Abstract
Chordomas and chondrosarcomas are two major malignant bone neoplasms located at the skull base. These tumors are rarely metastatic, but can be locally invasive and resistant to conventional chemotherapies and radiotherapies. Accordingly, therapeutic approaches for the treatment of these tumors can be difficult. Additionally, their location at the skull base makes them problematic. Although accurate diagnosis of these tumors is important because of their distinct prognoses, distinguishing between these tumor types is difficult due to overlapping radiological and histopathological findings. However, recent accumulation of molecular and genetic studies, including extracranial location analysis, has provided us clues for accurate diagnosis. In this report, we review the genetic aberrations and molecular biology of these two tumor types. Among the abundant genetic features of these tumors, brachyury immunohistochemistry and direct sequencing of IDH1/2 are simple and useful techniques that can be used to distinguish between these tumors. Although it is still unclear why these tumors, which have such distinct genetic backgrounds, show similar histopathological findings, comparison of their genetic backgrounds could provide essential information.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chordomas and chondrosarcomas are two major malignant bone neoplasms occurring at the skull base. Although most of these tumors are slow growing and rarely metastasize, they are locally invasive, highly recurrent, and potentially lethal. Because these tumors are mostly resistant to conventional chemotherapies and radiotherapies, surgical resection plays a crucial role in the treatment of these tumors. However, for chordoma and chondrosarcoma located at the skull base, radical resection is rarely achieved because of challenges associated with the location of the tumors.
In the clinical setting, it is important to distinguish between these two tumor types because they have different prognoses [1, 2]. However, this can be challenging owing to their overlapping radiological and histopathological findings. Indeed, chondroid chordoma, a subtype of chordoma, is a matrix-mimicking cartilaginous tumor [3]. Additionally, in a previous report, 37% (74/200 cases) of skull base chondrosarcomas were initially misdiagnosed as a chordoma [4].
Although chordoma and chondrosarcoma, including those located extracranially, are not as common as other types of cancer, researchers have begun to carry out molecular and genetic studies of these tumors. In this report, we review the genetic aberrations and molecular biology of chordoma and chondrosarcoma, including extracranially located tumors. Because these two types of tumors have distinct genetic backgrounds, molecular and genetic examinations of these tumors are expected to provide useful clues for distinguishing between the two tumor types.
Chordoma
Chordoma accounts for 1–4% of all bone malignancies [5] and 0.5% of primary intracranial central nervous system (CNS) tumors [6]. Chordoma frequently occurs in the sacrum, vertebral body, and skull base, with each of these locations accounting for approximately one-third of all chordomas [7]. Chordoma is thought to be derived from undifferentiated notochordal remnants; this is strongly supported by the fact that aberrations in the T gene, which encodes an important transcription factor involved in notochord development, were detected in the germ-line of familial chordoma [8]. The current common therapeutic strategy for the treatment of chordomas is maximal surgical resection followed by unconventional radiotherapy, such as proton beam therapy and carbon ion radiotherapy [9]. Incomplete surgical resection and the absence of postoperative irradiation significantly contribute to the poor prognosis of chordoma [10–13]. Because chordomas are resistant to conventional chemotherapies, researchers have attempted using molecular targeting agents for chordoma treatment.
T (brachyury)
In genetic analysis of chordoma, the T gene, located on chromosome 6q27, and its protein product brachyury, have been extensively studied. T was originally discovered as a site of mutation in the short-tailed mouse and has been shown to act as a transcription factor in notochord development [14]. Additionally, a recent study showed that brachyury is an essential factor for maintaining notochord cell fate and function [15]. Although brachyury is thought to function in the epithelial-to-mesenchymal transition (EMT) of malignant tumors [16, 17], a recent report showed that T function is dispensable for the EMT [15].
Brachyury expression is most frequently observed in chordomas (81–100%); it is less frequent in germ cell tumors and small cell lung cancers, but is rarely present in other types of tumor [12, 18–22]. Immunohistochemical analysis of brachyury is a useful biomarker for distinguishing among similar tumors, such as chondrosarcomas, chordoid meningiomas, and carcinomas, because of its high sensitivity and specificity. Furthermore, in case of skull base tumors, brachyury expression is a prognostic factor; patients having chordomas with brachyury expression exhibit significantly shorter progression-free survival (PFS) than patients having chordomas without brachyury expression [12].
Notably, researchers identified a duplication at the T site in the germ-line of familial chordomas [8]. Furthermore, Pillay et al. reported that the common nonsynonymous single nucleotide polymorphism (SNP) rs2305089 on the T gene was strongly associated with chordoma risk [23]. These reports suggested that T plays an important role in the tumorigenesis of chordoma. Presneau et al. reported that chromosomal aberrations resulting in gain of T were common in some sporadic chordomas, and that the downregulation of T using short hairpin RNA (shRNA) in chordoma cell lines decreases cell proliferation and enhances morphological features consistent with a senescence-like phenotype [24]. Additionally, Hsu et al. reported that silencing of brachyury using shRNA led to complete growth arrest in other cell lines [25]. Nelson et al. showed that brachyury activated oncogenic transcription through binding directly to 99 target genes and indirectly affecting the expression of 64 other genes [26]. Based on these studies, the T gene (encoding brachyury) is now a promising therapeutic target for the treatment of chordomas [17, 27].
SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily B, member 1 (SMARCB1)/INI
Generally, aberrations in SMARCB1/INI1, located on chromosome 22q11.2, predispose patients to rhabdoid tumors and schwannomatosis. Mutations and deletions in SMARCB1/INI1 have been detected in several pediatric chordoma cases, accompanied by brachyury expression [28–31]. Recently, Hasselblatt et al. reported that these chordomas had a distinct molecular background from conventional chordomas [32], showing poor differentiation, lack of SMARCB1 expression, lack of complex chromosomal alterations as found in conventional chordomas, onset at an early age (young children), and poor prognoses. Furthermore, these tumors had methylation profiles distinct from those of atypical teratoid/rhabdoid tumors, which are commonly found in young children, and lacked SMARCB1 expression. These observations strongly supported the existence of a subtype of chordoma.
Receptor tyrosine kinases (RTKs)
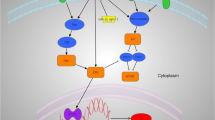
RTKs play an important role in malignant transformation and tumor proliferation in cancers. Many reports have shown that RTKs, such as epidermal growth factor receptor (EGFR) [33–37], platelet-derived growth factor receptor-α (PDGFRα) [33, 37–39], PDGFRβ [35, 37–39], fibroblast growth factor receptor (FGFR) [40], hepatocyte growth factor receptor (MET) [33], KIT [39], p75 receptor [41], tropomyosin-related kinase A (TrkA) [41], and insulin-like growth factor-1 receptor (IGF-1R) [42, 43], are frequently overexpressed and/or activated in chordoma. Additionally, vascular endothelial growth factor (VEGF) [44] and nerve growth factor (NGF) were shown to be overexpressed in chordoma [41]. Although polysomy and amplification of EGFR were frequently detected in chordomas by fluorescence in situ hybridization [36], no EGFR mutation was detected [36, 45]. Scheipl et al. reported the efficacy of EGFR inhibitors, such as sapitinib, gefitinib, and erlotinib, by demonstrating that these compounds suppressed the phosphorylation of EGFR and activation of its downstream pathways in chordoma cell lines [45]. Sommer et al. reported that patients with chordoma who were positive for phospho-IGF-1R had significantly shorter median disease-free survival [43].
Major downstream pathways
Akt/phosphoinositide 3-kinase (PI3K)/mammalian target of rapamycin (mTOR) pathway
The Akt/PI3K/mTOR pathway is activated in various cancers, resulting in hyperproliferation and tumor growth. Phosphoinositide-dependent kinase-1 (PDK1) [46], Akt [34, 35, 43, 46, 47], tuberous sclerosis complex 2 (TSC2) [47], mTOR [47], s6 ribosomal protein (s6) [47], and phosphatase and tensin homolog (PTEN) [48] are dysregulated frequently in chordomas. Schwab et al. reported that PI-103, an inhibitor of Akt and mTOR, blocked proliferation and induced apoptosis in a chordoma cell line [46]. Recently, Tauziède-Espariat reported that PIK3CA, which is commonly mutated or amplified in various malignant neoplasms but never detected in chordoma, was mutated in two chordoma cases [44].
Mitogen-activated protein kinase (MAPK)/extracellular signal-regulated kinase (ERK) pathway
Activation of the MAPK pathway is common in various types of cancers and has been detected in several studies of chordoma [35, 40, 43]. However, no mutations in KRAS, NRAS, HRAS, and BRAF have been detected [36, 40]. Long et al. reported that miR-149-3p, miR-663a, miR-1908, miR-2861, and miR-3185 were likely to play important roles in dysregulation of the MAPK signaling pathway, leading to chordoma development [49].
Janus kinase (JAK)/signal transducer and activator of transcription (STAT) pathway
The JAK/STAT pathway is known to be active in several human cancers and associated with a poor prognosis. Activation of components of the JAK/STAT pathway, such as STAT3 and the proto-oncogene tyrosine kinase Src, has been detected in chordomas [35, 50, 51]. Yang et al. reported that STAT3 inhibitors strongly blocked cell growth and proliferation in chordoma cell lines [50] and suggested that inhibition of the JAK/STAT pathway may represent a potential therapeutic strategy for the treatment of chordoma.
