Abstract
The present study highlights the apoptotic activity of magnetic Fe3O4 nanoparticulates functionalized by glutamic acid and 2-pyridinecarboxaldehyde thiosemicarbazone (PTSC) toward human lung epithelial carcinoma A549 cell line. To this aim, the Fe3O4 nanoparticulates were prepared using co-precipitation method. Then, the glutamic acid and Fe3O4 nanoparticulates were conjugated to each other. The product was further functionalized with bio-reactive PTSC moiety. In addition, the synthesized Fe3O4@Glu/PTSC nanoparticulates were characterized by physico-chemical techniques including scanning electron microscope (SEM), energy dispersive X-ray (EDX), X-ray diffraction (XRD), Fourier-transform infrared (FT–IR) spectroscopy and zeta potential analysis. The effects of in vitro cell viability in Fe3O4@Glu/PTSC nanoparticulate indicated the anti-proliferative properties in a dose-dependent manner (IC50 = 135.6 µM/mL). The high selectivity for tumor cells and far below of activity in HEK293 non-tumorigenic cells is considered as an important feature for this complex (SI, 3.48). Based on the results, PTSC failed to reveal any activity against A549 cells alone. However, Fe3O4 nanoparticulates had some effects in inhibiting the growth of lung cancer cell. Furthermore, Bax and Bcl-2 gene expressions were quantified by real-time PCR method. The expression of Bax increased 1.62-fold, while the expression of Bcl-2 decreased 0.76-fold at 135.6 µM/mL concentration of Fe3O4@Glu/PTSC compared to untreated A549 cells. Furthermore, the Fe3O4@Glu/PTSC nanoparticulate-inducing apoptosis properties were evaluated by Hoechst 33258 staining, Caspase-3 activation assay and Annexin V/propidium iodide staining. The results of the present study suggest that Fe3O4@Glu/PTSC nanoparticulates exhibit effective anti-cancer activity against lung cancer cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung carcinoma (NSCLC) is a heterogeneous class of tumors which accounts for approximately 85% of all lung cancer. However, the conventional treatment process has limited influence due to the occurence of most diagnostic cases in advanced stages [1]. Thiosemicarbazones (TSCs) are considered as the substantial ligands used as potential therapeutic agents against lung cancer [2]. Thiosemicarbazones have extensive and long-term biomedical applications due to their potential biological activity including antitumor, antimicrobial, antifungal, antiviral and antimalarial activity [3,4,5]. Thiosemicarbazones induce apoptosis (programmed cell death) based on caspase activation and BAX/Bcl-2 regulation mechanisms [6, 7]. Because cancer chemotherapy is always accompanied by some side effects such as fatigue, nausea/vomiting, diarrhea, febrile neutropenia and rash [8]. Thus, reducing side effects and increasing drug efficiency should be considered for developing tumor treatment [9].
Some recent studies demonstrated the significant higher anti-cancer influence of thiosemicarbazone-metal complexes compared to the ligand alone [10, 11]. In another in vivo and in vitro study, the anti-tumor activity of 2-pyridinecarboxaldehyde thiosemicarbazone (PTSC) was confirmed [12]. The Fe2+ ion can form stable complexes with thiosemicarbazone chelating agents like other elements such as Cu, Mn, Zn and Ga [13, 14]. Iron is an essential element for deference cellular processes including cell proliferation and growth [15]. Cancer cells are sensitive to iron exclusion due to their energy generation and DNA synthesis, compared to normal cells [16]. Magnetic iron oxide (Fe3O4) nanoparticulates (MIONs) have been widely used as diagnostic fields, biomedical and drug delivery agents [17]. MIONs like other nanoparticulates such as liposomes, polymeric micelles, dendrimers and colloidal gold have been used in targeted cancer therapies. The MIONs based on active targeting bind their ligand to receptors expressed by tumor cells [18]. A large body of research was conducted to develop new thiosemicarbazone compounds and their metal complexes as antitumor agents [19,20,21]. To the best of our knowledge, this is the first report to conduct the simple route for synthesizing novel Fe3O4@Glu/PTSC nanoparticulate as an effective agent against lung cancer cells. In the present study, magnetic thiosemicarbazone-Fe3O4 nanoparticulate (Fe3O4@Glu/PTSC) was synthesized through anchoring 2-pyridinecarboxaldehyde thiosemicarbazone (PTSC) on the surface of glutamic acid functionalized iron oxide nanoparticulates. The magnetic Fe3O4@Glu/PTSC was characterized by several physico-chemical techniques. The inhibitory concentration (IC50) for A549 cancer cells and cytotoxicity concentration (CC50) for HEK293 non-tumorigenic cells were separately determined by methylthiazol tetrazolium (MTT) assay. Finally, the apoptotic activities of Fe3O4@Glu/PTSC were analyzed by Hoechst33258 staining, Caspase-3 activation assay, annexin V/propidium iodide staining and real-time PCR method.
