Abstract
A new group of anaerobic thermophilic bacteria was isolated from enrichment cultures obtained from deep sea sediments of Peru Margin collected during Leg 201 of the Ocean Drilling Program. A total of ten isolates were obtained from cores of 1–2 m below seafloor (mbsf) incubated at 60°C: three isolates came from the sediment 426 m below sea level with a surface temperature of 9°C (Site 1227), one from 252 m below sea level with a temperature of 12°C (Site 1228), and six isolates under sulfate-reducing condition from the lower slope of the Peru Trench (Site 1230). Strain JW/IW-1228P from the Site 1228 and strain JW/YJL-1230-7/2 from the Site 1230 were chosen as representatives of the two identified clades. Based on the 16S rDNA sequence analysis, these isolates represent a novel group with Thermovenabulum and Caldanaerobacter as their closest relatives. The temperature range for growth was 52–76°C with an optimum at around 68°C for JW/IW-1228P and 43–76°C with an optimum at around 64°C for JW/YJL-1230-7/2. The pH25C range for growth was from 6.3 to 9.3 with an optimum at 7.5 for JW/IW-1228P and from 5 to 9.5 with an optimum at 7.9–8.4 for JW/YJL-1230-7/2. The salinity range for growth was from 0% to 6% (w/v) for JW/IW-1228P and from 0% to 4.5% (w/v) for JW/YJL-1230-7/2. The G+C content of the DNA was 50 mol% for both JW/IW-1228P and JW/YJL-1230-7/2. DNA–DNA hybridization yielded 52% similarity between the two strains. According to 16S rRNA gene sequence analysis, the isolates are located within the family, Thermoanaerobacteriaceae. Based on their morphological and physiological properties and phylogenetic analysis, it is proposed that strain JW/IW-1228PT is placed into a novel taxa, Thermosediminibacter oceani, gen. nov., sp. nov. (DSM 16646T=ATCC BAA-1034T), and JW/YJL-1230-7/2T into Thermosediminibacter litoriperuensis sp. nov. (DSM 16647T =ATCC BAA-1035T).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
It is estimated that the number of prokaryotes in the deep subsurface sediment can make up more than 60% of the global number of prokaryotes (Whitman et al. 1998). They also represent 10–35% of the total biomass on the Earth (Parkes et al. 2000; Whitman et al. 1998). Recent studies showed microbial community in deep subsurface sediments may affect atmospheric carbon stocks and climate change (D’Hondt et al. 2002 and literature cited therein). Despite their significant impacts on Earth’s surface chemistry and climate (Dickens 2001), and oceanic alkalinity (D’Hondt et al. 2002), little is known about the phylogenetic, metabolic, and physiological diversity of the deep subsurface microbiota.
Due to the low culturability and viability (Cragg et al. 1990), the study of individual microorganisms in deep subsurface biosphere is a challenge. Temperature, one of the limiting factors also affects bacterial distributions in the deep subsurface sediments. Temperature rises as the depth increases, and may be responsible for the limitation of the organic matters. However, the presence of a significant bacterial population has been reported in deep sea sediments (Parkes et al. 1994; Cragg et al. 1996). Besides their potential for environmental and biotechnological applications, the isolation and characterization of thermophilic microorganisms from marine deep subsurface sediments can provide better understanding of metabolic and biogeochemical influences of the indigenous microorganisms on the ecosystem.
The Ocean Drilling Program (ODP) Leg 201 primarily focused on the microbial communities in deep sea sediments, at a series of sites in the eastern equatorial Pacific, the Peru Basin, and the Peru Margin. To determine whether thermophilic microorganisms can survive at suboptimal temperatures in marine sediments over long periods of time, an attempt was made to isolate thermophilic anaerobes from sediment samples collected at various depths and thus of increasing age at the equatorial Pacific sites and at the Peru Margin sites. Here, we report the identification of several isolates representing two novel thermophilic anaerobes from the upper layers of the Peru Margin sediment.
Methods
Collection of inocula
Core samples were collected from Eastern Equatorial Pacific and Peru Margin during the ODP cruise Leg 201 in February/March 2002 as described in detail in D’Hondt et al. (2003). The testing for drilling fluid contamination is described by House et al. (2003). The characteristics of different sites and drilling holes from which core samples were used and the schemes for microbial analysis are given in detail by Shipboard Scientific Party (sections “Microbiology”, 2003b, c, d, e). The incubations of interest for this report are those denoted by the incubation temperature 60°C. Basically after the cores were retrieved on board and cut in about 1.5-m sections, subsections were made in the cold room from which subsamples were aseptically cored from the center using sterile 60-ml syringes or 5-ml syringes (for smaller subsamples) from which the tips were cut off. These cores were extruded into sterile glass vials under a stream of sterile anaerobic nitrogen gas, the vials were closed with a butyl rubber stopper and stored until use.
