Abstract
Thermus thermophilus is an extremely thermophilic bacterium that grows between 50 and 80 °C and is an excellent model organism not only for understanding life at high temperature but also for its biotechnological and industrial applications. Multiple molecular capabilities are available including targeted gene inactivation and the use of shuttle plasmids that replicate in T. thermophilus and Escherichia coli; however, the ability to disrupt gene function randomly by transposon insertion has not been developed. Here we report a detailed method of transposon mutagenesis of T. thermophilus HB27 based on the EZ-Tn5 system from Epicentre Biotechnologies. We were able to generate insertion mutations throughout the chromosome by in vitro transposition and transformation with mutagenized genomic DNA. We also report that an additional step, one that fills in single stranded gaps in donor DNA generated by the transposition reaction, was essential for successful mutagenesis. We anticipate that our method of transposon mutagenesis will enable further genetic development of T. thermophilus and may also be valuable for similar endeavors with other under-developed organisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The thermophilic bacterium T. thermophilus has become a model organism for understanding life at extreme temperatures (Cava et al. 2009). It is also a rich source of proteins and macromolecular complexes for which structures have been determined including RNA polymerase (Murakami and Darst 2003), the 70S ribosome (Schmeing and Ramakrishnan 2009), and the Cmr complex of the CRISPR-Cas host-defense system (Staals et al. 2013). Thermophilic bacteria are currently being developed as hosts for the production of biotechnologically relevant products and enzymes, and the expansion of available genetic tools for such organisms is instrumental in advancing their full potential (reviewed in Taylor et al. 2011). T. thermophilus HB27 is particularly well suited as a model thermophile as its genome is completely sequenced (Henne et al. 2004), it is easily cultivated aerobically in the laboratory, it is sensitive to a wide array of antibiotics (Gregory et al. 2005) and the rate of DNA uptake and transformation are particularly unique. T. thermophilus HB27 is naturally competent for transformation with genomic or plasmid DNA with an astonishing transformation efficiency of 10−2 transformants per cell (Koyama et al. 1986) and DNA binding and uptake rates of 40 kb s−1 per cell (Schwarzenlander and Averhoff 2006; reviewed in Averhoff 2009). Additionally, several genetic tools are in place including an E. coli-T. thermophilus shuttle vector (de Grado et al. 1999) and the ability to make targeted gene knockouts by homologous recombination and gene replacement with any of three thermostable antibiotic resistance genes (Hashimoto et al. 2001; Nakamura et al. 2005; Brouns et al. 2005).
A particularly powerful addition to the genetic toolbox would be transposon mutagenesis, as such a capability would enable genome-wide gene disruptions as well as multiple downstream applications. Transposons provide convenient drug-resistance markers greatly facilitating strain construction and genetic mapping of spontaneous mutations (Kleckner et al. 1991; Miller 1992; Berg and Berg 1996). Transposon mutagenesis has for many years served as one of the most powerful tools in bacterial genetics (reviewed in Hayes 2003). Most modern synthetic transposons used for insertional mutagenesis have been engineered to lack a transposase gene, which is carried externally on a transposon delivery vehicle (Way et al. 1984). Such delivery vehicles are not stably maintained, preventing unwanted subsequent transposition. Although in vivo transposition of the naturally-occurring Tn916 via conjugation has been demonstrated for Thermus aquaticus (Sen and Oriel 1990), and active transposition of endogenous IS elements in T. thermophilus has been documented (Gregory and Dahlberg 2008; Swarts et al. 2014), an effective system for in vivo transposon mutagenesis of Thermus spp. has not been described. Thus, a major impediment to progress in the genetic analysis of thermophiles is the lack of characterized transposons that function at high temperatures.
A potentially effective way to circumvent the need for a thermostable transposase is to perform transposition of transposase-deficient elements in vitro, with transposase enzyme provided in trans (Morisato and Kleckner 1987). Several commercially available transposase systems have been developed. For this study, we utilized the customizable EZ-Tn5 system from Epicentre Biotechnologies. This system, based on Tn5 (Reznikoff 2002), is comprised of donor DNA flanked by a pair of inverted repeats, hyperactive 19-bp mosaic ends (ME), which are recognized in vitro by transposase (Goryshin et al. 2000; reviewed in Reznikoff 2008). This in vitro approach yields insertions that are stably integrated in the chromosome since there is no transposase expressed in the cell. Importantly, Tn5 shows no target sequence bias (Reznikoff 2002) which is ideal for random transposon mutagenesis. Here we report a detailed method for transposon mutagenesis of T. thermophilus HB27 using the EZ-Tn5 system, with a modification to the manufacturer’s protocol that was absolutely essential for successful implementation. We believe this will be extremely useful for expanding the capacity of T. thermophilus as a model organism and that our protocol variation may be advantageous for other organisms for which transposon mutagenesis has thus far been recalcitrant.