Materials and methods
Synthesis of PTSC ligand
PTSC ligand was synthesized using the established method reported by Kovacevic et al. [22]. In brief, the commercially available thiosemicarbazide (0.91 g) and 2-pyridinecarboxaldehyde (0.95 mL) were added to ethanol (100 mL). The reaction mixture was stirred at 60–70 °C for 2 h. Two drops of glacial acetic acid were added to the mixture and refluxed for 6 h. Then, the solution was cooled to 5 °C to give a yellow PTSC precipitate. Finally, the precipitate was separated by centrifugation and washed twice with 5–10 mL of diethyl ether.
Fe3O4 and Fe3O4@Glu/PTSC nanoparticulate synthesis
Fe3O4 nanoparticulates were prepared using the described co-precipitation method by Zhang [23] and Yazdani et al. [24]. FeCl3·6H2O (7.57 g) and FeCl2·4H2O (3.17 g) were mixed in double-distilled water (300 mL) and was stirred at 75–80 °C for 1 h. Next, NH3 (40 mL) was added to this mixture and stirred at 75–80 °C for 3 h under nitrogen gas. The resulting sediment was separated using the magnet and washed 2–3 times with distilled water and dried at 70 °C. To functionalize Fe3O4 nanoparticulates with l-glutamic acid, IONs were synthesized in the presence of l-glutamic acid (0.625 g), following the procedure described above. Finally, glutamic acid functionalized IONs (1.829 g) and PTSC (0.625 g) were added to ethanol (20 mL) and the mixture was stirred at 75–80 °C for 3 h under nitrogen gas to provide Fe3O4@Glu/PTSC nanoparticulate. After separation and washing procedure, the resulting precipitate was dried at 70 °C for 8 h.
Determination of Fe3O4@Glu/PTSC nanoparticulates
Fourier-transform infrared (FT–IR) spectra of synthesized compounds were analyzed by Nicolet IR 100 spectrophotometer. The powder X-ray diffraction (PXRD) data were collected with an X-ray diffractometer (JEOL JDX-803). The size, shape and metal amount of modified Fe3O4@Glu/PTSC nanoparticulate were determined by scanning electron microscopy (MIRA3 FEG-SEM) and energy dispersive X-ray (EDX) analysis. The physical stability of Fe3O4 nanoparticulates coated with PTSC was determined using Zeta potential analyzer.
Cell culture
The human lung epithelial carcinoma A549 cell line and HEK293 kidney immortalized cell line were purchased from Iranian Biological Resource Center. Using established protocol by Cree [25], A549 cells were cultured in RPIM1640 containing 10% fetal bovine serum (FBS) including 0.05 mg/mL penicillin G and 0.08 mg/mL streptomycin at 37 °C and 5% CO2 in a humidified incubator. In addition, according to the protocol reported by Kirschner et al. [26], the HEK293 cells were grown in Dulbecco’s modified Eagle medium and prepared for various tests.
Cell viability assay
MTT assay was conducted to evaluate the cytotoxicity effects of PTSC, Fe3O4 nanoparticulates and Fe3O4@Glu/PTSC complex against human lung cancer cell line (A549) according to the protocol of Zhang et al. [27]. The HEK293 kidney immortalized cell line was used as a control in the selectivity analysis for non-tumorigenic origin [28]. To calculate the selectivity index (SI) of the Fe3O4@Glu/PTSC, the equation: CC50 (no cancer cells)/IC50 (cancer cells) was used [29]. In brief, the cells (2 × 106 cells/well) were seeded in 96-well plates. Next, the cells were treated with different concentrations (62.5, 125, 255, 500, 1000 μM/mL) of PTSC, Fe3O4 nanoparticulates and Fe3O4@Glu/PTSC nanoparticulate and incubated at 37 °C with 5% CO2 for 24 h. Afterward, 0.5 mg/mL of the MTT was added to each well and the A549 cells were incubated for 4 h. The medium in each well was replaced by 200 μL of dimethyl sulfoxide (DMSO) and gentle shaking for 30 min to dissolve the MTT formazan crystals and the absorbance of the converted dye in living cells was measured at 590 nm with a microplate reader (Bio-Rad, Hercules, CA, USA).
Cell death assays
Caspase 3 activity assay
The caspase-3 enzyme activity was measured using the caspase-3 colorimetric Protease Assay Kit (Invitrogen™ Novex™) [30]. In this test, the free chromophore p-nitroanilide (PNA) is generated following the effect of caspase-3 on tetrapeptide substrates DEVD-PNA [31]. The reaction mixture includes 50 μL of cell extract protein, 50 μL of 2 × reaction buffers (containing 0.5 μL dithiothreitol) and 5 μL of 4 mM DEVD-PNA substrate in a total volume of 105.5 µL. The reaction mixture was incubated at 37 °C for 24 h and the absorbance of the product was measured using microplate reader (Biotek Epoch, USA) at a wavelength of 405 nm according to manufacturer’s instruction.
Flow cytometry
To distinguish the apoptosis and necrosis of the cancer cells, a fluorescein isothiocyanate annexin V and propidium iodide (Annexin V-FITC Apoptosis Detection Kit, Merk) was assessed according to the manufacturer’s protocol [32]. Briefly, the A549 cells (~ 106 cells/well) were grown in six-well plates at 37 °C in a humidified atmosphere containing 5% CO2 for 24 h. Afterwards, the cells were exposed to Fe3O4@Glu/PTSC nanoparticulate with IC50 concentration with an un-treated A549 cells as a positive control.