Enrichment and isolation
Two different approaches were used to obtain the here described new isolates (a) inoculation and incubation of enrichments and MPNs (five tenfold dilutions in triplicates) performed immediately after recovering the core sample on board of the ship including using heterotrophic glycolytic media (b) inoculation and incubation on shore after obtaining the samples (2.5–4 months after collection and storing at about 4°C under an oxygen-free nitrogen atmosphere) using a sulfate-reducing media for enrichments.
-
(a)
On board procedure: After the core samples were collected, samples were either suspended in oxygen-free saline under an atmosphere of nitrogen which then were used for inoculation or samples were transferred via a sterile spatula into the incubation tubes using basically the Hungate procedure to keep samples and media anaerobic (Ljungdahl and Wiegel 1986). The equivalence of 0.8–1 ml solid core sample was inoculated in Balch tubes containing 9 ml of pre-reduced anaerobic heterotrophic sea salt media of pH60C (Wiegel 1998) 8.0 and 8.8 and supplemented with 0.05% yeast extract and 0.2% each of glucose, fructose and mannose (hexose media), or xylose and ribose (pentose media) with or without 25 mM thiosulfate as additional electron acceptor, respectively. Basic Sea Salt media (full strength SSM) contained 40 g Sigma sea salt, 2 mM Na2HPO4, 2.5 ml/l modified (Ni, W) Wolfe’s mineral and 5 ml Wolfe’s vitamin solution (Freier et al. 1988), 0.1% NH4Cl and 25 mM Na2CO3. 2 mM each of Na2S and cystein HCl were used as reducing agents (Ljungdahl and Wiegel 1986). From the second subculture made on board, inocula were made on shore into the half diluted sea salt media containing only one of the carbon sources, and the subcultures were also incubated at 60°C.
-
(b)
On shore procedure: Aseptically collected core samples (Shipboard party 2003a) were inoculated into 150 ml serum bottles containing either a phosphate-buffered basal medium or a carbonate-buffered medium (Widdel and Bak 1992) under anaerobic condition by using the modified Hungate technique (Ljungdahl and Wiegel 1986). The phosphate-buffered basal media contained the following (gram per liter of deionized water unless otherwise indicated): NaH2PO4, 5 mM; Na2HPO4, 15 mM; NH4Cl, 0.5; (NH4)2SO4, 0.5; NaCl, 10; MgSO4 7H2O, 0.01; CaCl2, 0.01; trace element solution, 5 ml; vitamin solution, 1 ml; yeast extract, 5; resazurin, 1 mg; cysteine HCl, 0.05. The carbonate-buffered medium was amended by adding 1% NaCl and 0.1 mM ferric citrate. The pH25C (Wiegel 1998) was adjusted to 7.3 (at 25°C) for a phosphate-buffered basal media and to either 7.0 or 8.0 (at 25°C) for a carbonate-buffered medium before degassing, and cysteine was added after degassing with N2.
Pure cultures were isolated by using agar-shake-roll technique and were usually grown in 10 ml medium in Balch tubes closed with black butyl rubber and aluminum crimps. All incubations were done at 60°C and 80°C. Pure cultures were usually grown in the sea salt medium containing only 1% sea salt or NaCl.
Determination of growth
The growth of isolates was determined by direct cell count using microscopy and by measuring the optical density (OD) at 600 nm using Spectronic 21 spectrophotometer (Bausch and Lomb, Rochester, NY, USA).
Microscopy
The morphology was studied by light and electron microscopy using an Olympus VANOX phase-contrast microscope and JEM-1210 Transmission Electron Microscope (JEOL Inc., Tokyo, Japan), respectively. Phase-contrast micrographs of bacteria were taken using agar-coated slides. Cells used for negative staining were from both early exponential growth phase and stationary growth phase.
Effect of temperature, pH, and salinity
The temperature-gradient incubator (Scientific Industries Inc., Bohemia, NY, USA) was used to determine the temperature range for growth of each isolate. To determine the pH optimum, the various pH values were determined at the optimum growth temperature (Topt) as described by Wiegel (1998). Media for the pH range determination were buffered with 10 mM each of MES, HEPES, and TAPS in combination with 2 mM phosphate. Various NaCl and KCl (ratio of 9:1) concentrations were added to the basal medium (minus NaCl) to obtain the ranges of salinity supporting growth.
Range of substrate utilization
The ability of the isolates to grow on potential carbon sources was assayed using the phosphate-buffered basal medium as described above but with only 0.02% yeast extract. The cultures were incubated and observed for more than 2 weeks, and the utilization was judged positive if the OD of the culture was twice above the value of control culture containing only yeast extract.
