Abstract
Tuberculosis (TB) is a global health problem and the situation has become more precarious due to the advent of HIV infections and continuous rise in the number of multi-drug resistant strains of Mycobacterium tuberculosis (M. tb). Biochemical studies on Fatty Acyl-CoA Synthetases (FadD13), one of the gene products of mymA operon, have provided insights into the involvement of this protein in the activation of fatty acids. Due to non-availability of the crystal structure of FadD13, we have employed in silico approaches to resolve and characterize the structure of this important protein of M. tb. A three dimensional model of M. tb FadD13 was predicted by a de novo structure prediction server that integrates fragment assembly with SimFold energy function. With the aid of molecular mechanics and dynamics methods, the final model was obtained and assessed subsequently for global and local accuracy by various assessment programs. With this model, a flexible docking study with the substrates was performed. Results of ligand interactions with key amino acids in the binding site are also summarized. The molecular model for the M. tb FadD13 obtained sheds light on the topographical features of the binding pocket of the protein and provides atomic insight into the possible modes of substrate recognition. The three-dimensional model of FadD13 presented here would be helpful in guiding both enzymatic studies as well as design of specific inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tuberculosis (TB) remains one of the most important global health problems causing the loss of 2–3 million lives every year [1]. One-third of the world’s population is asymptomatically infected with Mycobacterium tuberculosis (M. tb), the etiologic agent of TB [2, 3]. Treatment of the active cases of TB includes simultaneous therapy with two or more of the frontline drugs: isoniazid, ethambutol, rifampicin and pyrazinamide [4]. Recent outbreaks of TB caused by multidrug-resistant (MDR) strains, mainly in the individuals infected with HIV, have created a precarious situation worldwide and have generated a significant interest in expanding the current programs of developing antitubercular drugs. Greater efforts are needed to investigate the molecular basis of pathogenicity and to develop high efficacy drugs against the key targets of M. tb. The determination of the M. tb genome sequence [5] has provided an enormous boost to these efforts and subsequent studies have attempted to identify genes that are likely to be required for the establishment and progression of TB leading to the identification of novel drug targets [6]. The rapid expansion of TB-related genomic data sources provides considerable opportunities to apply advanced computational analyses for the prediction of potential drug targets [7, 8]. TB Structural Genomics Consortium has significantly promoted these endeavors [9, 10]. Several proteins or pathways of M. tb have been demonstrated to be valid targets for drug design. The fatty acid biosynthesis pathway represents one such target for the development of new antimycobacterial agents [11, 12]. M. tb has ∼250 genes for lipid metabolism [5, 13]. Many of them are involved in the synthesis of cell envelope and required for the virulence of M. tb. Figure 1 shows mymA operon (Rv3083–Rv3089), which is involved in maintaining appropriate cell wall architecture of M. tb on exposure to acidic pH faced in macrophages [14]. This operon has been shown to be upregulated at acidic pH [15]. Functional loss of mymA operon results in increased drug sensitivity and killing of the pathogen. Therefore, gene products of mymA operon can be employed as effective drug targets [14]. One of the crucial genes of mymA operon, the fadD13, encodes a Fatty Acyl-CoA Synthetase (FACS) which is involved in the activation of fatty acids and catalyzes the following reaction [16]:
Schematic organization of the structural genes of mymA operon of M. tuberculosis [14] mymA operon consist of seven genes with fadD13 being the last one, which is involved in activation of fatty acids
In the present study, we model the structure of M. tb Fatty Acyl-CoA Synthetases (FadD13) by using a de novo structure prediction server by fragment assembly with SimFold energy function, which is subjected to molecular dynamics simulations in an explicit solvent environment. Mutational studies were carried out on the refined structure in order to evaluate the possible binding modes reported earlier by experimental investigations [17–21].
Materials and methods
All computations were performed on Sun Fire X4600 M2 server, dual-core AMD Opteron with eight processors running SUSE Linux Enterprise Server Edition 10.0.