Hoechst 33258 staining
The apoptotic cells were detected by Hoechst 33258 staining as described previously [33]. The A549 cells were seeded in six-well plates (2 × 106 cells/well) and treated with IC50 concentration (135.6 µM/mL) of Fe3O4@Glu/PTSC followed by overnight incubation. Then, the cells were collected and washed with PBS. In the next procedure, the cells were stained with Hoechst 33258 dye at 37 °C for 15 min. Finally, apoptotic and live cells were expanded on slides and were distinguished under fluorescence microscope.
Quantitative real-time PCR
In the present study, mRNA expression levels of Bcl-2 and Bax genes were evaluated using SYBR Green real-time quantitative PCR analysis [34,35,36]. Total cellular RNA was extracted using TransZol reagent (Transgen Biotech) according to the manufacturer’s protocol. The cells were cultured in a 12-well plate. To separate the adherent cells, washing was done with PBS for three times. The PBS was discarded and 1 mL TransZol was added into each well and incubated for a few minutes. After scrapping off, the cells were transferred to an RNase-free microfuge tube. The cells were dispersed with pipetting up and down and incubated for 5 min at room temperature. To provide cellular emulsion, 0.1 mL chloroform was added to the microfuge tube and the cells were shaken for 15 s. Incubation was done for 3 min at the room temperature of 4 °C and centrifuged at 10,000 rpm/min, for 10 min and the supernatant were transferred to a new microfuge tube. Then, isopropyl alcohol of 0.25 mL was added to microfuge tube and was entirely mixed. The microfuge tube was incubated for 10 min at the room temperature of 4 °C and centrifuged at 10,000 rpm/min for 10 min. After observing some colloidal precipitations at the bottom of the microfuge tube, the supernatant was discarded and 1 mL 96% ethanol was added into the tube. After vortexing, the microfuge tube was centrifuged at 8000 rpm/min, 4 °C for 15 min. The supernatant was discarded and the ethanol air-dried at the room temperature for several minutes. The precipitates were dissolved with 1–2 mL of RNA lysate and incubated at the room temperature for 1 min. Then, the microfuge tube was centrifuged at 12,000 rpm/min for 1 min. The liquid was divided into small-volume aliquots and stored at − 80 °C. Finally, the concentration of extracted RNA was detected at A260/A230 ratio using a nanodrop spectrophotometer. In addition, cDNA was synthesized using EasyScript® First-Strand cDNA Synthesis SuperMix (Transgen Biotech) based on manufacturer’s instructions and was amplified within target and control sequences. In a microtube, 5 µg total RNA, 1 µL Anchored oligo (DT) 18 primer (0.5 µg/µL) and 20 µL DNase-free water were mixed and incubated at 65 °C for 5 min and put on ice for 2 min. In addition, they were added to other components including 1 µL EasyScript RT/RI Enzyme Mix and 10 µL 2 × ES Reaction Mix. The resultant mixture was incubated at 42 °C for 15 min. To inactivate Enzymes, the mixture was incubated at 85 °C for 5 s and stored in − 80 °C. Primer sequences for the reverse transcription reactions were as follows: Bax forward: 5′-TTGCTTCAGGGTTTCATCCA-3′ and Bax reverse: 5′-AGACACTCGCTCAGCTTCTTG-3′, Bcl-2 forward: 5′-TGGCCAGGGTCAGAGTTAAA-3′ and Bcl-2 reverse: 5′-TGGCCTCTCTTGCGGAGTA-3′. The glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was utilized as the housekeeping gene to normalize within a given sample. Primer sequences for the reverse transcription reactions were GAPDH forward: 5′-CCCACTCCTCCACCTTTGAC-3′ and GAPDH reverse: 5′-CATACCAGGAAATGAGCTTGACAA-3′. qRT-PCR reactions were performed using SYBR® Premix Ex Taq™ kit (TaKaRa, Japan) in an ABI 7500 real-time PCR system (Applied Biosystems, USA) after testing the primer pairs by Premix Ex Taq (TaKaRa, Japan) and gene length analysis via electrophoresis on a 2.5% agarose gel. The thermal cycling conditions according to manufacturer’s instructions were 30 at 95 °C, 60 at 65 °C and 60 s at 72 °C for 40 cycles. Finally, the reaction was terminated at 4 °C. The reactions were conducted in triplicate. The relative quantifications of Bax and Bcl-2 mRNA expression levels were compared with the GAPDH mRNA level using the 2−ΔΔCt method.
Statistical analysis
ANOVA was used for statistical analysis using Graph Pad Prism 5.0 software. P < 0.05 was considered as the statistical significance. All measurements in the present study were evaluated in triplicate.