Electron acceptors
The potential use of various electron acceptors was studied using the basal medium (1% NaCl) containing 0.3% yeast extract as an electron donor. Cultures in the exponential growth phase in the basal medium without any additional electron acceptors were used as the inocula (2% v/v). The electron acceptors tested were fumarate (20 mM), sulfate (20 mM), sulfite (2 mM), thiosulfate (20 mM), elemental sulfur (20 mM), nitrate (20 mM), amorphous Fe (III) oxide (90 mM), Fe (III) citrate (20 mM), AQDS (10 mM), and MnO2 (10 mM). The use of electron acceptors was determined by measuring growth (OD600), sulfide, ammonium or nitrite production, or color-change, respectively.
Analytical techniques
The concentration of dissolved and precipitated sulfides was determined by the CuSO4 spectrophotometric assay (Cord-Ruwisch 1985). Nitrate reduction was performed as previously described (Finegold and Baron 1986). Ferric ion was monitored by measuring ferrous ion production using the Ferrozine assay (Dailey and Lascelles 1977).
Phospholipid fatty acid analysis
Samples were extracted by a single-phase organic solvent system comprised of chloroform, methanol, and aqueous 50 mM PO4 buffer (pH 7.4) in the ratio of 1:2:0.8 (v/v/v; White et al. 1979). After extracting overnight, equal volumes of chloroform and nanopure water were added to the extractant, resulting in a two-phase system. The lower organic (lipid-containing) phase was collected and concentrated to yield the total lipid extract. The concentrated lipid extract was fractionated on a silicic acid column into neutral lipids, glycolipids, and polar lipids (Guckert et al. 1985). The phospholipid fatty acid (PLFA) in the polar lipid fraction were subjected to a mild alkaline methanolysis to produce fatty acid methyl esters and then the hydroxyl groups converted to the silyl ethers prior to GC and GC-MS analyses.
G+C content of genomic DNA
The DNA was extracted from each isolate using DNeasy Tissue Kit (Qiagen Inc., Valencia, CA, USA). The guanine plus cytosine (G+C) content was measured by HPLC as described previously (Mesbah et al. 1989) with the modification of using S1 nuclease (Invitrogen Co., Carlsbad, CA, USA) and 0.3 M sodium acetate (pH 5.0).
DNA–DNA hybridization
DNA–DNA hybridization was performed by the German culture collection (DSMZ). DNA was isolated using a French pressure cell (Thermo Spectronic) and was purified by chromatography on hydroxyapatite as described by Cashion et al. (1977). DNA–DNA hybridization was carried out as described by De Ley et al. (1970), with the modifications described by Huss et al. (1983), using a model Cary 100 Bio UV/VIS-spectrophotometer equipped with a Peltier-thermostatted 6×6 multicell changer and a temperature controller with in-situ temperature probe (Varian).
16S rRNA gene sequence determination and phylogenetic analyses
The DNA was extracted as described above and amplified with bacterial domain-specific primer set for 16S rDNA, 27 forward and 1492 reverse (Lane 1991). The PCR amplification was carried out as described previously (Wise et al. 1999). PCR products were purified using QIAquick PCR Purification Kit (Qiagen) and sequenced by Macrogen Inc. (Seoul, Korea). The Similarities of partial sequences were determined using the Sequencher v4.0.5 (Gene Codes Co., Ann Arbor, MI, USA). Retrieved 16S rDNA sequences, 1,386 for JW/IW-1228P and 1,402 for JW/YJL-1230-7/2 were analyzed using BLAST (basic local alignment search tool) and then aligned manually using ClustalX v1.81 (Thompson et al. 1997) to create a multiple sequence alignment. Phylogenetic trees were inferred by the neighbor-joining method (Saitou and Nei 1987) using the model of Jukes and Cantor (Jukes and Cantor 1969), with the phylogenetic analysis package PHYLIP v3.6a2.1 (Felsenstein 2001).
Nucleotide sequence accession number
The 16S rDNA sequences of both strains JW/IW-1228P and JW/YJL-1230-7/2 were submitted to GenBank and assigned accession number AY703478 and AY703479, respectively.
