Abstract
Tuberculosis (TB) is a devastating disease of worldwide importance. The availability of the genome sequence of Mycobacterium tuberculosis (Mtb), the causative agent, has stimulated a large variety of genome-scale initiatives. These include international structural genomics efforts which have the dual aim of characterising potential new drug targets and addressing key aspects of the biology of Mtb. This review highlights the various ways in which structural analysis has illuminated the biological activities of Mtb gene products, which were previously of unknown or uncertain function. Key information comes from the protein fold, from bound ligands, solvent molecules, ions etc. or from unexpectedly modified amino acid residues. Most importantly, the three dimensional structure of a protein permits the integration of data from many sources, both bioinformatic and experimental, to develop testable functional hypotheses. This has led to many new insights into TB biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
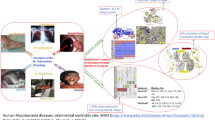
The availability of complete genome sequences for large numbers of microbial species presents an unprecedented opportunity to understand how living species can exploit and adapt to different environments. In the case of those that are human pathogens this also makes it possible to gain new understanding of the origins and mechanisms of disease. No bacterial species is more significant in this regard than Mycobacterium tuberculosis (Mtb), which is the causative agent of tuberculosis (TB). Worldwide deaths from TB total 2–3 million annually [1, 2], more than for any other infectious disease [3]. Although effective drugs exist, current therapy requires prolonged treatment using 3–4 drugs over a period of 6–9 months, leading to compliance problems and the emergence of multi-drug resistance [4]. Moreover, the phenomenon of persistence (see below) means that a huge reservoir of latent TB exists, with one-third of the world’s population infected and at risk of reactivation [1, 5]. This has led to a deadly synergy with HIV/AIDS [2].
The organism itself is extremely slow-growing, with a doubling time of ∼24 h, and has a number of features that make it particularly difficult to combat. It has a thick, waxy, cell wall that is rich in novel lipids, glycolipids and polysaccharides [6], many of which are important in host pathogenesis, and which provide a challenging barrier to drugs and other small molecules. Most importantly, it can enter a persistent state after engulfment by activated macrophages in the lung [7], and is able to survive in this state for many years, to be reactivated as active tuberculosis later in life [5]. In this state of non-replicating persistence (NRP), sometimes referred to as dormancy, it remains metabolically active but is believed to undergo a switch in metabolism, utilising host lipids as an energy source [8]. Crucially, current drugs, most of which target only actively growing bacteria, are largely ineffective against bacteria in the NRP state [9]. This makes it of the utmost importance to understand how Mtb survives the initial onslaught inside macrophages, how it enters the persistent state, and what metabolic processes it depends upon.
The publication of the complete genome sequence for Mtb (H37Rv strain) in 1998 [10], with the identification of the ∼3900 open reading frames (ORFs) that encode proteins within the organism, and functional annotation of many of them, transformed TB research worldwide. Many distinctive and unusual features were noted. In addition to genes encoding the biosynthetic apparatus for the diverse array of lipids used by Mtb—including the characteristic mycolic acids—a large array of ∼250 distinct enzymes involved in fatty acid degradation testified to the importance of lipids in the Mtb lifestyle. Other features of the genome included a relative paucity of two-component regulatory systems, possibly offset by a family of eukaryotic-like Ser/Thr protein kinases; a large number of polyketide synthase systems; and an extraordinary proportion (∼10%) of the coding capacity of the genome dedicated to two mycobacteria-specific protein families of unknown function, the so-called PE and PPE families [10].
In the initial functional annotation of the Mtb genome, functions were attributed to ∼40% of gene products, some information or similarity could be found for another 44%, many of which were conserved hypotheticals, and 16% were described as unknowns, being found only in Mtb or in other mycobacteria. As for other genome annotations, the biological activities of gene products were mostly inferred from sequence similarities with homologous proteins from other organisms, and in many cases only “low-resolution” functions could be deduced, meaning that a protein could be described as an acyl-CoA dehydrogenase, or lipase/esterase or transcription factor, without knowing what its specific substrate or role is. A conservative estimate is that at least 65% of gene products are of unknown or uncertain function. Lastly, there are clearly many novel biochemical pathways that have not yet been delineated, and known pathways that differ in detail from those in other organisms.