Results
Synthesized complex data analysis
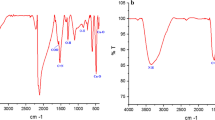
To synthesis magnetic thiosemicarbazone-Fe3O4 nanoparticulate, the glutamic acid and Fe3O4 nanoparticulates were first conjugated to each other. The product was further functionalized with bio-reactive PTSC moiety by covalent bonding of carboxylic functional groups of the glutamic acid and NH2-terminuses of PTSC. Figure S1 (supporting information) represents the FT-IR spectra of the PTSC and final Fe3O4@Glu/PTSC nanoparticulate, respectively. The significant absorption at 567 cm−1 and 621 cm−1 is attributed to stretching frequency of Fe–O [37]. The functionalization of Fe3O4 nanoparticulates with PTSC ligand can be identified by the band of 1641 cm−1, which is related to (C=N) stretching vibration [38]. The band at 813 cm−1 was assigned to (C=S) vibration in PTSC structure [39]. The emerged band at nearby 1700 cm−1 in Fe3O4@Glu/PTSC spectrum refers to the asymmetric vibrations of glutamate COO-groups [40]. The peaks at 3407 and 3469 cm−1 can be attributed to the asymmetric (N–H) vibration of the terminal NH2 group [39]. Furthermore, the peaks at 1537 and 1598 cm−1 are related to the stretching vibration of C=C bonds in the unsaturated aromatic structure. Therefore, a comparison between FT–IR peaks in commercial PTSC and synthesized Fe3O4@Glu/PTSC nanoparticulates revealed the complex successful formation.
The powder XRD pattern for Fe3O4@Glu/PTSC nanoparticulates is shown in Fig. 1. Based on the XRD analysis, the typical diffraction peaks correspond to the reflection planes (111), (220), (311), (222), (400), (422), (511) and (440) at 2θ degree of 18.63, 29.53, 35.52, 43.04, 52.68, 56.73 and 62.55, respectively, which can be indexed to the face-centered cubic phase of ferrite. The diffraction patterns of the characteristic peaks are in good agreement with the data presented previously [41]. The size of the Fe3O4@Glu/PTSC nanoparticulate was calculated from the most intense peak (311) by the Scherrer’s equation: D = kλ/βcosθ, where D indicates the average crystalline size, k shows the Scherrer constant (0.89), λ represents the X-ray wavelength used, β is considered as the angular line width at half maximum intensity and θ means the Bragg’s angle in degree unit [42]. The average calculated crystallite size was found to be about 8 nm for Fe3O4@Glu/PTSC nanoparticulate.
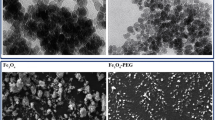
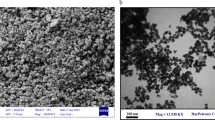
The SEM images of Fe3O4@Glu/PTSC nanoparticulate (Fig. 2) revealed that the approximately spherical, along with some moderately agglomerated particles are available in the sample. In addition, the spherical nanoparticulates were in the size range of about 20–50 nm. The main reason for the difference between the measured values in the XRD and the Scherrer equation and the observed values in the SEM images can be attributed to the relative accumulation of synthesized nanoparticulate. As shown in EDX analysis (Fig. 3), the Fe3O4@Glu/PTSC is composed of Fe, S, O, C, N atoms, without observing any other impurities. Based on characterization results, the PTSC ligand was successfully conjugated on the surface of IONs. The zeta potential measurement for the Fe3O4 nanoparticulate-coated PTSC dissolved in deionized water was about − 23.4 mV which supplies electrostatic repulsion forces between the nanoparticulates (Fig. S2: Supporting information). In a good physical colloidal stability, a zeta potential value between − 30 and + 30 mV is appropriate for providing a sufficient repulsive force between the particles [43].
Fe3O4@Glu/PTSC inhibits the growth of A549 cells
The growth inhibitory concentration (IC50), cytotoxic concentration (CC50) of Fe3O4@Glu/PTSC nanoparticulate against A549 cells and human embryonic kidney 293 cells (HEK293) were defined using MTT assay. The CC50 for HEK293 cells was 472.28 μg/mL, while the IC50 for A549 cells was 135.6 μM/mL. In vitro cell viability effects of Fe3O4@Glu/PTSC nanoparticulate indicated the anti-proliferative properties in a dose-dependent manner. The Fe3O4@Glu/PTSC nanoparticulates were significantly more toxic than PTSC and Fe3O4 nanoparticle alone (P < 0.001) (Fig. 4a). Furthermore, an increased survival rate was observed in HEK293 cells at low concentrations after treatment for 24 h (Fig. 4b). The calculated selectivity index against the A549 cell line was 3.48.
Anti-proliferative activity of PTSC, Fe3O4 NPs and Fe3O4@Glu/PTSC cell lines. Cells were grown for 24 h in the presence above iron complex and viability was determined by MTT assay. ***P < 0.001 means a significant difference between A549 cells treated with Fe3O4@Glu/PTSC. Fe3O4@Glu/PTSC nanoparticulates indicate the anti-proliferative properties in a dose-dependent manner (red bars, IC50 = 135.6 µM/mL). However, PTSC indicated no activity against A549 cells alone (blue bars), while Fe3O4 nanoparticulates present some effects in inhibiting lung cancer cell growth (pale green bars) (a). Dose–response of HEK293 after 24-h treatment with different concentrations of Fe3O4@Glu/PTSC (CC50 = 472.28). Thus, an increased survival rate can be observed in HEK293 cells at low concentrations. Finally, the cytotoxic effects of higher concentrations were observed in the Fe3O4@Glu/PTSC (b)
Enhancing Fe3O4@Glu/PTSC through activating caspase-3
The activation of apoptotic properties of nanoparticulate was evaluated by colourimetric caspase-3 assay [33, 44]. Based on the results, the caspase-3 activity of A549 cells increased 1.6 times after 24 h Fe3O4@Glu/PTSC treatment compared to untreated cells (P < 0.05) (Fig. 5).