Retinoblastoma (RB) pathway
The RB pathway plays an important role in the control of cell proliferation. Loss of CDKN2A/p16 with or without the loss of CDKN2B has frequently been observed in chordoma [48, 52], and one case with promoter methylation of CDKN2A/p16 has been reported [48].
Other biomarkers
Triana et al. reported that the downregulation of E-cadherin and upregulation of N-cadherin were correlated with the worse prognosis of clival chordomas [53]. Schoenfeld et al. reported that positive expression of chondroitin sulfate proteoglycan 4 (CSPG4), a membrane-bound proteoglycan expressed in several types of malignant tumors, was associated with a higher risk of mortality and an increased risk of metastasis in chordomas [54]. The expression of fragile histidine triad protein (FHIT), a potential tumor suppressor, was absent or reduced in 98% of sacral chordomas and 67% of skull base chordomas, and the authors suggested that chromosome 3 aneuploidy and epigenetic regulation of FHIT contributed to loss of the FHIT tumor suppressor in chordoma [55]. Sa et al. reported recurrent somatic variants, including single nucleotide variations in MUC4, NBPF1, and NPIPB15 and the gene fusion of SAMD5-SASH1, in whole-exome and whole-transcriptome sequencing of 13 chordomas [56]. Rinner et al. reported that 20 genes were hyper-/hypomethylated in samples from patients with chordoma compared with those in normal blood; among these epigenetically regulated genes, C3, XIST, TACSTD2, FMR1, DLEC1, RARB, HIC1,, KL, and RASSF1 were suggested to be the most promising candidate genes [57], with the latter three identified as tumor-suppressor genes. Alholle et al. reported several genes (FAM181B, KANK2, NPR3, PON3, RAB32, RAI1, SLC16A5, and ZNF397OS) that were differentially methylated between recurrent cases and nonrecurrent cases [58]; among these genes, KANK2 has been shown to be a candidate tumor-suppressor gene.
Cytogenetics
Many researchers have evaluated chromosomal copy numbers in patients with chordoma. These studies have reported similar findings, including losses on chromosomes 1p, 3, 4, 9, 10, 13, 14, and 18 and gains on chromosomes 1q and 7 [12, 48, 56, 59–62]. Of these chromosomes, 1p has been the most intensely investigated chromosome arm, and several studies have suggested that 1p36 may be a putative tumor-suppressive locus in chordomas, as supported by the finding that loss of 1p36 is associated with poor prognosis [63–65]. Longoni et al. listed tumor necrosis factor (TNF) receptor superfamily member 8 (TNFRSF8) as a candidate gene on 1p36. Sawyer et al. reported that isochromosome 1q and monosomy 13 were frequent structural abnormalities in skull base chordomas by spectral karyotyping, supporting the hypothesis that chromosome 1p was a tumor-suppressive locus [66]. Horbinski et al. suggested that loss of heterozygosity (LOH) on chromosome 9p was significantly associated with shorter survival in patients with skull base chordomas [11]. We previously conducted whole genome analyses by comparative genomic hybridization and performed multivariate analyses with genetic and clinical factors; these studies suggested that gain on chromosome 2p was correlated with the prognosis [12].
MicroRNAs (miRNAs)
Several studies have suggested that aberrant miRNAs may play roles in tumorigenesis or progression of chordomas. Duan et al. reported that miR-1 and miR-206 were downregulated in chordoma-derived cell lines and chordoma tissue, and miR-1 expression was inversely correlated with MET expression [67]. In a follow-up study, they showed that miR-1 expression was correlated with poor prognosis and that induction of miRNA expression suppressed MET expression and inhibited the growth of chordoma cells [68]. Moreover, Osaka et al. reported that overexpression of Slug, a target of miR-1, promoted cell proliferation in chordomas [69]. The same team reported that miR-155 was downregulated in chordomas, resulting poor prognosis [70]. Moreover, Zou et al. reported that overexpression of miR-140-3p and downregulation of miR-1237-3p were associated with chordoma invasion and poor prognosis in spinal chordomas [71, 72]. Gulluoglu et al. assessed the functions of miR-31, miR-140-3p, miR-148a, and miR-222 by transfecting these miRNAs into chordoma cell lines transiently. These miRNAs were found to target proteins such as MET, MAPK1(ERK2), BCL2L11, and KIT, suggesting that these miRNAs play roles in cell viability, cell cycle, and apoptosis in chordomas [73]. Additionally, as described above, miR-149-3p may facilitate chordoma development through the dysregulation of the MAPK signaling pathway [49].
Immune system biomarkers
Checkpoints of programmed cell death protein 1 (PD-1), programmed death-ligand 1 (PD-L1), and PD-L2 have been studied in various malignant tumors, and the expression of these proteins has been implicated in promoting tumor progression. Mathios et al. showed that PD-L1 and PD-L2 were not constitutively expressed in chordoma cell lines, but could be induced by pro-inflammatory cytokines [74]. Using paraffin-embedded tissues, researchers also demonstrated that PD-1 expression could be detected in tumor-infiltrating lymphocytes (TILs) in some cases of chordoma and that PD-L1 was not expressed in chordoma cells but was expressed in tumor-infiltrating macrophages and TILs [74, 75]. Zou et al. reported that the expression of PD-L1 in TILs was associated with a favorable prognosis in spinal chordoma [76]. However, further studies are needed to elucidate the detailed mechanisms and roles of immune checkpoint molecules in chordomas.
Molecular-targeted therapy
Target therapies for these proteins have been carried out using cetuximab (an EGFR inhibitor) [77, 78], gefitinib (an EGFR inhibitor) [77, 78], erlotinib (an EGFR inhibitor) [79–81], lapatinib (an inhibitor of EGFR and HER2) [82], imatinib (an inhibitor of PDGFR, c-Kit, and ABL) [83–85], sirolimus (rapamycin; an mTOR inhibitor) [85, 86], dasatinib (an inhibitor of Src, c-Kit, and ABL) [87], and bevacizumab (a VEGF inhibitor) [79]. Among these reports, the study with the largest number of participants was a phase II study of imatinib in 50 PDGFβ/PDGFRβ-positive patients with advanced chordoma [84]. The results showed that one patient (2%) had a partial response (PR), whereas 35 patients (70%) had stable disease (SD) at 6 months. Hindi et al. reported a retrospective series of 48 PDGFβ/PDGFRβ-positive patients with advanced chordoma treated with imatinib [83]; no patients achieved PR, and 34 patients (74%) showed SD. Stacchiotti et al. tested the efficacy of imatinib plus sirolimus for nine patients with imatinib-resistant advanced chordoma [85]; one patient achieved PR, and seven patients showed SD. Additionally, a phase II trial of lapatinib with 18 patients with EGFR-positive advanced chordoma showed that six patients (33%) achieved PR, whereas seven patients (39%) showed SD at 6 months [82]. A phase II trial of dasatinib in patients with advanced chordoma showed that six patients had an objective tumor response based on Choi criteria [87]. Furthermore, bevacizumab plus erlotinib was administered to three patients with chordoma, resulting in SD for 2–4.5 years [79]. Clinical trials using new targets, including therapeutic vaccines targeting brachyury, PD-1, PD-L1 inhibitors, are planned or underway [88].
Chondrosarcoma
Chondrosarcoma is the third most frequent primary malignancy of the bone after myeloma and osteosarcoma [3]; it is systemically more frequent than chordoma. However, intracranial chondrosarcomas comprise only approximately 1% of all chondrosarcomas [89]; consequently, it is less frequent than chordoma in cases of intracranial location [90]. Chondrosarcomas are graded on a scale of I–III based on histological findings [3], reflecting the rates of local recurrence and metastasis. Almost all chondrosarcomas are grades I or II, and grade III is extremely rare (grade I: 61%, grade II: 36%, grade III: 3%) [91]. A similar distribution has been observed in skull base chondrosarcoma (grade I: 50.5%, contained areas of grade I and II: 28.5%, grade II: 21%, grade III: 0%) [4]. Generally, low-grade chondrosarcomas are locally invasive, but rarely metastasize; thus, surgical resection is the best approach for managing this disease [92]. For chondrosarcomas located in the skull base, however, radical resection is rarely achieved because of the difficulties associated with resection of tumors in this location [4]. Because conventional chemotherapy and radiotherapy are usually ineffective for chondrosarcomas, systemic therapies, including molecular-targeted therapies, have been attempted.
Mutations in isocitrate dehydrogenase 1 (IDH1) and IDH2
Recent molecular studies showed that there are two major groups of chondrosarcomas with distinct genetic backgrounds: central and periosteal chondrosarcomas, which are related to mutations in IDH1 or IDH2; and peripheral chondrosarcoma, which is related to exostosin-1 (EXT1) or EXT2 inactivation [3]. Because IDH1/2 mutations have been detected in many cases of skull base chondrosarcoma [93, 94], these chondrosarcomas are thought to be molecularly consistent with a subset of central chondrosarcomas.