Results and discussion
Enrichment and isolation
Samples from the Site 1225, 1226, 1227, and 1228 were inoculated shortly after collecting the core samples into the basal sea salt medium containing 0.05% (w/v) yeast extract and 0.2% each of glucose, fructose and mannose, or xylose and ribose at 60°C. Samples from the Site 1230 and 1231 were inoculated into the bicarbonate-buffered medium containing acetate or lactate with 28 mM sulfate and incubated at 60°C and 80°C, respectively. Positive enrichment cultures were obtained only at 60°C, but at various pHs (7.0, 7.3, 7.8, 8.0). After three rounds of purification procedures using the agar-shake-roll tube technique (Ljungdahl and Wiegel 1986) a total of ten pure isolates were obtained. Besides the below-described pure isolates, at the shipboard positive enrichments were additionally obtained from site 201-1228-2H (565–582 cm below seafloor) and from site 201-1226E-1H-2 (75–80 cm below sea floor). The first subculture of the latter enrichment had produced after about 8 days 40 mM lactate, 10 mM acetate, and 5 mM formate (Arthur Spivack, personal communication). However these two enrichments were not any longer viable after returning to the University of Georgia. Three of the ten purified isolates had been obtained from the heterotrophic media originally inoculated with sediment from core 201–1227D-1H-1 (ca. 131–138 cm below seafloor) (Site 201-1227, Trujillo Basin on the Peru continental shelf, sea floor 426 m below sea level and a mudline temperature of 9°C) and one from core 201-1228E-1H-1 (136–143 cm below seafloor; Site 201-1228 outer shelf edge of the Peruvian high productivity upwelling system with a sea floor 252 m below sea level and a mud line temperature of 12°C). The strains were designated as JW/IW-1227G (glucose-supplemented media), JW/IW-1227M (mannose- supplemented media), JW/IW-1227X (xylose-supplemented media) and JW/IW1228P (pyruvate-supplemented media). Growth was obtained only in the first two dilution tubes from MPN experiments, indicating less than 100 viable cells/ml sediments. Six isolates were obtained from the medium for sulfate reducers inoculated on shore with samples from Site 201-1230 (lower slope of the Peru Trench, with a mudline temperature of 2°C, 5,086 m below sea level). The strains were designated as JW/YJL-1230-7/1, JW/YJL-1230-7/2, JW/YJL-1230-7/3 (all isolated from media with a pH25C 7.0), JW/YJL-1230-8/1, JW/YJL-1230-8/2, and JW/YJL-1230-8/3 (isolated from media with a pH25C 8.0). Post cruise enrichments using the medium for sulfate reducers were also set for the samples from 201-1227A-2H-5 (section 64–78 cm), 201 1229-2H-2 (section 64–78 cm), 201 1230A 2H –2 (section 82–75 cm), 201-1230A 11H-2 (section 60–67 cm), 201-1230A 12H-3 (section 64–78 cm), and 201-1230A 15H-3 (section 27–34 cm). Incubations of enrichment cultures at 60°C and 80°C yielded no visible growth.
Colony and cell morphology
Based on 16S rDNA sequence analysis and growth parameters during isolation, two of the isolates were chosen for more detailed characterizations: JW/IW-1228P and JW/YJL-1230-7/2. In agar-roll-tube cultures, the colonies appeared after 2–3 days. The colonies were irregular shaped with 0.1–1.5 mm in diameter. Vegetative cells of strain JW/IW-1228P grown in liquid cultures were straight, sometimes highly elongated rods with 0.2–0.7 μm in diameter and 1.5–16 μm in length, which occurred singly, in pairs or in chains (Fig. 1a) and staining Gram-negative. Without agitation, cells grown in liquid cultures had the tendency to elongate, form chains or/and aggregates and to flocculate. In the late-exponential or stationary growth phase, cells started to yield swollen ends and bulging sections throughout the elongated cells, and the cytoplasm became granular and heterogeneous, and eventually formed autoplasts (L-shaped cells) (Fig. 1b). Cells of JW/YJL-1230-7/2 isolated under sulfate-reducing conditions, were straight rods, with a diameter of 0.3–0.5 μm and 2.0–10.0 μm in length (Fig. 1c), and thus were less elongated than the JW/IW-1228P. Cells occurred singly, in pairs, or in chains and stained Gram-negative. Strain JW/YJL-1230-7/2 also produced swollen ends, but infrequently formed autoplasts (Fig. 1d). Electron microscopy revealed both strains were flagellated. Strain JW/YJL-1230-7/2 had 2–4 long peritrichous flagella with a periodicity (wavelength) of ~1–1.3 μm (Fig. 1e). Less than 1% cells of strain JW/IW-1228P and up to 5% cells of strain JW/YJL-1230-7/2 exhibited branched cell morphology (Fig. 1). The occurrence of spores was not detected by microscopy or by heat treatment (10 min at 100°C). Despite flagellation no motility besides tumbling was detected by microscopy for any of the strains.