Fe3O4@Glu/PTSC induce apoptosis in A549 cells
To evaluate the effect of Fe3O4@Glu/PTSC on apoptosis and necrosis, A549 cells were exposed to Annexin V/PI staining followed by flow cytometry analysis. In the viable cells (> 58%), both Annexin V and PI were negative (down left), while Annexin V-positive/PI-negative (down right) or Annexin V/PI double-positive (upright) occurred in the apoptotic cells, indicating the cells tolerated early (> 33%) or late (nearly 7%) apoptosis, respectively. Increasing the percentage of early and late apoptotic cells treated with Fe3O4@Glu/PTSC nanoparticulate was considerably higher than that of the untreated cells (P < 0.05) (Fig. 6). Annexin V-negative and PI-positive stained cells refer to necrotic cells (less than 3%) (upleft).
Staining the untreated A549 cells and Fe3O4@Glu/PTSC treated cells was performed with Annexin V/PI, followed by FACS analysis. The untreated primary living cells were over 94%. The cell groups included (down left), dual-negative stained cells indicating live cell group (down right), Annexin V-positive/PI-negative stained cells referring to early apoptosis (upright), Annexin V/PI dual-positive stained cells representing late apoptosis (upleft) and Annexin V-negative and PI-positive stained cells displaying dead cells
Fe3O4@Glu/PTSC increase morphological changes in A549 cells
In the present study, Hoechst 33258 staining assay was used to study cell death from another aspect. Based on the result, the untreated A549 cells stayed with a natural morphology of lung cancer cells. However, the treated cells indicated a significant change with different concentrations of Fe3O4@Glu/PTSC nanoparticulate. Furthermore, the morphological changes in the apoptotic cells were observed with an increasing reduction of cellular volume, staining bright and condensed or fragmented nucleus gradually in a dose-dependent manner (Fig. 7).
The morphology of apoptotic A549 cells was recognized with Hoechst 33258 staining and observed using fluorescence microscope (×400). Based on the results, control cells indicate normal morphology and 24-h exposure to 135.6 µM/mL Fe3O4@Glu/PTSC (IC50 dose) shows decreased cell volume, condensed chromatin, fragmented nuclei and brightly stained cells (scale bar 50 µm)
Fe3O4@Glu/PTSC upregulate BCL-2 gene and downregulate BAX gene
Based on the results, the qRT PCR indicated that the magnetic Fe3O4 nanoparticulate and PTSC play a significant synergistic role on Bax over-expression. The Bax mRNA expression in the treated A549 cells increased to 1.62-fold in 24 h treatments compared to the untreated control cells (Fig. 8a). However, the Bcl-2 mRNA expression indicated a significant decrease of 0.76-fold at IC50 concentration of Fe3O4@Glu/PTSC nanoparticulate compared to untreated control cells (Fig. 8b).
Discussion
Some studies reported a strong relationship between thiosemicarbazones and their metal complexes for inhibiting the growth of lung cancer cell. The results of another study reported the synergistic activity of the copper complexes of Dp4e4mT and DpC in combination with gemcitabine or cisplatin against lung cancer cells [45]. In addition, a series of novel nickel (II) thiosemicarbazone complexes (1–4) inhibited the migration and metastasis in lung cancer cells [46]. The toxicity of 2-acetylpyridine-thiosemicarbazones Ga (III) complex against human fetal lung fibroblast cells (MRC-5) was shown via Topo-I inhibition and distinct DNA cleavage [47]. Some of palladium thiosemicarbazone complexes exhibited better activity than their parent thiosemicarbazone analogues on A549 cell line [48]. By considering these studies, the present research aimed to determine the effect of Fe3O4 magnetite nanoparticulates (MNPs) Pyridinecarboxaldehyde thiosemicarbazone complex called as Fe3O4@Glu/PTSC on A549 cell line. The present project was conducted to identify the anti-proliferative activity of this compound on these cells and compare the inhibitory concentration (IC50) of the drug with its cytotoxic concentration (CC50) on cancer cell line and normal cells. In addition, the mechanism of inhibiting this complex was evaluated, along with the changes in gene expression of BAX and BCL-2 in treated cells with Fe3O4@Glu/PTSC. The results indicated that Fe3O4@Glu/PTSC complex has a proper anti-proliferative activity (IC50, 135.6 µM/mL) against the growth of A549 lung cancer cells after 24-h incubation (Fig. 4a). However, less toxic effect on the non-tumorigenic HEK293 cells (CC50, 472.28 µM/mL) was observed in the compound (Fig. 4b). Furthermore, the drug selectivity index of Fe3O4@Glu/PTSC exhibited a high degree of cytotoxic selectivity (SI, 3.48), because the SI values greater than 2 can be recommended in developing anticancer drugs [49]. The computed SI is higher than that of Matesanz et al. (max, 2.46) for other metal heterocyclic thiosemicarbazone complex on the A549 cells [50]. Unexpectedly, the thiosemicarbazone revealed no activity against A549 cells alone, indicating that this compound is unable to penetrate cells. The finding is consistent with that of Hall et al. [51]. Furthermore, Fe3O4 nanoparticulates could prevent from growing lung cancer cell, which is in line with that of Zhang et al. [52]. The results can provide further support