Materials and methods
Bacterial strains, plasmids and growth conditions
The EZ-Tn5 pMOD-3 <R6Kγori/MCS> transposon construction vector (catalog number TNP10623) and electrocompetent E. coli TransforMax EC100D pir-116 cells (catalog number EC6P095H) were obtained from Epicentre Biotechnologies (Madison, WI). T. thermophilus HB27 (ATCC BAA-163) was grown in ATCC medium 1598 (Thermus enhanced medium, TEM) at 65 °C and was supplemented with 2.8 % BD Bacto Agar for growth on solid media. T. thermophilus HB8 was obtained from ATCC (27634) and plasmid pTT8 was isolated using a Qiagen Miniprep kit from 6 ml of culture. Kanamycin was purchased from Fisher Scientific and used at 30 or 50 μg/ml for T. thermophilus and E. coli, respectively. Hygromycin B was from Sigma and used at 50 μg/ml. Cultures were archived in 25 % glycerol and stored at −80 °C.
Preparation of T. thermophilus chromosomal DNA
T. thermophilus chromosomal DNA was isolated from a 3 ml saturated overnight culture as follows. Pelletted cells were lysed with 25 μl of lysozyme solution (5 mg lysozyme dissolved in 0.5 ml 250 mM Tris–HCl, pH 8) on ice for 15 min. The lysate was treated with 50 μl STEP solution (1 % SDS; 50 mM Tris–HCl, pH 7.6; 50 mM EDTA; 1 mg/ml proteinase K) at 65 °C for 15 min, followed by RNase treatment (5 μl; Sigma-Aldrich R4642) at 37 °C for 30 min. Tubes were chilled on ice 5 min before adding 110 μl Protein Precipitation Solution (Promega catalog number A7951), vortexed vigorously and centrifuged. These large fragments of DNA were precipitated with isopropanol, washed with 70 % ethanol and resuspended in TE buffer.
Transposon generation and transposition
The slpA promoter and kat gene were amplified as a unit from plasmid pMK18 (de Grado et al. 1999) with primers slpA-kat-2 and slpA-kat-3 (see Online Resource 1 for primers used in this study). This was inserted using PstI and BamHI into the MCS of EZ-Tn5 pMOD-3 <R6Kγori/MCS> (Epicentre Biotechnologies, Madison, WI) an E. coli plasmid containing both the Tn5 mosaic ends and the R6Kγori. The transposon was generated by PCR amplification with primers PCRFP and PCRRP in a 50 μl reaction using high-fidelity Phusion polymerase, then purified with a Qiagen PCR cleanup kit and estimated to be 100 ng/μl by gel electrophoresis. The in vitro transposition reaction was carried out with 1 μl EZ-Tn5 transposase according to the manufacturer’s instructions using 1 μg each of purified transposon DNA and fragmented T. thermophilus HB27 DNA in a 20 μl volume. The reaction was incubated at 37 °C for 2 h, followed by addition of 1X Stop Solution and incubation at 70 °C for 10 min. The entire reaction volume was used to transform T. thermophilus. In the case of pTT8, we used 500 ng each of pTT8 and transposon DNA in a 50 μl transposition reaction.
Transformation of T. thermophilus
For gap repair of DNA, drop dialysis was performed on nitrocellulose filters (Millipore, 0.025 μm) for 15 min at room temperature against 200 ml distilled water. This was incubated with the PreCR Repair Mix from New England Biolabs (catalog number M0309) at 37 °C for 20 min. Transformation of T. thermophilus HB27 with chromosomal DNA was then performed as described (Koyama et al. 1986) using 1.5 ml culture, allowing 3 h of outgrowth after addition of DNA, and plating all cells on selective media. Standard molecular biology techniques and reagents were employed for additional procedures.
Rescue Cloning
Genomic DNA from transposon mutants was isolated and 1 μg was fragmented by restriction digestion with either AatII or SacII in a 20 μl volume at 37 °C for 2 h. Enzymes were inactivated at 65 °C for 20 min prior to adding T4 DNA ligase and 10X buffer in a 23 μl reaction volume to allow for self-ligation and circularization of plasmid clones. Ligase was inactivated at 70 °C for 10 min and the plasmid was desalted by drop dialysis, as described above, prior to transforming electrocompetent E. coli TransforMax EC100D pir-116 cells.