Sequence homology and conserved domain search
The 503-long amino acid sequence of mycobacterial FadD13 (Entry name: Probable chain fatty acid-CoA ligase FadD13, [Mycobacterium tuberculosis H37Rv]; [GenBank: CAA16147]) was retrieved from NCBI databank. Sequence of M. tb FadD13 was subjected to pairwise alignment using NCBI-BLAST [22, 23] against PDB [24]. Multiple sequence alignment using CLUSTAL W [25] was carried out with FadD in Escherichia coli (E. coli FadD) and long chain-Fatty Acyl-CoA Synthetase in Thermus thermophilus (ttLC-FACS) for the search of conserved regions in M. tb FadD13. Domain search was carried out by using Pfam database [26] and validated through literature survey.
Initial model generation
The model of M. tb FadD13 was predicted by using comparative modeling, fold recognition and de novo methods. Homology models were generated using ESyPred3D [27], SWISS-MODEL [28], Prime (Schrödinger) [29] and Modeller9v5 [30]. The sequence of FadD13 was also submitted to PHYRE (Protein Homology/analogY Recognition Engine) [31]; a threading based method and Rokky-P [32]; structure prediction server that integrates PDB-BLAST, 3D-Jury, and the SimFold fragment assembly simulator. Five models were generated from Rokky-P server. Structures generated by all these methods were assessed using various protein structure evaluation servers like PROCHECK [33], WHAT IF [34] and VERIFY-3D [35] to select a final model for further refinement by molecular dynamics (MD) simulations.
Model refinement
With the aim of evaluating the stability and folding, conformational changes and getting insights into the natural dynamics on different timescales of protein in solution, 12 nano-seconds (ns) MD simulations were performed in the study. Simulations were carried out by using Desmond 2.0 (Schrödinger) [36] by employing OPLS-AA force field [37]. All production-phase MD simulations were run with a time step of 2.0 femto-seconds (fs) with far time step size of 6.0 fs using RESPA integrator under the NPT ensemble (300 K and 1.01325 bar pressure) with explicit solvent by using the TIP4P [38] model for water and by using Na+ and Cl− for ion placement, periodic boundary conditions, the particle mesh Ewald (PME) [39] method for electrostatics, a 10 Å cutoff for Lennard-Jones interactions, and the use of SHAKE [40] for restricting motion of all covalent bonds involving hydrogen atoms. The temperature was maintained by the Nosé-Hoover coupling algorithm [41] while the pressure was maintained by using Martyna-Tobias-Klein method [42]. Energy and trajectory were recorded after every 100 ps and 5 ps, respectively.
The equilibration process is comprised of six stages: the solvated structure was minimized first with solute restrained and then again minimized without restraints by using hybrid method of steepest descent and the LBFGS (limited-memory Broyden-Fletcher-Goldfarb-Shanno) algorithm [43] with a maximum of 2000 steps including initial 10 steps of steepest-descent. The system was heated at a temperature of 10 K in two stages in NVT and NPT ensemble, respectively by using Berendson thermostat and barostat [44] for a time period of 12 ps keeping non-hydrogen solute atoms restrained. The system was then heated for a time of 24 ps to 300 K in NPT ensemble again keeping non-hydrogen solute atoms restrained. In the last stage, the system was simulated for 24 ps in NPT ensemble by using Berendson thermostat and barostat [44] with no atoms restrained. Frames from the trajectory were extracted every 1 ns and were energy minimized by using Prime (Schrödinger) [29]. The geometric correctness of the minimized frames was assessed using VERIFY-3D [35], PROCHECK [33], WHAT IF [34], ProQ [45], ProSA [46] and ERRAT [47].
Binding site prediction
Binding site was characterized by using Q-SiteFinder [48] and CASTp [49] and these were validated by using the information on binding sites in other homologous proteins [17, 50].