Micrographs of JW/IW-1228-P and JW/YJL-1230-7/2. Cells from mid-exponential growth phase of JW/IW-1228P (a), late-exponential growth phase of JW/IW-1228P exhibiting partly swollen cells and L-form-like cells (b), mid-exponential growth phase of JW/YJL-1230-7/2 (c), late-exponential growth phase of JW/YJL-1230-7/2 with primary branches (d), TEM of JW/YJL-1230-7/2, Arrows point to branched cells. e Arrowhead indicates periodicity. Bars, 10 (a, b, c, d) and 1 (e) μm
Temperature, pH, and salinity ranges
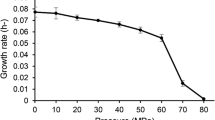
The temperature range for growth at pH25C 7.8 of strain JW/IW-1228P was 52–76°C, with an optimum at 68°C and 1.6 h doubling time. No growth was detected at above 78°C or below 50°C. Strain JW/YJL-1230-7/2 grew at 45–75°C, with an optimum at 64–65°C. No growth was detected at above 76°C or below 43°C. Due to bioturbation of the sediment it is difficult to exactly determine how long the bacteria have survived in the sediment at temperatures below the minimal growth temperatures determined under laboratory conditions. The minimal growth temperature determined under laboratory conditions is probably higher than in vivo where bacteria could sustain at doubling times of many months, but those growth rates are usually not measured in the laboratory. The age of the sediments from which the samples were taken (Skilbeck, personal communication) is estimated to be around 30,000–50,000 years for 201-1226-1H-1 (no pure isolate obtained), 2,000 years for 201-1228E-1H-1 (strain JW/IW-1228P), and 201-1228E-2H-1 around 50,000 years (no pure isolate obtained). The age of sediments from 201-1230A-1H-1 from which strain JW/YJL-1230-7/2 came is estimated to be between 10,000 years and 15,000 years. Considering that the samples came from sediments hundreds to several tens of thousands of years old (Pleistocene) one could postulate minimal maintenance metabolisms or/and extremely slow growth aiding in survival at temperatures below the lower temperature limit determined in the laboratory. At 60°C, the pH range for JW/IW-1228P was 6.3–9.3, with an optimum at pH25C 7.5. Strain JW/YJL-1230-7/2 grew at pH25C 6.2–9.1, with an optimum at pH 7.9–8.4. The salinity range for JW/IW-1228P was from 0% to 6.0% (w/v), with an optimum at 1%. No growth was detected at 7% (w/v) and above. Strain JW/YJL-1230-7/2 has salinity range from 0% to 4.5% (w/v), with an optimum at 0.5–2%. No growth was detected at 5% (w/v) and above.
Substrate utilization
Strain JW/IW-1228P grew well on Casamino acids, fructose, glucose, mannose, sucrose and xylose (0.2%, w/v). It showed very weak growth on Difco Beef extract, tryptone, lactate, pyruvate, methanol, inositol, manitol, sorbitol, cellobiose, maltose, raffinose and trehalose (0.2%, w/v). However, strain JW/YJL-1230-7/2 utilized tryptone, acetate, lactate, inositol, manitol, xylitol, fructose, galactose, glucose, mannose, raffinose, sucrose and xylose (0.2%, w/v) in the presence of 0.1% yeast extract, but showed only little growth on this relatively broad substrate spectrum. This might indicate the survival strategy of the isolate in unfavorable oligotrophic environment. Both strains required yeast extract for growth. There was no indication of growth either under aerobic condition or under chemolithoautotrophic conditions using H2/CO2 (80:20, v/v) in the presence of 0.02% yeast extract and in the presence or absence of Fe(III). The main fermentation end product from glucose was acetate in both strains. Propionate, isobutyrate, and isovalerate were also detected in small amounts.
Electron acceptors
In the presence of yeast extract (0.3%, w/v) as a sole carbon source and electron donor, both strains reduced thiosulfate and elemental sulfur to sulfide, and Mn(IV)O2. There was no indication of Fe(III) reduction, however, the supplementation of 0.1 mM of ferric citrate enhanced growth of both strains. The selective utilization of Mn(IV) ions as an e− acceptor, together with the 4–6% NaCl tolerance can be taken as a hint that the isolates are indeed marine bacteria, although it is very possible that the strains originated from the terrestrial geothermal features of Peru.
Phospholipid fatty acid composition
Both strains were differentiated clearly by the composition of their PLFA profiles (Table 1). While the most abundant fatty acid in strain JW/IW-1228P was i15:0, strain JW/YJL-1230-7/2 contained the four major fatty acids, i15:0, 16:1w9c, 16:0 and 18:1w9c. The polyunsaturated PFLA 18:2w6 was found in minor amounts only in strain JW/YJL-1230-7/2. Cyclopropane fatty acids, a possible biomarker for Gram-type negative bacteria (Zelles 1997), were not observed in either strain, which is in agreement that the isolates are Gram-type positive bacteria (Wiegel 1981).
DNA base composition
The G+C contents of the genomic DNA of both strain JW/IW-1228P and JW/YJL-1230-7/2 were 50 mol% (HPLC). The G+C mol% contents of the 16S rDNA of strain JW/IW-1228P and JW/YJL-1230-7/2 were 60 and 61, respectively. The DNA–DNA hybridization tests resulted in a re-association value of around 52%, which confirmed that strains JW/IW-1228P and JW/YJL-1230-7/2 are not related at the species level (Wayne et al. 1987).