Somatic mutations in IDH1/2 have been identified in gliomas and acute myeloid leukemia (AML). IDH1/2 mutations are thought to play a tumorigenic role in the development of these tumors. Amary et al. detected mutant IDH1/2 in central chondrosarcoma, periosteal chondrosarcoma, and enchondroma [95]. The frequency of IDH1/2 mutations was reported to be 55–66% in central chondrosarcoma [95, 96]. In skull base chondrosarcomas, these mutations were detected in 10 of 20 cases (50%) [93, 94]. Although the vast majority (>90%) of total IDH1 mutations detected in gliomas involved the R132H substitution, R132H was detected only in 17% of cartilaginous tumors; other substitutions are common in cartilaginous tumors, such as R132C, R132G, R132L, and R132S [93–95]. Accordingly, although immunohistochemical analysis of IDH1 R132H is a useful tool for detection of mutated IDH1 in glioma, this is not the case in chondrosarcoma [3]. Li et al. reported the efficiency of a mutant IDH1 inhibitor in human chondrosarcoma cell lines [97], and phase I/II trials of an IDH1/2 inhibitor for IDH-mutated chondrosarcomas are ongoing [98].
RTKs
RTKs and their ligands have also been extensively investigated in chondrosarcoma and have been shown to play important roles in the progression of chondrosarcoma, including EGFR [99, 100], PDGFR [101], IGF-1 [102, 103], and VEGF [104, 105]. EGFR is expressed in chondrosarcoma, and gefitinib markedly inhibits the growth of chondrosarcoma cell lines [99]. Sulzbacher et al. reported that PDGFR expression in conventional chondrosarcoma was positively correlated with aggressiveness and that PDGFR may be a potential therapeutic target [101]. IGF-1 has been reported to positively regulate mitotic and matrix synthetic activities in chondrosarcoma and to play a role in the progression from chondroma to chondrosarcoma [102, 103]. However, a recent study indicated that the IGF pathway was not expected to be an effective therapeutic target of chondrosarcoma because it was not essential for chondrosarcoma growth, migration, or chemoresistance [106].
Major downstream pathways
RB pathway
Homozygous deletion, methylation, and missense mutations in CDKN2A/p16 have been detected in central chondrosarcomas [96, 107]. Furthermore, aberrations in CDK4, CDK6, and Cyclin D1 were also reported [96, 108]. The absence of RB and CDKN2A/p16 expression is strongly correlated with higher grade of central chondrosarcoma, implying that loss of RB and CDKN2A/p16 function is an important event during central chondrosarcoma progression [108, 109]. Schrage et al. reported that overexpression of CDKN2A/p16 decreased cell viability and proliferation of chondrosarcoma in vitro [108].
Hedgehog signaling
Tarpery et al. reported that the Indian hedgehog (IHH) signaling pathway is involved in chondrosarcomas [96]. Tiet et al. showed that hedgehog signaling is activated in chondrosarcomas and it plays an important role in tumor cell proliferation. Moreover, treatment with triparanol, an inhibitor of hedgehog signaling, results in decreased tumor volume, cellularity, and proliferation rates in xenografts of human chondrosarcoma in mice [110]. Furthermore, IPI-926 (saridegib; a hedgehog inhibitor) inhibits the hedgehog pathway and blocks tumor growth in chondrosarcoma xenografts in mice [111]; however, in a phase II trial, saridegib did not show efficacy in patients with chondrosarcoma [112]. miR-30a has also been shown to be downregulated in chondrosarcoma, promote cell proliferation via the RUNX2 expression, and encode runt-related transcription factor 2, which is closely linked to IHH signaling [113]; accordingly, miR-30a/RUNX2 may also represent a therapeutic target in the treatment of chondrosarcoma.
Akt/PI3K/mTOR pathway
Akt, mTOR, and s6 are also activated in chondrosarcomas [100, 114, 115]. Treatment with BEZ235 (a PI3K/mTOR inhibitor) significantly reduced the growth of chondrosarcoma cell lines [100]. The efficacy of everolimus (an mTOR inhibitor) in the inhibition of cell proliferation and tumor progression has been demonstrated using rats [116].
JAK/STAT pathway
The Src kinase family is also activated in chondrosarcomas [115]. Additionally, dasatinib has been shown to decrease cell proliferation in seven of nine cell lines and primary cultures [115]. Hypoxia-inducible factor 1α (HIF1α), which is induced by Src and Akt, is expressed in high-grade central chondrosarcoma and may facilitate chemoresistance and radioresistance, leading to poor survival [117].
MAPK/ERK pathway
NRAS mutations have been identified in chondrosarcoma cell lines and six patient tissues (12%) [100]. Importantly, the MEK inhibitor ARRY-142886 was effective at inhibiting cell growth only in cell lines with NRAS mutations and was not effective in other cell lines.
Other biomarkers
Tarpery et al. reported that hypermutability of the major cartilage collagen gene COL2A1, i.e., insertions, deletions, and rearrangements, occurred in 37% of 49 chondrosarcoma cases [96]; it is the second most frequent mutation in chondrosarcoma. The authors also reported the TP53 was mutated in 20% of cases [96]. Oshiro et al. reported that overexpression and/or structural alterations in TP53 were observed in 38.1% of 158 chondrosarcomas and were correlated with aggressive behavior [118]. van Oosterwijk et al. reported that Bcl-2, which regulates cell death, was expressed in chondrosarcomas and caused chemoresistance to doxorubicin and cisplatin in vitro [119]. Hypomethylation of maspin and 14-3-3σ was detected in chondrosarcoma lines. These genes are epithelial-specific markers that play roles in the mesenchymal-to-epithelial transition during chondrosarcoma development [120]. Bui et al. reported that the 3-O-sulfotransferase gene, which encodes a heparan sulfate biosynthetic enzyme, was abnormally hypermethylated, resulting in altered heparan sulfate proteoglycan sulfation and the invasive phenotype in chondrosarcomas [121]. Jin et al. reported that RUNX3, which plays a role in both normal developmental processes and carcinogenesis of the bone, was downregulated in specimens from patients with chondrosarcoma due to hypermethylation of its promoter, resulting in higher proliferation and lower apoptosis rates [122].
Cytogenetics
Although the results of various studies have differed substantially, frequent chromosomal alterations in extracranial chondrosarcomas include gains on chromosomes 2p, 5p, 7, 8q, 14q, 19, 20, and 21q and losses on chromosomes 4q, 6q, 9p, 13q, and 17 [123–128]. Among these aberrations, 8q24.1-qter and 14q24-qter were found to be correlated with shorter overall survival in patients with chondrosarcomas [125]; loss on chromosome 6 and gain on chromosome 12q12 were associated with high-grade chondrosarcomas in one report [128], and losses on chromosomes 5q14.2-q21.3, 6q16-q25.3, 9p24.2-p12, and 9p21.3 were associated with high-grade chondrosarcomas in another [124]. In skull base chondrosarcomas, gains on chromosomes 2q22-q32, 5qcen-q14, 8q21-q22, 15qcen-q14, and 19 were detected, suggesting the absence of distinct chromosomal alterations specific to the skull base localization of the tumors [94].
miRNAs
Many reports have described the expression and roles of miRNAs in chondrosarcomas, most within the last few years. miR-30a and miR-335 are downregulated in chondrosarcomas, resulting in overexpression of SRY-related HMG box (SOX) 4, a member of the SOX gene family, which is related to metastasis and is a poor prognostic factor for low-grade chondrosarcoma [129, 130]. SOX9, which is overexpressed in chondrosarcomas, is induced by downregulation of miR-145 and miR-494 [131, 132]. Additionally, miR-181a, which plays a role in hypoxic regulation and enhances the expression of VEGF, is overexpressed in chondrosarcoma [104]. Tsai et al. reported that miR-519d expression was downregulated in chondrosarcoma, resulting in activation of p38, which is related to tumorigenesis and metastasis [133]. Aili et al. reported that miR-10b was significantly downregulated in chondrosarcomas, resulting in chondrosarcoma cell migration and invasion through the overexpression of brain-derived neurotrophic factor (BDNF) [134]; thus, miR-10b/BDNF may be potential therapeutic targets for chondrosarcoma.
Immune system biomarkers
PD-L1 expression was absent in conventional chondrosarcomas (n = 119), but was detected in some dedifferentiated chondrosarcomas [135]. In dedifferentiated chondrosarcomas, PR for anti-PD1 therapy with nivolumub was observed [136].
Molecular-targeted therapies
A tyrosine kinase inhibitor [137, 138], mTOR inhibitor [138, 139], hedgehog inhibitor [140], and Akt inhibitor [112] have been evaluated for applications in the treatment of chondrosarcomas. Additionally, clinical trials of IDH inhibitors and anti PD-1 antibodies are ongoing [98, 141]. Although imatinib has been tested for application in recurrent chondrosarcomas with PDGFRα or PDGFRβ expression as a phase II trial, it failed to show meaningful effects in terms of both obvious responses and blocking progression in patients [137]. A phase II trial of cixutumumab and temsirolimus (inhibitors of IGF-1R and mTOR, respectively) showed that longer PFS occurred in patients with chondrosarcomas showing IGF-1R expression than those without IGF-1R expression [138]. Two phase II studies using multikinase inhibitors (regorafenib [142] and pazopanib [143]) are ongoing. A phase II trial of sirolimus plus cyclophosphamide for 10 recurrent unresectable chondrosarcomas resulted in one patient achieving PR and six patients showing SD for at least 6 months [144]. A phase I/II study of temsirolimus and liposomal doxorubicin for sarcomas including chondrosarcoma is currently ongoing [139]. Furthermore, a phase II trial of perifosine (an Akt inhibitor) with 33 patients showed that 3% achieved PR (according to Choi criteria) and 25% showed SD [112]. A phase II trial of GDC-0449 (vismodegib; a hedgehog inhibitor) in 45 patients with progressive advanced chondrosarcoma showed no objective response and 10 patients with SD for at least 6 months [140], whereas IPI-926 did not show clinical benefit [112].
