Abstract
The Chinese perch (Siniperca chuatsi) is one of the most commercially important carnivorous fish species in aquaculture with its large-scale culture in China. Increasing evidence suggests that microRNAs (miRNAs) play an important role in muscle cell proliferation and differentiation. However, the knowledge of the identity of myogenic miRNAs and the effect of nutrient status on miRNA expression in teleost remains limited. In the present study, among the 21 miRNAs identified with high abundance in the fast muscle of adult Chinese perch, 19 miRNAs were differentially expressed in the adults and juveniles. The postprandial changes in the transcript abundance were determined for the 21 miRNAs following a single satiating meal in the juveniles after fasting for 1 week. The results showed that the seven miRNAs (miR-10c, miR-107a, miR-133a-3p, miR-140-3p, miR-181a-5p, miR-206, and miR-214) were sharply upregulated or downregulated within 1 h after refeeding. These miRNAs may be the promising candidate miRNAs involved in a fast-response signaling system that regulates fish skeletal muscle growth. Target prediction and expressional analysis suggested that four miRNAs (miR-10c, miR-107a, miR-140-3p, and miR-181a-5p) might play a role in regulating the translation of target gene transcripts such as myostatin following acute anabolic stimuli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Skeletal muscle, separated to white (fast) and red (slow) muscles, is the largest tissue and the main edible part in the fish body (Zhang et al. 2009). The maintenance of skeletal muscle mass is a complex and controlled process that is largely influenced by the nutritional and physiological states of the animals (Fuentes et al. 2012). Fasting-refeeding is a commonly used tool to investigate the regulation of muscle growth in fish that experiences the transition from catabolic to anabolic states (Valente et al. 2012). In carnivorous fish, nutrient restriction enhances the release of amino acids from muscle fibers, which are subsequently used by hepatocytes as the main gluconeogenic precursors (Ballantyne 2001). During refeeding, an accelerated turnover takes place, resulting in an increased protein synthesis ratio (Peragón et al. 1999; Carter et al. 2001; Liu et al. 2013). Molecular tools enable us to gain a deeper insight into how growth responses are regulated by dietary factors (Bower and Johnston 2010; Panserat and Kaushik 2010). Currently, growth factors, regulatory proteins, and transcription factors have been identified as participating in the regulation of the myogenesis in fish (Nihei et al. 2006; Steinbacher et al. 2006). However, the underlying molecular pathway elements remain poorly understood.
MicroRNAs (miRNAs) have emerged as a new class of key regulators of myogenesis. miRNAs are approximately 22-nt non-coding RNAs that act as negative regulators of gene expression either by inhibiting messenger RNA (mRNA) translation or promoting mRNA degradation through base pairing to the 3′-untranslated region (3′-UTR) of the target mRNAs (Xie et al. 2005; Liu 2008). It has been shown that miR-1, miR-133, and miR-206 are specifically expressed in muscle cells and play a vital role in muscle cell differentiation. Studies using the mouse myogenic C2C12 cell line demonstrated that miR-1 and miR-133 were involved in myoblast proliferation and differentiation by regulating the expression of HDAC4 and SRF, respectively (Chen et al. 2006). A previous study showed miR-206 promotes myogenesis by repressing the expression of a subunit of DNA polymerase, the polymerase responsible for DNA synthesis during cell proliferation (Kim et al. 2006). However, the effect of nutrient status on miRNA expression and the role of miRNAs in response to changes of skeletal muscle anabolic state from nutrient restriction to refeeding remain unknown.
The Chinese perch (Siniperca chuatsi) is one of the most commercially important carnivorous fish species in China as well as in eastern Asia (Chu et al. 2010; Zhang et al. 2011). We have constructed a miRNA library from the skeletal muscle of adult Chinese perch and identified 21 miRNAs with high abundance in the fast muscle (Chu et al. 2013). In the present study, we analyzed the expression profiles of the 21 miRNAs in juvenile Chinese perch skeletal muscle and characterized these miRNAs’ expressional signature during the transition from catabolic to anabolic states. Our results showed that seven miRNAs were significantly upregulated or downregulated within 1 h after refeeding. Four miRNAs and their predicted target gene myostatin may coordinately regulate the rapid resumption of myogenesis from a catabolic to anabolic state in Chinese perch skeletal muscle.