16S rRNA gene sequences and phylogenetic analyses
Almost complete 16S rRNA gene sequences of strain JW/IW-1228P and JW/YJL-1230-7/2 were determined, comprising 1,386 (47–1457 based on E. coli numbering) nucleotides and 1,402 (47–1475 based on E. coli numbering) nucleotides, respectively. When compared, both sequences were about 98.3% similar to each other. All partial sequences of JW/IW-strains and all of the JW/YJL-strains clustered together, respectively (data not shown). According to BLASTN search, these isolates represent a novel group with Thermovenabulum and Caldanaerobacter as their closest relatives. The closest relative of strain JW/IW-1228P is Thermovenabulum ferriorganovorum (AY033493) with a G+C mol% of 36 showing 98% similarity for the first 88 bp, 37 gaps, and then 93% similarity to the rest of 1,204 bp. The probability score (P score) from BLASTN results for strain JW/YJL-1230-7/2, however, showed anaerobic syntrophic bacterium OL (GenBank accession number; AB106354) as the closest relative. When compared to T. ferriorganovorum, the sequences of the novel isolates showed two gap regions (data not shown). When the gaps were included in this analysis, strain JW/IW-1228P and JW/YJL-1230-7/2 showed 90.3% and 89.7% similarity, respectively to T. ferriorganovorum. When unalignable regions eliminated, strain JW/IW-1228P and JW/YJL-1230-7/2 showed 93.7 and 94.4% similarity, respectively to T. ferriorganovorum. In a phylogenetic tree constructed by the neighbor-joining method (Fig. 2), both strains had the same closest relative, T. ferriorganovorum. The 16S rDNA sequences place the new isolates within the radius of the family Thermoanaerobacteriaceae. Based on the 16S rDNA sequences analysis and the difference of about 14 mol% in the G+C content of the DNA in respect to the closest identified relative, Thermovenabulum, the novel strains cannot be assigned to any of the genera in the family. In addition, the novel strains differ in morphology and physiology from the closest relative Thermovenabulum (Table 2). Thus, isolates JW/IW-1228P and JW/YJL-1230-7/2 together with the related strains are placed in the new genus Thermosediminibacter. Based on the phenotypic differences between the two strains, the DNA–DNA hybridization analysis, and the fatty acid composition, the strains are placed into two different species.
A phylogenetic dendrogram based on 16S rDNA sequence showing the positions of strains JW/IW-1228P and JW/YJL-1230-7/2 (boldface text) amongst members of the family Thermoanaerobacteriaceae. The tree was constructed using Neighbor-joining method with Jukes and Cantor distance corrections. Numbers at the nodesrepresent the bootstrap values (% of 1,000 replicates); values above 90% were considered significant. The scale barindicates two nucleotide substitutions per 100 nucleotides
Description of Thermosediminibacter gen. nov.
Thermosediminibacter (Ther.mo.se.di.mi.ni.bac’ter. Gr. Adj. thermos, hot; L. neut. n. sediment -inis, sediment; N. L. masc. n. bacter (from Gr. neut. n. bactron), a rod or staff; N. L. masc. n. thermosediminibacter, thermophilic rod from sediment, referring to its origin and growth temperature)
The genus Thermosediminiibacter belongs to the low G+C, Gram-type positive Bacillus-Clostridium subphylum. Habitat: so far only isolated from ocean subsurface sediments. The cells are straight rod to curved, and swollen and subsequently form autoplasts (L-shaped) in the late-exponential or stationary phase of growth. Anaerobic and thermophilic chemoorganotrophs. Yeast extract is required for growth. No growth on H2/CO2 (80:20, v/v). The G+C mol% of the DNA is around 50. The type species is Thermosediminibacter oceani.
Description of Thermosediminibacter oceani sp. nov.
Thermosediminibacter oceani (o.ce.a’ni. L. masc. n. oceanus, ocean; L. gen. masc. n. oceani, of an ocean, referring to its origin from the ocean)
The cells are straight to curved rods, 0.2–0.7 μm in diameter and 1.5–16 μm in length. Cells occur singly, in pairs, or in chains and stain Gram-negative. Cells tend to elongate and form aggregates. Flagella observed. The temperature range for growth is 52–76°C, with an optimum at around 68°C. The pH25C range for growth is from 6.3 to 9.3, with an optimum at 7.5. The salinity range for growth is from 0 to 6% (w/v), with an optimum at 1%. In the presence of 0.02% yeast extract, casamino acids, beef extracts, tryptone, cellobiose, fructose, galactose, glucose, maltose, mannose, raffinose, sucrose, trehalose, xylose, methanol, inositol, manitol, sorbitol, lactate, pyruvate serve as carbon and energy. Thiosulfate, elemental sulfur, and MnO2 can serve as e− acceptors. No indication of sulfate or Fe(III) reduction. The most abundant fatty acid is i15:0. The G+C content of the genomic DNA is 50 mol% (HPLC). The type strain is JW/IW-1228PT (DSM 16646T, ATCC BAA-1034T).