Distinguishing between chordoma and chondrosarcoma
The genetic differences between chordoma and chondrosarcoma are summarized in Table 1. Traditionally, immunohistochemical analysis of epithelial markers, such as cytokeratin and epithelial membrane antigen (EMA), has been used to distinguish between chordoma and chondrosarcoma. However, some cases are extremely difficult to judge because of intermediate staining or inconsistencies among these markers. As described above, immunohistochemical analysis of brachyury has become a useful tool for differentiation because of the high positive rate in chordoma (81–100%) and lack of expression in chondrosarcoma [12, 18–22]. Additionally, IDH1/2 mutations are useful biomarkers to distinguish between these tumors objectively. Because the R132H substitution in IDH1 is not frequently observed in chondrosarcoma [93–95], the usefulness of immunohistochemical analysis of IDH1 R132H is limited. As an alternative to immunohistochemical analysis, direct sequencing of IDH1/2 could be a simple and useful tool for distinguishing between these tumors because of the higher positive rate for detecting IDH1/2 mutations in central chondrosarcoma (55–66%) [95, 96]. In previous studies of chordomas, wild-type IDH1/2 was detected in all 89 cases [94, 95]. In gliomas, genetic findings often provide a better reflection of prognosis than morphological findings, and the importance of genetic findings has been emphasized in the latest World Health Organization (WHO) classification of CNS tumors [145]. Similarly, the results of immunohistochemical examination of brachyury and direct sequencing of IDH1/2 may provide a more accurate reflection of the prognosis than morphological findings.
Conclusions
In this review, we discussed the genetic profiles of the two major bone tumors of the skull base, chordoma and chondrosarcoma. It is still unclear why these tumors, which have such distinct backgrounds, exhibit highly similar histopathological findings, and comparisons of their genetic backgrounds could provide important insights into the distinct pathologies of these tumors.
References
Almefty K, Pravdenkova S, Colli BO, Al-Mefty O, Gokden M (2007) Chordoma and chondrosarcoma: similar, but quite different, skull base tumors. Cancer 110:2457–2467. doi:10.1002/cncr.23073
Bohman LE, Koch M, Bailey RL, Alonso-Basanta M, Lee JY (2014) Skull base chordoma and chondrosarcoma: influence of clinical and demographic factors on prognosis—a SEER analysis. World Neurosurg. doi:10.1016/j.wneu.2014.07.005
Fletcher CDM, Bridge JA, Hogendoorn P, Mertens F (2013) WHO classification of tumours of soft tissue and bone. Fourth Edition. IARC, Lyon
Rosenberg AE, Nielsen GP, Keel SB, Renard LG, Fitzek MM, Munzenrider JE, Liebsch NJ (1999) Chondrosarcoma of the base of the skull: a clinicopathologic study of 200 cases with emphasis on its distinction from chordoma. Am J Surg Pathol 23:1370–1378
Healey JH, Lane JM (1989) Chordoma: a critical review of diagnosis and treatment. Orthop Clin North Am 20:417–426
Brain Tumor Registry of Japan (2009) Report of Brain Tumor Registry of Japan (1984–2000). Neurol Medico-Chir 49 Suppl: PS1–96
McMaster ML, Goldstein AM, Bromley CM, Ishibe N, Parry DM (2001) Chordoma: incidence and survival patterns in the United States, 1973–1995. Cancer Causes Control 12:1–11
Yang XR, Ng D, Alcorta DA, Liebsch NJ, Sheridan E, Li S, Goldstein AM, Parry DM, Kelley MJ (2009) T (brachyury) gene duplication confers major susceptibility to familial chordoma. Nat Genet 41:1176–1178. doi:10.1038/ng.454
Walcott BP, Nahed BV, Mohyeldin A, Coumans JV, Kahle KT, Ferreira MJ (2012) Chordoma: current concepts, management, and future directions. Lancet Oncol 13:e69–e76. doi:10.1016/S1470-2045(11)70337-0
Favre J, Deruaz JP, Uske A, de Tribolet N (1994) Skull base chordomas: presentation of six cases and review of the literature. J Clin Neurosci 1:7–18
Horbinski C, Oakley GJ, Cieply K, Mantha GS, Nikiforova MN, Dacic S, Seethala RR (2010) The prognostic value of Ki-67, p53, epidermal growth factor receptor, 1p36, 9p21, 10q23, and 17p13 in skull base chordomas. Arch Pathol Lab Med 134:1170–1176. doi:10.1043/2009-0380-OA.1
Kitamura Y, Sasaki H, Kimura T, Miwa T, Takahashi S, Kawase T, Yoshida K (2013) Molecular and clinical risk factors for recurrence of skull base chordomas: gain on chromosome 2p, expression of brachyury, and lack of irradiation negatively correlate with patient prognosis. J Neuropathol Exp Neurol 72:816–823. doi:10.1097/NEN.0b013e3182a065d0
Takahashi S, Kawase T, Yoshida K, Hasegawa A, Mizoe JE (2009) Skull base chordomas: efficacy of surgery followed by carbon ion radiotherapy. Acta Neurochir (Wien) 151:759–769. doi:10.1007/s00701-009-0383-5
Clark FH (1934) Linkage studies of Brachyury (Short Tail) in the house mouse. Proc Natl Acad Sci USA 20:276–279
Zhu J, Kwan KM, Mackem S (2016) Putative oncogene Brachyury (T) is essential to specify cell fate but dispensable for notochord progenitor proliferation and EMT. Proc Natl Acad Sci USA 113:3820–3825. doi:10.1073/pnas.1601252113
Fernando RI, Litzinger M, Trono P, Hamilton DH, Schlom J, Palena C (2010) The T-box transcription factor Brachyury promotes epithelial-mesenchymal transition in human tumor cells. J Clin Invest 120:533–544. doi:10.1172/JCI38379
Palena C, Fernando RI, Litzinger MT, Hamilton DH, Huang B, Schlom J (2011) Strategies to target molecules that control the acquisition of a mesenchymal-like phenotype by carcinoma cells. Exp Biol Med 236:537–545. doi:10.1258/ebm.2011.010367
Jambhekar NA, Rekhi B, Thorat K, Dikshit R, Agrawal M, Puri A (2010) Revisiting chordoma with brachyury, a “new age” marker: analysis of a validation study on 51 cases. Arch Pathol Lab Med 134:1181–1187. doi:10.1043/2009-0476-OA.1
Miettinen M, Wang Z, Lasota J, Heery C, Schlom J, Palena C (2015) Nuclear brachyury expression is consistent in chordoma, common in germ cell tumors and small cell carcinomas, and rare in other carcinomas and sarcomas: an immunohistochemical study of 5229 cases. Am J Surg Pathol 39:1305–1312. doi:10.1097/PAS.0000000000000462
Oakley GJ, Fuhrer K, Seethala RR (2008) Brachyury, SOX-9, and podoplanin, new markers in the skull base chordoma vs chondrosarcoma differential: a tissue microarray-based comparative analysis. Mod Pathol 21:1461–1469. doi:10.1038/modpathol.2008.144
Sangoi AR, Karamchandani J, Lane B, Higgins JP, Rouse RV, Brooks JD, McKenney JK (2011) Specificity of brachyury in the distinction of chordoma from clear cell renal cell carcinoma and germ cell tumors: a study of 305 cases. Mod Pathol 24:425–429. doi:10.1038/modpathol.2010.196
Vujovic S, Henderson S, Presneau N, Odell E, Jacques TS, Tirabosco R, Boshoff C, Flanagan AM (2006) Brachyury, a crucial regulator of notochordal development, is a novel biomarker for chordomas. J Pathol 209:157–165. doi:10.1002/path.1969
Pillay N, Plagnol V, Tarpey PS, Lobo SB, Presneau N, Szuhai K, Halai D, Berisha F, Cannon SR, Mead S et al (2012) A common single-nucleotide variant in T is strongly associated with chordoma. Nat Genet 44:1185–1187. doi:10.1038/ng.2419
Presneau N, Shalaby A, Ye H, Pillay N, Halai D, Idowu B, Tirabosco R, Whitwell D, Jacques TS, Kindblom LG et al (2011) Role of the transcription factor T (brachyury) in the pathogenesis of sporadic chordoma: a genetic and functional-based study. J Pathol 223:327–335. doi:10.1002/path.2816
