Abstract
Pairs of PCR primers that targeted the archae/bacteriorhodopsin gene were used to clone the archaerhodopsin (aR) gene of Halorubrum xinjiangense strain BD-1T, and this gene was sequenced and functionally expressed in Escherichia coli. Recombinant E. coli cells harboring the plasmid carrying this gene became slightly purple or blue depending on whether they were supplemented with all- trans retinal or 3,4-dihydroretinal, respectively, during induction with IPTG. The purple and blue membranes from the recombinant E. coli showed maximal absorption at 555 and 588 nm, respectively, which are different from maximal absorption at 568 nm of the wild-type purple membrane. Purple membranes from the recombinant E. coli and from strain BD-1T were investigated in parallel. The E. coli purple membrane was fabricated into films and photoelectric responses were observed that depended on the light-on and light-off stimuli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteriorhodopsins (bR) are of interest because they are potentially useful for the fabrication of biomaterial-based devices such as artificial retinas (Frydrych et al. 2000) and optical memories (Wise et al. 2002). Khodonov et al. (1997) listed many advantages of bR as material for fabrication of devices, and those advantages could be further improved by modifications either of the protein (Wise et al. 2002; Weetall et al. 2000), or of the chromophore (Druzhko and Chamorovsky 1995; Jussila et al. 2001; Drachev et al. 1989). Modification of the chromophore has been done by the replacement of the retinal with various retinal analogs, e.g., 3,4-dihydroretinal (Khodonov et al. 1997) or 4-keto-retinal (Druzhko and Chamorovsky 1995). The procedure for such modification includes usually, bleaching of the purple membrane, and reconstitution in the presence of the retinal analog.
A bR-like protein, archaerhodopsin (aR) from the Halorubrum sp. was identified (Mukohata et al. 1988; Sugiyama et al. 1989; Uegaki et al. 1991). This protein shows about 56–59% homology to the bR protein from Halobacterium spp., and exhibits similar photochemical properties. But aR and bR molecules showed differences in their absorbance maxima, the kinetics of the photocycles, and especially in alkaline-induced red-shifted absorption. In this study, we attempted to identify the aR in Halorubrum xinjiangense strain BD-1T, and further functionally express the gene and modify the gene product in Escherichia coli.
Materials and methods
Bacterial strains, plasmids, medium, and cultivation
H. xinjiangense strain BD-1T (CGMCC 1.3527T =JCM 12388T) was grown at 40°C in the medium as described previously (Feng et al. 2004). E. coli strains (Table 1) were routinely grown at 37°C in Luria–Bertani (LB) medium. E. coli strains JM109 and BL21(DE3) (Novagen) and plasmids pGEM-T (Promega, WA, USA) and pET28a (Novagen) were used as hosts and vectors for gene sequencing and expression, respectively. When required, 100 mg of ampicillin per liter or 50 mg of kanamycin per liter was added to the culture medium.
DNA extraction and restriction enzyme treatment
Genomic DNA and plasmid DNA extraction, and restriction enzyme treatment on plasmid DNA and amplified DNA fragments were carried out according to Sambrook and Russell (2001).
Cloning and sequencing the aR gene from strain BD-1T
A fragment of and the entire aR gene were PCR-amplified with two pairs of primers, (Primers 1, 2 and 3, and 4, Table 1). The PCR was carried out at 30 cycles under the following conditions: denaturation for 1 min at 94°C, annealing for 1 min at 54°C, and extension for 1 min at 72°C. The amplified DNA fragments were inserted into pGEM-T easy vectors and the generated pGEMaR-1 and pGEMaR-2 were transformed into E. coli JM109. DNA sequences were determined by Beijing Genome Institute (Huada Corp., Beijing, China).
Functional expression of aR gene from strain BD-1T in E. coli in the presence of all- trans retinal and 3,4-dihydroretinal
The PCR-amplified aR gene from strain BD-1T was ligated into pET28a at the sites of NcoI and BamHI. The resulting plasmid, pET28aR, was transformed into E. coli BL21(DE3) by electroporation at the following conditions: 25 μF, 12.5 kV/cm, and 200 Ω (ECM630, BTX, USA). Synthesis of aR proteins in recombinant E. coli cells harboring pET28aR were induced by the addition of 1 mM IPTG and 10 μM all- trans retinal or 3,4-dihydroretinal when the culture reached OD600 of 0.4–0.6. After further continuous cultivation for 2 h, cells were harvested by centrifugation at 10,000 g for 5 min at 4°C.