The availability of genome sequence information has stimulated many genome-scale investigations of the biology of Mtb. These have included, for example, bioinformatic analyses to identify secreted proteins [11] or iron-regulated proteins [12]; microarray analyses of genes involved in the hypoxic response [13] (believed to model the onset of persistence) or genes whose expression is altered by exposure to drugs [14]; transcriptome analyses [15]; and genome-wide transposon mutagenesis studies to identify genes that are essential for growth [16], or for survival in a mouse model for TB [17], or for Mtb adaptation and survival in macrophages [18]. A feature of all of these studies is the large number of proteins which are implicated in key aspects of Mtb biology or are relevant to disease, but which are of unknown biochemical function.
It is in this context that the TB Structural Genomics Consortium (TBSGC) was formed in 2000, focusing on Mtb, as one of the seven original initiatives funded by the U.S. National Institutes of Health under their Protein Structure Initiative (PSI). The TBSGC (http://www.webtb.org/) differs from the other PSI consortia, however, in both its goals and its mode of operation. Given the global impact of TB, the TBSGC has chosen to operate in a globally inclusive manner, with a coordinated programme that currently involves approximately 400 members in 80 centres around the world, including national structural genomics efforts in Germany, India and New Zealand. The focus is firmly on function, targeting proteins that are potential new drug targets or are believed to play key roles in Mtb biology [19]. When European efforts within the SPinE project (Structural Proteomics in Europe) are added, together with the results of conventional structural biology efforts inspired by the importance of the organism and the availability of its genome sequence, the overall impact has been huge. Structures are now available for ∼200 unique Mtb proteins (Table 1), together with a further ∼250 ligand complexes or additional structures. Two-thirds come from structural genomics initiatives with the other third from individual structural biology efforts. This compares with only 8 Mtb protein structures in the Protein Data Bank at the time the genome sequence was published.
The general issue of inferring function from structure has been much discussed, for example [20], and several articles have reviewed the results from structural genomics initiatives [21–24]. This article focuses specifically on Mtb, taking selected examples that come mostly from Mtb structural genomics efforts. The choice is necessarily selective, with an emphasis on examples from our own laboratory, but these examples are representative of functional discoveries from many laboratories.
Functional clues from the protein fold
The fold of a protein provides a powerful guide to evolutionary relationships because of the need to retain a stable three-dimensional structure during protein evolution. Thus, the fold tends to be strongly conserved in protein families even when little or no sequence identity remains, and can provide strong clues to function. The nature of these functional clues spans a spectrum of possibilities. For proteins of completely unknown function, structural homology with already-characterised proteins can provide testable hypotheses as to potential substrates or even specific functional roles. Typical examples include a family of Mtb proteins with “hotdog” folds [25, 26] that are found to act on thiol ester substrates and are in some cases essential for Mtb viability, and Rv1155, where the structural analysis led to the recognition that this gene product is an enzyme that catalyses the terminal step in pyridoxal 5’-phosphate biosynthesis [27]. In favourable cases, as for the PIN-domain proteins [28], described below, this can open up a new area of biology of an organism. Even when a specific function is not clear, the fold provides a firm basis for further biological studies, as for the PE/PPE proteins [29]. Where a function is already proposed, the fold can provide confirmation and point to key mechanistic details, as for the Mtb cyclopropane synthases [30], or even provide completely unexpected and transformational insights, as for the fluoroquinolone resistance protein MfpA [31].
