Abstract
Objectives
This in vitro study was established to examine whether visfatin thought to be a link between periodontitis and obesity is produced by periodontal ligament (PDL) cells and, if so, whether its synthesis is modulated by microbial and/or biomechanical signals.
Materials and methods
PDL cells seeded on BioFlex® plates were exposed to the oral pathogen Fusobacterium nucleatum ATCC 25586 and/or subjected to biomechanical strain for up to 3 days. Gene expression of visfatin and toll-like receptors (TLR) 2 and 4 was analyzed by RT-PCR, visfatin protein synthesis by ELISA and immunocytochemistry, and NFκB nuclear translocation by immunofluorescence.
Results
F. nucleatum upregulated the visfatin expression in a dose- and time-dependent fashion. Preincubation with neutralizing antibodies against TLR2 and TLR4 caused a significant inhibition of the F. nucleatum-upregulated visfatin expression at 1 day. F. nucleatum stimulated the NFκB nuclear translocation. Biomechanical loading reduced the stimulatory effects of F. nucleatum on visfatin expression at 1 and 3 days and also abrogated the F. nucleatum-induced NFκB nuclear translocation at 60 min. Biomechanical loading inhibited significantly the expression of TLR2 and TLR4 at 3 days. The regulatory effects of F. nucleatum and/or biomechanical loading on visfatin expression were also observed at protein level.
Conclusions
PDL cells produce visfatin, and this production is enhanced by F. nucleatum. Biomechanical loading seems to be protective against the effects of F. nucleatum on visfatin expression.
Clinical relevance
Visfatin produced by periodontal tissues could play a major role in the pathogenesis of periodontitis and the interactions with obesity and other systemic diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Periodontitis is a chronic inflammatory disease caused by oral pathogenic microorganisms present in dental plaque. These periodontopathogens as well as their components and metabolic products can elicit local immunoinflammatory reactions in periodontal tissues. As a consequence of the exaggerated inflammatory processes and immune responses, periodontal soft and hard tissues are subjected to degradation and resorption, respectively, which can result in periodontal attachment and even tooth loss [1, 2]. Moreover, periodontitis has been shown to be associated with cardiovascular diseases, diabetes mellitus, rheumatoid arthritis, obesity, metabolic syndrome, and other systemic diseases and conditions [3].
The periodontal tissues are periodically exposed to biomechanical loading during mastication, speech, and dental habits. Since the periodontal ligament (PDL) represents a richly vascular and cellular connective tissue, it can absorb and distribute physiological forces, which are at the same time critical for remodeling and maintenance of the periodontium [4, 5]. However, dental overloading due to occlusal discrepancies, dental habits, or strong orthodontic forces can be harmful to periodontal tissues and contribute to progressive periodontal destruction [6–8]. It is well known from in vitro and clinical studies, that biomechanical loading can induce the synthesis of inflammatory mediators and proteases, thereby acting as proinflammatory and catabolic signals [9, 10]. For example, excessive mechanical stress due to hyperocclusion has been demonstrated to stimulate osteoclastogenesis and alveolar bone destruction [11]. In summary, both bacterial infection and biomechanical loading can cause immunoinflammatory reactions and, thereby, destructive processes in the periodontium. However, if and how microbial infection and biomechanical loading interact on the level of periodontal cells has yet to be elucidated.
Recently, an association between periodontal diseases and obesity has been reported [12]. Systemic low-grade inflammation present in obesity may be a pathomechanistic link between obesity and other chronic diseases, such as periodontitis [13]. In the adipose tissue, a number of cytokines, such as visfatin, leptin, resistin, and adiponectin, are produced and released [14]. These adipokines not only regulate insulin sensitivity and energy expenditure, but also proinflammatory and wound healing processes. Whereas adiponectin exerts anti-inflammatory effects, visfatin, leptin, and resistin possess proinflammatory characteristics [15, 16]. Interestingly, the serum levels of these adipokines are altered in obesity and overweight [15, 16]. It is thought that adipokines not only contribute to the subclinical inflammatory state in obesity, but that such molecules also represent the pathomechanistic link between type 2 diabetes, obesity, and periodontitis [17].