Docking studies
In this study, docking was performed with the substrates ATP, coenzymeA (CoA) and fatty acids—capric acid (10:0), palmitic acid (16:0), lignoceric acid (24:0) and cerotic acid (26:0) by using induced fit docking (IFD) protocol (Schrödinger) [51]. Initially, protein structure was prepared using protein preparation wizard and tautomers of substrates were generated by using LigPrep module (Schrödinger) [52]. Then IFD was performed according to the following three steps. First, the ligand was docked into a rigid receptor model with scaled-down vdW radii (Glide SP mode). Next, Prime was used to generate the induced-fit protein–substrate complexes and these complexes were then ranked by Prime energy. Finally, each ligand was redocked into every refined low-energy receptor structure produced in the second step using Glide XP [53] at default settings. An IFD score that accounts for both the protein–substrate interaction energy and the total energy of the system was calculated for ranking the IFD poses.
IFD protocol was then subsequently used to dock all three substrates (multiple ligands) into the binding site of the predicted model. The docking of substrates was performed in the order of ATP followed by fatty acid and CoA in the end. This order was decided according to the mechanistic action of the enzyme stated above. The docked complexes were subjected to molecular dynamics simulation for 6 ns using Desmond 2.2 [36].
Mutational studies
Based on structural information from firefly luciferase [18], propionyl CoA Synthetase (PrpE) enzyme of Salmonella enterica [19], human long-chain fatty acid-CoA ligase 4 (ACSL4) [20] and E. coli FadD [17], mutants at K487A and R397A were constructed which provided us with a clue to investigate substrate binding in these altered structures. Mutations were carried out by using Modeller9v5 [30] and then docked with the substrates (ATP and fatty acids) by using IFD protocol [51] as discussed above and the docked complexes were subjected to molecular dynamics simulation for 6 ns using Desmond 2.2 [36]. Protein-ligand contacts were generated by using LIGPLOT [54].
Results and discussion
Homology comparison
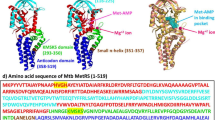
ttLC-FACS (PDB-ID: 1ULT) was identified as the best homologue with a reliable expectant value from BLAST [22, 23] analyses. Alignment of sequence from 1ULT with that of M. tb FadD13 resulted in 32% identity and an expectancy score of 2e-57. Pfam [26] database search revealed one AMP-binding domain. Figure 2 shows multiple sequence alignment of M.tb FadD13 with E. coli FadD and ttLC-FACS, which reveals three conserved regions: two ATP-AMP binding domains, residues 163–173 referred to as P-motif, 300–306 denoted as A-motif and one fatty-acid binding domain, residues 375–399 named as FACS signature motif. These domains are conserved within the superfamily of adenylate forming enzymes.
