Abstract
We have studied the effect of urea-induced unfolding on the electron transfer process of yeast iso-1-cytochrome c and its mutant K72AK73AK79A adsorbed on electrodes coated by mixed 11-mercapto-1-undecanoic acid/11-mercapto-1-undecanol self-assembled monolayers. Electrochemical measurements, complemented by surface enhanced resonance Raman studies, indicate two distinct states of the adsorbed proteins that mainly differ with respect to the ligation pattern of the haem. The native state, in which the haem is axially coordinated by Met80 and His18, displays a reduction potential that slightly shifts to negative values with increasing urea concentration. At urea concentrations higher than 6 M, a second state prevails in which the Met80 ligand is replaced by an additional histidine residue. This structural change in the haem pocket is associated with an approximately 0.4 V shift of the reduction potential to negative values. These two states were found for both the wild-type protein and the mutant in which lysine residues 72, 73 and 79 had been substituted by alanines. The analysis of the reduction potentials, the reaction enthalpies and entropies as well as the rate constants indicates that these three lysine residues have an important effect on stabilising the protein structure in the adsorbed state and facilitating the electron transfer dynamics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytochrome c is one of the most widely studied proteins in the past few decades. The strong interest in this small soluble haem protein is, on the one hand, related to its important physiological functions in the respiratory chain of aerobic organisms and in apoptotic pathways [1–4]. On the other hand, owing to its small size, its well-characterised structural and spectral properties as well as the availability of engineered protein variants, cytochrome c is frequently used as model protein for studying fundamental biophysical processes such as electron transfer [5–9] and protein folding [10–17]. Specifically, the elucidation of relationships between protein folds and dynamics and the electron transfer properties is of particular interest in view of its impact for understanding biological processes on a molecular level and for the design of novel tailor-made redox enzymes for potential biotechnological applications.
The present work is dedicated to contributing to the determination of those structural parameters that control the thermodynamics and kinetics of interfacial redox processes. We have focussed on the effect of protein structural changes induced by urea on the redox properties of cytochrome c immobilised on electrodes. So far, polypeptide unfolding of cytochrome c has been intensively studied in solution by a variety of techniques, monitoring changes of the secondary and the tertiary structure of the protein as well as structural alterations of the haem cofactor [10–17]. A variety of intermediates along the unfolding pathways have been identified, including those differing with respect to the haem ligation.
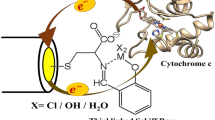
The most vulnerable part of the haem pocket of ferric cytochrome c has been shown to be the Met80 ligand, which is easily detached from haem iron, constituting the first step of the denaturant-induced structural perturbation of the haem pocket. As shown by Yeh et al. [17], the vacant coordination site is readily occupied by His33 or His26, constituting a kinetic trap in the unfolding process. However, 1H-NMR spectroscopic studies led to the conclusion that a Lys residue substitutes the Met80 ligand [15], which would imply a quite different unfolding mechanism. In the present work, experiments with the K72AK73AK79A cytochrome c variant helped to discriminate between these two ligation patterns. In fact, the substitution of those Lys residues (Scheme 1) involved in the coordination with the iron atom [15] excludes the existence of a Lys-coordinated species. Moreover, since the same Lys residues are also responsible for the electrostatic binding of cytochrome c on negatively charged self-assembled monolayers (SAMs) [18, 19], their substitution affects the kinetics of the interfacial electron transfer process, suggesting different haem orientations for recombinant non-trimethylated Saccharomyces cerevisiae iso-1-cytochrome c (ycc) and K72AK73AK79A with respect the electrode surface.