Preparation of aR membranes from strain BD-1T and recombinant E. coli cells
The aR membranes of strain BD-1T were fractionated by sucrose density gradient centrifugation, according to the method described by Oesterhelt and Stoecknius (1974). Purification of aR-membrane fractions from recombinant E. coli were performed in the same way except that the recombinant E. coli cells were broken up by sonification at 4°C (160 W, 3 s sonifying vs. 5 s break, 99 cycles) in Tris–HCl buffer (50 mM Tris–HCl, 5 mM MgCl2, pH 8.0).
Measurement of absorption spectra
The membranes were suspended in a buffer of 50 mM Tris–HCl (pH 8.0). Before recording the absorption of spectra, membrane suspensions were light-adapted for 20 min. Absorption spectra were recorded with a Vis–UV spectrophotometer (Beckman Coulter DU800) with wavelength interval of 0.5 nm in the visible range (400–700 nm). Data were treated using origin 6.0 Software (Microcal Software Inc., Northampton, MA, USA).
Determination of photoelectric properties of fabricated films made from aR-membrane fractions of strain BD-1T and recombinant E. coli
Indium-tin-oxide (ITO) glass slide after negative-charged treatment was immersed into Poly(allylamine hydrochloride) (PAH) aqueous solution (2 mg/mL, pH 6.4) for 5 min, rinsed with doubly distilled water, and then dried by nitrogen flow. This modified glass slide was immersed in aR-membrane suspension (pH 9.4) for 5 min, rinsed with doubly distilled water, and then dried with nitrogen again. In this way, we obtained one bilayer of aR-membrane/PAH (M/PAH) films that were marked as (M/PAH)1. This process was repeated six times. The ITO glass with M/PAH films was used as a working electrode and platinum wire as a counter electrode. The electrolyte solution was 0.5 M KCl, with pH 7.3. To test the photoelectric property of the fabricated M/PAH films, the films were irradiated with light from a 150 W xenon lamp and through the filter (560 nm). Photocurrent generated by the film was measured by using a picoamper with the sensitivity of 0.1 nA and digitalized by using a digital storage oscilloscope (20 MHz, Gold Star) (Chu et al. 2003).
Chemicals and reagents
DNA restriction enzymes, DNA ligase, and DNA polymerase were purchased from Takara or Promega. All- trans retinal was purchased from Sigma. The 3,4-dihydroretinal was synthesized from all- trans retinal by following the method of Drachev et al. (1989). PAH was purchased from Aldrich Chemicals.
The Genbank accession numbers
The Genbank accession number for the aR gene of strain BD-1T is AY510709. Other Genbank numbers of aR or bR proteins are given in Fig. 1.
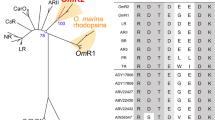
The alignment of nine archaerhodopsin and bacteriorhodopsin proteins that function as proton pump. The names of the bacterial species that the archaerhodopsin and bacteriorhodopsin isolated are given at the beginning and Genbank accession numbers of these proteins are provided at the end. All conserved amino acid residues are shaded in gray
Results
Genetic cloning and characterization of aR gene from strain BD-1T
With Primers 1 and 2 (Table 1), a DNA fragment of 388 bp was PCR-amplified from genomic DNA of strain BD-1T. This fragment showed 93% identity to a part of the aR gene of Halorubrum aus-1, indicating that an aR gene existed in strain BD-1T. A second pair of primers (Primers 3 and 4, Table 1) according to the aR gene sequence of Halorubrum aus-1 was synthesized and used for amplification of the entire aR gene from strain BD-1T. The amplified entire aR gene of strain BD-1T was 777 bp in length, and encoded a protein of 258 amino acids. When this protein was aligned to other known aR or bR proteins, all the amino acid residues that were previously revealed to be essential for proton transport and linkage to retinal were completely conserved (Fig. 1). The aR protein from strain BD-1T showed high identity (86–95%) to the aRs of Halorubrum aus-1, aus-2, and Halorubrum sodomense, and showed relatively low identities to the bacteriorhodopsin (55%) of Halobacterium salinarum(Dunn et al. 1981), to the cruxrhodopsins of Haloarcula spp. (38–47%, Yatsunami et al. 1997; Kitajima et al. 1996; Tateno et al. 1994; Otomo et al. 1992), and to the archaerhodopsin of Haloterrigena (49%, Ihara et al. 1999).