Multiple sequence alignment of M. tb FadD13 with E. coli fadD and ttLC-FACS. The identical residues in the aligned sequences are indicated with an asterisk (*). P-motif is phosphate-binding site colored in blue, A-motif is adenine-binding site colored in purple, L-motif is linker motif colored in yellow and fatty acid binding site is indicated in green
Model refinement and assessment
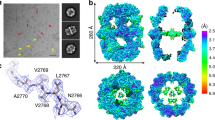
Based on the results obtained from various protein structure evaluation servers, model 3 generated by Rokky-P was selected as the final model (Table 1). A 12 ns molecular dynamics simulation was performed on the final model by using Desmond 2.0 [36]. Frames were collected after every 1 ns, energy minimized and then checked with various protein-evaluation servers. A plot of the total energy versus MD time shows that the total energy reaches equilibrium by 10 ns. The analysis of the RMSD map obtained in Fig. 3a during the 12 ns MD simulation shows that after a small rearrangement from the initial conformation (RMSD of ∼2 Å between the starting conformation and the first stabilized ∼500 ps), the structure is relatively stable during the whole MD. The final model obtained was tested with the ProSA [46] program by examining whether the interaction of each residue with the remainder of the protein is maintained in a favorable manner. Figure 3b shows that ProSA [46] of the model gave a Z score of −9.81; which is within the acceptable range (−10 to 10, good ProSA scores are negative and depend on length of protein). Figure 3c shows that the energy remains negative for almost all amino acid residues indicating the acceptability of the predicted model. The six graphs on the main chain parameters (Fig. 3d) plot shows the structure (represented by solid square) compared with well-refined structures at a similar resolution. The dark band in each graph represents the results from well-refined structures; the central line is a least squares fit to mean trend as a function of resolution, while the width of the band on either side of it corresponds to a variation of one standard deviation about this mean. The results show that the model lies within the allowed region for all six parameters checked. Ramachandran plot of the model shows that 99.8% of the residues lie in the allowed region as shown in Fig. 3e with only 1 residue in disallowed region for the same structure. The VERIFY-3D [35] analysis showed the compatibility 3D-1D score >0.2 to be 99.40% corresponding to acceptable side chain environments. ProQ [45] gave a very good LGScore of 6.03 and a good MaxSub of 0.17 for the model while ERRAT [47] showed the overall quality factor to be 79.59% for the model. The ‘what-if quality report’ [34] results summarized in Table 2 indicate that the refined model showed a Z-score of −2.16 which falls in the acceptable range for a valid structure. The Z-score of ≤−5.0 denotes a poorly refined molecule.
Analysis of the final model after molecular dynamics simulation. a RMSD plot of the MD simulation as a function of timescale b z-plot of final model generated by ProSA. The z-plot shows only chains with less than 1000 residues and a z-score ≤ 10. The z-score of M. tb FadD13 is highlighted as large dot c Energy plot of the final model obtained by ProSA d) Main-chain parameters of the final model as predicted by PROCHECK e) Ramachandran plot of the final model obtained by PROCHECK
Description of the model
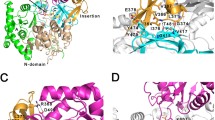
The predicted model for M. tb FadD13 consist of two domains—a large N-terminal domain (residues 1–395) and a small C-terminal domain (402–503) that are connected by a six-amino acid peptide linker, the L motif (residues 396–401). Secondary structure analysis of the model by iMolTalk [55], shows that the structure contains 12 α-helices, eight 310 helices and 26 β-strands (Fig. 4a). The protein belongs to the family of adenylate-forming enzymes that depicts the presence of an A-motif (adenine-binding site; residues 300–306) and P-motif (phosphate-binding site; residues 163–173) which forms the AMP/ATP binding domain, as predicted by Q-SiteFinder [48]. Another conserved 25-amino acid long segment, a fatty-acid binding region (residues 375–399; FACS signature motif), which is common to the family of FACS, was also identified and predicted by CASTp [49]. The motifs were designated based on the structural studies of ttLC-FACS [50] and E. coli FadD [17]. Analysis of the electrostatic potential mapped on to the solvent accessible surface of the model presented in Fig. 4b reveals that the electrostatic potential distribution is markedly different in the binding domains of ATP, CoA and fatty acid.
Three-dimensional model of M. tb FadD13. a Schematic representation of M. tb FadD13. Red color cylinders represent α-helix and blue arrows represent β-sheets. N and C terminals are represented in white color b Electrostatic potential surface map of the protein with the A-motif, P-motif and fatty-acid binding site. Positive potentials are shown in blue, negative potentials in red and neutral in white
Docking studies with substrates
Substrates ATP, CoA and various fatty acids were docked to M. tb FadD13 by using IFD protocol of Schrödinger [51]. ATP and CoA gave a XP Gscore of −9.06 and −9.88, respectively. Fatty acids bind to M. tb FadD13 in the following order of decreasing binding: cerotic acid> lignoceric acid> palmitic acid> capric acid as can be seen from their scores in Table 3. M. tb FadD13 has higher affinity for very long chain fatty acids especially cerotic (26:0) and lignoceric (24:0) acid as compared to palmitic (16:0) / capric (10:0) acid as also observed through experimental studies [21].