Results
In vitro transposition and transformation
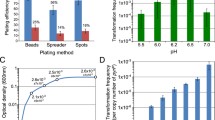
Our overall approach for transposon mutagenesis (Fig. 1) exploits the natural competence of T. thermophilus and its highly efficient natural transformation and homologous recombination system using chromosomal DNA (Koyama et al. 1986). The transposon is generated by PCR to fuse the mosaic ends and an E. coli plasmid replication origin to a selectable marker as described in the “Materials and methods” (Fig. 2a). For this study, our transposon contained the thermostable kat gene controlled by the slpA promoter, which is known to confer kanamycin resistance in both E. coli and T. thermophilus (Lasa et al. 1992). The resulting transposon is incubated with T. thermophilus chromosomal DNA and in vitro transposition is catalyzed with purified transposase. The in vitro transposition reaction generates insertions at random with little or no sequence preference (Reznikoff 2002). T. thermophilus is transformed with mutagenized DNA selecting antibiotic resistance and generates mutants that have undergone recombination with the chromosome via double cross over events at regions of homology flanking the transposon (Fig. 2b). The recombination events generate insertional inactivation of the target gene and could potentially affect expression of downstream genes. Sites of insertion are identified by rescue cloning, in which the transposon and surrounding genomic sequences are extracted and cloned in E. coli by virtue of the plasmid replication origin. Insertion sites are then readily identified by sequencing.
Overall approach for transposon mutagenesis and rescue cloning. PCR-generated transposon shown as a box with blue arrows indicating the inverted repeat mosaic ends (see Fig. 2 for additional details). Transposon and genomic DNA (gDNA) were incubated with transposase in vitro; this was used to transform T. thermophilus. DNA isolated from these mutants was digested using a restriction enzyme and was self-ligated and used to transform pir + E. coli. Rescued plasmid clones containing the transposon and surrounding genomic DNA were isolated and sequenced
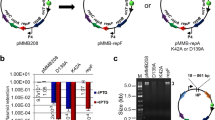
Schematic diagram of EZ-Tn5 transposon generation and mutagenesis. a Orientation of thermostable kanamycin resistance (kat) gene and primers used for PCR amplification of transposon and for sequencing rescued clones. Mosaic ends (ME) in blue; R6Kγori, replication origin in pir + E. coli cells including TransforMax EC100D pir-116. b Insertion of transposon in T. thermophilus by homologous recombination. In vitro-generated mutant chromosomal DNA was used to transform T. thermophilus, selecting kanamycin resistance. Double crossover events at genomic regions of homology generate a collection of insertion mutants in vivo. c Insertion site duplication event. The sequence of the 19-bp mosaic ends recognized by EZ-Tn5 transposase is shown in blue. Numerals 1 through 9 indicate the 9-bp direct repeat target site duplication. The 9-bp gap is illustrated
Initial attempts at transposon mutagenesis were unsuccessful following the manufacturer’s protocol. Although KmR colonies were obtained, rescue cloning (described below) was inefficient and, when possible, sequencing of some clones (primers SqFP and SqRP; Fig. 2a) gave conflicting results indicating insertion at discontiguous regions of the T. thermophilus chromosome. This made reliable identification of true insertion sites impossible. Transposon mutant JC852 lacked the expected 9-bp target site duplication (Fig. 2c), while mutant JC848 also lacked a 9-bp duplication and, instead, carried a 4-bp deletion. The mechanism by which EZ-Tn5 transposase inserts into target DNA involves a 9-bp staggered cut and, in E. coli, these single stranded gaps are repaired in vivo by the DNA replication and repair machinery. We speculated that the 9-bp single stranded gap might be recombinogenic in T. thermophilus and that the overhangs may have been removed by nuclease activity.
This problem was solved by filling in the 9-bp gaps after the in vitro transposition reaction to eliminate any single stranded DNA prior to transformation. To this end, we drop-dialyzed the transposon and initiated DNA repair using the PreCR Repair Mix which includes DNA polymerase and ligase activities. The reaction containing 1 μg of transposon-mutagenized, gap-repaired DNA was used to transform 1.5 ml of T. thermophilus and the entire cell volume was plated on one TEM kanamycin plate. Whereas no colonies arose on the no-DNA control plate over the course of 1 week, colonies were visible on the experimental plate after 1 day while slower growing mutants continued to appear over several days. Most of 186 total mutants were picked after 4 or 5 days of growth and each was streaked once on selective media, then once on non-selective media, and grown in liquid TEM. Saturated cultures were then used to prepare chromosomal DNA and frozen glycerol stocks.