Expression of aR gene and absorption spectra of aR-membranes from E. coli cells
The entire aR gene was PCR-amplified with primers 5 and 6 (Table 1) from strain BD-1T and was ligated to pET28a. The resulting plasmid, pET28aR, was electroporated into E. coli cells. Depending on the presence of all- trans retinal or 3,4-dihydroretinal, recombinant E. coli cells that harbored pET28aR became purple (with all- trans retinal) or grayish blue (with 3,4-dihydroretinal) during cultivation in LB broth and induction with IPTG. The membrane fractions were isolated from the recombinant E. coli cells and their absorption spectra were determined (Fig. 2). The maximal absorption of the purple membrane obtained with all- trans retinal was at 555 nm and of the blue membrane with 3,4-dihydroretinal was 588 nm (Fig. 2), which were different from the maximal absorption at 568 nm of purple membrane from halophilic archaea (Fig. 2 and also Lukashev et al. 1994). The difference in maximal absorption of purple membrane from recombinant E. coli and from Halorubrum sp. was attributed to the monomer state of archaerhodopsin in E. coli and the trimer state in wild membrane, as revealed by Corcelli et al. (2002).
Photoelectric response of film made from wild and recombinant purple membranes
To demonstrate that the aR proteins synthesized in recombinant E. coli are active for proton transport, the purple membranes from both wild strain BD-1 and recombinant E. coli were used to fabricate thin films (Chu et al. 2003). Upon illumination, the film generated electric currents due to proton movement across the film. Figure 3 shows the photoelectric response profiles of the films that were made of purple membranes from either strain BD-1T (Fig. 3a) or recombinant E. coli (Fig. 3b). Positive and anodic responses were corresponding to light-on and light-off photocurrents that were caused by proton release and uptake, respectively. This provided further evidence that the aR proteins in the recombinant E. coli membrane were correctly folded and functionally active.
Discussion
Heterologous expression of bR proteins in E. coli had been previously studied, e.g., Dunn et al. (1987) reported the synthesis and purification of retinal-free bacteriorhodopsin and Hohenfeld et al. (1999) reported the purification of histidine-tagged bR from recombinant E. coli. But to obtain active aR protein or functional membrane, it was necessary to refold the bR protein, in the presence of retinal, and reconstitute with polar lipid to form purple membrane (Dunn et al. 1987; Hohenfeld et al. 1999). Active expression of proteorhodopsin gene from uncultured proteobacteria and phoborhodopsin (a photosensory protein) gene from Natronobacterium pharaonis(NCIMB 2191) was reported (Béjà et al. 2000; Shimono et al. 1998). In this study, we had succeeded in the construction of an E. coli system that produces in one-step the active aR and purple membrane. Moreover, with this system, one aR with all- trans retinal and one aR analog with 3,4-dihydroretinal were synthesized in E. coli. Purple and blue membranes were obtained. We believe that preparation of other aR analogs with different retinal analogs, such as 3-hydroxyretinal and 4-ketoretinal, with this system is possible.
References
Béjà O, Aravind L, Koonin EV, Suzuki MT, Hadd A, Nguyen LP, Jovanovich SB, Gates CM, Feldman RA, Spudich JL, Spudich EN, DeLong EF (2000) Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science 289:1902–1906
Chu JF, Li XC, Zhang JP, Tang JA (2003) Fabrication and photoelectric response of ploly(allylamine hydrochloride)/PM thin films by layer-by-layer deposition technique. Biochem Biophys Res Commum 305:116–121
Corcelli A, Lattanzio VMT, Mascolo G, Papadia P, Fanizzi F (2002) Lipid-protein stoichiometries in a crystalline biological membrane: NMR quantitative analysis of the lipid extract of the purple membrane. J Lipid Res 43:132–140
Drachev LA, Drachev A, Chekulaeva LN, Evstigneeva RP, Kaulen AD, Khitrina LV, Khodonov AA, Lazarova ZR, Mitsner BI (1989) An investigation of the electrochemical cycle of bacteriorhodopsin analogs with the modified ring. Arch Biochem Biophys 270:184–197
Druzhko AB, Chamorovsky SK (1995) The cycle of photochromic reactions of a bacteriorhodopsin analog with 4-keto-retinal. Biosystems 35:133–136