Docking was also carried out with multiple ligands in the following order: ATP, fatty acid (lignoceric acid) followed by CoA (Fig. 5a) and the docked complex was refined using Desmond 2.2 [36]. The key amino acids interacting with the substrates were identified as: Gly166, Lys172, Thr304, Glu305, Thr485, Lys487 forming hydrogen bonds with ATP, Tyr362 and Asp371 with fatty acid and Thr167, Thr168, His170 and Tyr362 with CoA as analyzed by LIGPLOT [54] (Fig. 5b).
Docking of multiple ligands (ATP, fatty acid and CoA) to M. tb FadD13 by using induced fit docking. a M. tb FadD13 docked with ATP, lignoceric acid (24:0) and CoA with lignoceric acid shown in pink color, ATP in purple and CoA in blue b Ligplot showing the protein-ligand interactions in M. tb FadD13 complexed with ATP, lignoceric acid and CoA. ATP is represented by Atp 997, lignoceric acid by Faa 998 and CoA by Coa 999
Effect of the mutations at residue Lys487
Mutational studies involving Lys529 of firefly luciferase [18] and Lys592 of propionyl CoA Synthetase (PrpE) enzyme of Salmonella enterica [19] suggest that lysine at this position is a critical residue for effective substrate orientation and favorable interactions leading to efficient adenylate production. This prompted us to investigate the interactions and conformational changes upon ligand binding to the corresponding lysine (K487) in the predicted model of M. tb FadD13. Hence a model of mutant K487A was generated. Interestingly, K487A mutant showed a XP Gscore of −6.59 in comparison to the score of −9.06 in the wild type upon ATP docking as confirmed by experimental investigations that resulted in ∼95% loss of activity [21]. The RMSD calculation between Cα atoms of wild type and mutant model docked with ATP was 3.58 Å after 6 ns of MD, suggesting conformational changes upon ATP binding. The hydrogen bonding distances of ATP docked to the wild and the mutant structure were compared by using LIGPLOT [54]. Figure 6a shows that oxygen atom O3 of ATP is involved in hydrogen bonding with Thr167, O7 and O13 with Thr304 while O13 also forms H-bond with Lys487, O6 and O9 with Lys490 and O11 with Arg397 and Lys399 are involved in hydrogen bonding in the wild type while in the mutated structure, there is no hydrogen bonding between mutated residue (Ala487) and ATP, instead hydrophobic interactions exist, as seen in Fig. 6b. It is observed from protein-ligand interactions in wild and mutant structure that the substrate orientation has completely changed in the mutated protein which may affect the adenylation process.
Effect of the mutation at residue Arg397
Mutation of Arg397 in the fatty acid binding domain results in complete loss of activity as shown by studies carried out in E.coli FadD (R453A) [17] and human ACSL4 (R529S) [20]. We examined the effect of mutation of Arg397 to alanine in the predicted model of M. tb FadD13 by examining docking and binding interactions of the substrates. R397A mutant gave a XP Gscore of 3.26 on fatty acid (cerotic acid) binding in comparison to the score of −5.01 in the wild type suggesting significant decrease in binding. Superimposition of Cα atoms between the wild type and mutant model docked with cerotic acid gave RMS score of 3.83 Å after 6 ns of MD, suggesting a conformational change on the protein structure upon ligand binding. Analysis of protein-ligand contacts using LIGPLOT [54] shows the hydrogen bonding between oxygen atom (O2) of cerotic acid with nitrogen atom (NH1) of Arg397 and NZ atom of Lys399 and O1 with NZ atom of Lys490 as seen in Fig. 7a while only O2 participates in hydrogen bonding with NZ atom of Lys469 in the mutated structure (Fig. 7b) with no interaction with Ala397 signifying a change in substrate orientation.