PE and PPE proteins
The PE and PPE protein families, named for the presence of conserved Pro and Glu residues near their N-termini, represent one of the most striking discoveries from mycobacterial genomes; the M. tuberculosis H37Rv genome contains ∼100 PE and more than 60 PPE proteins [10]. The PE proteins have a conserved N-terminal domain of ∼110 residues, followed by a highly variable C-terminal region that often contains multiple copies of polymorphic repeat sequences. Likewise the PPE proteins have a conserved N-terminal domain of ∼180 residues, followed by variable C-terminal regions. Functions in immune evasion, antigenic variation, cell interactions and virulence have been suggested. However, structural studies of PE and PPE proteins have proved extremely difficult. In one attempt, of 28 PE or PPE proteins targeted, all except one were either insoluble or failed to express, and the one soluble protein was unfolded [29].
A key step forward was the realisation, based on the observation that PE and PPE proteins were often close together, in pairs, in the Mtb genome, that PE/PPE protein pairs may form complexes. Co-expression of one such pair, comprising Rv2430c (a small PPE protein) and Rv2431c (a small PE protein), gave soluble protein and crystals. The crystal structure [29] shows an extended α-helical structure in which the PE protein forms an antiparallel pair of helices that pack against two of the five helices of the PPE protein (Fig. 1a). It is immediately obvious why the proteins would be insoluble on their own, and the PE and PPE motifs are seen to be conserved for their structural roles. Prediction of function is more difficult. The most likely role is in signalling; the closest structural homologue for the PPE protein is the cytoplasmic domain of a serine chemotaxis receptor [32] and there are suggestions that some of the proteins are cell wall associated. At the very least, the structure is a major step forward to understanding proteins that are so dominant in the Mtb genome.
Functional inferences from protein folds. (a) The PE/PPE protein complex formed by the gene products of Rv2430c (PPE, in green) and Rv2431c (PE, in orange). The PE and PPE sequence motifs are close to the N-termini of the two polypeptides, and are probably conserved for structural reasons. (b) The dimer of the PIN-domain protein PAE2754, with residues conserved between the PIN-domain proteins in Mtb and P. aerophilum shown in stick mode. Four conserved acidic residues in each molecule form a metal-binding site (M) that is essential for their nuclease activity. (c) The dimer of the Mtb fluoroquinolone resistance protein MfpA, showing the polypeptide folding (above) and a surface representation (below) highlighting its strongly electronegative character. In size, shape and electrostatics, MfpA mimics DNA. (d) The cyclopropane synthase CmaA1 (Rv3392c), showing its characteristic SAM-dependent methyltransferase fold. S-adenosyl homocysteine is bound at the edge of the β-sheet, marking the cofactor binding site, and the detergent molecule cetyltrimethylammonium bromide binds in a long hydrophobic tunnel which is believed to be the binding site for the mycolic acid substrate
PIN-domain proteins
PIN domains are small proteins (∼16 kDa) that make up a very large family with representatives in all three domains of life. Structural analysis, undertaken in order to address their function, began with the recognition of four PIN domain proteins in the Mtb genome (Rv0065, Rv0549, Rv0960 and Rv1720) and another four in the P. aerophilum genome, one of which (PAE2754) was readily crystallized. The structure [28] revealed a tetramer, formed as a dimer of dimers, in which each monomer had an α/β fold (Fig. 1b). Initial searches of the protein structural database with DALI gave only weak hits (Z-scores ∼3) to proteins of disparate function. The key insight, however, came with recognition of a structural similarity to phage T4 ribonuclease H [33]. The DALI score was poor (Z = 2.8; 84 residues matching with an rms difference of 3.6 Å, and 10% sequence identity) but crucially four acidic residues that are conserved across all PIN domains matched four acidic residues that bound Mg2+ at the active site of ribonuclease H. This led to the hypothesis that PAE2754 was an Mg2+-dependent exonuclease—subsequently confirmed by biological assay [28]—and that all PIN domains may be functional nucleases.