for the hypothesis that the thiosemicarbazone is effective against A549 tumor cells and iron oxide nanoparticulates act as drug delivery compounds. Some examples were presented to illustrate the applications of Fe3O4 nanoparticulates as drug-delivery vehicles [42, 52, 53]. The ability of PTSC to inhibit the growth of tumor cells can be correlated with its efficacy to increase Fe efflux and prevent from uptaking Fe by cancer cells [54]. Accordingly, in the present study, Fe3O4@Glu/PTSC activated apoptotic pathways due to the PTSC iron chelator feature and tumor cell death. Therefore, we performed apoptotic confirmatory experiments to represent its inhibitory activity on growing and proliferating cancer cell. To evaluate the mode of apoptosis, the most effective enzyme activity was observed for caspase-3, which increased 1.6-fold compared with the control after 24-h incubation with Fe3O4@Glu/PTSC (Fig. 5). In line with the results of the present study, some demonstrated that some similar chelators with PTSC have proximity activity on caspase-3 in other cell lines. Shandiz et al. [40] demonstrated 1.7-fold activity of caspase 3 via thiosemicarbazide on the MCF-7 cell line and Qi et al. [55] reported an increase of 1.5–2.2-fold of caspase via 2-pyridinecarboxaldehyde thiosemicarbazones Ga(III) complexes. However, no study had focused on the effect of caspase assay on this complex. These minor differences are probably related to the structural properties of Fe3O4@Glu/PTSC and thiosemicarbazide. Since caspase activation is associated with h-cytochrome release from the mitochondrion [56], the enhanced caspase activity in treated cells compared to untreated cells, indicated that Fe3O4@Glu/PTSC induced the mitochondrial pathway of apoptosis. Apoptotic cells can be further determined using biochemical features including the expression of surface markers such as phosphatidylserine (PS), as well as the recognition ligand for phagocytosing dying cells [57]. PS is located inside the plasma membrane and is considered as a negatively charged phospholipid. PS exteriorization is a primary characteristic of the early apoptosis. The transposition of PS can be traced by Annexin V protein which has a high affinity [58]. Annexin V (+)/PI (−) staining in the A549 tumor cells after Fe3O4@Glu/PTSC treatment indicated induced early apoptosis (> 33%) compared to the control cells (0.23%) (Fig. 6). Thus, this thiosemicarbazone Fe3O4 nanoparticulate compound contains unique properties interpreting Fe3O4@Glu/PTSC as a potential anti-proliferative agent against A549 lung cancer cells. Therefore, both caspase activity and Annexin-V data refer to the fact that Fe3O4@Glu/PTSC activates intrinsic mitochondrial-mediated apoptosis in cultured lung cancer cells. Based on the results, PTSC activity conjugated with Fe3O4 is more than the case conjugated with metal ions such as Ga(III),as it was reported in the study of Qi et al. [55]. In addition, using 2-pyridyl derivatives alone can reduce early apoptosis and induce late apoptosis [54]. Thus, the functionalization of Fe3O4 nanoparticulates with thiosemicarbazones can promote early apoptosis in lung cancer cells, indicating that iron nanoparticulate enhances early apoptosis. Furthermore, it plays a role in drug delivery, which is consistent with the results of Namvar et al. [59].
The changes in Bax and BCL-2 gene expression are considered as another factor which may explain the relatively good correlation between Fe3O4@Glu/PTSC and apoptosis. BCL-2 is markedly localized in the outer membrane of mitochondria and is divided into pro-apoptotic BAX/BAK proteins, anti-apoptotic proteins, which are the same pro-survival BCL-2 family proteins (BCL-2, BCL-XL, BCL-W, MCL-1, A1/BFL-1) and pro-apoptotic BH3-only proteins [1, 60]. Some BH3-only proteins such as BIM, tBID and PUMA are directly connected to BAX/BAK and activate them, which subsequently inhibit the pro-survival BCL-2-like proteins. Furthermore, it becomes permeable outer mitochondrial membrane, which allows to release cytosolic apoptogenic factors such as cytochrome c Smac/DIABLO, leading to the activation of caspase cascade [60]. Finally, the extrinsic and intrinsic pathway meets together via activating caspase-3 and interacting through pro- and antiapoptotic proteins [61]. The tumor cells survive under undesirable stress conditions and overcome apoptotic pathways by their increased ratio of anti-apoptotic to pro-apoptotic Bcl-2 proteins [1]. Thus, regulating can be efficient for the lung cancer treatment by anti-apoptotic Bcl-2. In the present study, the modification of expressing BAX and Bcl-2 in A459 cell line was performed by Fe3O4@Glu/PTSC via qRT-PCR. This Nano-complex could reduce the expression of Bcl-2 mRNA and induce the expression of BAX mRNA, which is considered as one of the pro-apoptotic molecular pathways of Fe3O4@Glu/PTSC for preventing from growing lung cancer cells. In fact, Fe3O4@Glu/PTSC induces cancer cell apoptosis and inhibits the growth of NSCLC cells through regulating the Bcl-2 family proteins and triggering the caspase-3 activation. However, Fe3O4@Glu/PTSC effect on other cell lines and the expression of other genes lines, as well as its effect in vivo on health cells are suggested for further research.