In this work we employed electrochemical and spectroelectrochemical methods to gain further insight into the interplay between protein unfolding and electron transfer of yeast iso-1 cytochrome c immobilised on electrodes. The techniques of choice were cyclic voltammetry (CV), probing the thermodynamics and kinetics of the interfacial redox process, in combination with surface enhanced resonance Raman (SERR) spectroscopy, which is a powerful tool to identify the nature of the species involved. The aim of this work was to provide novel insights into the factors controlling the interfacial electron transfer of ycc immobilised on biocompatible surfaces in the presence of urea. In this respect, the experimental approach allows combination of the thermodynamic and kinetic electrochemical data with structural information obtained from SERR spectroscopy to gain comprehensive insight into the behaviour of ycc.
Materials and methods
Materials
Wild-type ycc and its variant K72AK73AK79A were expressed in Escherichia coli and purified following procedures described elsewhere [20–22]. In both cases, Cys102 was replaced by a threonine to avoid dimerisation and minimise autoreduction without affecting the spectral and the functional properties of the protein [23, 24]. All chemicals were of reagent grade. 11-Mercapto-1-undecanoic acid (MUA) and 11-mercapto-1-undecanol (MU) were purchased from Sigma-Aldrich and were recrystallised from hexane before use. Urea was purchased from Sigma-Aldrich. Nanopure water was used throughout.
Electrochemical measurements
A model 273A potentiostat/galvanostat (EG&G PAR, Oak Ridge, USA) was used to perform CV. Experiments were carried out at different scan rates (0.02–5 V s−1) using a cell for small-volume samples (0.5 mL) under argon. A 1-mm-diameter polycrystalline gold wire, a platinum sheet, and a saturated calomel electrode (SCE) were used as the working, counter, and reference electrodes, respectively. The electrical contact between the SCE and the working solution was achieved with a Vycor® (from PAR) set. Potentials were calibrated against the MV2+/MV+ couple (MV is methylviologen) [25]. All the redox potentials reported here are referred to the standard hydrogen electrode, unless otherwise specified. The working gold electrode was cleaned by flaming under oxidising conditions; afterwards, it was heated in concentrated KOH for 30 min, rinsed with water and subsequently cleaned with concentrated sulfuric acid for 30 min. To minimise residual adsorbed impurities, the electrode was subjected to 20 voltammetric cycles between +1.5 and −0.25 V (vs. SCE) at 0.1 V s−1 in 1 M H2SO4. Finally, the electrode was rinsed in water and anhydrous ethanol. The Vycor® set was treated in an ultrasonic pool for about 5 min. SAM coatings on the gold electrode were obtained by dipping the polished electrode into a 1 mM ethanol solution of both MUA and MU for 12 h and then rinsing it with Milli-Q water. Protein solutions were freshly prepared before use in 5 mM phosphate buffer at pH 7 and their concentration (typically 0.2 mM) was carefully checked spectrophotometrically (JASCO V-570 spectrophotometer). Protein adsorption on the SAM-coated gold electrode was achieved by dipping the functionalised electrode into a 0.2 mM protein solution at 277 K for 5 h. Standard electrolyte solutions included 5 mM sodium perchlorate and 5 mM phosphate buffer at pH 7. The urea concentration was varied between 0 and 8 M. The formal reduction potentials E°′ were taken to be the midpoint between the anodic and cathodic peak potentials [26] and were found to be almost independent of scan rate in the range 0.02–5 V s−1. For each species, the experiments were performed at least five times and the reduction potentials were found to be reproducible within ±2 mV. Cyclic voltammograms at different scan rates were also recorded to determine the electron transfer rate constant k s for the adsorbed protein. The k s values obtained from five measurements were found to be reproducible within 6%. The CV experiments at different temperatures were carried out with a cell in a “non-isothermal” setting [26], in which the reference electrode was kept at constant temperature (294 ± 0.1 K) whereas the half-cell containing the working electrode and the Vycor® junction to the reference electrode was under thermostatic control with a water bath. The temperature was varied from 278 to 323 K. With this experimental configuration, the standard entropy change for Fe(III) to Fe(II) cytochrome c reduction \( (\Updelta S^{\circ \prime}_{\text{rc}} ) \) is given by [27, 28]
Thus, \( \Updelta S^{\circ \prime}_{\text{rc}} \) was determined from the slope of the plot of E°′ versus T, which is linear under the assumption that \( \Updelta S^{\circ \prime}_{\text{rc}} \) is constant over the temperature range investigated. With the same assumption, the enthalpy change \( (\Updelta H^{\circ \prime}_{\text{rc}} ) \) was obtained from the Gibbs–Helmholtz equation, namely as the negative slope of the E°′/T versus 1/T plot. The non-isothermal behaviour of the cell was carefully checked by determining the \( \Updelta H^{\circ \prime}_{\text{rc}} \) and \( \Updelta S^{\circ \prime}_{\text{rc}} \) values of the ferricyanide/ferrocyanide couple [27, 28].