Dunn RJ, Mccoy JM, Simsek M, Majumdar A, Chang SH, Rajbhandary UL, Khorana HG (1981) The bacteriorhodopsin gene. Proc Natl Acad Sci USA 78:6744–6748
Dunn RJ, Hackett NR, McCoy JM, Chao BH, Kimura K, Khorana HG (1987) Structure-function studies on bacteriorhodopsin. I. Expression of the bacterio-opsin gene in Escherichia coli. J Biol Chem 262(19):9246–9254
Feng J, Zhou P, Liu SJ (2004) Halorubrum xinjiangense, a novel halophile from saline lakes of China. Int J Syst Evol Microbiol 54:1789–1791
Frydrych M, Silfsten P, Parkkinen S, Parkkinen J, Jaaskelainen T (2000) Color sensitive retina based on bacteriorhodopsin. Biosystems 54:131–140
Hohenfeld IP, Wegener AA, Engelhard M (1999) Purification of histidine tagged bacteriorhodopsin, pharaonis halorhodopsin and pharaonis sensory rhodopsin II functionally expressed in Escherichia coli. FEBS Lett 442:198–202
Ihara K, Umemura T, Katagiri I, Kitajima-Ihara T, Sugiyama Y, Kimura Y, Mukohata Y (1999) Evolution of the archaeal rhodopsins: evolution rate changes by gene duplication and functional differentiation. J Mol Biol 285:163–174
Jussila T, Tkachenko NV, Parkkinen S, Lemmetyinen H (2001) Kinetics of photo-active bacteriorhodopsin analog 3,4-didehydroretinal. J Photochem Photobiol B 62:128–132
Khodonov AA, Demina OV, Khitrina LV, Kaulen AD, Silfsten P, Parkkinen S, Parkkinen J, Jaaskelainen T (1997) Modified bacteriorhodopsin as a basis for new optical devices. Sens Actuators B 38–39:218–221
Kitajima T, Hirayama J, Ihara K, Sugiyama Y, Kamo N, Mukohata Y (1996) Novel bacterial rhodopsins from Haloarcula vallismortis. Biochem Biophys Res Commum 220:341–345
Lukashev EP, Govindjee R, Kono M, Ebrey TG, Sugiyama Y, Mukohata Y (1994) pH dependence of the absorption spectra and photochemical transformations of the archaerhodopsins. Photochem Photobiol 60:69–75
Mukohata Y, Sugiyama Y, Ihara K, Yoshida M (1988) An Australian halobacterium contains a novel proton pump retinal protein: archaerhodopsin. Biochem Biophys Res Commu 151:1339–1345
Oesterhelt D, Stoeckenius W (1974) Isolation of the cell membrane of Halobacterium halobium and its function into red and purple membrane. In: Fleischer S, Packer L (eds) Methods in enzymology, vol 31, pp 667–678
Otomo J, Urabe Y, Tomioka H, Sasabe H (1992) The primary structures of helices A to G of three new bacteriorhodopsin-like retinal proteins. J Gen Microbiol 138:2389–2396
Sambrook J, Russell DW (2001) Molecular cloning, 3rd edn. CSHL Press, Cold Spring Harbor, New York
Shimono K, Iwamoto M, Sumi M, Kamo N (1997) Functional expression of pharaonis phoborhodopsin in Escherichia coli. FEBS Lett 420:54–56
Sugiyama Y, Maeda M, Futai M, Mukohata Y (1989) Isolation of a gene that encodes a new retinal protein, archaerhodopsin, from Halobacterium sp. aus-1. J Biol Chem 264:20859–20862
Tateno M, Ihara K, Mukohata Y (1994) The novel ion pump rhodopsins from Haloarcula form a family independent from both the bacteriorhodopsin and archaerhodopsin families/tribes. Arch Biochem Biophys 315:127–132
Uegaki K, Sugiyama Y, Mukohata Y (1991) Archaerhodopsin-2, from Halobacterium sp. aus-2 futher reveals essential amino acid residues for light-driven proton pumps. Arch Biochem Biophys 286:107–110
Weetall HH, Druzhko A, Lera AR, Alvarez R, Robertson B (2000) Measurement of proton release and uptake by analogs of bacteriorhodopsin. Bioelectrochemistry 51:27–33
Wise KJ, Gillespie NB, Stuart JA, Krebs MP, Birge RR (2002) Optimization of bacteriorhodopsin for bioelectronic devices. Trends Biotechnol 20:387–394
Yatsunami R, Kawakami T, Ohtani H, Nakamura S (1997) Primary structure of the novel bacterial rhodopsin from extremely halophilic archaeon Haloarcula japonica strain TR-1. Nucleic Acids Symp Ser 37:111–112
Acknowledgements
This work was supported by grants from Chinese Academy of Sciences (KJCX1-SW-07) and from the Ministry of Science and Technology (2004CB719600). Careful reading and constructive suggestions by Prof. Dr. J. K. Lanyi at University of California, Irvine is greatly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by W. D. Grant
Rights and permissions
About this article
Cite this article
Feng, J., Liu, HC., Chu, JF. et al. Genetic cloning and functional expression in Escherichia coli of an archaerhodopsin gene from Halorubrum xinjiangense. Extremophiles 10, 29–33 (2006). https://doi.org/10.1007/s00792-005-0468-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-005-0468-x