A postscript to this analysis, with potentially profound implications for Mtb biology, comes from the recognition that in prokaryotes most PIN-domain proteins are encoded as the toxin component of toxin-antitoxin gene cassettes [34, 35]. Toxin-antitoxin protein pairs were first recognised for their role in plasmid maintenance during cell division, but current evidence points to a more general role in facilitating persistence in organisms that inhabit variable and frequently stressful environments. Remarkably, the Mtb genome contains no fewer than 48 PIN-domain proteins, of which 38 are encoded adjacent to antitoxin-like genes [36]. This remarkable expansion points to an important biological role in growth or survival, perhaps in persistence within activated macrophages.
Cyclopropane synthases
The mycolic acids, which are a characteristic feature of the mycobacterial cell wall, possess long hydrocarbon chains whose structural diversity is achieved by post-synthetic modifications. One of these which appears to be strongly correlated with persistence is the formation of cis and trans cyclopropanes by methyl transfer to double bonds in the unsaturated meromycolate chain [9]. The Mtb genome sequence led to the identification of a family of homologous genes that were believed to encode cyclopropane synthases or related methyltransferases that catalyse these important modifications. The structural analyses of three of these proteins, PcaA, CmaA1 and CmaA2, revealed a highly conserved fold (Fig. 1d) that clearly identified them as S-adenosyl-L-methionine (SAM)-dependent methyltransferases [30]. Importantly, the crystal structures also revealed two other intriguing features: a long hydrophobic tunnel extending from the protein surface to the SAM binding site, and a putative bicarbonate ion at the active site of each protein. For both CmaA1 and CmaA2, the hydrophobic tunnel is shown to bind lipid-like detergents which model the mycolic acid lipid chain, suggesting a common mechanism of action and the possibility that a single drug could be developed against all three enzymes [30].
The fluoroquinolone resistance protein, MfpA
A spectacular example of the way in which protein fold can illuminate function comes from the structural analysis of MfpA, an Mtb protein implicated in resistance to fluoroquinolone antibiotics [31]. The amino acid sequence of MfpA has a pentapeptide repeat, in which every fifth amino acid is either Leu or Phe, and the structure revealed a β-helix fold (Fig. 1c) that has striking similarities in size, shape and charge distribution to B-form DNA. This led to the hypothesis that fluroquinolone resistance arises from the ability of this protein to bind to DNA gyrase, thus depriving fluroquinolones of their normal target, the DNA gyrase-DNA complex.
Discovery of bound ligands or other species
All crystallographic analyses carry with them the possibility that other species may be found in the crystal structure. These may be cofactors, substrate or product molecules, metal ions, anions such as phosphate or sulfate, buffer molecules, crystallization additives or cryoprotectant molecules (glycerol is particularly common). They may be carried over from the cellular environment from which the protein was isolated, or have been introduced during purification or crystallization. In our own laboratory at least 50% of solved structures have such “extra” density. There is still a challenge in identifying what these bound species are and what their significance is, but they can offer very important insights into function.
The lipid transfer protein, LipB
The post-translational modification of proteins with lipoyl moieties is important for the function of some multicomponent enzyme complexes and is also strongly implicated in host-pathogen interactions for some pathogenic bacteria. In Mtb, the expression of a putative lipid transfer protein LipB is strongly up-regulated in the lungs of patients with multi-drug resistant TB [15]. This protein was known from sequence similarities to be a distant member of a family of ligases that includes biotin ligase (BirA) and lipoyl protein ligase A (LplA).
The structure of LipB, determined at 1.08 Å resolution [37], showed that it is indeed homologous with LplA and BirA. A completely unexpected finding, however, was the presence of decanoic acid, covalently attached to the thiol of Cys176 (Fig. 2a). The decanoic acid moiety, whose identity was confirmed by mass spectrometry, was assumed to be derived from the E. coli host cells used for expression; use of the expression host M. smegmatis, which, like Mtb, has a different population of endogenous lipids, led to no such modification. Importantly, the bound decanoic acid provided a model for octanoic acid, the natural substrate of LipB, identifying the hydrophobic tunnel inside which the lipid moiety binds, and showing that the head group would be perfectly placed alongside Cys176 to form a thioester acyl intermediate. The structure further pointed to an unprecedented cysteine/lysine acyltransferase activity for LipB, and a model in which a lysine residue from an acceptor protein would insert into the active site to form the final amide-like octanoyl-protein linkage.
