative target gene—a location unique among all the other known plant and animal miRNA-binding sites (Sunkar 2004). This case is even more exceptional in that there are multiple copies of this miRNA target sequence in the 5′-UTR.
From these observations, both the number of miRNA-binding sites and their location appear to reflect an important mechanistic difference between plant and animal miRNAs.
Mechanistic action of miRNAs
miRNAs appear to operate through two main mechanisms, mRNA cleavage or translational repression. The mode of the mechanism appears to depend largely on the degree of complementarity between the miRNA and its binding site within the target mRNA. miRNAs with high complementarity to the target mediate cleavage, those with lower or partial complementarity mediate translational repression (Doench et al. 2003; Zeng et al. 2003). Most animal miRNAs have low complementarity to their target mRNA, suggesting that translational repression is the predominant form of miRNA regulation in animals and this is supported by limited experimental studies (Olsen and Ambros 1999). However, one mammalian miRNA (miR-196) is known to have near-perfect complementarity to its target mRNA (HOX8B), and it mediates cleavage of this mRNA (Yekta et al. 2004). While this raises the question of how many other metazoan miRNA targets might be down regulated by cleavage, it seems likely to be uncommon because no other animal miRNA with such extensive complementarity to its target mRNA has yet been found.
Most plant miRNA have high complementarity (less than four mismatches; G:U pairing permissible) to their target mRNAs and regulate gene expression via mRNA cleavage. In vitro or in vivo assays have demonstrated this cleavage for nearly 50 such target genes (Llave et al. 2002b; Kasschua et al. 2003; Tang et al. 2003; Palatnik et al. 2003; Jones-Rhoades and Bartel 2004). However, APETALA2 (AP2) has one to zero mismatches with members of the miR172 family, yet appears to be regulated predominantly by translational repression, although some mRNA cleavage also occurs (Aukerman and Sakai 2003). Currently, detection of cleavage is interpreted as diagnostic of regulation by mRNA degradation. However, its been suggested that the cleavage and translational repression pathways overlap, which raises the possibility that in some cases, where cleavage has shown to be occurring, the primary mode of regulation is translation repression followed by mRNA cleavage.
In plants, there are large families of miRNAs (with 1–3 nt variation) and large families of target genes with variable target sites (1–3 nt variation). Therefore, combinatorial regulation may be occurring, in which low-complementarity miRNA–mRNA interactions give repression of translation and high-complementarity miRNA–mRNA interactions result in mRNA cleavage. This combinatorial regulation could be occurring within a cell or differentially across cells types or tissues. For example, the AP2 transcript could be translationally regulated in some cells by low-affinity members of the miR172 family, while in other cells, it is cleaved by high-affinity family members.
The operation of these different mechanisms may be related to the number and location of miRNA-binding sites in the target genes. In animals, where there are multiple miRNA-binding sites within 3′-UTRs, their number and complementarity may be related to the extent to which translation is attenuated. If only one site is occupied expression is lowered, but if all the sites are occupied translation is fully repressed. Indeed, synergistic translational repression has been directly demonstrated by the addition of multiple binding sites into a 3′-UTR. This resulted in more efficient inhibition of translation than that expected from the sum of the effect of each binding site individually (Doench et al. 2003). Translation repression has the attributes of variable regulation and reversibility: removal of the miRNA from the sites may allow the mRNA to be transcribed. In contrast, a single miRNA site cleavage within a coding region destroys the mRNA molecule permanently, giving efficient control that can only be reversed by further transcription of the mRNA.
Translational repression occurs in both plants and animals, but do they operate by the same mechanism? For both cases, a decrease in protein level without a decrease in mRNA level has been termed translational repression (Wrightman et al. 1991; Aukerman and Sakai 2003; Chen 2004). However, little is known how miRNAs exert this repression in either system. Biochemical analysis revealed that the repressed mRNAs remain in polysomes, suggesting that the block in expression occurs after translation initiation (Olsen and Ambros 1999; Seggerson et al. 2002). In animals, translation regulation through the 3′-UTR is required for many important developmental processes including tissue patterning, embryonic axes formation, and mammalian spermatogenesis (Kuersten and Goodwin 2003); thus, miRNAs may be utilizing similar machinery that are involved in those processes. The fact that animal miRNA-binding sites are within the 3′-UTR, compared to the AP2 example, where the binding motif is within the coding region, may suggest that there could be mechanistic differences in translational repression in plants and animals.
Conclusions
There are many obvious similarities between plant and animal miRNA systems; both systems play fundamental roles in development and appear to predominantly exert their influence by controlling regulatory genes. However, there are also many differences (Table 3). In animals, the first step of miRNA biogenesis involves Drosha, but this role is performed by DCL1 in plants. The majority of plant miRNAs are each derived from single primary transcript from loci found in the intergenic regions, whereas many of animal miRNAs are generated from polycistronic transcripts from intergenic regions of the chromosome and many are produced from introns. In plants, miRNAs mainly regulate their targets by cleavage in the coding region of the RNA, whereas animal miRNAs mainly operate by translational repression using targets at the 3′-UTR. Although these differences between the animal and plant miRNA systems are clear-cut, in a general sense, there is almost always an exception that breaks the rule.
One possible reason for the general differences between the two systems is that they evolved separately, although probably from common ancient components, after the divergence of animals and plants. If this is the case, their functional similarity and mechanistic differences exemplifies convergent evolution.