Visfatin is predominantly secreted by cells from human visceral adipose tissue and was originally denominated as pre-B cell colony enhancing factor. This protein has also an enzymatic nicotinamide phosphoribosyltransferase activity responsible for the synthesis of nicotinamide adenine dinucleotide which is essential for cell metabolism. Visfatin induces the synthesis of proinflammatory cytokines and acts as a chemotactic factor [18–20]. High serum levels of visfatin have been found in a number of inflammatory diseases and conditions, such as obesity, type 2 diabetes, metabolic syndrome, atherosclerosis, cancer, rheumatoid arthritis, and sepsis [21–27]. Recently, it has been demonstrated that visfatin is present in high levels in gingival crevicular fluid (GCF) and serum from periodontally diseased patients, indicating that visfatin might also be produced in the periodontium and regulated by bacterial infection and/or inflammation [28–30]. However, it is as yet unknown whether PDL cells produce this proinflammatory adipokine, and if so, whether its synthesis is regulated by oral bacteria and biomechanical loading. Hence, the aim of this study was to examine in vitro whether visfatin is produced in PDL cells and, if so, whether its synthesis is modulated by microbial and/or biomechanical signals.
Materials and methods
Culture and treatment of cells
PDL cells from periodontally healthy donors, who underwent tooth extraction for orthodontic reasons, were used. Written informed parental consent and approval of the Ethics Committee of the University of Bonn were obtained. Cells from nine donors were collected from the middle third of the tooth roots and cultured in Dulbecco’s Modified Eagle’s Medium (Invitrogen, Karlsruhe, Germany) supplemented with 10 % fetal bovine serum (FBS, Invitrogen), 100 units of penicillin and 100 μg/ml of streptomycin (both from Biochrom, Berlin, Germany) at 37 °C in a humidified atmosphere of 5 % CO2. Cells of passages 3–5 were seeded (1 × 105 cells/well) on six-well BioFlex® collagen-coated culture plates (Flexcell International, Hillsborough, NC, USA) and grown to 80 % confluence. One day prior to experiments, the FBS concentration was reduced to 1 %. The medium was changed every other day during the course of the experiment. In order to mimic bacterial infection in vitro, cells were stimulated with the inactivated oral pathogen Fusobacterium nucleatum ATCC 25586 (optical density 0.025, 0.05, and 0.1). The strain was precultivated 48 h on Schaedler agar plates (Oxoid, Basingstoke, UK) in an anaerobic atmosphere. Thereafter, bacteria were suspended in phosphate-buffered saline (PBS) (OD660nm = 1, equivalent to 1.2 × 109 bacterial cells/ml) and exposed two times to ultrasonication (160 W for 15 min) resulting in a complete killing. Moreover, biomechanical loading conditions were simulated in vitro by application of cyclic tensile strain (CTS) of low (CTSL, 3 %) and high (CTSH, 20 %) magnitudes at a rate of 0.05 Hz to cells. Like in previous experiments, a strain device (CESTRA) developed at the University of Bonn was used to apply biomechanical loading [31–34]. In the present study, cells were exposed to F. nucleatum ATCC 25586, CTS, and their combination for 1 and 3 days, respectively.
Real-time RT-PCR
Total RNA was extracted using the RNeasy Mini Kit (Qiagen, Hilden, Germany) following manufacturer’s protocol. By using the iScript™ Select cDNA Synthesis Kit (Bio-Rad, Munich, Germany), 500 μg of total RNA was reverse transcribed at 42 °C for 90 min followed by 85 °C for 5 min, following manufacturer’s instructions. Gene expression of visfatin, toll-like receptor (TLR) 2 and 4, and glyceraldehyde-3-phosphate dehydrogenase as housekeeping gene was analyzed by real-time RT-PCR using the iCycler iQ detection system (Bio-Rad), SYBR Green (Qiagen), and specific primers (QuantiTect Primer Assay, Qiagen). One microliter of cDNA was amplified as a template in a 25-μl reaction mixture containing 12.5 μl of 2x QuantiFast SYBR Green PCR Master Mix (Qiagen), 2.5 μl of primers and RNase free water. The mixture was heated initially at 95 °C for 5 min and then followed by 50 cycles of denaturation at 95 °C for 10 s and combined annealing/extension at 60 °C for 30 s. Data were analyzed using the comparative threshold cycle method.
ELISA
Protein levels of visfatin in cell supernatants were measured using a commercially available ELISA kit (R&D Systems, Minneapolis, MN, USA) according to the manufacturer’s instructions. The absorbance was determined using a microtiter plate reader (POWERWAVE X; BioTek Instruments, Winooski, VT, USA) at 450 nm. Data were normalized by the cell number.