Materials and Methods
Tissue Sampling
All Chinese perch individuals were reared at the Xingda Chinese perch hatchery, Changde, Hunan, China. The fast skeletal muscles were dissected from the dorsal myotome of juvenile (average body weight 150 g, 90 days post-hatching (dph)) and adult (average body weight 500 g, 150 dph) fish. Five individuals were sampled at each stage. All tissue samples were snap-frozen in liquid nitrogen and stored at −80 °C for further processing.
Fasting-Refeeding Experiments and Sampling
Two homogeneous groups of Chinese perch juveniles (average body weight 150 g, 90 dph) were reared in two net cages (5 m × 5 m × 2 m) with 60 fish per cage. All the juveniles were fed mud carp Cirrhinus molitorella (average body weight 10 g) for 1 week. The juveniles were fasted for 1 week, and then fed with a single meal (mud carp) to all individuals to satiation. Tissue sampling was carried out at 0 h (before the recovery meal) and at 1, 3, 6, 12, 24, 48, and 96 h after the single meal feeding. Five fish were sampled at each time point. Fast muscles were dissected from the dorsal myotome of individuals. All samples were snap-frozen in liquid nitrogen and stored at −80 °C until further processing.
Quantitative Real-Time PCR for the miRNAs and mRNA
Tissue samples were ground in liquid nitrogen. The total RNAs were extracted with the TRIzolR Reagent (Invitrogen, USA) and then treated with RNAse-free DNAse I (Promega, USA) in the presence of RNAse inhibitor (Sigma, China branch). The RNAs were precipitated by ethanol precipitation. The obtained RNAs were polyadenylated by poly(A) polymerase and then reverse-transcribed using the One Step PrimeScript miRNA cDNA synthesis Kit (TaKaRa, Dalian, China) with the universal adaptor primer (a poly(T) primer ligated with an adapter) for miRNA quantitative assays (Table 1). The expression levels of miRNA were quantified using real-time PCR with RPL-13 gene as the internal control.
The expression level of the myostatin gene was quantified using real-time PCR with RPL-13 as the internal control. Primers for the selected gene were designed according to the gene sequence provided by the muscle transcriptome of S. chuatsi (Table 1). Total RNAs were extracted from the muscle tissues as described earlier. Complementary DNAs (cDNAs) were synthesized by Super-Script III RNase H-Reverse Transcriptase (Invitrogen, USA) according to the manufacturer’s instructions. Negative controls contained either no cDNA template or reverse transcriptase. The cDNA samples were used as templates for quantitative RT-PCR assays with SYBR Green PCR reaction kit (Stratagene, Shanghai, China) and its amplification reaction was carried out with Stratagene Mx3005 system (Stratagene, CA, USA).
The relative expression ratio (R) of the target miRNAs and mRNA was calculated by R = 2-ΔΔCt (Livak and Thomas 2001; Bustin et al. 2009), where Ct is the cycle threshold. The basic equation employed was ΔΔCt = (Cttarget gene − Cthousekeeping gene)experiment − (Cttarget gene − Cthousekeeping gene)control.
The expression levels were analyzed by one-way ANOVA procedures and regression analysis of SPSS 17.0 (SPSS Inc., Chicago, USA). Duncan’s multiple range tests were used to compare the control (0 h before the recovery meal) and experimental (hours after refeeding) groups. The differences were considered statistically significant when p < 0.05. Data are shown as means ± SE (n = 5).
Target Prediction and Hierarchical Clustering Analysis
Target prediction was performed using Watson-Crick base-pairing between the miRNA and mRNA 3′-UTR database of Chinese perch. A heat map summary and hierarchical clustering analysis were performed using the Cluster3 software according to the similarity in their expression across different postprandial times (0 to 96 h). Correlation of the miRNA expression was analyzed by the Spearman rank order correlation test.