Protein-ligand contacts of M. tb FadD13 docked with cerotic acid. a Ligplot showing the protein-ligand contacts in M. tb FadD13-cerotic acid complex with cerotic acid represented by Fac 999 b Ligplot showing the protein-ligand contacts in mutant FadD13-cerotic acid complex with cerotic acid represented by Fac 999
Summary
The emergence of multi-drug resistant M. tb strains, coupled with the increasing overlap of the AIDS and TB pandemics has brought TB to the forefront as a major worldwide health concern. No new classes of drugs for TB have been developed in the past 40 years and there is an urgent need for discovering new drug targets.
FadD13, a key gene product of mymA operon of M. tb offers to be a promising drug target for the development of antitubercular agents. In this study, we propose the three-dimensional structure of M. tb FadD13 generated by using a de novo method and further refined by MD simulations. The structure predicted was shown to conform to experimental data revealing the conserved motifs corresponding to ATP/AMP and fatty acid binding. The ATP-AMP binding domain (Y163TSGTTGHPKG173, G300YALTES306) and fatty acid signature motif (N375GWFRTGDIGEIDDEGYLYIKDRLK399) are the conserved motifs of FACS. Key amino acid interactions between substrate and protein were in agreement with the experimentally determined important residues. It is known that residues Lys (487) and Arg (397) are significant for ATP and fatty acid binding, respectively and are conserved throughout the family of Fatty Acyl-CoA Synthetases. Mutations at K487A and R397A resulted in the reduction of binding affinity of ATP and fatty acids, respectively as seen from docking analysis. It also caused change in substrate orientation as observed from protein-ligand interactions in wild and mutant structure. These observations support that these mutations can affect adenylation process and loss of activity as known for other homologues. We hope that the validated model of FadD13 presented in this study will be a step forward toward the design and the development of novel therapeutics against tuberculosis.
References
World Health Organization (2007) Tuberculosis facts http://www.who.int/tb/publications/2007/factsheet_2007.pdf
World Health Organization (2002) Factsheet No 104: tuberculosis. WHO, Geneva http://www.who.int/mediacentre/factsheets/fs104/en/
Snider DE Jr, Raviglione M, Kochi A (1994) Global burden of tuberculosis. In: Bloom BR (ed) Tuberculosis: pathogenesis, protection, and control. American Society for Microbiology, Washington, DC, pp 3–11
Dunlap NE, Bass J, Fujiwara P, Hopewell P, Horsburgh CR Jr, Salfinger M, Simione PM (2000) Diagnostic standards and classification of tuberculosis in adults and children. Am J Respir Crit Care Med 161:1376–1395
Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE III, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Krogh A, McLean J, Moule S, Murphy L, Oliver K, Osborne J, Quail MA, Rajandream MA, Rogers J, Rutter S, Seeger K, Skelto J, Squares R, Squares S, Sulston JE, Taylor K, Whitehead S, Barrell BG (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393:537–544
Mdluli K, Spigelman M (2006) Novel targets for tuberculosis drug discovery. Curr Opin Pharmacol 6:459–467
Hasan S, Daugelat S, Rao PS, Schreiber M (2006) Prioritizing genomic drug targets in pathogens: application to Mycobacterium tuberculosis. PLoS Comput Biol 2:e61
Baker EN (2007) Structural genomics as an approach towards understanding the biology of tuberculosis. J Struct Funct Genomics 8:57–65