Immunofluorescence
Cells attached to the flexible bottom of the culture plates were fixed with 4 % paraformaldehyde in PBS pH 7.4 (Sigma-Aldrich, Munich, Germany) for 10 min at room temperature (RT), washed with PBS, and treated with 0.1 % Triton X-100 (Sigma-Aldrich) for 5 min at RT. Then cells were washed again with PBS and blocked with a blocking buffer (nonfat dry milk; Bio-Rad) for 1 h at RT. After washing, the cells were incubated with primary rabbit antibody NFκB p65 (1:400; Cell Signaling Technology, Danvers, MA, USA) for 90 min and with secondary antibody CY3 (1:2,000; Abcam, Cambridge, MA, USA) for 45 min. Cells were observed under a × 20 objective using an Axioplan 2 imaging microscope (Carl Zeiss MicroImaging, Jena, Germany). The images were captured with a PVCAM camera and the VisiView capturing software (Visitron Systems, Puchheim, Germany).
Immunocytochemistry
Cells were cultured on glass coverslips in a 24-well plate. Cells were stimulated or not with F. nucleatum ATCC 25586 and stained for the presence of visfatin. After 3 days, cells were fixed in 4 % paraformaldehyde at pH 7.4 and RT for 15 min and then permeabilized in 0.1 % Triton X-100 for 5 min. Nonspecific antigens were blocked by incubation with serum block (LSAB System; Santa Cruz Biotechnology, Santa Cruz, CA, USA) for 20 min. Subsequently, cells were incubated overnight at 4 °C with rabbit polyclonal antibody to visfatin (Santa Cruz Biotechnology). Afterwards, cells were labeled with goat anti-rabbit IgG-HRP secondary antibody (Cell Signaling Technology) for 45 min. For staining, cells were exposed to DAB chromogen (Dako, Hamburg, Germany) for 3 min at RT. After each incubation step, cells were washed twice with PBS. Counterstaining was performed with Mayer’s Hematoxylin (Merck Eurolab, Dietikon, Switzerland) for 2 min. Coverslips were mounted in DePex mounting medium (Serva Electrophoresis, Heidelberg, Germany). Standardized photomicrographs were taken using an Axioplan 2 imaging microscope (Carl Zeiss MicroImaging).
Statistical analysis
Statistical analysis of the data was performed using SPSS 20.0 software (SPSS Inc., Chicago, IL, USA). For quantitative analysis, values were expressed as mean and standard errors of the mean. One-way analysis of variance followed by the post hoc Dunnett’s or Tukey’s tests was applied. Differences between groups were considered significant at p < 0.05. All experiments were performed in triplicate and repeated at least twice.
Results
Time- and dose-dependent stimulation of visfatin production by F. nucleatum ATCC 25586
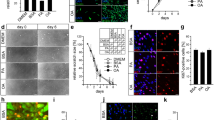
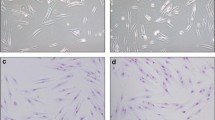
First, we sought to examine whether PDL cells produce visfatin and, if so, whether this production is regulated by bacteria. Our experiments revealed that visfatin was constitutively expressed by PDL cells and upregulated by F. nucleatum ATCC 25586 (Fig. 1a,b). As shown in Fig. 1a, F. nucleatum ATCC 25586 had no significant effect on visfatin mRNA expression up to 12 h, but it stimulated significantly the visfatin expression at 24 h (3.8-fold; p < 0.05) and 72 h (3.9-fold; p < 0.05). Furthermore, the F. nucleatum-induced increase in visfatin expression was dose-dependent. At 24 h, low concentration of F. nucleatum ATCC 25586 enhanced the visfatin expression by 3.8-fold (p < 0.05), whereas higher concentrations caused a more than 6-fold increase in its expression (p < 0.05) (Fig. 1b). The stimulatory effect of F. nucleatum ATCC 25586 on visfatin was also observed at protein level by immunocytochemistry and ELISA (Fig. 1c and Table 1).