Ligand binding to Mtb proteins provides important functional insights. (a) A decanoic acid molecule, derived from the E. coli expression host, was found bound to the lipoyl transferase protein LipB (Rv2217), where it is covalently attached to Cys176 and models the natural octanoic acid substrate. (b) The active site of the mycothiol deacetylase MshB (Rv1170) showing the metal ion (magenta sphere) bound to three invariant residues, His13, Asp16 and His147, adjacent to an N-acetylglucosamine substrate molecule, modelled into the position of the glucosyl moiety of the detergent β-octylglucoside, used in crystallization. (c) The active site of PAE2307 showing the binding site for the adenosine substrate, adjacent to His85, and (in the inset) electron density showing that His85 was phosphorylated in the native structure. (d) The Mtb acyltransferase Rv1347c, identified as being responsible for acylation of the N-hydroxylysine side chain of the siderophore mycobactin. The density shows the binding site for a long, hydrophobic acyl group, which was occupied by the detergent β-octylglucoside in the native structure
Rv1170 from M. tuberculosis
The gene product of the open reading frame (ORF) Rv1170 was originally annotated as “conserved hypothetical”, having homologues of unknown function in a number of bacteria of the order actinomycetes, including mycobacteria and streptomycetes. Subsequent genetic studies identified it as being involved in the biosynthesis of mycothiol. Mycothiol is a novel disaccharide, 1-D-myo-inosityl-2-(N-acetyl-l-cysteinyl)amido-2-deoxy-α-d-glucopyranoside, which is an antioxidant and key protective agent that maintains a reducing environment inside cells [38]; it is the mycobacterial equivalent of glutathione. Rv1170 was shown to function as a metal-dependent deacetylase, cleaving the acetyl group from the N-acetylglucosamine (NAG) moiety of mycothiol [39].
The structure, determined at 1.9 Å resolution [40], revealed an α/β fold in which helices pack against a 7-stranded mostly parallel β-sheet. Structure comparisons identified superficial similarities to many Rossmann-fold domains, but it is undoubtedly significant that the two closest structural homologues were UDP-N-acetylglucosamine 2-epimerase and the glycosyltransferase MurG, both of which act on substrates with NAG moieties. Three key observations enabled the active site and a plausible catalytic mechanism to be identified. Firstly, a sequence motif AHPDDE that is invariant in all homologues was located at the bottom of a deep cavity formed by large loops emanating from the C-termini of the β-strands. Secondly, a bound Hg2+ ion, used in structure determination, was coordinated by three invariant residues, His13 and Asp16 from the AHPDDE motif, and His147; this modelled the position of the essential Zn2+ ion, which had been lost due to the use of EDTA during protein expression. Thirdly, the glucosyl moiety of the detergent β-octylglucoside, used in crystallization, was also found to be bound adjacent to the AHPDDE motif and the metal binding site (Fig. 2b). This enabled a testable mechanism to be developed.
A novel adenosine-specific kinase
The ORF Rv3735 from Mtb encodes a member of a family of highly conserved proteins found in bacterial and archaeal species. The high level of conservation suggested some important, but as yet uncharacterised, function. Whereas the Rv3735 protein was insoluble when expressed in E. coli, the homologous protein from the hyperthermophile Pyrobaculum aerophilum, PAE2307, was soluble and its structure could be solved at 1.45 Å resolution [41]. A striking and unexpected discovery in the crystal structure was the presence of a phosphorylated histidine, pHis85 (Fig. 2c). Phosphorylated histidines are rarely observed because of the lability of the P–N bond, and this observation for PAE2307 suggested a phosphoryl transfer function. Adjacent to pHis85 was a pocket lined with more conserved residues, Phe28, Trp95 and Tyr165, into which the adenine base of adenosine could be neatly docked. Subsequent binding studies with a variety of nucleotides and nucleosides showed a strong preference for adenosine over guanosine, thymidine and deoxyuridine, and for adenosine or AMP (KD values of 15 μM and 26 μM respectively) over adenine, ADP or ATP [41]. The crystal structure of PAE2307 in complex with adenosine confirmed the binding mode and led to the conclusion that PAE2307 and Rv3735 are representative of a previously unknown family of adenosine-specific kinases in which a phosphate group is passed from an unknown donor, via His85, to AMP or adenosine [41].