Results
Differential Expression of miRNAs in Adults and Juveniles
Our previous studies have shown that the 21 miRNAs are expressed with high abundance in the fast muscles of adult Chinese perch (Chu et al. 2013). To determine whether the expression of these miRNAs changes during fish growth, we compared their expression levels in fast muscles of adult and juvenile Chinese perch. The results showed that all the 21 miRNAs were detected in the fast muscle of the juvenile Chinese perch. Among them, 18 miRNAs showed significantly higher levels of expression in juveniles than adults (Fig. 1). In contrast, only one miRNA (miR-499) showed a twofold increase in adults relative to juveniles (Fig. 1). The expression of the two miRNAs, miR-23a and miR-101a, were found to be stable, with little changes between these two stages. We found that the expression of miR-1a, miR-206, miR-133a-3p, and miR-133b-3p were significantly higher than other miRNAs in the fast muscle of the juveniles (Fig. 2), while the expression of miR-499, miR-146a, miR-140-3p, miR-103, and miR-107a represented much lower levels in the fast muscle of the juveniles (Fig. 2).
The contrastive expression of miRNAs in fast muscle between juveniles and adults. RPL-13 expression was detected as the internal control. The data was expressed as the relative change compared with the expression level of each miRNAs in adults. An asterisk indicates difference more than five times when compared with the expression in adult fish. Values are the mean ± SE, n = 5
Effect of Fasting and Refeeding on the Expression of the miRNAs
It is known that fasting and refeeding has a significant impact on fish muscle growth. To determine the effect of the fasting and refeeding on miRNA expression in skeletal muscles, we analyzed the expression pattern of the 21 miRNAs at various time points after refeeding. A significant upregulation was observed for miR-10c, miR-107a, miR-133a-3p, miR-140-3p, miR-181a-5p, and miR-206 at 1 h after the single meal (p < 0.05). miR-107a, miR-133a-3p, and miR-206 reached the maximal levels at 24 h postprandially, and then decreased from 24 to 96 h (Fig. 3a–c). Interestingly, miR-140-3p reached the maximal level at 1 h after the meal administration, and then decreased from 1 to 6 h (Fig. 3f). However, the expression of miR-214 was significantly decreased 1 h after the single meal (p < 0.05), and then sustained at a low level till 96 h (Fig. 3g).
The relative expression of miRNAs with significant up- or downregulation immediately within 1 h of refeeding after fasted for 1 week (a miR-107a, b miR-133a-3p, c miR-206, d miR-10c, e miR-181a-5p, f miR-140-3p, and g miR-214). RPL-13 expression was detected as the internal control. Values are the mean ± SE, n = 5. The different letters indicate significant differences between columns (p < 0.05)
Among the 21 miRNAs analyzed, 14 showed little response to refeeding within 1 h (Fig. 4). These miRNAs include miR-1a, miR-133b-3p, miR-143, miR-22a, miR-23a, miR-26a, miR-103, miR-199-3p, miR-152a, miR-101a, miR-146a, miR-199-5p, miR-499, and miR-21 that responded slowly to refeeding, with slightly increasing or decreasing at 1 h (p > 0.05), and reached the maximal levels at 48 or 96 h postprandially (Fig. 4). Collectively, these data indicate that fasting and refeeding have different effects on the expression of various miRNAs.
Heat Map Summary of Hierarchical Clustering of miRNAs in Skeletal Muscle During Fasting-Refeeding Periods
A heat map summary and hierarchical clustering analysis were performed according to the similarity in their expression across different postprandial times (0 to 96 h). Two clades were showed by hierarchical clustering of the miRNAs throughout the trial (Fig. 5). The first clade showed six pairs of closely linked miRNAs (miR-103 and miR-107a; miR-21 and miR-1a; miR-181a-5p and miR-10c; miR-133a-3p and miR-22a; miR-26a and miR-199-3p; miR-199-5p and miR-152a). The second clade showed a close relation and covariation of miR-140-3p and miR-146a, which was subsequently clustered with miR-101a and miR-214 (Fig. 5).
Heat map summary of hierarchical clustering of miRNAs in skeletal muscle during fasting-refeeding periods. In the heat map, the green and red colors, respectively, indicate a decrease and increase. The absolute signal intensity ranges from −2.08 to +2.08, with corresponding color changes from green to red
Target Gene Analysis
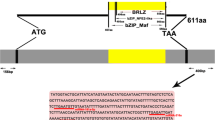
To better understand the biological significance behind the sharply up- or downregulation of miRNA expression, we conducted a search on the miRNA and mRNA database of Chinese perch by Watson-Crick base-pairing to determine whether the seven differentially expressed miRNAs targeted any of the muscle growth-related genes. We found that miR-10c, miR-107a, miR-140-3p, and miR-181a-5p were predicted to target the myostatin gene (Fig. 6). Interestingly, we showed that myostatin mRNA expression decreased within 12 h following an anabolic stimulus of refeeding in Chinese perch (Fig. 7).