Terwilliger TC, Park MS, Waldo GS, Berendzen J, Hung LW, Kim CY, Smith CV, Sacchettini JC, Bellinzoni M, Bossi R, De Rossi E, Mattevi A, Milano A, Riccardi G, Rizzi M, Roberts MM, Coker AR, Fossati G, Mascagni P, Coates AR, Wood SP, Goulding CW, Apostol MI, Anderson DH, Gill HS, Eisenberg DS, Taneja B, Mande S, Pohl E, Lamzin V, Tucker P, Wilmanns M, Colovos C, Meyer-Klaucke W, Munro AW, McLean KJ, Marshall KR, Leys D, Yang JK, Yoon HJ, Lee BI, Lee MG, Kwak JE, Han BW, Lee JY, Baek SH, Suh SW, Komen MM, Arcus VL, Baker EN, Lott JS, Jacobs W Jr, Alber T, Rupp B (2003) The TB structural genomics consortium: a resource for Mycobacterium tuberculosis biology. Tuberculosis (Edinb) 83:223–249
Celia W, Goulding L, Jeanne P, Daniel A, Michael RS, Duilio C, Marcin IA, Sum C, Angineh P, Shui-Shu W, Yim W, Vicent C, Harindarpal SG, David E (2003) Structural genomics of Mycobacterium tuberculosis: a preliminary report of progress at UCLA. Biophys Chem 105:361–370
Payne DJ (2004) The potential of bacterial fatty acid biosynthetic enzymes as a source of novel antibacterial agents. Drug News Perspect 17:187–194
Wright HT, Reynolds KA (2007) Antibacterial targets in fatty acid biosynthesis. Curr Opin Microbiol 10:447–453
Daffe M, Draper P (1998) The envelope layers of mycobacteria with reference to their pathogenicity. Adv Microb Physiol 39:131–203
Singh A, Gupta R, Vishwakarma RA, Narayanan PR, Paramasivan CN, Ramanathan VD, Tyagi AK (2005) Requirement of the mymA operon for appropriate cell wall ultrastructure and persistence of Mycobacterium tuberculosis in the spleens of guinea pigs. J Bacteriol 187:4173–4186
Fisher MA, Plikaytis BB, Shinnick TM (2002) Microarray analysis of the Mycobacterium tuberculosis transcriptional response to the acidic conditions found in phagosomes. J Bacteriol 184:4025–4032
Groot PHE, Scholte HR, Hulsmann WC (1976) Fatty acid activation: specificity, localization, and function. Adv Lipid Res 14:75–126
Black PN, Zhang Q, Weimar JD, DiRusso CC (1997) Mutational analysis of a fatty acyl-coenzyme A synthetase signature motif identifies seven amino acid residues that modulate fatty acid substrate specificity. J Biol Chem 272:4896–4903
Branchini BR, Murtiashaw MH, Magyar RA, Anderson SM (2000) The role of lysine 529, a conserved residue of the acyl-adenylate-forming enzyme superfamily, in firefly luciferase. Biochemistry 39:5433–5440
Horswill AR, Escalante-Semerena JC (2002) Characterization of the propionyl-CoA synthetase (PrpE) enzyme of Salmonella enterica: residue Lys592 is required for propionyl-AMP synthesis. Biochemistry 41:2379–2387
Meloni I, Muscettola M, Raynaud M, Longo I, Bruttini M, Moizard MP, Gomot M, Chelly J, des Portes V, Fryns JP, Ropers HH, Magi B, Bellan C, Volpi N, Yntema HG, Lewis SE, Schaffer JE, Renieri A (2002) FACL4, encoding fatty acid-CoA ligase 4, is mutated in nonspecific X-linked mental retardation. Nat Genet 30:436–440
Khare G, Gupta V, Gupta RK, Gupta R, Bhat R, Tyagi AK (2009) Dissecting the role of critical residues and substrate preference of a fatty acyl-CoA synthetase (FadD13) of mycobacterium tuberculosis. PLoS ONE 4:e8387. doi:10.1371/journal.pone.0008387
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 50:3389–3402
Bernstein FC, Koetzle TF, Williams GJ, Meyer EF Jr, Brice MD, Rodgers JR, Kennard O, Shimanouchi T, Tasumi M (1977) The protein data bank: a computer-based archival file for macromolecular structures. J Mol Biol 112:535–542
Higgins D, Thompson J, Gibson T, Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Finn RD, Tate J, Mistry J, Coggill PC, Sammut JS, Hotz HR, Ceric G, Forslund K, Eddy SR, Sonnhammer EL, Bateman A (2008) The Pfam protein families database. Nucleic Acids Res 36:D281–D288