Rescue cloning and identification of chromosomal mutants
One of the advantages of the EZ-Tn5 transposon system is the incorporation of the R6Kγ origin of replication. This plasmid ori is active only in E. coli strains expressing the Π protein, the product of the pir gene. This arrangement allows the excision of the transposon and surrounding genomic DNA by restriction digest, circularization by ligation, and replication in E. coli. The transposition site can then be identified by sequencing the cloned genomic fragment. From the 186 transformants obtained, 66 were examined and successfully cloned in E. coli.
Electrocompetent E. coli TransforMax EC100D pir-116 cells were transformed by electroporation with 5 μl circularized, rescued plasmid clones, selected on LB kanamycin plates and grown in liquid LB kanamycin. Plasmids were isolated using a Qiagen Miniprep kit and digested with KpnI or HindIII to estimate their size for optimal sequencing results. Clones were sequenced with primers SqFP and SqRP which anneal within the transposon and sequence out toward the cloned chromosomal insertion site in either direction (Fig. 2a). Sequencing indicated that the gap-filled transposon reaction worked as anticipated, having generated both the 19-bp ME and the 9-bp target site duplication. Sequence results were used to search the T. thermophilus HB27 genome (GenBank accession number NC_005835.1; Henne et al. 2004) using BLAST (Altschul et al. 1990) to identify sites of transposon insertion. Importantly, results from both sequencing primers were in agreement in all cases and clearly indicated a single insertion site for each transposition event, in contrast to our initial efforts using non-repaired DNA. Insertions at 44 distinct genetic loci were identified among the 66 clones including, in some cases, multiple insertional events in the same gene (Table 1). All clones resulted from independent transposition events, having different insertion sites within the particular gene (see Online Resource 2 for all insertion site sequences).
The mutants listed in Table 1 highlight several interesting insertional events including the disruption of rpsI encoding ribosomal protein S9 (JC1094). S9 is characterized by an N-terminal globular domain and an extended C-terminal tail, which winds deeply through the internal structure of the 30S subunit and makes extensive contact with 16S rRNA. Insertion of the transposon generates an altered S9 C-terminus including an extension of the protein by 58 amino acids; thus we expect that the extended form of S9 will most likely fail to assemble into 30S subunits due to both loss of sequence-specific contacts with 16S rRNA and steric clashes involving the tail extension (Online Resource 3). It is likely that ribosomes from this strain lack S9 which is consistent with S9 being a nonessential protein (Bubunenko et al. 2007). In wild-type ribosomes the C-terminal residues of S9 contact the anticodon of the tRNA in the P site of the 30S (Selmer et al. 2006) and help to maintain reading frame. Alterations of the S9 C-terminus increase translational frameshifting (Näsvall et al. 2009) and these ribosomes exhibit altered translational initiation (Arora et al. 2013). Thus, the absence of functional S9 in this strain is likely to result in erroneous translation, albeit at non-lethal levels. A similar type of insertion was identified (JC1063) in which the C-terminal 18 amino acids of RF-2 (protein release factor 2, prfB) have been swapped for 8 different amino acids. However, RF-2 is an essential protein involved in stop codon recognition and release of the peptide chain from the ribosome (reviewed in Youngman et al. 2008); therefore, this short truncation and alteration is not anticipated to interfere with overall function of RF-2.
Another mutant, JC1077, was found to have a transposon insertion at the Shine Dalgarno (SD) sequence (Shine and Dalgarno 1974), or ribosome binding site of infA encoding translation initiation factor IF-1. IF-1 is an essential protein (Cummings and Hershey 1994) which binds to the ribosomal A site during initiation of protein synthesis in order to direct initiator tRNA binding to the P site (reviewed in Laursen et al. 2005). T. thermophilus infA is imbedded in an operon containing downstream ribosomal protein and RNA polymerase genes that are essential for growth. As a consequence of the mechanism of Tn5 transposition, the target site sequence that contains the SD of infA in JC1077 is duplicated downstream of the transposon, providing a SD for infA. It is interesting nonetheless that this large insertion between the native promoter and the SD apparently does not confer any significant polar effects on IF-1 nor the essential genes downstream.