Description of Thermosediminibacter litoriperuensis sp. nov.
Thermosediminibacter litoriperuensis (li.to.ri.pe.ru.en’sis. L. neut. n. litus–oris, the seashore, seaside, beach, coast; N. L. masc. adj. peruensis, pertaining to Peru; N. L. masc. adj. litoriperuensis, of a Peruvian coast, referring to its origin from the coast of Peru)
The cells are straight to slightly curved rod, 0.3–0.5 μm in diameter and 2.0–10.0 μm in length. Cells occur singly, in pairs, or in chains and stain Gram-negative. Retarded peritrichous flagella detected. The temperature range for growth is 43–76°C with an optimum at around 64°C. The pH25C range for growth is from 5 to 9.5 with an optimum at 7.9–8.4. The salinity range for growth is from 0 to 4.5% (w/v), with an optimum at 0.5–2%. Substrates utilized include yeast extract, tryptone, acetate, lactate, inositol, manitol, xylitol, fructose, galactose, glucose, mannose, raffinose, sucrose and xylose. The major fatty acids are i15:0, 16:1w9c, 16:0 and 18:1w9c with small amount of the polyunsaturated PFLA, 18:2w6. Thiosulfate, elemental surfur, MnO2 can function as e− acceptors. There was no indication of sulfate or Fe(III) reduction. The G+C content of the genomic DNA was 50 mol% (HPLC-method). The type strain is JW/YJL-1230-7/2T (DSM 16647T, ATCC BAA-1035T).
References
Cashion P, Hodler-Franklin MA, McCully J, Franklin M (1977) A rapid method for base ratio determination of bacterial DNA. Anal Biochem 81:461–466
Cord-Ruwisch R (1985) A quick method for the determination of dissolved and precipitated sulfides in cultures of sulfide-reducing bacteria. J Microbiol Methods 4:33–36
Cragg BA, Parkes RJ, Fry JC, Herbert RA, Wimpenny JWT, Getliff JM (1990) Bacterial biomass and activity profiles within deep sediment layers. Proc Ocean Drilling Program Scientific Results 112:607–619
Cragg BA, Parkes RJ, Fry JC, Weightman AJ, Rochelle PA, Maxwell JR (1996) Bacterial populations and processes in sediments containing gas hydrates (ODP Leg 146:Cascadia Margin). Earth Planetary Sci Lett 139:497–507
Dailey HA Jr, Lascelles J (1977) Reduction of iron and synthesis of protoheme by Spirillum itersonii and other organisms. J Bacteriol 129:815–820
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
D’Hondt S, Rutherford S, Spivack, AJ (2002) Metabolic activity of subsurface life in deep-sea sediments. Science 295:2067–2070
D’Hondt SL, Jørgensen BB, Miller DJ et al (2003) Proc ODP Init Repts 201 http://www-odp.tamu.edu/publications/201_IR/201ir.htm. http://www-odp.tamu.edu/publications/201_IR/chap_01/chap_01.htm
Dickens G (2001) On the fate of past gas: what happens to methane released from a bacterially mediated gas hydrate capacitor? Geochem Geophys Geosyst 2:2000GC000131
Felsenstein J (2001) PHYLIP (Phylogeny Inference Package) version 3.6a2.1. Department of Genome Sciences, University of Washington, Seattle
Finegold SM, Baron EJ (1986) Conventional and rapid microbiological methods for identification of bacteria and fungi. In: Finegold SM, Baron EJ (eds) Bailey and Scott’s diagnostic microbiology. The CV Mosby Co, St. Louis, pp 117–118
Freier D, Mothershed CP, Wiegel J (1988) Characterization of Clostridium thermocellum JW20. Appl Environ Microbiol 54:204–211
Guckert JB, Antworth CB et al (1985) Phospholipid ester-linked fatty acid profiles as reproducible assays for changes in prokaryotic community structure of estuarine sediments. FEMS Microbiol Ecol 31:147–158
House CH, Cragg BA, Teske A, The Leg 201 Scientific Party (2003) Drilling contamination tests during ODP Leg 201 using chemical and particulate tracers. In: D’Hondt SL, Jørgensen BB, Miller DJ et al. Proc ODP Init Repts 201:1–19 http://www-odp.tamu.edu/publications/201_IR/chap_05/chap_05.htm
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrometric determination of DNA hybridization from renaturation rate. Syst Appl Microbiol 4:184–192
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism, Academic, New York, pp 21–132
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Ljungdahl LG, Wiegel J (1986) Anaerobic fermentations. In: Demain AL, Solomon NA (eds) Manual of industrial microbiology and biotechnology. American Society for Microbiology, Washington DC, pp 84–96
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Parkes RJ, Cragg BA, Bale SJ, Getliff JM, Goodman K, Rochelle PA, Fry JC, Weightman AJ, Harvey SM (1994) Deep bacterial biosphere in Pacific-ocean sediments. Nature 371:410–413
Parkes RJ, Cragg BA, Wellsbury P (2000) Recent studies on bacterial populations and processes in subseafloor sediments: a review. Hydrogeol J 8:11–28
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shipboard Scientific Party (2003a) Explanatory notes. In: D’Hondt SL, Jørgensen BB, Miller DJ et al. Proc ODP Init Repts 201 http://www-odp.tamu.edu/publications/201_IR/chap_01/chap_01.htm.