a Stimulation of visfatin expression by F. nucleatum ATCC 25586 (OD, 0.025) in PDL cells over time. *: significantly (p < 0.05) different from unstimulated cells (n = 9); b Stimulation of visfatin expression by various concentrations of F. nucleatum ATCC 25586 in PDL cells at 24 h. *: significantly (p < 0.05) different from unstimulated cells (n = 9); c Visfatin protein synthesis in PDL cells in the presence and absence of F. nucleatum ATCC 25586 (OD, 0.025) at 3 days, as analyzed by immunocytochemistry. Images from one representative experiment are shown; d and e Regulation of visfatin expression in the presence of F. nucleatum ATCC 25586 (Fn; OD, 0.025) and/or biomechanical loading (CTS) of low (3 %) and high (20 %) magnitudes in PDL cells at 1 (d) and 3 days (e). *: significant (p < 0.05) difference between groups (n = 9)
Regulation of F. nucleatum-induced effects on visfatin by biomechanical loading
Next, we studied whether the response of PDL cells to F. nucleatum ATCC 25586 is modulated by biomechanical loading, because the periodontium is often subjected simultaneously to microbial infection and biomechanical, i.e., occlusal, loading. Biomechanical loading alone slightly decreased the constitutive visfatin mRNA expression in PDL cells at 1 and 3 days, but these effects were not significant (Fig. 1d and e). However, biomechanical loading modulated the actions of F. nucleatum ATCC 25586 on visfatin in PDL cells. CTSL abrogated the stimulatory effect of F. nucleatum ATCC 25586 on visfatin by 17 % (p > 0.05) and by 70 % (p < 0.05) at 1 and 3 days, respectively. CTSH reduced the F. nucleatum-induced stimulation of visfatin expression by 86 % (p < 0.05) at 1 day and completely blocked (p < 0.05) the visfatin upregulation at 3 days (Fig. 1d,e). The counterregulatory effects of CTSL and CTSH were also observed at protein level, as analyzed by ELISA (Table 1).
Involvement of TLRs in the actions of F. nucleatum ATCC 25586 on visfatin
We then sought to unravel the mechanisms underlying the stimulatory effect of F. nucleatum ATCC 25586 on visfatin expression. PDL cells were preincubated with neutralizing antibodies against TLR2 and TLR4 and, subsequently, stimulated with F. nucleatum ATCC 25586. Our experiments revealed that preincubation with antibodies against TLR2 and TLR4 caused a significant (p < 0.05) inhibition of the F. nucleatum-induced visfatin stimulation by 20 and by 24 %, respectively, at 1 day (Fig. 2a).
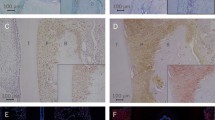
a Stimulation of visfatin expression by F. nucleatum ATCC 25586 (OD, 0.025) in the presence and absence of blocking TLR2 or TRL4 antibodies in PDL cells at 1 day. Unstimulated PDL cells in the presence and absence of blocking antibodies served as controls. *: significant (p < 0.05) difference between groups (n = 9). b Stimulation of NFκB nuclear translocation by F. nucleatum ATCC 25586 (OD, 0.025) over time, as analyzed by immunofluorescence. Images from one representative experiment are shown; c Inhibition of F. nucleatum (Fn; OD, 0.025)-stimulated p65 nuclear translocation by biomechanical loading of low (CTSL; 3 %) and high (CTSH; 20 %) magnitudes in PDL cells at 60 min. Images from one representative experiment are shown; d and e Expression of TLR2 (d) and TLR4 (e) in PDL cells subjected to biomechanical loading of low (3 %) and high (20 %) magnitudes at 1 and 3 days. *: significant (p < 0.05) difference between groups (n = 9)
Exploitation of the NFkB pathway by F. nucleatum ATCC 25586
Upon ligand binding, TLRs can trigger the NFκB signaling pathway. We therefore examined whether F. nucleatum ATCC 25586 activates this pathway in PDL cells. As evidenced by immunofluorescence microscopy, F. nucleatum ATCC 25586 stimulated the nuclear translocation of NFκB and caused a maximal NFκB accumulation within the nucleus at 60 and 90 min (Fig. 2b). Then, we examined whether this F. nucleatum-stimulated NFκB transactivation is regulated by biomechanical loading. As demonstrated in Fig. 2c, CTSL and CTSH remarkably inhibited the F. nucleatum-induced NFκB nuclear translocation at 60 min.
Regulation of TLRs by biomechanical loading
We then examined whether the inhibitory effects of biomechanical loading on F. nucleatum ATCC 25586 actions might be caused by downregulation of TLRs. Although CTSL had no regulatory effect on TLR2 at 1 day, CTSH reduced significantly the constitutive TLR2 mRNA expression by 35 % (p < 0.05) at this time point (Fig. 2d). At 3 days, the TLR2 expression was significantly inhibited by 12 % (p < 0.05) by CTSL and by 69 % (p < 0.05) by CTSH (Fig. 2d). The TLR4 expression was not significantly affected by biomechanical loading at 1 day (Fig. 2e). At 3 days, the expression of TLR4 was inhibited by 12 % (p < 0.05) by CTSL and by 52 % (p < 0.05) by CTSH (Fig. 2e).