Hsu W, Mohyeldin A, Shah SR, ap Rhys CM, Johnson LF, Sedora-Roman NI, Kosztowski TA, Awad OA, McCarthy EF, Loeb DM et al (2011) Generation of chordoma cell line JHC7 and the identification of Brachyury as a novel molecular target. J Neurosurg 115:760–769. doi:10.3171/2011.5.JNS11185
Nelson AC, Pillay N, Henderson S, Presneau N, Tirabosco R, Halai D, Berisha F, Flicek P, Stemple DL, Stern CD et al (2012) An integrated functional genomics approach identifies the regulatory network directed by brachyury (T) in chordoma. J Pathol 228:274–285. doi:10.1002/path.4082
Nibu Y, Jose-Edwards DS, Di Gregorio A (2013) From notochord formation to hereditary chordoma: the many roles of Brachyury. Biomed Res Int 2013: 826435 doi:10.1155/2013/826435
Antonelli M, Raso A, Mascelli S, Gessi M, Nozza P, Coli A, Gardiman MP, Arcella A, Massimino M, Buttarelli FR et al (2017) SMARCB1/INI1 involvement in pediatric chordoma: a mutational and immunohistochemical analysis. Am J Surg Pathol 41:56–61. doi:10.1097/PAS.0000000000000741
Chavez JA, Nasir Ud D, Memon A, Perry A (2014) Anaplastic chordoma with loss of INI1 and brachyury expression in a 2-year-old girl. Clin Neuropathol 33:418–420. doi:10.5414/NP300724
Mobley BC, McKenney JK, Bangs CD, Callahan K, Yeom KW, Schneppenheim R, Hayden MG, Cherry AM, Gokden M, Edwards MS et al (2010) Loss of SMARCB1/INI1 expression in poorly differentiated chordomas. Acta Neuropathol (Berl) 120:745–753. doi:10.1007/s00401-010-0767-x
Yadav R, Sharma MC, Malgulwar PB, Pathak P, Sigamani E, Suri V, Sarkar C, Kumar A, Singh M, Sharma BS et al (2014) Prognostic value of MIB-1, p53, epidermal growth factor receptor, and INI1 in childhood chordomas. Neuro Oncol 16:372–381. doi:10.1093/neuonc/not228
Hasselblatt M, Thomas C, Hovestadt V, Schrimpf D, Johann P, Bens S, Oyen F, Peetz-Dienhart S, Crede Y, Wefers A et al (2016) Poorly differentiated chordoma with SMARCB1/INI1 loss: a distinct molecular entity with dismal prognosis. Acta Neuropathol 132:149–151. doi:10.1007/s00401-016-1574-9
Akhavan-Sigari R, Abili M, Gaab MR, Rohde V, Zafar N, Emami P, Ostertag H (2015) Immunohistochemical expression of receptor tyrosine kinase PDGFR-alpha, c-Met, and EGFR in skull base chordoma. Neurosurg Rev 38: 89–98; discussion 98–89 doi:10.1007/s10143-014-0579-x
Dewaele B, Maggiani F, Floris G, Ampe M, Vanspauwen V, Wozniak A, Debiec-Rychter M, Sciot R (2011) Frequent activation of EGFR in advanced chordomas. Clin Sarcoma Res 1: 4 doi:10.1186/2045-3329-1-4
Fasig JH, Dupont WD, LaFleur BJ, Olson SJ, Cates JM (2008) Immunohistochemical analysis of receptor tyrosine kinase signal transduction activity in chordoma. Neuropathol Appl Neurobiol 34:95–104. doi:10.1111/j.1365-2990.2007.00873.x
Shalaby A, Presneau N, Ye H, Halai D, Berisha F, Idowu B, Leithner A, Liegl B, Briggs TR, Bacsi K et al (2011) The role of epidermal growth factor receptor in chordoma pathogenesis: a potential therapeutic target. J Pathol 223:336–346. doi:10.1002/path.2818
Tamborini E, Virdis E, Negri T, Orsenigo M, Brich S, Conca E, Gronchi A, Stacchiotti S, Manenti G, Casali PG et al (2010) Analysis of receptor tyrosine kinases (RTKs) and downstream pathways in chordomas. Neuro Oncol 12:776–789. doi:10.1093/neuonc/noq003
Orzan F, Terreni MR, Longoni M, Boari N, Mortini P, Doglioni C, Riva P (2007) Expression study of the target receptor tyrosine kinase of imatinib mesylate in skull base chordomas. Oncol Rep 18:249–252
Tamborini E, Miselli F, Negri T, Lagonigro MS, Staurengo S, Dagrada GP, Stacchiotti S, Pastore E, Gronchi A, Perrone F et al (2006) Molecular and biochemical analyses of platelet-derived growth factor receptor (PDGFR) B, PDGFRA, and KIT receptors in chordomas. Clin Cancer Res 12:6920–6928. doi:10.1158/1078-0432.CCR-06-1584
Shalaby AA, Presneau N, Idowu BD, Thompson L, Briggs TR, Tirabosco R, Diss TC, Flanagan AM (2009) Analysis of the fibroblastic growth factor receptor-RAS/RAF/MEK/ERK-ETS2/brachyury signalling pathway in chordomas. Mod Pathol 22:996–1005. doi:10.1038/modpathol.2009.63
Park JB, Lee CK, Koh JS, Lee JK, Park EY, Riew KD (2007) Overexpressions of nerve growth factor and its tropomyosin-related kinase A receptor on chordoma cells. Spine (Phila Pa 1976) 32: 1969–1973 doi:10.1097/BRS.0b013e318133fbb5
Scheipl S, Froehlich EV, Leithner A, Beham A, Quehenberger F, Mokry M, Stammberger H, Varga PP, Lazary A, Windhager R et al (2012) Does insulin-like growth factor 1 receptor (IGF-1R) targeting provide new treatment options for chordomas? A retrospective clinical and immunohistochemical study. Histopathology 60:999–1003. doi:10.1111/j.1365-2559.2012.04186.x
Sommer J, Itani DM, Homlar KC, Keedy VL, Halpern JL, Holt GE, Schwartz HS, Coffin CM, Kelley MJ, Cates JM (2010) Methylthioadenosine phosphorylase and activated insulin-like growth factor-1 receptor/insulin receptor: potential therapeutic targets in chordoma. J Pathol 220:608–617. doi:10.1002/path.2679
Tauziede-Espariat A, Bresson D, Polivka M, Bouazza S, Labrousse F, Aronica E, Pretet JL, Projetti F, Herman P, Salle H et al (2016) Prognostic and therapeutic markers in chordomas: a study of 287 tumors. J Neuropathol Exp Neurol 75:111–120. doi:10.1093/jnen/nlv010
Scheipl S, Barnard M, Cottone L, Jorgensen M, Drewry DH, Zuercher WJ, Turlais F, Ye H, Leite AP, Smith JA et al (2016) EGFR inhibitors identified as a potential treatment for chordoma in a focused compound screen. J Pathol 239:320–334. doi:10.1002/path.4729
Schwab J, Antonescu C, Boland P, Healey J, Rosenberg A, Nielsen P, Iafrate J, Delaney T, Yoon S, Choy E et al (2009) Combination of PI3K/mTOR inhibition demonstrates efficacy in human chordoma. Anticancer Res 29:1867–1871
Presneau N, Shalaby A, Idowu B, Gikas P, Cannon SR, Gout I, Diss T, Tirabosco R, Flanagan AM (2009) Potential therapeutic targets for chordoma: PI3K/AKT/TSC1/TSC2/mTOR pathway. Br J Cancer 100:1406–1414. doi:10.1038/sj.bjc.6605019
Le LP, Nielsen GP, Rosenberg AE, Thomas D, Batten JM, Deshpande V, Schwab J, Duan Z, Xavier RJ, Hornicek FJ et al (2011) Recurrent chromosomal copy number alterations in sporadic chordomas. PLoS ONE 6:e18846. doi:10.1371/journal.pone.0018846
Long C, Jiang L, Wei F, Ma C, Zhou H, Yang S, Liu X, Liu Z (2013) Integrated miRNA-mRNA analysis revealing the potential roles of miRNAs in chordomas. PloS one 8:e66676. doi:10.1371/journal.pone.0066676
Yang C, Hornicek FJ, Wood KB, Schwab JH, Choy E, Mankin H, Duan Z (2010) Blockage of Stat3 with CDDO-Me inhibits tumor cell growth in chordoma. Spine (Phila Pa 1976) 35: 1668–1675 doi:10.1097/BRS.0b013e3181c2d2b4
Yang C, Schwab JH, Schoenfeld AJ, Hornicek FJ, Wood KB, Nielsen GP, Choy E, Mankin H, Duan Z (2009) A novel target for treatment of chordoma: signal transducers and activators of transcription 3. Mol Cancer Ther 8:2597–2605. doi:10.1158/1535-7163.MCT-09-0504
Wang L, Zehir A, Nafa K, Zhou N, Berger MF, Casanova J, Sadowska J, Lu C, Allis CD, Gounder M et al (2016) Genomic aberrations frequently alter chromatin regulatory genes in chordoma. Genes Chromosomes Cancer 55:591–600. doi:10.1002/gcc.22362
Triana A, Sen C, Wolfe D, Hazan R (2005) Cadherins and catenins in clival chordomas: correlation of expression with tumor aggressiveness. Am J Surg Pathol 29:1422–1434
Schoenfeld AJ, Wang X, Wang Y, Hornicek FJ, Nielsen GP, Duan Z, Ferrone S, Schwab JH (2016) CSPG4 as a prognostic biomarker in chordoma. Spine J 16:722–727. doi:10.1016/j.spinee.2015.11.059
Diaz RJ, Guduk M, Romagnuolo R, Smith CA, Northcott P, Shih D, Berisha F, Flanagan A, Munoz DG, Cusimano MD et al (2012) High-resolution whole-genome analysis of skull base chordomas implicates FHIT loss in chordoma pathogenesis. Neoplasia 14:788–798