Discussion
MicroRNAs (miRNAs) are a class of small, endogenous non-coding RNA molecules that repress gene expression by inhibiting translation of target mRNAs (Eisenberg et al. 2009). miRNAs play a key role in diverse biological processes such as cell proliferation, tumor genesis, fat metabolism, gonad development, and embryogenesis (Fu et al. 2011; Škugor et al. 2014). Several hundred mammalian miRNAs have been identified in vertebrate genomes, and many of them show a tissue-specific and/or temporally regulated pattern of expression (Bartel 2004). Recently, hundreds of miRNAs also have been identified in aquaculture species by Solexa deep sequencing (Salem et al. 2010; Yan et al. 2012a, b; Jiao et al. 2014). A small number of miRNAs have been shown to have a role in skeletal muscle development and growth (McCarthy and Esser 2007).
In this study, we choose 21 miRNAs with high abundance in the fast muscle of Chinese perch adults as our interest miRNAs to investigate their role in muscle development and growth. We compared the expressional levels of the 21 miRNAs at the two stages (90 and 150 dph) in Chinese perch muscle. Most of these miRNAs have a similar expression pattern in the two stages. We also found that 18 of the 21 miRNAs have higher abundance in juveniles than in adults. However, 8 of the identified 11 miRNAs were found to be stably expressed in different developmental stages in tilapia muscle (Yan et al. 2012a). The expression of miR-1, miR-133a-3p, miR-206, and miR-21 was increased in the carp skeletal muscles from 30 dph to 2 years old (Yan et al. 2012b). These researches suggested that developmental expression pattern of miRNAs was unique to the different fish species. Further investigations are needed to answer the question of the biological significance behind the different developmental expressional patterns of the miRNAs in Chinese perch, carp, and tilapia. It was striking that the Chinese perch miR-214 showed a more than 20-fold higher levels of expression in juvenile muscles than adults muscles (Fig. 2). In carp, miR-214 also expressed at relatively higher level in the skeletal muscles of the 30-dph juveniles than the 1 year adults (Yan et al. 2012b). miR-214 was first shown to play an essential role in specifying slow muscle development in zebrafish (Flynt et al. 2007). In a mouse model of cardiac hypertrophy, miR-214 was found to be expressed at an elevated level, but forced expression of miR-214 did not lead to increased muscle growth (Watanabe et al. 2008). Juan et al. (2009) identified MyoD and myogenin as the transcription factors that control the pri-miR-214 transcription, thus suggesting a positive feedback loop, in which activation of MyoD and myogenin turns on the expression of miR-214, which in turn reinforces MyoD and myogenin expression by negatively feeding back to Ezh2. Using the in vitro differentiation of mouse C2C12 cells as a model, Liu et al. (2010) found that the miR-214 promoted myogenic differentiation in mouse C2C12 myoblasts at a step preceding the induction of p21 and myogenin. The two studies demonstrate that miR-214 can function through multiple mechanisms to promote myogenic differentiation. The higher level of miR-214 in juveniles indicated that miR-214 might perform a strong effect on the muscle development of Chinese perch and carp.