The activation enthalpy ΔH # was obtained using the Arrhenius equation assuming ΔH # = ΔG #. This approximation implies that the contribution of the activation entropy is negligibly small [28–30].
SERR spectroscopy measurements
SERR spectra were obtained with 413-nm excitation using the experimental set-ups described previously [31, 32]. A detailed description of the preparation of the SERR-active surface, SAM formation, and protein adsorption is given elsewhere [32, 33].
Results and discussion
Formal reduction potentials of the immobilised cytochromes
The electrochemical response of cytochrome c immobilised on MUA/MU-coated electrodes exposed to increasing urea concentrations is qualitatively the same for both ycc and K72AK73AK79A immobilised either on polycrystalline gold (Fig. 1) or on roughened silver electrodes (Fig. S1).
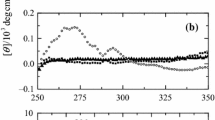
a Cyclic voltammetry (CV) curves for recombinant non-trimethylated Saccharomyces cerevisiae iso-1-cytochrome c (ycc) immobilised on an 11-mercapto-1-undecanoic acid (MUA)/11-mercapto-1-undecanol (MU)-modified gold electrode in absence of urea (lightest trace), in 2 M urea (darker trace) and in 8 M urea (darkest trace) at scan rate of 0.05 V s−1. b CV curves for ycc immobilised on an MUA/MU-modified gold electrode in absence of urea (lighter trace) and in 8 M urea (darker trace) at scan rate of 0.5 V s−1. The solution contained 5 mM sodium perchlorate and 5 mM phosphate buffer (pH 7, T = 278 K)
The CV traces of ycc obtained at urea concentrations between 0 and 2 M are very similar (Fig. 1a, wave I, Table 1), featuring two voltammetric peaks ascribed to the monoelectronic oxidation and reduction of the haem group. Importantly, experiments performed in a broader potential window did not display further features (Fig. S2). Given the quasi-reversibility of the electron transfer process, as inferred from the linear plot of the peak current versus the scan rate (not shown), the formal potential E°′ of the immobilised ycc was determined as the average of the anodic and the cathodic peak. The value of E°′ = 0.209 V is in agreement with previous findings and is indicative of native ycc [34].
At urea concentration between 3 and 8 M, a new cathodic peak (Fig. 1a, wave II, Table 1) is detected at more negative potentials. The anodic counterpart of wave II is observed only at scan rates v ≥ 0.05 V s−1, allowing us to determine the formal potential of −0.233 V for this new redox couple. Evidence that wave II originates from a non-native protein species immobilised on the electrode is derived from the following considerations. Cyclic voltammograms recorded under identical experimental conditions on the bare SAM, i.e. in the absence of immobilised protein, did not feature voltammetric peaks in the potential region of wave II, which hence must be attributed to the protein. When the electrode featuring wave II is exposed to buffer solution without urea for 36 h, the partial recovery of the native wave I and the concomitant decrease of wave II were observed (Fig. S2). In this range of urea concentration the “non-native peaks” coexist with the “native voltammetric signal”, which is still present, albeit with diminished current intensity and an E°′ slightly shifted to negative values (see Table 1). Thus, the cathodic current intensities of waves I and II are correlated as shown in Fig. 2. The results reveal that with increasing urea concentration the magnitude of wave II increases at the expense of that of wave I, although the sum of the peak currents is constant. This finding reflects the distribution of two ycc species present on the electrode at the different urea concentrations.