Structure as a means of integrating evidence from other sources
Discovery of function from structure seldom comes from one piece of evidence alone, but the protein structure provides a unique platform for integrating evidence from a diverse array of sources. The following provides just such an example.
Recognition of a “missing” enzyme in mycobactin biosynthesis—Rv1347c
This protein was originally annotated as an aminoglycoside N-acetyltransferase (AAC), with a supposed activity of acetylating amino groups on aminoglycoside antibiotics such as streptomycin and kanamycin, thereby inactivating them. Its sequence identity with known enzymes of this type was less than 15%, however, and no such activity could be demonstrated [42]. Moreover, transposon mutagenesis identified Rv1347c as essential for growth [16], pointing to some other, unknown, function; it is difficult to see why a protein would be essential if its function was to inactivate antibiotics that might never be encountered.
The crystal structure showed that the fold clearly identified Rv1347c as belonging to the GCN5-related N-acetyltransferase (GNAT) family [43], to which the AACs belong (Fig. 1d). These enzymes have the general function of transferring an acyl group from acyl-CoA to a suitable acceptor molecule. Further investigation showed that Rv1347c expression is regulated by iron [44], that its closest homologues are other bacterial proteins involved in the biosynthesis of siderophores (small molecule chelators used to acquire iron), and that neighbouring genes in the Mtb genome are also iron-regulated and are implicated in the biosynthesis of mycobactin, the siderophore used by Mtb to take up iron [45]. Crucially, in the crystal structure “extra” density attributed to a detergent molecule marked a hydrophobic channel leading to the active site and capable of binding a long hydrocarbon chain. The conclusion, which has since been verified experimentally [46], was that Rv1347c is a “missing” enzyme of mycobactin biosynthesis that adds a long-chain acyl group to the N-hydroxylysine side chain of mycobactin.
The problem of incorrect annotations
The annotation of genomes by homology carries with it the risk of incorrect annotations; an incorrect annotation in one genome as a result of a weak sequence similarity or misleading functional data can be propagated through many subsequent genome annotations. Several examples of incorrect annotations in the Mtb genome have been clarified by structural analysis of the gene products [47, 48]. One such an example is given by the Mtb ORF Rv3853. The gene product was originally annotated as MenG, a SAM-dependent methyltransferase in the biosynthetic pathway for menaquinone. The gene was remote from other menaquinone biosynthesis genes in the genome, however, and no methyltransferase activity could be demonstrated. The crystal structure analysis [48] showed that the fold of Rv3853 is completely different from that of known methyltransferases, such as the cyclopropane synthases (Fig. 3), and the conclusion is that this gene has been annotated wrongly. This incorrect annotation is indeed propagated through many bacterial genomes in which homologues exist. The true function of MenG is not clear however. Its E. coli homologue has been identified as an inhibitor of ribonuclease E [49], but MenG homologues are also found in organisms which lack ribonuclease E. Its sequence shows similarity to a family of aldolases, but although the Mtb MenG structure has several binding sites for small molecules, these do not point to any testable function.