Although much attention has been given to the function of miRNAs in control of muscle development, recent studies suggested that miRNAs, such as miR-499, miR-208b, and miR-23a, also had an important role in human muscle growth (Drummond et al. 2009). Huang et al. (2012) detected differentially expressed miRNA between two strains of Nile tilapia and found miR-140, miR-192, miR-204, miR-218a, miR-218b, miR-301c, miR-460, miR-133, miR-152, miR-15a, miR-193a, miR-30b, and miR-34 were associated with body growth in tilapia. Nutrient availability is among the most important environmental variable affecting muscle growth (Valente et al. 2012). Starvation and refeeding experiments have been used as the model system to study the regulation of muscle growth in fish. The effects of fasting and subsequent refeeding on mRNA transcript abundance have been studied in several fish species (Bower et al. 2008; Fuentes et al. 2012). However, very little information is available regarding the changes of early miRNA transcriptional during the postprandial period. In this study, we explored the postprandial regulation of growth-related miRNAs shortly after feeding a single meal in Chinese perch. The primary finding of this study was that miR-10c, miR-107a, miR-133a-3p, miR-140-3p, miR-181a-5p, and miR-206 were significantly elevated at 1 h after refeeding in fast muscles (Fig. 3a–f). This is consistent with a previous report by Drummond et al. (2009) showing a rapid upregulation of miR-1, miR-208b, miR-23a, and miR-499 following the amino acid ingestion in humans. These miRNAs may represent candidate miRNAs involved in a fast-response signaling system that regulates fish skeletal muscle growth. Interesting, miR-214 was the only one shown to be significantly decreased at 1 h after refeeding (Fig. 3g). Blocking miR-214 function helped C2C12 cells maintain an active cell cycle progression and contribute to myoblast proliferation (Liu et al. 2010). The short-term fasting could suppress initial proliferation of juvenile rainbow trout muscle satellite cells, after refeeding the initial proliferation rate and the changes in proliferation rates were completely restored (Fauconneau and Paboeuf 2000). Therefore, the downregulation of the Chinese perch miR-214 after refeeding might facilitate satellite cell proliferation to restore muscle cell loss during fasting stage.
Furthermore, we conducted a search on the miRNA and mRNA database of Chinese perch to determine whether the seven differentially expressed miRNAs targeted any of the muscle growth-related genes. We found that miR-10c, miR-107a, miR-140-3p, and miR-181a-5p were predicted to target the myostatin gene (Fig. 6). Myostatin is a member of the TGF-β family that is expressed nearly exclusively in the skeletal muscle, which is a powerful inhibitor of muscle growth (Amthor et al. 2007). Luciferase reporter assay verified the direct interaction between miR-181a-5p and myostatin gene in Chinese perch (Chu et al. 2013). Interestingly, we showed that myostatin mRNA expression decreased within 12 h following an anabolic stimulus of refeeding in Chinese perch (Fig. 7). These miRNAs may downregulate the myostatin gene expression and stimulate the muscle growth during 24 h after refeeding. However, making inferences between miRNA and their targets is a limitation and needs verification. MiRNA regulations of gene targets may not only occur by means of degradation but some of the gene targets may also be regulators of miRNA expression through a novel feedback mechanism (Pillai et al. 2007). In the present study, the myostatin mRNA showed a spike at 48 h after the meal, it suggested that there may be another signaling pathway (such as a feedback mechanism) to improve the expression of myostatin that limited excessive regulation of muscle growth to ensure normal growth of muscle.
The second finding of this study was the dramatic and simultaneous upregulation of miR-107a and miR-103 expression after feeding a single meal. MiR-103 and miR-107 are located within the introns of the pantothenate kinase (PANK) genes (Wilfred et al. 2007). It may suggest a congenerous function of miR-103 and miR-107 with the PANK genes to regulate fatty acid synthesis and utilization (Wilfred et al. 2007). In the results of postprandial regulation of hepatic miRNAs in rainbow trout, transient postprandial increases of miR-107a and miR-103 were consistent with a stimulatory role on the gluconeogenic pathway (Mennigen et al. 2012). These novel findings suggest that a cluster of miR-103 and miR-107a may play an important role in regulating the expression of metabolic pathways of Chinese perch muscle.
In conclusion, the present results showed that the transcription of several miRNAs in the fast skeletal muscle of Chinese perch responded quickly to a single meal after 7 days of fasting. Our findings indicate that refeeding induced a coordinated expression of several miRNAs involved in a strong resumption of myogenesis with feeding. These miRNAs may be involved in regulating fish myotomal muscle growth.