T. thermophilus HB27 also contains a ~230-kb megaplasmid pTT27 (Accession number NC_005838.1) that is generally characterized as harboring genes that aid growth at high temperature or are involved in UV protection and DNA repair (Brüggemann and Chen 2006). It also contains eight of the ten CRISPR loci in the HB27 genome. We identified four insertions into pTT27 (designated “TT_P”, Table 1). Mutant JC1071 has an insertion in the bgl gene, encoding β-glucosidase (TT_P0042), and has the great potential to serve as a host strain when combined with a plasmid-encoded bgl gene acting as a reporter (Ohta et al. 2006).
Transposon mutagenesis of plasmid pTT8
T. thermophilus HB8 harbors the 9322 bp cryptic plasmid pTT8 (Hishinuma et al. 1978), the sequence of which has been determined (Accession number NC_006463.1; Takayama et al. 2004), and that is absent from the HB27 strain. It is predicted to contain fourteen open reading frames, most of which are annotated as hypothetical proteins. Thus, it would be useful to generate mutants of these genes to gain a better understanding of their function. We also wished to demonstrate, as a proof of principle, that we could mutagenize pTT8 with the EZ-Tn5 transposon and simultaneously create a plasmid that replicates in T. thermophilus and E. coli by virtue of the R6Kγori within the transposon.
It has been shown that a fragment of pTT8 corresponding to the replication origin is active in the HB27 strain (Takayama et al. 2004; Fujita et al. 2012). We amplified the hph gene (primers hphKpnI and hphPstI) from plasmid pMHPnqosGFP (Cava et al. 2008), which confers resistance to hygromycin B in T. thermophilus and E. coli, cloned it into pMOD-3 <R6Kγori/MCS> and selected transformants on LB hygromycin B plates. The transposon was generated by PCR as described in the “Materials and methods”. Unlike the chromosomal mutagenesis we did not repair the single stranded gaps as we directly transformed E. coli TransforMax EC100D pir-116 cells; the gaps are expected to be repaired in vivo in E. coli. Transformants were selected on LB hygromycin B plates, purified once by streaking on selective media, grown in liquid LB with hygromycin B, and plasmids were isolated from 4.5 ml culture using a Qiagen Miniprep kit.
Sequencing of two clones demonstrated that the transposition reaction worked as anticipated and included the 9-bp target site duplication. These mutants were determined to be in gene TTHC004 and between open reading frames TTHC009 and TTHC010, all of which are annotated as hypothetical proteins. Both plasmids were used to successfully transform T. thermophilus HB27, selecting hygromycin B resistance, thus demonstrating their ability to replicate in both E. coli and T. thermophilus. Each plasmid was isolated from T. thermophilus HB27, subjected to restriction digestion, and was determined to be identical to that isolated from E. coli. Notably a lower plasmid yield was obtained from T. thermophilus HB27 compared to E. coli, consistent with the known copy number of eight per cell in T. thermophilus HB8 (Hishinuma et al. 1978).
Discussion
Here, we have described in detail the successful transposon mutagenesis of the extremely thermophilic bacterium T. thermophilus HB27 based on the Epicentre EZ-Tn5 mutagenesis kit. Most notably, we found that modification of the protocol to include a step to repair single stranded DNA gaps in the transposon was absolutely essential. We hypothesize that the presence of single stranded regions of DNA in this instance may have been recombinogenic in T. thermophilus during transformation. DNA uptake in T. thermophilus occurs initially with the binding of double stranded DNA followed by translocation to the inner membrane channel where one strand of DNA is degraded while the other strand proceeds to the cytoplasm where it may recombine with the genome (reviewed in Averhoff 2009). The competence protein DprA is required for high transformation frequency in T. thermophilus (Friedrich et al. 2002) and the DprA homolog in S. pneumoniae is known to promote loading of RecA onto ssDNA (Mortier-Barrière et al. 2007). Thus, the presence of internal regions of single strandedness on donor DNA may promote more than one recombination event per transformation, consistent with our observation that discontiguous regions of DNA were identified in initial rescue cloning attempts.
Our method enabled the identification of transposon insertions at sites throughout the chromosome and megaplasmid including several genes that had been targeted more than once. Specifically, the hypothetical protein TTC0051 was identified four times; mannose-6-phosphate isomerase was identified twice; and both copies of the 16S rRNA gene, rrsA and rrsB, were identified twelve and eight times, respectively. In all cases these were independent, non-clonal isolates with distinct insertion sites (Online Resource 2). As there is no apparent sequence motif common to the insertion sites, we speculate this may relate to the manner of preparation of genomic DNA for in vitro transposition and/or that these regions may have a more accessible structure in vivo if they are highly expressed, as would be expected for rRNA genes.