Towards an improved understanding of TB biology
The examples described above illustrate a number of ways in which knowledge of protein structure can lead to new functional insights. Nevertheless, the Mtb genome, like those of other organisms, encodes a large number of proteins of unknown or uncertain function, currently estimated at ∼1000 conserved hypotheticals, ∼300 unknowns, and many others that can only be placed in general functional classes [10, 50]. This raises questions as to how priorities should best be established. In the case of Mtb, the growing database of whole-genome microarray studies, proteomic analyses and gene-disruption studies suggests a way forward. By specifically targeting small- or large-scale structural genomics efforts at uncharacterised proteins implicated in key aspects of TB biology—for example, non-replicating persistence—we can expect major advances in understanding and learning to combat this serious human pathogen.
References
Bloom BR, Murray CJ (1992) Science 257:1055
Corbett EL, Watt CJ, Walker N, Maher D, Williams BG, Raviglione MC, Dye C (2003) Arch Intern Med 163:1009
Bloom BR, Small PM (1998) New Engl J Med 338:677
Espinal MA (2003) Tuberculosis 83:44
O’Regan A, Joyce-Brady M (2001) Brit Med J 323:635
Brennan PJ, Nikaido H (1995) Annu Rev Biochem 64:29
Stewart GR, Robertson BD, Young DB (2003) Nature Rev Microbiol 1:97
Munoz-Elias EJ, McKinney JD (2005) Nature Med 11:638
Smith CV, Sharma V, Sacchettini JC (2004) Tuberculosis 84:45
Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE III, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Barrell BG (1998) Nature 393:537
Gomez M, Johnson S, Gennaro ML (2000) Infect Immun 68:2323
Makita Y, Terai G, Mitaku S, Takagi T, Nakai K (2002) Genome Inform 13:297
Sherman DR, Voskuil M, Schnappinger D, Liao R, Harrell MI, Schoolnik GK (2001) Proc Natl Acad Sci USA 98:7534
Wilson M, DeRisi J, Kristensen HH, Imboden P, Rane S, Brown PO, Schoolnik GK (1999) Proc Natl Acad Sci USA 96:12833
Rachmann H, Strong M, Ulrichs T, Grode L, Schuchhardt J, Mollenkopf H, Kosmiadi GA, Eisenberg D, Kaufmann SH (2006) Infect Immun 74:1233
Sassetti CM, Boyd DH, Rubin EJ (2003) Mol Microbiol 48:77
Sassetti CM, Rubin EJ (2003) Proc Natl Acad Sci USA 100:12989
Rengarajan J, Bloom BR, Rubin EJ (2005) Proc Natl Acad Sci USA 102:8327
Terwilliger TC, Park MS, Waldo GS, Berendzen J, Hung L-W, Kim C-Y, Smith CV, Sacchettini JC, Bellinzoni M, Bossi R, De Rossi E, Mattevi A, Milano A, Riccardi G, Rizi M, Roberts MM, Coker AR, Fossati G, Mascagni P, Coates ARM, Wood SP, Goulding CW, Apostol MI, Anderson DH, Gill HS, Eisenberg DS, Taneja B, Mande S, Pohl E, Lamzin V, Tucker P, Wilmanns M, Colovos C, Meyer-Klaucke W, Munro AW, McLean KJ, Marshall KR, Leys D, Yang JK, Yoon H-J, Lee BI, Lee MG, Kwak JE, Han BW, Lee JY, Baek S-H, Suh SW, Komen MM, Arcus VL, Baker EN, Lott JS, Jacobs W, Alber T, Rupp B (2003) Tuberculosis 83:223
Watson JD, Laskowski RA, Thornton JM (2005) Curr Opin Struct Biol 15:275
Eisenstein E, Gilliland GL, Herzberg O, Moult J, Orban J, Poljak RJ, Banerjei L, Richardson D, Howard AJ (2000) Curr Opin Biotechnol 11:25
Teichmann SA, Murzin AG, Chothia C (2001) Curr Opin Struct Biol 11:354