Discussion
This study shows for the first time that the proinflammatory adipokine visfatin is produced by PDL cells. In addition, visfatin was upregulated by the oral pathogen F. nucleatum ATCC 25586, and this increase was significantly counterregulated by biomechanical loading. These findings suggest that visfatin may be produced at sites of periodontal infection with F. nucleatum ATCC 25586 and, thereby, contributes to periodontal inflammation and destruction. By contrast, biomechanical loading of the PDL might be at least partially protective against the stimulatory effects of F. nucleatum ATCC 25586 on visfatin.
Recently, the role of visfatin in inflammatory diseases and conditions has been widely investigated. Plasma visfatin levels have been found to be increased in patients with obesity, metabolic syndrome, diabetes mellitus, cardiovascular diseases, and insulin resistance [21–27]. Furthermore, it has also been demonstrated that body-weight reduction by physical exercise can lead to decreased visfatin levels [35]. In recent years, evidence has accumulated that obese individuals have an increased risk of periodontitis, and it is thought that adipokines might represent the pathomechanistic link between both diseases [12, 17]. Visfatin has been shown to stimulate a variety of cells to produce inflammatory mediators and proteases by using different intracellular pathways [36]. Moreover, this adipokine can inhibit apoptosis of inflammatory cells [37, 38]. However, if and how visfatin affects periodontal cells is as yet unclear. Interestingly, Pradeep and coworkers have recently reported increased visfatin levels in GCF from gingivitis and periodontitis patients, as compared to periodontally healthy individuals [28–30]. Furthermore, periodontal treatment caused a decrease in visfatin levels [28–30]. These reports suggest that visfatin might also be produced in the periodontium and that its synthesis is enhanced by periodontal infection. Our study demonstrates for the first time that PDL cells produce visfatin and, thereby, suggests that PDL cells may contribute to the increased visfatin levels in GCF in periodontitis. However, if and to what extent other immunoinflammatory or structural cells of the periodontium produce visfatin has yet to be elucidated.
In the present study, F. nucleatum ATCC 25586, which was used to mimic a bacterial infection, increased the visfatin production in a dose- and time-dependent manner. F. nucleatum is a gram-negative, anaerobic microorganism, acts as a bridge bacterium between early and late colonizers during plaque development, and is associated with both gingivitis and periodontitis [39, 40]. In vitro studies have shown that F. nucleatum ATCC 25586 can invade epithelial cells, fibroblasts, and PDL cells [41–43]. In addition, F. nucleatum ATCC 25586 supports other periodontal pathogens to invade host cells [44]. Nevertheless, periodontitis is caused by a complex bacterial biofilm. Further studies should clarify whether other microorganisms associated with periodontitis are also capable of stimulating the synthesis of visfatin.
Although biomechanical loading alone had only minor effects on the visfatin production in PDL cells, they inhibited the F. nucleatum-induced upregulation of visfatin in a magnitude-dependent manner. These findings indicate that biomechanical loading, for example, caused by occlusal loading or orthodontic treatment may not necessarily augment the stimulatory action of F. nucleatum ATCC 25586 on visfatin. At least for the strain regimens used in our experiments, biomechanical loading had a protective role against the actions of F. nucleatum ATCC 25586 on this proinflammatory adipokine. Our results concur with previous reports, which have also demonstrated that biomechanical loading can exert anti-inflammatory effects [31, 45, 46].
Our study revealed that one of the pathways by which F. nucleatum ATCC 25586 upregulates visfatin are TLR2 and TLR4. Interestingly, biomechanical loading decreased time- and magnitude-dependently the expression of these TLRs, suggesting that the counterregulatory effects of CTS on the action of F. nucleatum ATCC 25586 might be mediated at least in part by downregulation of these receptors, especially at 3 days. TLRs are strong activators of the NFκB pathway [47]. F. nucleatum LPS was shown to be a strong ligand for TLR2 and a weak ligand for TLR4 for activation of NFκB [48]. Interestingly, we could observe that CTS inhibited the NFκB nuclear translocation stimulated by F. nucleatum ATCC 25586, indicating that both F. nucleatum ATCC 25586 and CTS exploit this pathway for their opposite effects.
In our experiments, a suspension of F. nucleatum ATCC 25586 was used. Since this suspension was exposed to intensive ultrasonication, it can be assumed that the suspension contained disrupted cell wall particles with a high amount of LPS. However, other bacterial components also present in the suspension may interact with the receptors. The F. nucleatum ATCC 25586 concentration used in this study was determined by dose–response experiments. A low F. nucleatum concentration, as can be found in subgingival plaque, was chosen for the subsequent experiments, because this concentration exerted a pronounced stimulatory effect on visfatin, while also allowing to study the modulatory effects of biomechanical loading on the F. nucleatum-stimulated visfatin synthesis [49].