Cathodic peak currents measured for ycc and its variant K72AK73AK79A immobilised on an MUA/MU-modified gold electrode at different urea concentrations. ycc, wave I filled circles; ycc, wave II open circles; K72AK73AK79A, wave I inverted triangles; K72AK73AK79A, wave II upright triangles. The solution contained 5 mM sodium perchlorate and 5 mM phosphate buffer (pH 7, T = 278 K)
At urea concentrations higher than 6 M, the voltammetric trace is dominated by the non-native signal, although a minor peak at +0.110 V is still present (Fig. 1a). The cathodic peak of wave II is readily detectable, whereas its anodic counterpart is observed only at scan rates v ≥ 0.05 V s−1, although the intensities of the two peaks become comparable at higher scan rates (Fig. 1b). Moreover, whereas at low scan rates the anodic currents of waves I and II are comparable (Fig. 1a, darkest trace), at high scan rates only the counterpart of wave II is observed (Fig. 1b, darker trace). This behaviour can be explained in terms of the low stability of the reduced non-native species, which rapidly converts to the native one, such that its oxidation is detectable only at high scan rates (vide infra).
The outstanding negative potentials of the non-native species (wave II) are in good agreement with those reported in solution for cytochrome c having a Met80 replaced by a His or a Lys residue [35, 36]. Solely on the basis of electrochemistry measurements, we cannot discriminate between these two different ligation patterns. The nature of the immobilised species at high urea concentrations was therefore further investigated with SERR spectroscopy.
SERR spectroscopic characterisation of the immobilised cytochromes
In the absence of urea in the electrolyte solution, the immobilised ycc affords SERR spectra at +0.1 and −0.2 V (vs. SCE) that are essentially identical to the resonance Raman spectra of the oxidised and reduced protein solution, respectively (data not shown). This agreement implies that under these conditions the native protein structure is preserved in the immobilised state, thereby confirming previous results [32]. Very similar SERR spectra were obtained for K72AK73AK79A (Fig. S5).
After equilibration of the electrode with a solution containing 8 M urea, the SERR spectrum of ycc obtained at +0.1 V (vs. SCE) displayed the band pattern characteristic of a ferric six-coordinated low-spin haem (Fig. 3, spectrum B), but all bands in the so-called marker band region are upshifted by 2–4 cm−1 compared with those of the native ferric form (Fig. 3, spectrum A). Essentially the same upshifts of the marker bands are observed for ferric cytochrome c in solution containing 6 M guanidinium hydrochloride [17, 37] or after binding to negatively charged surfaces at high electric fields (i.e., state B2) [32, 37–39]. Under these conditions, the Met80 ligand is replaced by His33 (or His26) [17, 37], and this replacement evidently also takes place for ycc immobilised on the SAM-coated electrode in the presence of 8 M urea. This conclusion is also true for K72AK73AK79A as judged from the similarities of the SERR spectra (Fig. S5).
Surface enhanced resonance Raman spectra of ycc immobilised on an MUA/MU-coated silver electrode at +0.1 V (vs. the saturated calomel electrode) in the absence of urea in the electrolyte solution (A, C) and in the presence of 8 M urea (B, D). The solution contained 5 mM sodium perchlorate and 5 mM phosphate buffer (pH 7, T = 278 K). The spectra were obtained with 413-nm excitation
This ligand exchange is further reflected by the SERR spectrum between 280 and 700 cm−1 which displays a unique vibrational signature for the specific haem–protein interactions in cytochrome c. Thus, the quite drastic spectral changes in this region (Fig. 3, spectra C, D) indicate the structural rearrangement of the haem pocket associated with the ligand exchange. Again, these spectral changes are very similar to those observed for cytochrome c bound to negatively charged surfaces [40].