We believe that the transposon mutagenesis described here will help to expand the development of T. thermophilus as a model thermophile. This opens the door to classic genetic approaches such as promoter probing which could, as an example, be done under variable growth conditions using a promoterless version of the bgl gene as a reporter. Adding to this is the ability to generate by transposition transcriptional or translational fusions that would shed light on specific gene expression or regulation. Transposon mutagenesis of the chromosome may also enable a more comprehensive understanding of how frequently or likely polar inactivation occurs in this organism. Also promising is the potential to explore the biotechnologically relevant CRISPR-Cas systems (Shen 2013) that primarily reside on pTT27, since the megaplasmid is also targeted with this approach. Owing to the natural competence of T. thermophilus with chromosomal DNA there is the opportunity to combine transposon insertions with mutations in other strains, simply by transformation. Similarly, we have the capacity to investigate the functions of the hypothetical genes of plasmid pTT8, as our hygromycin B-resistant derivatives may be combined with various chromosomal mutants and screened for phenotypic interactions.
Significantly, the approach and methods described here have potential for use with other extremophiles. In particular, additional genetic tools are desired for engineering the biotechnologically relevant Thermoanaerobacter and Thermoanaerobacterium spp., which have recently been shown to be naturally competent and transformable with genomic DNA (Shaw et al. 2010). The method may also aid in the genetic development of archaea that are transformable, such as Methanococcus voltae (Bertani and Baresi 1987), Pyrococcus furiosus (Lipscomb et al. 2011), Sulfolobus spp. (reviewed in Atomi et al. 2012), and Halobacterium spp. (Cline et al. 1989). Although transformation with genomic DNA has not been formally demonstrated for some of these organisms, the general approach and the requirement for gap repair may prove insightful in future applications.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Arora S, Bhamidimarri SP, Bhattacharyya M, Govindan A, Weber MH, Vishveshwara S, Varshney U (2013) Distinctive contributions of the ribosomal P-site elements m2G966, m5C967 and the C-terminal tail of the S9 protein in the fidelity of initiation of translation in Escherichia coli. Nucleic Acids Res 41:4963–4975
Atomi H, Imanaka T, Fukui T (2012) Overview of the genetic tools in the Archaea. Front Microbiol 3:337
Averhoff B (2009) Shuffling genes around in hot environments: the unique DNA transporter of Thermus thermophilus. FEMS Microbiol Rev 33:611–626
Berg CM, Berg DE (1996) Transposable element tools for microbial genetics. In: Neidhardt FC (ed) Escherichia coli and Salmonella: cellular and molecular biology. ASM Press, Washington, DC, pp 2588–2612
Bertani G, Baresi L (1987) Genetic transformation in the methanogen Methanococcus voltae PS. J Bacteriol 169:2730–2738
Brouns SJJ, Wu H, Akerboom J, Turnbull AP, de Vos WM, van der Oost J (2005) Engineering a selectable marker for hyperthermophiles. J Biol Chem 280:11422–11431
Brüggemann H, Chen C (2006) Comparative genomics of Thermus thermophilus: plasticity of the megaplasmid and its contribution to a thermophilic lifestyle. J Biotechnol 124:654–661
Bubunenko M, Baker T, Court DL (2007) Essentiality of ribosomal and transcription antitermination proteins analyzed by systematic gene replacement in Escherichia coli. J Bacteriol 189:2844–2853
Cava F, de Pedro MA, Blas-Galindo E, Waldo GS, Westblade LF, Berenguer J (2008) Expression and use of superfolder green fluorescent protein at high temperatures in vivo: a tool to study extreme thermophile biology. Environ Microbiol 10:605–613
Cava F, Hidalgo A, Berenguer J (2009) Thermus thermophilus as biological model. Extremophiles 13:213–231