Conclusion
Based on the results, the novel Fe3O4@Glu/PTSC indicated promising an anti-proliferation activity against cancer cells. In addition, the results suggested that the synergistic effect of PTSC and Fe3O4 nanoparticulates trigger BAX up-regulation and Bcl 2 down-regulation, which can lead to the activation of intrinsic apoptosis pathway.
References
Prabhu VV, Elangovan P, Devaraj SN, Sakthivel KM (2018) Gene 679:352–359
Sever B, Akalin Çiftci G, Özdemir A, Altintop MD (2019) J Res Pharm 23:16–24
Ganim MA, Baloglu MC, Aygun A, Altunoglu YC, Sayiner HS, Kandemirli F, Sen F (2018) Int J Biol Macromol 122:1271–1278
Altıntop MD, Atlı Ö, Ilgın S, Demirel R, Özdemir A, Kaplancıklı ZA (2016) Eur J Med Chem 108:406–414
Da Silva Mesquita R, Tadei WP, Bastos AMB (2018) Brazil J Entomol Nematol 10:37–42
Liu B, Sun R, Luo H, Liu X, Jiang M, Yuan C, Yang L, Hu J (2017) Immunobiology 222(2):198–205
Zheng JH, Follis AV, Kriwacki RW, Moldoveanu T (2016) FEBS J 283(14):2690–2700
Schirrmacher V (2019) FEBS J 283:2690–2700
Li-Weber M (2009) Cancer Treat Rev 35(1):57–68
Akladios FN, Andrew SD, Parkinson CJ (2016) JBIC J Biol Inorg Chem 21(8):931–944
Pelosi G (2010) Open Crystallogr J 3:16–28
Gou Y, Wang J, Chen S, Zhang Z, Zhang Y, Zhang W, Yang F (2016) Eur J Med Chem 123:354–364
Jansson PJ, Sharpe PC, Bernhardt PV, Richardson DR (2010) J Med Chem 53(15):5759–5769
El Metwally NM, Arafa R, El-Ayaan U (2014) Therm Anal Calorim 115(3):2357–2367
Babu KR, Muckenthaler MU (2016) J Mol Med 94(3):347–359
Whitnall M, Howard J, Ponka P, Richardson DR (2006) Proc Natl Acad Sci 103(40):14901–14906
Liu Y, Li J, Xu K, Gu J, Huang L, Zhang L, Liu N, Kong J, Xing M, Zhang L (2018) Toxicol Lett 292:151–161
Wang J, Sui M, Fan W (2010) Curr Drug Metab 11(2):129–141
Lisic EC, Rand VG, Ngo L, Kent P, Rice J, Gerlach D, Papish ET, Jiang X (2018) Open J Med Chem 8(2):30
Prakash Kinthada M, Gust R (2018) Int J Drug Des Dev 1(1):1001
Akam EA, Tomat E (2016) Bioconjug Chem 27(8):1807–1812
Kovacevic Z, Chikhani S, Lovejoy DB, Richardson DR (2011) Mol Pharmacol 80:598–609
Zhang H, Thomas R, Oupicky D, Peng F (2008) J Biol Inorg Chem 13(1):47–55
Yazdani F, Seddigh M (2016) Mater Chem Phys 184:318–323
Cree IA (2011) Cancer cell culture: methods and protocols. Springer, Berlin
Kirschner KM, Wagner N, Wagner K-D, Wellmann S, Scholz H (2006) J Biol Chem 281(42):31930–31939
Zhang B, Luo H, Xu Q, Lin L, Zhang B (2017) Oncotarget 8(8):13620
Convertini P, Tramutola F, Iacobazzi V, Lupattelli P, Chiummiento L, Infantino V (2015) Chem Biol Interact 237:1–8
Peña-Morán O, Villarreal M, Álvarez-Berber L, Meneses-Acosta A, Rodríguez-López V (2016) Molecules 21(8):1013
Cabrera M, Gomez N, Lenicov FR, Echeverría E, Shayo C, Moglioni A, Fernández N, Davio C (2015) PLoS One 10(9):e0136878
Suliman YAO, Ali D, Alarifi S, Harrath AH, Mansour L, Alwasel SH (2015) Environ Toxicol 30(2):149–160
Wang Y, Gao W, Shi X, Ding J, Liu W, He H, Wang K, Shao F (2017) Nature 547(7661):99
Zhao Y, Guo C, Wang L, Wang S, Li X, Jiang B, Wu N, Guo S, Zhang R, Liu K (2017) Biochem Biophys Res Commun 491(1):65–71
Pearce MC, Gamble JT, Kopparapu PR, O’Donnell EF, Mueller MJ, Jang HS, Greenwood JA, Satterthwait AC, Tanguay RL, Zhang X-K (2018) Oncotarget 9(40):26072