References
Altuvia S (2004) Regulatory small RNAs: the key to co-ordinating global regulatory circuits. J Bacteriol 186:6679–6680
Ambros V (2004) The functions of animal microRNAs. Nature 431:244–350
Ambros V, Lee RC, Lavanway A, Williams PT, Jewell D (2003) MicroRNAs and other tiny endogenous RNAs in C. elegans. Curr Biol 13:807–818
Aravin AA, Lagos-Quintana M, Yalcin A, Zavolan M, Marks D, Snyder B, Gaasterland T, Meyer J, Tuschl T (2003) The small RNA profile during Drosophila melanogaster development. Dev Cell 5:337–350
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bartel B, Bartel DP (2003) MicroRNAs—at the root of plant development? Plant Physiol 132:709–717
Bonnet E, Wuyts J, Rouze P, Van de Peer Y (2004) Detection of 91 potential conserved plant microRNAs in Arabidopsis thaliana and Oryza sativa identifies important target genes. Proc Natl Acad Sci U S A 101:11511–11516
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Doench JG, Sharp PA (2004) Specificity of microRNA target selection in translational repression. Genes Dev 18:504–511
Doench JG, Peterson CP, Sharp PA (2003) siRNAs can function as miRNAs. Genes Dev 17:438–442
Enright AJ, John B, Gaul U, Tuschl T, Sander C, Marks DS (2003) MicroRNA targets in Drosophila. Genome Biol 5:R1
Floyd SF, Bowman JL (2004) Ancient microRNA target sequences in plants. Nature 428:485–486
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human microRNA targets. PLOS Biol 2:e363
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 4:205–217
Kuersten S, Goodwin EB (2003) The power of the 3′ UTR-translational control and development. Nat Rev Genet 4:626–637
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294:853–858
Lau NC, Lim LP, Weinstein EG, Bartel DP (2001) An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294:858–862
Lee RC, Feinbaum R, Ambros V (1993) The heterochronic gene lin-4 of C. elegans encodes two small RNAs with antisense complementarity to lin41. Cell 75:843–854
Lewis BP, Shih I, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115:787–798
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20
Lim LP, Lau NC, Weinstein EG, Abdelhakim A, Yekta S, Rhoades MW, Burge CB, Bartel DP (2003a) The microRNAs of Caenorhabditis elegans. Genes Dev 17:991–1008
Lim LP, Glasner ME, Yekta S, Burge CB, Bartel DP (2003b) Vertebrate microRNA genes. Science 299:1540
Llave C, Kasschau KD, Rector MA, Carrington JC (2002a) Endogenous and silencing-associated small RNAs in plants. Plant Cell 14:1605–1619
Llave C, Xie Z, Kasschau KD, Carrington JC (2002b) Cleavage of scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 297:2053–2056
Meyerowitz EM (2002) Plants compared to animals: the broadest comparative study of development. Science 295:1482–1485
Millar AA, Gubler F (2005) The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 17:705–721
Olsen PH, Ambros V (1999) The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev Biol 216:671–680
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263
Parizotto EA, Dunoyer P, Rahm N, Himber C, Voinnet O (2004) In vivo investigation of the transcription, processing, endonucleolytic activity, and functional relevance of the spatial distribution of a plant miRNA. Genes Dev 18:2237–2242
Park W, Li J, Song R, Messing J, Chen X (2002) CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 12:1484–1495
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda M, Maller B, Srinivasan A, Fishman M, Hayward D, Ball E et al (2000) Conservation across animal phylogeny of the sequence and temporal regulation of the 21 nucleotide let-7 heterochronic regulatory RNA. Nature 408:86–89
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP (2002) MicroRNAs in plants. Genes Dev 16:1616–1626
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110:513–520
Seggerson K, Tang L, Moss EG (2002) Two genetic circuits repress the Caenorhabditis elegans heterochronic gene lin-28 after translation initiation. Dev Biol 243:215–225
Seitz H, Royo H, Bortolin ML, Lin SP, Ferguson-Smith AC, Cavaille J (2004) A large imprinted microRNA gene cluster at the mouse Dlk1–Gtl2 domain. Genome Res 9:1741–1748
Stark A, Brennecke J, Russell RB, Cohen SM (2003) Identification of Drosophila microRNA targets. PLOS Biol 1:E60
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Tang G, Reinhart BJ, Bartel DP, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17:49–63
Voinnet O (2003) RNA silencing bridging the gaps in wheat extracts. Trends Plant Sci 8:307–309
Wrightman B, Burglin TR, Gatto J, Arasu P, Ruvkun G (1991) Negative regulatory sequences in the lin-14 3′-untranslated region are necessary to generate a temporal switch during Caenorhabditis elegans development. Genes Dev 5:1813–1824
Xie Z, Kasschau KD, Carrington JC (2003) Negative feedback regulation of Dicer-like1 in Arabidopsis by microRNA-guided mRNA degradation. Curr Biol 13:784–789
Yekta S, Shih I-H, Bartel DP (2004) MicroRNA-directed cleavage of HOXB8 mRNA. Science 304:594–596
Zeng Y, Yi R, Cullen BR (2003) MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc Natl Acad Sci U S A 100:9779–9784
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Millar, A.A., Waterhouse, P.M. Plant and animal microRNAs: similarities and differences. Funct Integr Genomics 5, 129–135 (2005). https://doi.org/10.1007/s10142-005-0145-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-005-0145-2