Sa JK, Lee IH, Hong SD, Kong DS, Nam DH (2016) Genomic and transcriptomic characterization of skull base chordoma. Oncotarget. doi:10.18632/oncotarget.13616
Rinner B, Weinhaeusel A, Lohberger B, Froehlich EV, Pulverer W, Fischer C, Meditz K, Scheipl S, Trajanoski S, Guelly C et al (2013) Chordoma characterization of significant changes of the DNA methylation pattern. PLoS ONE 8:e56609. doi:10.1371/journal.pone.0056609
Alholle A, Brini AT, Bauer J, Gharanei S, Niada S, Slater A, Gentle D, Maher ER, Jeys L, Grimer R et al (2015) Genome-wide DNA methylation profiling of recurrent and non-recurrent chordomas. Epigenetics 10: 213–220 doi:10.1080/15592294.2015.1006497
Brandal P, Bjerkehagen B, Danielsen H, Heim S (2005) Chromosome 7 abnormalities are common in chordomas. Cancer Genet Cytogenet 160:15–21. doi:10.1016/j.cancergencyto.2004.11.016
Hallor KH, Staaf J, Jonsson G, Heidenblad M, Vult von Steyern F, Bauer HC, Ijszenga M, Hogendoorn PC, Mandahl N, Szuhai K et al (2008) Frequent deletion of the CDKN2A locus in chordoma: analysis of chromosomal imbalances using array comparative genomic hybridisation. Br J Cancer 98:434–442. doi:10.1038/sj.bjc.6604130
Scheil S, Bruderlein S, Liehr T, Starke H, Herms J, Schulte M, Moller P (2001) Genome-wide analysis of sixteen chordomas by comparative genomic hybridization and cytogenetics of the first human chordoma cell line, U-CH1. Genes Chromosomes Cancer 32: 203–211
Tallini G, Dorfman H, Brys P, Dal Cin P, De Wever I, Fletcher CD, Jonson K, Mandahl N, Mertens F, Mitelman F et al (2002) Correlation between clinicopathological features and karyotype in 100 cartilaginous and chordoid tumours. A report from the Chromosomes and Morphology (CHAMP) Collaborative Study Group. J Pathol 196:194–203. doi:10.1002/path.1023
Longoni M, Orzan F, Stroppi M, Boari N, Mortini P, Riva P (2008) Evaluation of 1p36 markers and clinical outcome in a skull base chordoma study. Neuro Oncol 10:52–60. doi:10.1215/15228517-2007-048
Miozzo M, Dalpra L, Riva P, Volonta M, Macciardi F, Pericotti S, Tibiletti MG, Cerati M, Rohde K, Larizza L et al (2000) A tumor suppressor locus in familial and sporadic chordoma maps to 1p36. Int J Cancer 87:68–72
Riva P, Crosti F, Orzan F, Dalpra L, Mortini P, Parafioriti A, Pollo B, Fuhrman Conti AM, Miozzo M, Larizza L (2003) Mapping of candidate region for chordoma development to 1p36.13 by LOH analysis. Int J Cancer 107:493–497. doi:10.1002/ijc.11421
Sawyer JR, Husain M, Al-Mefty O (2001) Identification of isochromosome 1q as a recurring chromosome aberration in skull base chordomas: a new marker for aggressive tumors? Neurosurg Focus 10:E6
Duan Z, Choy E, Nielsen GP, Rosenberg A, Iafrate J, Yang C, Schwab J, Mankin H, Xavier R, Hornicek FJ (2010) Differential expression of microRNA (miRNA) in chordoma reveals a role for miRNA-1 in Met expression. J Orthop Res 28:746–752. doi:10.1002/jor.21055
Duan Z, Shen J, Yang X, Yang P, Osaka E, Choy E, Cote G, Harmon D, Zhang Y, Nielsen GP et al (2014) Prognostic significance of miRNA-1 (miR-1) expression in patients with chordoma. J Orthop Res 32:695–701. doi:10.1002/jor.22589
Osaka E, Yang X, Shen JK, Yang P, Feng Y, Mankin HJ, Hornicek FJ, Duan Z (2014) MicroRNA-1 (miR-1) inhibits chordoma cell migration and invasion by targeting slug. J Orthop Res 32:1075–1082. doi:10.1002/jor.22632
Osaka E, Kelly AD, Spentzos D, Choy E, Yang X, Shen JK, Yang P, Mankin HJ, Hornicek FJ, Duan Z (2015) MicroRNA-155 expression is independently predictive of outcome in chordoma. Oncotarget 6:9125–9139. doi:10.18632/oncotarget.3273
Zou MX, Huang W, Wang XB, Li J, Lv GH, Wang B, Deng YW (2015) Reduced expression of miRNA-1237-3p associated with poor survival of spinal chordoma patients. Eur Spine J 24:1738–1746. doi:10.1007/s00586-015-3927-9
Zou MX, Huang W, Wang XB, Lv GH, Li J, Deng YW (2014) Identification of miR-140-3p as a marker associated with poor prognosis in spinal chordoma. Int J Clin Exp Pathol 7:4877–4885
Gulluoglu S, Tuysuz EC, Kuskucu A, Ture U, Atalay B, Sahin F, Bayrak OF (2016) The potential function of microRNA in chordomas. Gene 585:76–83. doi:10.1016/j.gene.2016.03.032
Mathios D, Ruzevick J, Jackson CM, Xu H, Shah S, Taube JM, Burger PC, McCarthy EF, Quinones-Hinojosa A, Pardoll DM et al (2015) PD-1, PD-L1, PD-L2 expression in the chordoma microenvironment. J Neurooncol 121:251–259. doi:10.1007/s11060-014-1637-5
Feng Y, Shen J, Gao Y, Liao Y, Cote G, Choy E, Chebib I, Mankin H, Hornicek F, Duan Z (2015) Expression of programmed cell death ligand 1 (PD-L1) and prevalence of tumor-infiltrating lymphocytes (TILs) in chordoma. Oncotarget 6:11139–11149. doi:10.18632/oncotarget.3576
Zou MX, Peng AB, Lv GH, Wang XB, Li J, She XL, Jiang Y (2016) Expression of programmed death-1 ligand (PD-L1) in tumor-infiltrating lymphocytes is associated with favorable spinal chordoma prognosis. Am J Transl Res 8:3274–3287
Hof H, Welzel T, Debus J (2006) Effectiveness of cetuximab/gefitinib in the therapy of a sacral chordoma. Onkologie 29:572–574. doi:10.1159/000096283
Linden O, Stenberg L, Kjellen E (2009) Regression of cervical spinal cord compression in a patient with chordoma following treatment with cetuximab and gefitinib. Acta Oncol 48:158–159. doi:10.1080/02841860802266672
Asklund T, Sandstrom M, Shahidi S, Riklund K, Henriksson R (2014) Durable stabilization of three chordoma cases by bevacizumab and erlotinib. Acta Oncol 53:980–984. doi:10.3109/0284186X.2013.878472
Launay SG, Chetaille B, Medina F, Perrot D, Nazarian S, Guiramand J, Moureau-Zabotto L, Bertucci F (2011) Efficacy of epidermal growth factor receptor targeting in advanced chordoma: case report and literature review. BMC Cancer 11:423. doi:10.1186/1471-2407-11-423
Singhal N, Kotasek D, Parnis FX (2009) Response to erlotinib in a patient with treatment refractory chordoma. Anticancer Drugs 20:953–955. doi:10.1097/CAD.0b013e328330c7f0
Stacchiotti S, Tamborini E, Lo Vullo S, Bozzi F, Messina A, Morosi C, Casale A, Crippa F, Conca E, Negri T et al (2013) Phase II study on lapatinib in advanced EGFR-positive chordoma. Ann Oncol 24:1931–1936. doi:10.1093/annonc/mdt117
Hindi N, Casali PG, Morosi C, Messina A, Palassini E, Pilotti S, Tamborini E, Radaelli S, Gronchi A, Stacchiotti S (2015) Imatinib in advanced chordoma: a retrospective case series analysis. Eur J Cancer 51:2609–2614. doi:10.1016/j.ejca.2015.07.038
Stacchiotti S, Longhi A, Ferraresi V, Grignani G, Comandone A, Stupp R, Bertuzzi A, Tamborini E, Pilotti S, Messina A et al (2012) Phase II study of imatinib in advanced chordoma. J Clin Oncol 30:914–920. doi:10.1200/JCO.2011.35.3656
Stacchiotti S, Marrari A, Tamborini E, Palassini E, Virdis E, Messina A, Crippa F, Morosi C, Gronchi A, Pilotti S et al (2009) Response to imatinib plus sirolimus in advanced chordoma. Ann Oncol 20:1886–1894. doi:10.1093/annonc/mdp210
Ricci-Vitiani L, Runci D, D’Alessandris QG, Cenci T, Martini M, Bianchi F, Maira G, Stancato L, De Maria R, Larocca LM et al (2013) Chemotherapy of skull base chordoma tailored on responsiveness of patient-derived tumor cells to rapamycin. Neoplasia 15:773–782
Schuetze SM, Bolejack V, Choy E, Ganjoo KN, Staddon AP, Chow WA, Tawbi HA, Samuels BL, Patel SR, von Mehren M et al (2017) Phase 2 study of dasatinib in patients with alveolar soft part sarcoma, chondrosarcoma, chordoma, epithelioid sarcoma, or solitary fibrous tumor. Cancer 123:90–97. doi:10.1002/cncr.30379