Like in previous experiments, tensile forces were applied to the cells [31–34]. However, during mastication, dental habits, and orthodontic treatment, the tooth-supporting periodontal tissues are subjected to complex forces. Whether compressive, hydrostatic, and shear forces as well as their combinations exert similar effects, as observed for tensile strain in the present study, needs to be examined. As in our previous investigations, cells were exposed to biomechanical loading of low and high magnitudes, which have been shown to occur in the periodontium [50, 51].
In summary, the present study demonstrates for the first time that visfatin is produced by PDL cells. Moreover, the oral pathogen F. nucleatum ATCC 25586 caused an upregulation of visfatin, whereas biomechanical loading counterregulated the F. nucleatum-induced stimulation of this adipokine. Within the limits of this study, we conclude that visfatin may be produced by the PDL in the presence of periodontal infection with F. nucleatum ATCC 25586. In addition, biomechanical loading of the PDL might be at least partially protective against the F. nucleatum-induced stimulation of visfatin synthesis.
References
Pihlstrom BL, Michalowicz BS, Johnson NW (2005) Periodontal diseases. Lancet 366:1809–1820
Sbordone L, Bortolaia C (2003) Oral microbial biofilms and plaque-related diseases: microbial communities and their role in the shift from oral health to disease. Clin Oral Investig 7:181–188
Seymour GJ, Ford PJ, Cullinan MP, Leishman S, Yamazaki K (2007) Relationship between periodontal infections and systemic disease. Clin Microbiol Infect 13(Suppl 4):3–10
Naveh GR, Lev-Tov Chattah N, Zaslansky P, Shahar R, Weiner S (2012) Tooth-PDL-bone complex: response to compressive loads encountered during mastication—a review. Arch Oral Biol 57(12):1575–1584
McCulloch CA, Lekic P, McKee MD (2000) Role of physical forces in regulating the form and function of the periodontal ligament. Periodontol 24:56–72
Harrel SK, Nunn ME (2001) The effect of occlusal discrepancies on periodontitis. II. Relationship of occlusal treatment to the progression of periodontal disease. J Periodontol 72:495–505
Nunn ME, Harrel SK (2001) The effect of occlusal discrepancies on periodontitis. I. Relationship of initial occlusal discrepancies to initial clinical parameters. J Periodontol 72:485–494
Wennström JL, Stokland BL, Nyman S, Thilander B (1993) Periodontal tissue response to orthodontic movement of teeth with infrabony pockets. Am J Orthod Dentofacial Orthop 103:313–319
Bildt MM, Bloemen M, Kuijpers-Jagtman AM, Von den Hoff JW (2009) Matrix metalloproteinases and tissue inhibitors of metalloproteinases in gingival crevicular fluid during orthodontic tooth movement. Eur J Orthod 31:529–535
Ren Y, Vissink A (2008) Cytokines in crevicular fluid and orthodontic tooth movement. Eur J Oral Sci 116:89–97
Goto KT, Kajiya H, Nemoto T, Tsutsumi T, Tsuzuki T, Sato H, Okabe K (2011) Hyperocclusion stimulates osteoclastogenesis via CCL2 expression. J Dent Res 90:793–798
Chaffee BW, Weston SJ (2010) Association between chronic periodontal disease and obesity: a systematic review and meta-analysis. J Periodontol 81:1708–1724
Johnson AR, Justin Milner J, Makowski L (2012) The inflammation highway: metabolism accelerates inflammatory traffic in obesity. Immunol Rev 249:218–238
Rasouli N, Kern PA (2008) Adipocytokines and the metabolic complications of obesity. J Clin Endocrinol Metab 93:64–73
Conde J, Scotece M, Gómez R, López V, Gómez-Reino JJ, Lago F, Gualillo O (2011) Adipokines: biofactors from white adipose tissue. A complex hub among inflammation, metabolism, and immunity. Biofactors 37:413–420
Inadera H (2008) The usefulness of circulating adipokine levels for the assessment of obesity-related health problems. Int J Med Sci 5:248–262