Lambert C, Leonard N, De Bolle X, Depiereux E (2002) ESyPred3D: prediction of proteins 3D structures. Bioinformatics 18:1250–1256
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Prime 2.0 (2008) Schrödinger, LLC, New York, NY
Eswar N, Marti-Renom MA, Webb B, Madhusudhan MS, Eramian D, Shen M, Pieper U, Sali A (2006) Comparative protein structure modeling with MODELLER. In: Curr Protoc Bioinformatics, Supplement 15, pp 5.6.1–5.6.30
Bennett-Lovsey RM, Hebert AD, Sternberg MJE, Kelley LA (2008) Exploring the extremes of sequence/structure space with ensemble fold recognition in the program Phyre. Proteins 70:611–625
Fujitsuka Y, Chikenji G, Takada S (2005) SimFold energy function for de novo protein structure prediction: consensus with Rosetta. Proteins 62:381–398
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Vriend G (1990) WHAT IF: a molecular modeling and drug design program. J Mol Graph 8:52–56
Lüthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356:83–85
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossvary I, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In: Proceedings of the ACM/IEEE Conference on Supercomputing (SC06), Nov 11–17, Florida
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc 118:11225–11236
Jorgensen WL, Madura JD (1985) Temperature and size dependence for monte carlo simulations of TIP4P water. Mol Phys 56:1381–1392
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N.log (N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comp Phys 23:327–341
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics method. J Chem Phys 81:511–519
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101:4177–4189
Liu DC, Nocedal J (1989) On the limited memory BFGS method for large scale minimization. Math Program 45:503–528
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Wallner B, Elofsson A (2003) Can correct protein models be identified? Protein Sci 12:1073–1086
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519
Laurie ATR, Jackson RM (2005) Q-SiteFinder: an energy-based method for the prediction of protein-ligand binding sites. Bioinformatics 21:1908–1916
Dundas J, Ouyang Z, Tseng J, Binkowski A, Turpaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acid Res 34:W116–W118
Hisanaga Y, Ago H, Nakagawa N, Hamada K, Ida K, Yamamoto M, Hori T, Arii Y, Sugahara M, Kuramitsu S, Yokoyama S, Miyano M (2004) Structural basis of the substrate-specific two-step catalysis of long chain fatty acyl-CoA synthetase dimer. J Biol Chem 279:31717–31726
Schrödinger Suite 2009 Induced Fit Docking protocol. Glide 5.5 (2009) Schrödinger, LLC, New York, NY. Prime 2.1 (2009) Schrödinger, LLC, New York, NY
LigPrep 2.2 (2005) Schrödinger, LLC, New York, NY
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49:6177–6196
Wallace AC, Laskowski RA, Thornton JM (1995) LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng 8:127–134
Diemand AV, Scheib H (2004) iMolTalk: an interactive, internet-based protein structure analysis server. Nucleic Acids Res 32:W512–W516
Acknowledgments
This work is supported by funding from Department of Biotechnology, Ministry of Science and Technology, Govt. of India. NJ is thankful to University Grants Commission for providing fellowship. GK is grateful to the Council of Scientific and Industrial Research for providing fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jatana, N., Jangid, S., Khare, G. et al. Molecular modeling studies of Fatty acyl-CoA synthetase (FadD13) from Mycobacterium tuberculosis—a potential target for the development of antitubercular drugs. J Mol Model 17, 301–313 (2011). https://doi.org/10.1007/s00894-010-0727-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0727-3