References
Amthor H, Macharia R, Navarrete R, Schuelke M, Brown SC, Otto A, Voit T, Muntoni F, Vrbova G, Partridge T, Zammit P, Bunger L, Patel K (2007) Lack of myostatin results in excessive muscle growth but impaired force generation. Proc Natl Acad Sci U S A 104:1835–1840
Ballantyne JS (2001) Amino acid metabolism. In: Wright PA, Anderson AJ (eds) Nitrogen excretion, vol 20, Fish physiology. Academic Press, San Diego, pp 77–107
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bower NI, Johnston IA (2010) Discovery and characterization of nutritionally regulated genes associated with muscle growth in Atlantic salmon. Physiol Genomics 42A:114–130
Bower NI, Li X, Taylor R, Johnston IA (2008) Switching to fast growth: the insulin-like growth factor (IGF) system in skeletal muscle of Atlantic salmon. J Exp Biol 211:3859–3870
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Carter C, Houlihan D, Kiessling A, Médale F, Jobling M (2001) Physiological effects of feeding. In Food intake in fish. Ed. by D.Houlihan., T. Boujard. and M.Jobling. Blackwell Science, UK. vol. 1, pp 297–331
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM, Conlon FL, Wang DZ (2006) The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 38:228–233
Chu WY, Fu GH, Chen J, Chen DG, Meng T, Zhou RX, Xj X, Zhang JS (2010) Gene expression profiles of the muscle tissues of the commercial important teleost, Siniperca chuatsi L. Aquacult Int 18:667–678
Chu WY, Liu LS, Li YL, Chen L, Wang KZ, Li HH, Du SJ, Zhang JS (2013) Systematic identification and differential expression profiling of microRNAs from white and red muscles of Siniperca chuatsi. Curr Mol Med 13:1397–1407
Drummond MJ, Glynn EL, Fry CS, Dhanani V, Shaheen E, Asmussen BB (2009) Essential amino acids increase microRNA-499, -208b, and -23a and downregulate myostatin and myocyte enhancer factor 2C mRNA expression in human skeletal muscle. J Nutr 139:2279–2284
Eisenberg I, Alexander MS, Kunkel LM (2009) miRNAs in normal and diseased skeletal muscle. J Cell Mol Med 13:2–11
Fauconneau B, Paboeuf G (2000) Effect of fasting and refeeding on in vitro muscle cell proliferation in rainbow trout (Oncorhynchus mykiss). Cell Tissue Res 301:459–463
Flynt AS, Li N, Thatcher EJ, Solnica-Krezel L, Patton JG (2007) Zebrafish miR-214 modulates Hedgehog signaling to specify muscle cell fate. Nat Genet 39:259–263
Fu YS, Shi ZY, Wu ML, Zhang JL, Jia L, Chen XW (2011) Identification and differential expression of microRNAs during metamorphosis of the Japanese flounder (Paralichthys olivaceus). PLoS One 6:e22957
Fuentes EN, Ruiz P, Valdes JA, Molina A (2012) Catabolic signaling pathways, atrogenes, and ubiquitinated proteins are regulated by the nutritional status in the muscle of the fine flounder. PLoS One 7:e44256
Huang CW, Li YH, Hu SY, Chi JR, Lin GH, Lin CC, Gong HY, Chen JY, Chen RH, Chang SJ, Liu FG, Wu JL (2012) Differential expression patterns of growth-related microRNAs in the skeletal muscle of Nile tilapia(Oreochromis niloticus). J Anim Sci 90:4266–4279
Jiao Y, Zheng Z, Du X, Wang Q, Huang R, Deng Y, Shi S, Zhao X (2014) Identification and characterization of microRNAs in pearl oyster Pinctada martensii by Solexa deep sequencing. Mar Biotechnol (NY) 16:54–62
Juan AH, Kumar RM, Marx JG, Young RA, Sartorelli V (2009) Mir-214-dependent regulation of the polycomb protein Ezh2 in skeletal muscle and embryonic stem cells. Mol Cell 36:61–74
Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A (2006) Muscle specific microRNA miR-206 promotes muscle differentiation. J Cell Biol 174:677–687
Liu J (2008) Control of protein synthesis and mRNA degradation by microRNAs. Curr Opin Cell Biol 20:214–221
Liu J, Luo XJ, Xiong AW, Zhang ZD, Yue S, Zhu MS, Cheng SY (2010) MicroRNA-214 promotes myogenic differentiation by facilitating exit from mitosis via down-regulation of proto-oncogene N-ras. J Biol Chem 285:26599–26607