Cline SW, Lam WL, Charlebois RL, Schalkwyk LC, Doolittle WF (1989) Transformation methods for halophilic archaebacteria. Can J Microbiol 35:148–152
Cummings HS, Hershey JW (1994) Translation initiation factor IF1 is essential for cell viability in Escherichia coli. J Bacteriol 176:198–205
de Grado M, Lasa I, Berenguer J (1998) Characterization of a plasmid replicative origin from an extreme thermophile. FEMS Microbiol Lett 165:51–57
de Grado M, Castan P, Berenguer J (1999) A high-transformation-efficiency cloning vector for Thermus thermophilus. Plasmid 42:241–245
Friedrich A, Prust C, Hartsch T, Henne A, Averhoff B (2002) Molecular analyses of the natural transformation machinery and identification of pilus structures in the extremely thermophilic bacterium Thermus thermophilus strain HB27. Appl Environ Microbiol 68:745–755
Fujita A, Misumi Y, Koyama Y (2012) Two versatile shuttle vectors for Thermus thermophilus-Escherichia coli containing multiple cloning sites, lacZα gene and kanamycin or hygromycin resistance marker. Plasmid 67:272–275
Goryshin IY, Jendrisak J, Hoffman LM, Meis R, Reznikoff WS (2000) Insertional transposon mutagenesis by electroporation of released Tn5 transposition complexes. Nat Biotechnol 18:97–100
Gregory ST, Dahlberg AE (2008) Transposition of an insertion sequence, ISTth7, in the genome of the extreme thermophile Thermus thermophilus HB8. FEMS Microbiol Lett 289:187–192
Gregory ST, Carr JF, Rodriguez-Correa D, Dahlberg AE (2005) Mutational analysis of 16S and 23S rRNA genes of Thermus thermophilus. J Bacteriol 187:4804–4812
Hashimoto Y, Yano T, Kuramitsu S, Kagamiyama H (2001) Disruption of Thermus thermophilus genes by homologous recombination using a thermostable kanamycin-resistant marker. FEBS Lett 506:231–234
Hayes F (2003) Transposon-based strategies for microbial functional genomics and proteomics. Annu Rev Genet 37:3–29
Henne A, Bruggemann H, Raasch C, Wiezer A, Hartsch T, Liesegang H, Johann A, Lienard T, Gohl O, Martinez-Arias R, Jacobi C, Starkuviene V, Schlenczeck S, Dencker S, Huber R, Klenk HP, Kramer W, Merkl R, Gottschalk G, Fritz HJ (2004) The genome sequence of the extreme thermophile Thermus thermophilus. Nature Biotechnol 22:547–553
Hishinuma F, Tanaka T, Sakaguchi K (1978) Isolation of extrachromosomal deoxyribonucleic acids from extremely thermophilic bacteria. J Gen Microbiol 104:193–199
Kleckner N, Bender J, Gottesman S (1991) Uses of transposons with emphasis on Tn10. Methods Enzymol 204:139–180
Koyama Y, Hoshino T, Tomizuka N, Furukawa K (1986) Genetic transformation of the extreme thermophile Thermus thermophilus and of other Thermus spp. J Bacteriol 166:338–340
Lasa I, Castón JR, Fernández-Herrero LA, de Pedro MA, Berenguer J (1992) Insertional mutagenesis in the extreme thermophilic eubacteria Thermus thermophilus HB8. Mol Microbiol 6:1555–1564
Laursen BS, Sørensen HP, Mortensen KK, Sperling-Petersen HU (2005) Initiation of protein synthesis in bacteria. Microbiol Mol Biol Rev 69:101–123
Lipscomb GL, Stirrett K, Schut GJ, Yang F, Jenney FE Jr, Scott RA, Adams MW, Westpheling J (2011) Natural competence in the hyperthermophilic archaeon Pyrococcus furiosus facilitates genetic manipulation: construction of markerless deletions of genes encoding the two cytoplasmic hydrogenases. Appl Environ Microbiol 77:2232–2238
Miller JH (1992) A short course in bacterial genetics: a laboratory manual and handbook for Escherichia coli and related bacteria. Cold Spring Harbor Laboratory Press, New York
Miyazaki T, Miyazaki J, Yamane H, Nishiyama M (2004) alpha-Aminoadipate aminotransferase from an extremely thermophilic bacterium, Thermus thermophilus. Microbiology 150:2327–2334
Morisato D, Kleckner N (1987) Tn10 transposition and circle formation in vitro. Cell 51:101–111