The Chordoma Foundation (2017) Clinical Trials Program https://www.chordomafoundation.org/clinical-trials-program/?utm_source=Chordoma+Foundation+Newsletter&utm_campaign=3bad75d02c-2017_CF_Newsletter_01_Jan_1_26_20171_30_2017&utm_medium=email&utm_term=0_288d805c80-3bad75d02c-86990177. Accessed 2/2/2017
Dorfman HD, Czerniak B (1995) Bone cancers. Cancer 75:203–210
Rigau V, Zouaoui S, Mathieu-Daude H, Darlix A, Maran A, Tretarre B, Bessaoud F, Bauchet F, Attaoua R, Fabbro-Peray P et al (2011) French brain tumor database: 5-year histological results on 25 756 cases. Brain Pathol 21:633–644. doi:10.1111/j.1750-3639.2011.00491.x
Bjornsson J, McLeod RA, Unni KK, Ilstrup DM, Pritchard DJ (1998) Primary chondrosarcoma of long bones and limb girdles. Cancer 83:2105–2119
Bovee JV, Hogendoorn PC, Wunder JS, Alman BA (2010) Cartilage tumours and bone development: molecular pathology and possible therapeutic targets. Nat Rev Cancer 10:481–488. doi:10.1038/nrc2869
Arai M, Nobusawa S, Ikota H, Takemura S, Nakazato Y (2012) Frequent IDH1/2 mutations in intracranial chondrosarcoma: a possible diagnostic clue for its differentiation from chordoma. Brain Tumor Pathol 29:201–206. doi:10.1007/s10014-012-0085-1
Kanamori H, Kitamura Y, Kimura T, Yoshida K, Sasaki H (2015) Genetic characterization of skull base chondrosarcomas. J Neurosurg 123:1036–1041. doi:10.3171/2014.12.JNS142059
Amary MF, Bacsi K, Maggiani F, Damato S, Halai D, Berisha F, Pollock R, O’Donnell P, Grigoriadis A, Diss T et al (2011) IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. J Pathol 224:334–343. doi:10.1002/path.2913
Tarpey PS, Behjati S, Cooke SL, Van Loo P, Wedge DC, Pillay N, Marshall J, O’Meara S, Davies H, Nik-Zainal S et al (2013) Frequent mutation of the major cartilage collagen gene COL2A1 in chondrosarcoma. Nat Genet 45:923–926. doi:10.1038/ng.2668
Li L, Paz AC, Wilky BA, Johnson B, Galoian K, Rosenberg A, Hu G, Tinoco G, Bodamer O, Trent JC (2015) Treatment with a small molecule mutant IDH1 inhibitor suppresses tumorigenic activity and decreases production of the oncometabolite 2-hydroxyglutarate in human chondrosarcoma cells. PLoS ONE 10:e0133813. doi:10.1371/journal.pone.0133813
Dang L, Yen K, Attar EC (2016) IDH mutations in cancer and progress toward development of targeted therapeutics. Ann Oncol 27:599–608. doi:10.1093/annonc/mdw013
Song J, Zhu J, Zhao Q, Tian B (2015) Gefitinib causes growth arrest and inhibition of metastasis in human chondrosarcoma cells. J BUON 20:894–901
Zhang YX, van Oosterwijk JG, Sicinska E, Moss S, Remillard SP, van Wezel T, Buhnemann C, Hassan AB, Demetri GD, Bovee JV et al (2013) Functional profiling of receptor tyrosine kinases and downstream signaling in human chondrosarcomas identifies pathways for rational targeted therapy. Clin Cancer Res 19:3796–3807. doi:10.1158/1078-0432.CCR-12-3647
Sulzbacher I, Birner P, Trieb K, Muhlbauer M, Lang S, Chott A (2001) Platelet-derived growth factor-alpha receptor expression supports the growth of conventional chondrosarcoma and is associated with adverse outcome. Am J Surg Pathol 25:1520–1527
Ho L, Stojanovski A, Whetstone H, Wei QX, Mau E, Wunder JS, Alman B (2009) Gli2 and p53 cooperate to regulate IGFBP-3- mediated chondrocyte apoptosis in the progression from benign to malignant cartilage tumors. Cancer Cell 16:126–136. doi:10.1016/j.ccr.2009.05.013
Matsumura T, Whelan MC, Li XQ, Trippel SB (2000) Regulation by IGF-I and TGF-beta1 of Swarm-rat chondrosarcoma chondrocytes. J Orthop Res 18:351–355. doi:10.1002/jor.1100180305
Sun X, Charbonneau C, Wei L, Chen Q, Terek RM (2015) miR-181a targets RGS16 to promote chondrosarcoma growth, angiogenesis, and metastasis. Mol Cancer Res 13:1347–1357. doi:10.1158/1541-7786.MCR-14-0697
Tzeng HE, Chen PC, Lin KW, Lin CY, Tsai CH, Han SM, Teng CL, Hwang WL, Wang SW, Tang CH (2015) Basic fibroblast growth factor induces VEGF expression in chondrosarcoma cells and subsequently promotes endothelial progenitor cell-primed angiogenesis. Clinical Sci 129:147–158. doi:10.1042/CS20140390
Peterse EF, Cleven AH, De Jong Y, Briaire-de Bruijn I, Fletcher JA, Danen EH, Cleton-Jansen AM, Bovee JV (2016) No preclinical rationale for IGF1R directed therapy in chondrosarcoma of bone. BMC Cancer 16:475. doi:10.1186/s12885-016-2522-8
Asp J, Sangiorgi L, Inerot SE, Lindahl A, Molendini L, Benassi MS, Picci P (2000) Changes of the p16 gene but not the p53 gene in human chondrosarcoma tissues. Int J Cancer 85:782–786
Schrage YM, Lam S, Jochemsen AG, Cleton-Jansen AM, Taminiau AH, Hogendoorn PC, Bovee JV (2009) Central chondrosarcoma progression is associated with pRb pathway alterations: CDK4 down-regulation and p16 overexpression inhibit cell growth in vitro. J Cell Mol Med 13:2843–2852. doi:10.1111/j.1582-4934.2008.00406.x
van Beerendonk HM, Rozeman LB, Taminiau AH, Sciot R, Bovee JV, Cleton-Jansen AM, Hogendoorn PC (2004) Molecular analysis of the INK4A/INK4A-ARF gene locus in conventional (central) chondrosarcomas and enchondromas: indication of an important gene for tumour progression. J Pathol 202:359–366. doi:10.1002/path.1517
Tiet TD, Hopyan S, Nadesan P, Gokgoz N, Poon R, Lin AC, Yan T, Andrulis IL, Alman BA, Wunder JS (2006) Constitutive hedgehog signaling in chondrosarcoma up-regulates tumor cell proliferation. Am J Pathol 168:321–330. doi:10.2353/ajpath.2006.050001
Campbell VT, Nadesan P, Ali SA, Wang CY, Whetstone H, Poon R, Wei Q, Keilty J, Proctor J, Wang LW et al (2014) Hedgehog pathway inhibition in chondrosarcoma using the smoothened inhibitor IPI-926 directly inhibits sarcoma cell growth. Mol Cancer Ther 13:1259–1269. doi:10.1158/1535-7163.MCT-13-0731
Speetjens FM, de Jong Y, Gelderblom H, Bovee JV (2016) Molecular oncogenesis of chondrosarcoma: impact for targeted treatment. Curr Opin Oncol 28:314–322. doi:10.1097/CCO.0000000000000300
Jiang D, Zheng X, Shan W, Shan Y (2016) The overexpression of miR-30a affects cell proliferation of chondrosarcoma via targeting Runx2. Tumour Biol 37:5933–5940. doi:10.1007/s13277-015-4454-3
Brown RE (2004) Morphoproteomic portrait of the mTOR pathway in mesenchymal chondrosarcoma. Ann Clin Lab Sci 34:397–399
Schrage YM, Briaire-de Bruijn IH, de Miranda NF, van Oosterwijk J, Taminiau AH, van Wezel T, Hogendoorn PC, Bovee JV (2009) Kinome profiling of chondrosarcoma reveals SRC-pathway activity and dasatinib as option for treatment. Cancer Res 69:6216–6222. doi:10.1158/0008-5472.CAN-08-4801
Perez J, Decouvelaere AV, Pointecouteau T, Pissaloux D, Michot JP, Besse A, Blay JY, Dutour A (2012) Inhibition of chondrosarcoma growth by mTOR inhibitor in an in vivo syngeneic rat model. PLoS ONE 7:e32458. doi:10.1371/journal.pone.0032458
Kubo T, Sugita T, Shimose S, Matsuo T, Arihiro K, Ochi M (2008) Expression of hypoxia-inducible factor-1alpha and its relationship to tumour angiogenesis and cell proliferation in cartilage tumours. J Bone Joint Surg Br 90:364–370. doi:10.1302/0301-620X.90B3.19806
Oshiro Y, Chaturvedi V, Hayden D, Nazeer T, Johnson M, Johnston DA, Ordonez NG, Ayala AG, Czerniak B (1998) Altered p53 is associated with aggressive behavior of chondrosarcoma: a long term follow-up study. Cancer 83:2324–2334
van Oosterwijk JG, Herpers B, Meijer D, Briaire-de Bruijn IH, Cleton-Jansen AM, Gelderblom H, van de Water B, Bovee JV (2012) Restoration of chemosensitivity for doxorubicin and cisplatin in chondrosarcoma in vitro: BCL-2 family members cause chemoresistance. Ann Oncol 23:1617–1626. doi:10.1093/annonc/mdr512
Fitzgerald MP, Gourronc F, Teoh ML, Provenzano MJ, Case AJ, Martin JA, Domann FE (2011) Human chondrosarcoma cells acquire an epithelial-like gene expression pattern via an epigenetic switch: evidence for mesenchymal-epithelial transition during sarcomagenesis. Sarcoma 2011: 598218 doi:10.1155/2011/598218