Shipboard Scientific Party (2003b) Site 1227. In: D’Hondt SL, Jørgensen BB, Miller DJ et al. Proc ODP Init Repts 201 http://www-odp.tamu.edu/publications/201_IR/chap_08/chap_08.htm
Shipboard Scientific Party (2003c) Site 1228. In: D’Hondt SL, Jørgensen BB, Miller DJ et al. Proc ODP Init Repts 201 www-odp.tamu.edu/publications/201_IR/chap_09/chap_09.htm
Shipboard Scientific Party (2003d) Site 1229. In: D’Hondt SL, Jørgensen BB, Miller DJ et al. Proc ODP Init Repts 201 www-odp.tamu.edu/publications/201_IR/chap_10/chap_10.htm
Shipboard Scientific Party (2003e) Site 1230. In: D’Hondt SL, Jørgensen BB, Miller DJ, et al. Proc ODP Init Repts 201 www-odp.tamu.edu/publications/201_IR/chap_11/chap_11.htm
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wayne LG, Brenner DJ, Colwell RR et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
White DC, Davis WM et al (1979) Determination of the sedimentary microbial biomass by extractable lipid phosphate. Oecologia 40:51–62
Whitman WB, Coleman DC, Wiebe WJ (1998) Prokaryote: the unseen majority. Proc Natl Acad Sci USA 95:6578–6583
Widdel F, Bak F (1992) Gram-negative mesophilic sulfate-reducing bacteria. In: Baloes A, Trüper HG, Dworkin M, Harder W, Schleifer K-H (eds) The procaryotes. Springer, Berlin Heidelberg New York, pp 583–624
Wiegel J (1981) Distinction between the Gram reaction and the Gram type of bacteria. Int J Syst Bacteriol 31:88
Wiegel J (1998) Anaerobic alkalithermophiles, a novel group of extremophiles. Extremophiles 2:257–267
Wise MG, McArthur JV, Shimkets LJ (1999) Methanotroph diversity in landfill soil: isolation of novel type I and type II methanotrophs whose presence was suggested by culture-independent 16S ribosomal DNA analysis. Appl Environ Microbiol 65:4887–4897
Xue Y, Xu Y, Liu Y, Ma Y, Zhou P (2001) Thermoanaerobacter tengcongensis sp nov, a novel anaerobic, saccharolytic, thermophilic bacterium isolated from a hot spring in Tengcong, China. Int J Syst Evol Microbiol 51:1335–1341
Zavarzina DG, Tourova TP, Kuznetsov BB, Bonch-Osmolovskaya EA, Slobodkin AI (2002) Thermovenabulum ferriorganovorum gen nov, sp nov, a novel thermophilic anaerobic, endospore-forming bacterium. Int J Syst Evol Microbiol 52:1737–1743
Zelles L (1997) Phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere 35:275–294
Acknowledgements
This study was supported by a shipboard grant for Ocean Drilling Program (ODP) Leg 201 and post-cruise grant (USSSP-Grant# 201-F002587 and 201-F001649) to J.W. ODP is sponsored by the U.S. National Science Foundation (NSF) and participating countries under management of Joint Oceanographic Institutions (JOI), Inc. We would like to thank all members of the shipboard party for their great hospitality and help in collecting samples. We also thank J.P. Euzeby for his help regarding the Latin nomenclature, M.A. Eiteman for his fermentation end products analysis, and W.B. Whitman for his help with the G+C mol% determination. Y.L. was supported by a graduate fellowship from the Savannah River Ecology Laboratory, which is funded by the Environmental Remediation Sciences Division of the Office of Biological and Environmental Research, U.S. Department of Energy, through Financial Assistance Award No. DE-FC09-96-SR18546 to the University of Georgia Research Foundation. I.D.W. was supported by the NSF-REU site program (NSF-0139083).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Horikoshi
An erratum to this article can be found at http://dx.doi.org/10.1007/s00792-006-0521-4
Rights and permissions
About this article
Cite this article
Lee, YJ., Wagner, I.D., Brice, M.E. et al. Thermosediminibacter oceani gen. nov., sp. nov. and Thermosediminibacter litoriperuensis sp. nov., new anaerobic thermophilic bacteria isolated from Peru Margin. Extremophiles 9, 375–383 (2005). https://doi.org/10.1007/s00792-005-0453-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-005-0453-4