Chen B, Liang Y, Wu W, Cheng J, Xia G, Gao F, Ding J, Gao C, Shao Z, Li G (2009) Int J Nanomed 4:251
Eimani BG, Sanati MH, Houshmand M, Ataei M, Akbarian F, Shakhssalim N (2014) Cell J (Yakhteh) 15(4):356
Lopez JA, González F, Bonilla FA, Zambrano G, Gómez ME (2010) Revista Latinoamericana de Metalurgia y Materiales 30(1):60–66
Tadjarodi A, Ferdowsi SM, Zare-Dorabei R, Barzin A (2016) Ultrason Sonochem 33:118–128
Bamgboye TT, Bamgboye OA (1985) Inorg Chim Acta 105(3):223–226
Shandiz SAS, Montazeri A, Abdolhosseini M, Shahrestani SH, Hedayati M, Moradi-Shoeili Z, Salehzadeh A (2018) J Cluster Sci 29(6):1107–1114
Zargoosh K, Zilouei H, Mohammadi MR, Abedini H (2014) CLEAN Soil Air Water 42(9):1208–1215
El-Boubbou K (2018) Nanomedicine 13(8):929–952
Joseph E, Singhvi G (2019) Nanomaterials for Drug Delivery and Therapy 1:91–116
Liang W, Li X, Li Y, Li C, Gao B, Gan H, Li S, Shen J, Kang J, Ding S (2014) Int J Oncol 44(1):91–98
Lovejoy DB, Sharp DM, Seebacher N, Obeidy P, Prichard T, Stefani C, Basha MT, Sharpe PC, Jansson PJ, Kalinowski DS (2012) J Med Chem 55(16):7230–7244
Kalaivani P, Saranya S, Poornima P, Prabhakaran R, Dallemer F, Padma VV, Natarajan K (2014) Eur J Med Chem 82:584–599
Qi J, Zheng Y, Qian K, Tian L, Zhang GX, Cheng Z, Wang Y (2017) J Inorg Biochem 177:110–117
Prabhakaran R, Kalaivani P, Poornima P, Dallemer F, Huang R, Vijaya Padma V, Natarajan K (2013) Bioorg Med Chem 21(21):6742–6752
de Oliveira PF, Alves JM, Damasceno JL, Oliveira RAM, Dias Júnior H, Crotti AEM, Tavares DC (2015) Revista Brasileira de Farmacognosia 25(2):183–188
Matesanz AI, Jimenez-Faraco E, Ruiz MC, Balsa LM, Navarro-Ranninger C, León IE, Quiroga AG (2018) Inorg Chem Front 5(1):73–83
Hall MD, Brimacombe KR, Varonka MS, Pluchino KM, Monda JK, Li J, Walsh MJ, Boxer MB, Warren TH, Fales HM (2011) J Med Chem 54(16):5878–5889
Zhang G, Ding L, Renegar R, Wang X, Lu Q, Huo S, Chen YH (2011) Cancer Sci 102(6):1216–1222
Wu L, Zhang F, Wei Z, Li X, Zhao H, Lv H, Ge R, Ma H, Zhang H, Yang B (2018) Biomater Sci 6(10):2714–2725
Yuan J, Lovejoy DB, Richardson DR (2004) Blood 104(5):1450–1458
Qi J, Deng J, Qian K, Tian L, Li J, He K, Huang X, Cheng Z, Zheng Y, Wang Y (2017) Eur J Med Chem 134:34–42
Deus CM, Santos GL, Loureiro R, Vega-Naredo I, Faneca H, Oliveira PJ (2015) Curr Med Chem 22(20):2438–2457
Elmore S (2007) Toxicol Pathol 35(4):495–516
Hingorani R, Deng J, Elia J, McIntyre C, Mittar D (2011) BD Biosciences 1–11
Namvar F, Rahman HS, Mohamad R, Baharara J, Mahdavi M, Amini E, Chartrand MS, Yeap SK (2014) Int J Nanomed 9:2479–2488
Delbridge A, Strasser A (2015) Cell Death Differ 22(7):1071
Grobmyer SR, Moudgil BM (2010) Biomed Eng Online 9:55–56
Acknowledgements
We express gratefulness to the biology department, Islamic Azad University, Rasht, Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflict of interest related to this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Habibi, A., Sadat Shandiz, S.A., salehzadeh, A. et al. Novel pyridinecarboxaldehyde thiosemicarbazone conjugated magnetite nanoparticulates (MNPs) promote apoptosis in human lung cancer A549 cells. J Biol Inorg Chem 25, 13–22 (2020). https://doi.org/10.1007/s00775-019-01728-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-019-01728-4