Liu L, Li C, Su B, Beck BH, Peatman E (2013) Short-term feed deprivation alters immune status of surface mucosa in channel catfish (Ictalurus punctatus). PLoS One 8:e74581
Livak KJ, Thomas DS (2001) Analysis of relative gene expression data using real time quantitative PCR and the 2-ΔΔCt method. Methods 25:402–408
McCarthy JJ, Esser KA (2007) MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J Appl Physiol 102:306–313
Mennigen JA, Pansera TS, Larquier M, Plagnes-Juan E, Medale F (2012) Postprandial regulation of hepatic microRNAs predicted to target the insulin pathway in rainbow trout. PLoS One 7:e38604
Nihei Y, Kobiyama A, Ikeda D, Ono Y, Ohara S, Cole NJ, Johnston IA, Watabe S (2006) Molecular cloning and mRNA expression analysis of carp embryonic, slow and cardiac myosin heavy chain isoforms. J Exp Biol 209:188–198
Panserat S, Kaushik SJ (2010) Regulation of gene expression by nutritional factors in fish. Aquacult Res 41:751–762
Peragón J, Barroso JB, García-Salguero L, Aranda F, de la Higuera M, Lupiáñez JA (1999) Selective changes in the protein-turnover rates and nature of growth induced in trout liver by long-term starvation followed by re-feeding. Mol Cell Biochem 201:1–10
Pillai RS, Bhattacharyya SN, Filipowicz W (2007) Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell Biol 17:118–126
Salem M, Xiao C, Womack J, Rexroad CE 3rd, Yao J (2010) A microRNA repertoire for functional genome research in rainbow trout (Oncorhynchus mykiss). Mar Biotechnol (NY) 12:410–429
Škugor A, Slanchev K, Torgersen JS, Tveiten H, Andersen Ø (2014) Conserved mechanisms for germ cell-specific localization of nanos3 transcripts in teleost species with aquaculture significance. Mar Biotechnol (NY) 16:256–264
Steinbacher P, Haslett JR, Six M, Gollmann HP, Sänger AM, Stoiber W (2006) Phases of myogenic cell activation and possible role of dermomyotome cells in teleost muscle formation. Dev Dyn 235:3132–3143
Valente LM, Bower NI, Johnston IA (2012) Postprandial expression of growth related genes in Atlantic salmon (Salmo salar L.) juveniles fasted for 1 week and fed a single meal to satiation. Br J Nutr 108:2148–2157
Watanabe T, Sato T, Amano T, Kawamura Y, Kawamura N, Kawaguchi H, Yamashita N, Kurihara H, Nakaoka T (2008) Dnm3os, a non-coding RNA, is required for normal growth and skeletal development in mice. Dev Dyn 237:3738–3748
Wilfred BR, Wang WX, Nelson PT (2007) Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol Genet Metab 91:209–217
Xie X, Lu J, Kulbokas EJ (2005) Systematic discovery of regulatory motifs in human promoters and 3′UTRs by comparison of several mammals. Nature 434:338–345
Yan B, Guo JT, Zhao LH, Zhao JL (2012a) MicroRNA expression signature in skeletal muscle of Nile tilapia. Aquaculture 364–365:240–246
Yan XC, Ding L, Li YC, Zhang XF, Liang Y, Sun XW, Teng CB (2012b) Identification and profiling of microRNAs from skeletal muscle of the common carp. PloS One 7:e30925
Zhang JS, Fu GH, Chu WY, Chen J, Liu Z, Liu F, Lu SQ, Liang P (2009) cDNA cloning and expression analysis of myosin heavy chain gene (MHC) of the Mandarin fish, Sniperca kneri. Aquac Res 40:412–418
Zhang QG, Chu WY, Hu SN, Meng T, Pan LL, Zhou RX, Liu Z, Zhang JS (2011) Identification and analysis of muscle-related protein isoforms expressed in the white muscle of the mandarin fish (Siniperca chuatsi). Mar Biotechnol (NY) 13:151–162
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 31230076; 31340054) and the Natural Science Foundation of Hunan province (14JJ2135).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhu, X., Chen, D., Hu, Y. et al. The microRNA Signature in Response to Nutrient Restriction and Refeeding in Skeletal Muscle of Chinese Perch (Siniperca chuatsi). Mar Biotechnol 17, 180–189 (2015). https://doi.org/10.1007/s10126-014-9606-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-014-9606-8