Bui C, Ouzzine M, Talhaoui I, Sharp S, Prydz K, Coughtrie MW, Fournel-Gigleux S (2010) Epigenetics: methylation-associated repression of heparan sulfate 3-O-sulfotransferase gene expression contributes to the invasive phenotype of H-EMC-SS chondrosarcoma cells. FASEB J 24:436–450. doi:10.1096/fj.09-136291
Jin Z, Han YX, Han XR (2013) Loss of RUNX3 expression may contribute to poor prognosis in patients with chondrosarcoma. J Mol Histol 44:645–652. doi:10.1007/s10735-013-9511-x
Hallor KH, Staaf J, Bovee JV, Hogendoorn PC, Cleton-Jansen AM, Knuutila S, Savola S, Niini T, Brosjo O, Bauer HC et al (2009) Genomic profiling of chondrosarcoma: chromosomal patterns in central and peripheral tumors. Clin Cancer Res 15:2685–2694. doi:10.1158/1078-0432.CCR-08-2330
Hameed M, Ulger C, Yasar D, Limaye N, Kurvathi R, Streck D, Benevenia J, Patterson F, Dermody JJ, Toruner GA (2009) Genome profiling of chondrosarcoma using oligonucleotide array-based comparative genomic hybridization. Cancer Genet Cytogenet 192:56–59. doi:10.1016/j.cancergencyto.2009.03.009
Larramendy ML, Mandahl N, Mertens F, Blomqvist C, Kivioja AH, Karaharju E, Valle J, Bohling T, Tarkkanen M, Rydholm A et al (1999) Clinical significance of genetic imbalances revealed by comparative genomic hybridization in chondrosarcomas. Hum Pathol 30:1247–1253
Larramendy ML, Tarkkanen M, Valle J, Kivioja AH, Ervasti H, Karaharju E, Salmivalli T, Elomaa I, Knuutila S (1997) Gains, losses, and amplifications of DNA sequences evaluated by comparative genomic hybridization in chondrosarcomas. Am J Pathol 150:685–691
Ozaki T, Wai D, Schafer KL, Lindner N, Bocker W, Winkelmann W, Dockhorn-Dworniczak B, Poremba C (2004) Comparative genomic hybridization in cartilaginous tumors. Anticancer Res 24:1721–1725
Rozeman LB, Szuhai K, Schrage YM, Rosenberg C, Tanke HJ, Taminiau AH, Cleton-Jansen AM, Bovee JV, Hogendoorn PC (2006) Array-comparative genomic hybridization of central chondrosarcoma: identification of ribosomal protein S6 and cyclin-dependent kinase 4 as candidate target genes for genomic aberrations. Cancer 107:380–388. doi:10.1002/cncr.22001
Lu N, Lin T, Wang L, Qi M, Liu Z, Dong H, Zhang X, Zhai C, Wang Y, Liu L et al (2015) Association of SOX4 regulated by tumor suppressor miR-30a with poor prognosis in low-grade chondrosarcoma. Tumour Biol 36:3843–3852. doi:10.1007/s13277-014-3026-2
Yoshitaka T, Kawai A, Miyaki S, Numoto K, Kikuta K, Ozaki T, Lotz M, Asahara H (2013) Analysis of microRNAs expressions in chondrosarcoma. J Orthop Res 31:1992–1998. doi:10.1002/jor.22457
Li J, Wang L, Liu Z, Zu C, Xing F, Yang P, Yang Y, Dang X, Wang K (2015) MicroRNA-494 inhibits cell proliferation and invasion of chondrosarcoma cells in vivo and in vitro by directly targeting SOX9. Oncotarget 6:26216–26229. doi:10.18632/oncotarget.4460
Mak IW, Singh S, Turcotte R, Ghert M (2015) The epigenetic regulation of SOX9 by miR-145 in human chondrosarcoma. J Cell Biochem 116:37–44. doi:10.1002/jcb.24940
Tsai CH, Tsai HC, Huang HN, Hung CH, Hsu CJ, Fong YC, Hsu HC, Huang YL, Tang CH (2015) Resistin promotes tumor metastasis by down-regulation of miR-519d through the AMPK/p38 signaling pathway in human chondrosarcoma cells. Oncotarget 6:258–270. doi:10.18632/oncotarget.2724
Aili A, Chen Y, Zhang H (2016) MicroRNA10b suppresses the migration and invasion of chondrosarcoma cells by targeting brain-derived neurotrophic factor. Mol Med Rep 13: 441–446 doi:10.3892/mmr.2015.4506
Kostine M, Cleven AH, de Miranda NF, Italiano A, Cleton-Jansen AM, Bovee JV (2016) Analysis of PD-L1, T-cell infiltrate and HLA expression in chondrosarcoma indicates potential for response to immunotherapy specifically in the dedifferentiated subtype. Mod Pathol 29:1028–1037. doi:10.1038/modpathol.2016.108
Paoluzzi L, Cacavio A, Ghesani M, Karambelkar A, Rapkiewicz A, Weber J, Rosen G (2016) Response to anti-PD1 therapy with nivolumab in metastatic sarcomas. Clin Sarcoma Res 6: 24 doi:10.1186/s13569-016-0064-0
Grignani G, Palmerini E, Stacchiotti S, Boglione A, Ferraresi V, Frustaci S, Comandone A, Casali PG, Ferrari S, Aglietta M (2011) A phase 2 trial of imatinib mesylate in patients with recurrent nonresectable chondrosarcomas expressing platelet-derived growth factor receptor-alpha or -beta: an Italian Sarcoma Group study. Cancer 117:826–831. doi:10.1002/cncr.25632
Schwartz GK, Tap WD, Qin LX, Livingston MB, Undevia SD, Chmielowski B, Agulnik M, Schuetze SM, Reed DR, Okuno SH et al (2013) Cixutumumab and temsirolimus for patients with bone and soft-tissue sarcoma: a multicentre, open-label, phase 2 trial. Lancet Oncol 14:371–382. doi:10.1016/S1470-2045(13)70049-4
Thornton KA, Chen AR, Trucco MM, Shah P, Wilky BA, Gul N, Carrera-Haro MA, Ferreira MF, Shafique U, Powell JD et al (2013) A dose-finding study of temsirolimus and liposomal doxorubicin for patients with recurrent and refractory bone and soft tissue sarcoma. Int J Cancer 133:997–1005. doi:10.1002/ijc.28083
Italiano A, Le Cesne A, Bellera C, Piperno-Neumann S, Duffaud F, Penel N, Cassier P, Domont J, Takebe N, Kind M et al (2013) GDC-0449 in patients with advanced chondrosarcomas: a French Sarcoma Group/US and French National Cancer Institute Single-Arm Phase II Collaborative Study. Ann Oncol 24:2922–2926. doi:10.1093/annonc/mdt391
The U.S. National Institutes of Health (2017) SARC028: A phase II study of the anti-PD1 antibody pembrolizumab (MK-3475) in patients with advanced sarcomas https://clinicaltrials.gov/ct2/show/NCT02301039. Accessed 30 Jan 2017
The U.S. National Institutes of Health (2017) A phase II study evaluating efficacy and safety of regorafenib in patients with metastatic bone sarcomas (REGOBONE) https://clinicaltrials.gov/ct2/show/NCT02389244. Accessed 2 Feb 2017
The U.S. National Institutes of Health (2017) Study of pazopanib in the treatment of surgically unresectable or metastatic chondrosarcoma https://clinicaltrials.gov/ct2/show/NCT01330966. Accessed 2 Feb 2017
Bernstein-Molho R, Kollender Y, Issakov J, Bickels J, Dadia S, Flusser G, Meller I, Sagi-Eisenberg R, Merimsky O (2012) Clinical activity of mTOR inhibition in combination with cyclophosphamide in the treatment of recurrent unresectable chondrosarcomas. Cancer Chemother Pharmacol 70:855–860. doi:10.1007/s00280-012-1968-x
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK (2016) World Health Organization histological classification of tumours of the central nervous system. International Agency for Research on Cancer, Lyon
Lagonigro MS, Tamborini E, Negri T, Staurengo S, Dagrada GP, Miselli F, Gabanti E, Greco A, Casali PG, Carbone A et al (2006) PDGFRalpha, PDGFRbeta and KIT expression/activation in conventional chondrosarcoma. J Pathol 208:615–623. doi:10.1002/path.1945
Zhu Z, Wang CP, Zhang YF, Nie L (2014) MicroRNA-100 resensitizes resistant chondrosarcoma cells to cisplatin through direct targeting of mTOR. Asian Pac J Cancer Prev 15: 917–923
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors have no personal, financial, or institutional interest in any of the drugs, materials, or devices described in this article.
Rights and permissions
About this article
Cite this article
Kitamura, Y., Sasaki, H. & Yoshida, K. Genetic aberrations and molecular biology of skull base chordoma and chondrosarcoma. Brain Tumor Pathol 34, 78–90 (2017). https://doi.org/10.1007/s10014-017-0283-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-017-0283-y