Mortier-Barrière I, Velten M, Dupaigne P, Mirouze N, Piétrement O, McGovern S, Fichant G, Martin B, Noirot P, Le Cam E, Polard P, Claverys JP (2007) A key presynaptic role in transformation for a widespread bacterial protein: DprA conveys incoming ssDNA to RecA. Cell 130:824–836
Murakami KS, Darst SA (2003) Bacterial RNA polymerase: the wholo story. Curr Opin Struct Biol 13:31–39
Nakamura A, Takakura Y, Kobayashi H, Hoshino T (2005) In vivo directed evolution for thermostabilization of Escherichia coli hygromycin B phosphotransferase and the use of the gene as a selection marker in the host-vector system of Thermus thermophilus. J Biosci Bioeng 100:158–163
Näsvall SJ, Nilsson K, Björk GR (2009) The ribosomal grip of the peptidyl-tRNA is critical for reading frame maintenance. J Mol Biol 385:350–367
Nishida H, Nishiyama M (2012) Evolution of lysine biosynthesis in the phylum Deinococcus-Thermus. Int J Evol Biol 2012:745931
Ohta T, Tokishita S, Imazuka R, Mori I, Okamura J, Yamagata H (2006) Beta-Glucosidase as a reporter for the gene expression studies in Thermus thermophilus and constitutive expression of DNA repair genes. Mutagenesis 21:255–260
Reznikoff WS (2002) Tn5 Transposition. In: Craig NL, Cragie R, Gellert M, Lambowitz AM (eds) Mobile DNA II. ASM Press, Washington DC, pp 403–422
Reznikoff WS (2008) Transposon Tn5. Annu Rev Genet 42:269–286
Schmeing TM, Ramakrishnan V (2009) What recent ribosome structures have revealed about the mechanism of translation. Nature 461:1234–1242
Schwarzenlander C, Averhoff B (2006) Characterization of DNA transport in the thermophilic bacterium Thermus thermophilus HB27. FEBS J 273:4210–4218
Selmer M, Dunham CM, Murphy FV 4th, Weixlbaumer A, Petry S, Kelley AC, Weir JR, Ramakrishnan V (2006) Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313:1935–1942
Sen S, Oriel P (1990) Transfer of transposon Tn916 from Bacillus subtilis to Thermus aquaticus. FEMS Microbiol Lett 55:131–134
Shaw AJ, Hogsett DA, Lynd LR (2010) Natural competence in Thermoanaerobacter and Thermoanaerobacterium species. Appl Environ Microbiol 76:4713–4719
Shen H (2013) CRISPR technology leaps from lab to industry. Nature. doi:10.1038/nature.2013.14299
Shine J, Dalgarno L (1974) The 3′-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Nat Acad Sci USA 71:1342–1346
Staals RH, Agari Y, Maki-Yonekura S, Zhu Y, Taylor DW, van Duijn E, Barendregt A, Vlot M, Koehorst JJ, Sakamoto K, Masuda A, Dohmae N, Schaap PJ, Doudna JA, Heck AJ, Yonekura K, van der Oost J, Shinkai A (2013) Structure and activity of the RNA-targeting Type III-B CRISPR-Cas complex of Thermus thermophilus. Mol Cell 52:135–145
Swarts DC, Jore MM, Westra ER, Zhu Y, Janssen JH, Snijders AP, Wang Y, Patel DJ, Berenguer J, Brouns SJJ, van der Oost J (2014) DNA-guided DNA interference by a prokaryotic Argonaute. Nature. doi:10.1038/nature12971
Takayama G, Kosuge T, Maseda H, Nakamura A, Hoshino T (2004) Nucleotide sequence of the cryptic plasmid pTT8 from Thermus thermophilus HB8 and isolation and characterization of its high-copy-number mutant. Plasmid 51:227–237
Taylor MP, van Zyl L, Tuffin IM, Leak DJ, Cowan DA (2011) Genetic tool development underpins recent advances in thermophilic whole-cell biocatalysts. Microb Biotechnol 4:438–448
Way JC, Davis MA, Morisato D, Roberts DE, Kleckner N (1984) New Tn10 derivatives for transposon mutagenesis and for construction of lacZ operon fusions by transposition. Gene 32:369–379
Youngman EM, McDonald ME, Green R (2008) Peptide release on the ribosome: mechanism and implications for translational control. Annu Rev Microbiol 62:353–373
Acknowledgments
This work was supported by grant GM19756 (to AED) from the National Institutes of Health. We thank Sunthorn Pond-Tor for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Carr, J.F., Gregory, S.T. & Dahlberg, A.E. Transposon mutagenesis of the extremely thermophilic bacterium Thermus thermophilus HB27. Extremophiles 19, 221–228 (2015). https://doi.org/10.1007/s00792-014-0663-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-014-0663-8