Zhang C, Kim S-H (2003) Curr Opin Chem Biol 7:1
Yakunin AF, Yee AA, Savchenko A, Edwards AM, Arrowsmith CH (2004) Curr Opin Chem Biol 8:1
Castell A, Johansson P, Unge T, Jones TA, Backbro K (2005) Protein Sci 14:1850
Johansson P, Castell A, Jones TA, Backbro K (2006) Protein Sci 15:2300
Biswal BK, Cherney MM, Wang M, Garen C, James MNG (2005) Acta Crystallogr sect D 61:1492
Arcus VL, Backbro K, Roos A, Daniel EL, Baker EN (2004) J Biol Chem 279:16471
Strong M, Sawaya MR, Wang S, Phillips M, Cascio D, Eisenberg D (2006) Proc Natl Acad Sci USA 103:8060
Huang C-C, Smith CV, Glickman MS, Jacobs WR, Sacchettini JC (2002) J Biol Chem 277:11559
Hegde SS, Vetting MW, Roderick SL, Mitchenall LA, Maxwell A, Takiff HE, Blanchard JS (2005) Science 308:1480
Kim KK, Yokota H, Kim S-H (1999) Nature 400:787
Mueser TC, Nossal NG, Hyde CC (1996) Cell 85:1101
Clissold PM, Ponting CP (2000) Curr Biol 10:R888
Gerdes K, Christensen SK, Lobner-Olesen A (2005) Nat Rev Microbiol 3:371
Arcus VL, Rainey PB, Turner SJ (2005) Trends Microbiol 13:360
Ma Q, Zhao X, Eddine AN, Geerlof A, Li X, Cronan JE, Kaufmann SHE, Wilmanns M (2006) Proc Natl Acad Sci USA 103:8662
Newton GL, Fahey RC (2002) Arch Microbiol 178:388
Newton GL, Av-Gay Y, Fahey RC (2000) J Bacteriol 182:6958
McCarthy AA, Peterson NA, Knijff R, Baker EN (2004) J Mol Biol 335:1131
Lott JS, Paget B, Johnston JM, Delbaere LTJ, Sigrell-Simon JA, Banfield MJ, Baker EN (2006) J Biol Chem 281:22131
Draker K-A, Boehr DD, Elowe NH, Noga TJ, Wright GD (2003) J Antibiot (Tokyo) 56:135
Vetting MW, de Carvalho LPS, Yu M, Hegde SS, Magnet S, Roderick SL, Blanchard JS (2005) Arch Biochem Biophys 433:212
Rodriguez GM, Voskuil MI, Gold B, Schoolnik GK, Smith I (2002) Infect Immun 70:3371
LaMarca BBD, Zhu W, Arceneaux JEL, Byers BR, Lundrigan MD (2004) J Bacteriol 186:374
Krithika R, Marathe U, Saxena P, Ansari MZ, Mohanty D, Ghokhale RS (2006) Proc Natl Acad Sci USA 103:2069
Watkins HA, Baker EN (2006) J Bacteriol 188:3589
Johnston JM, Arcus VL, Morton CJ, Parker MW, Baker EN (2003) J Bacteriol 185:4057
Monzingo AF, Gao J, Qiu J, Georgiou G, Robertus JD (2003) J Mol Biol 332:1015
Camus JC, Pryor MJ, Medigue C, Cole ST (2002) Microbiology 148:2967
Acknowledgements
I gratefully acknowledge Heather Baker and Tom Caradoc-Davies for help with the illustrations used in this article; Jim Sacchettini, David Eisenberg, Tom Terwilliger and Matthias Wilmanns for helpful suggestions; and the wider membership of the International TB Structural Genomics Consortium for the stimulating environment in which this research has taken place. Research at the University of Auckland is funded by the Health Research Council of New Zealand, the Foundation for Research, Science and Technology and the Maurice Wilkins Centre, through the Centres of Research Excellence fund, and is the work of many outstanding postdoctoral and graduate student researchers.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Baker, E.N. Structural genomics as an approach towards understanding the biology of tuberculosis. J Struct Funct Genomics 8, 57–65 (2007). https://doi.org/10.1007/s10969-007-9020-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10969-007-9020-9