Preshaw PM, Foster N, Taylor JJ (2007) Cross-susceptibility between periodontal disease and type 2 diabetes mellitus: an immunobiological perspective. Periodontol 2000 45:138-157
Kanda N, Hau CS, Tada Y, Tatsuta A, Sato S, Watanabe S (2011) Visfatin enhances CXCL8, CXCL10, and CCL20 production in human keratinocytes. Endocrinology 152:3155–3164
Lee WJ, Wu CS, Lin H, Lee IT, Wu CM, Tseng JJ, Chou MM, Sheu WH (2009) Visfatin-induced expression of inflammatory mediators in human endothelial cells through the NF-kappaB pathway. Int J Obes (Lond) 33:465–472
Moschen AR, Kaser A, Enrich B, Mosheimer B, Theurl M, Niederegger H, Tilg H (2007) Visfatin, an adipocytokine with proinflammatory and immunomodulating properties. J Immunol 178:1748–1758
Taşkesen D, Kirel B, Us T (2012) Serum visfatin levels, adiposity and glucose metabolism in obese adolescents. J Clin Res Pediatr Endocrinol 4:76–81
Chang YH, Chang DM, Lin KC, Shin SJ, Lee YJ (2011) Visfatin in overweight/obesity, type 2 diabetes mellitus, insulin resistance, metabolic syndrome and cardiovascular diseases: a meta-analysis and systemic review. Diabetes Metab Res Rev 27:515–527
Zhang LQ, Heruth DP, Ye SQ (2011) Nicotinamide phosphoribosyltransferase in Human Diseases. J Bioanal Biomed 3:13–25
Saddi-Rosa P, Oliveira CS, Giuffrida FM, Reis AF (2010) Visfatin, glucose metabolism and vascular disease: a review of evidence. Diabetol Metab Syndr 2:21
Chen MP, Chung FM, Chang DM, Tsai JC, Huang HF, Shin SJ, Lee YJ (2006) Elevated plasma level of visfatin/pre-B cell colony-enhancing factor in patients with type 2 diabetes mellitus. J Clin Endocrinol Metab 91:295–299
Haider DG, Schindler K, Schaller G, Prager G, Wolzt M, Ludvik B (2006) Increased plasma visfatin concentrations in morbidly obese subjects are reduced after gastric banding. J Clin Endocrinol Metab 91:1578–1581
Berndt J, Klöting N, Kralisch S, Kovacs P, Fasshauer M, Schön MR, Stumvoll M, Blüher M (2005) Plasma visfatin concentrations and fat depot-specific mRNA expression in humans. Diabetes 54:2911–2916
Pradeep AR, Raghavendra NM, Prasad MV, Kathariya R, Patel SP, Sharma A (2011) Gingival crevicular fluid and serum visfatin concentration: their relationship in periodontal health and disease. J Periodontol 82:1314–1319
Pradeep AR, Raghavendra NM, Sharma A, Patel SP, Raju A, Kathariya R, Rao NS, Naik SB (2012) Association of serum and crevicular visfatin levels in periodontal health and disease with type 2 diabetes mellitus. J Periodontol 83:629–634
Raghavendra NM, Pradeep AR, Kathariya R, Sharma A, Rao NS, Naik SB (2012) Effect of non-surgical periodontal therapy on gingival crevicular fluid and serum visfatin concentration in periodontal health and disease. Dis Markers 32:383–388
Nokhbehsaim M, Deschner B, Winter J, Reimann S, Bourauel C, Jepsen S, Jäger A, Deschner J (2010) Contribution of orthodontic load to inflammation-mediated periodontal destruction. J Orofac Orthop 71:390–402
Nokhbehsaim M, Deschner B, Bourauel C, Reimann S, Winter J, Rath B, Jäger A, Jepsen S, Deschner J (2011) Interactions of enamel matrix derivative and biomechanical loading in periodontal regenerative healing. J Periodontol 82:1725–1734
Nokhbehsaim M, Deschner B, Winter J, Bourauel C, Rath B, Jäger A, Jepsen S, Deschner J (2011) Interactions of regenerative, inflammatory and biomechanical signals on bone morphogenetic protein-2 in periodontal ligament cells. J Periodontal Res 46:374–381
Nokhbehsaim M, Deschner B, Winter J, Bourauel C, Jäger A, Jepsen S, Deschner J (2012) Anti-inflammatory effects of EMD in the presence of biomechanical loading and interleukin-1β in vitro. Clin Oral Investig 16:275–283
Lee KJ, Shin YA, Lee KY, Jun TW, Song W (2010) Aerobic exercise training-induced decrease in plasma visfatin and insulin resistance in obese female adolescents. Int J Sport Nutr Exerc Metab 20:275–281
Dahl TB, Holm S, Aukrust P, Halvorsen B (2012) Visfatin/NAMPT: a multifaceted molecule with diverse roles in physiology and pathophysiology. Annu Rev Nutr 32:229–243
Li Y, Zhang Y, Dorweiler B, Cui D, Wang T, Woo CW, Brunkan CS, Wolberger C, Imai S, Tabas I (2008) Extracellular Nampt promotes macrophage survival via a nonenzymatic interleukin-6/STAT3 signaling mechanism. J Biol Chem 283:34833–34843
Jia SH, Li Y, Parodo J, Kapus A, Fan L, Rotstein OD, Marshall JC (2004) Pre-B cell colony-enhancing factor inhibits neutrophil apoptosis in experimental inflammation and clinical sepsis. J Clin Invest 113:1318–1327
Signat B, Roques C, Poulet P, Duffaut D (2011) Fusobacterium nucleatum in periodontal health and disease. Curr Issues Mol Biol 13:25–36
He J, Huang W, Pan Z, Cui H, Qi G, Zhou X, Chen H (2011) Quantitative analysis of microbiota in saliva, supragingival, and subgingival plaque of Chinese adults with chronic periodontitis. Clin Oral Investig 16(6):1579–8
Dabija-Wolter G, Cimpan MR, Costea DE, Johannessen AC, Sørnes S, Neppelberg E, Al-Haroni M, Skaug N, Bakken V (2009) Fusobacterium nucleatum enters normal human oral fibroblasts in vitro. J Periodontol 80:1174–1183
Ji S, Shin JE, Kim YS, Oh JE, Min BM, Choi Y (2009) Toll-like receptor 2 and NALP2 mediate induction of human beta-defensins by Fusobacterium nucleatum in gingival epithelial cells. Infect Immun 77:1044–1052
Han YW, Shi W, Huang GT, Kinder Haake S, Park NH, Kuramitsu H, Genco RJ (2000) Interactions between periodontal bacteria and human oral epithelial cells: Fusobacterium nucleatum adheres to and invades epithelial cells. Infect Immun 68:3140–3146
Saito A, Inagaki S, Kimizuka R, Okuda K, Hosaka Y, Nakagawa T, Ishihara K (2008) Fusobacterium nucleatum enhances invasion of human gingival epithelial and aortic endothelial cells by Porphyromonas gingivalis. FEMS Immunol Med Microbiol 54:349–355
Agarwal S, Long P, Seyedain A, Piesco N, Shree A, Gassner R (2003) A central role for the nuclear factor-kappaB pathway in anti-inflammatory and proinflammatory actions of mechanical strain. FASEB J 17:899–901
Long P, Hu J, Piesco N, Buckley M, Agarwal S (2001) Low magnitude of tensile strain inhibits IL-1beta-dependent induction of pro-inflammatory cytokines and induces synthesis of IL-10 in human periodontal ligament cells in vitro. J Dent Res 80:1416–1420
Doyle SL, O'Neill LA (2006) Toll-like receptors: from the discovery of NFkappaB to new insights into transcriptional regulations in innate immunity. Biochem Pharmacol 72:1102–1113
Yoshimura A, Kaneko T, Kato Y, Golenbock DT, Hara Y (2002) Lipopolysaccharides from periodontopathic bacteria Porphyromonas gingivalis and Capnocytophaga ochracea are antagonists for human toll-like receptor 4. Infect Immun 70:218–225
Eick S, Straube A, Guentsch A, Pfister W, Jentsch H (2011) Comparison of real-time polymerase chain reaction and DNA-strip technology in microbiological evaluation of periodontitis treatment. Diagn Microbiol Infect Dis 69:12–20
Ziegler A, Keilig L, Kawarizadeh A, Jäger A, Bourauel C (2005) Numerical simulation of the biomechanical behaviour of multi-rooted teeth. Eur J Orthod 27:333–339
Toms SR, Dakin GJ, Lemons JE, Eberhardt AW (2002) Quasi-linear viscoelastic behavior of the human periodontal ligament. J Biomech 35:1411–1415
Acknowledgments
This study was supported by a grant from São Paulo Research Foundation (FAPESP: 2010/07771-4, 2011/13752-5), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES: 2385-11-2), the German Research Foundation (DFG: KFO208/TP4), and the Medical Faculty of the University of Bonn. We would like to thank Ms. Ramona Hömig, Dr. Susanne Reimann, and Prof. Werner Götz for their great support.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nogueira, A.V.B., Nokhbehsaim, M., Eick, S. et al. Regulation of visfatin by microbial and biomechanical signals in PDL cells. Clin Oral Invest 18, 171–178 (2014). https://doi.org/10.1007/s00784-013-0935-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-013-0935-1