Identification of the adsorbed species
The SERR spectroscopic data allow the two waves in the cyclic voltammograms to be assigned to two distinct states of the immobilised cytochrome c which differ with respect to the haem ligation. Whereas wave I corresponds to the state including the native axial ligand pair Met80 and His18, wave II originates from a state in which Met80 is replaced by His33 (or His26). This latter state is only formed at high concentrations of the denaturant, consistent with previous results obtained for the protein in solution where this state, denoted as U[6cLS] (where 6cLS is “six-coordinated low spin”), prevails in the presence of 6 M guanidinium hydrochloride at pH 7 [17, 37]. The haem pocket structure of U[6cLS] is very similar to that of the bishistidine-ligated B2 state (B2[6cLS]) induced by high electric fields. The differences refer to the protein structure since the formation of U[6cLS] involves a partial unfolding of the polypeptide chain as demonstrated by circular dichroism spectroscopy, whereas in B2[6cLS] structural changes are largely restricted to the level of the tertiary structure [37]. For the sake of simplicity, we adopt this nomenclature and assign wave II to the U[6cLS] state. The appearance of the anodic counterpart of wave II only for high scan rates is probably related to the instability of the reduced bishistidine form which transforms rapidly into the corresponding His-Met form, denoted as the B1 state [38]. This is not surprising since the reduced form of cytochrome c in solution retains its His-Met ligation up to 9 M urea [41, 42]. The transition from the ferrous U[6cLS] to the ferric B1 state requires the removal of the His33 (His26) ligand from the haem, rebinding of Met80 and the movement of the peptide segment including the His ligand away from the haem pocket to its “native” position. It may be that in the presence of 8 M urea this tertiary structure change is slowed down such that the anodic peak at +0.110 V (Fig. 1a) may correspond to a B1-like species which possesses the same axial ligation pattern as the native protein but differs with respect to the arrangement of the His-carrying peptide segment.
In contrast, the slight decrease in the E°′ value of wave I upon increasing the urea concentration from 0 to 5 M points to moderate conformational perturbations beyond the level of a ligand exchange and a major rearrangement of the tertiary structure [43, 44]. Previous interpretation stressing a Met80 → Lys substitution [15] can be ruled out since K72AK73AK79A lacking all potential Lys ligands reveals essentially the same electrochemical behaviour as ycc. Instead, the increasing negative shift of the reduction potential, which is the result of large and opposing contributions of \( \Updelta S^{\circ \prime}_{\text{rc}} \) and \( \Updelta H^{\circ \prime}_{\text{rc}} \) (Table 1) [45], may be due to increasing exposure of the redox centre to the solvent [43, 44].
Thermodynamics of the interfacial redox process
The variation of the peak currents with urea concentration allows determination of the transition from the B1 to the U[6cLS] state, revealing a midpoint of 3.9 and 3.5 M for the immobilised ycc and K72AK73AK79A, respectively (Fig. 2). Conversely, the respective transitions in solution are found at 3.5 and 3.2 M for ycc and K72AK73AK79A, respectively [44]. Thus, we conclude that Lys72, Lys73 and Lys79 stabilise the protein structure in the adsorbed state. This can be rationalised in terms of the involvement of these residues in the electrostatic binding to the SAM surface [46], thereby reducing the mobility of these Lys residues and the respective peptide segments.
The E°′ values of U[6cLS] (wave II) obtained for ycc and K72AK73AK79A are nearly the same at the different urea concentrations (Table 1). This suggests that U[6cLS], once it is formed, does not undergo conformational changes at increasing concentration of the unfolding agent. This is consistent with the previous conclusion that U[6cLS] constitutes a metastable trap along the unfolding pathway [17]. For the U[6cLS] state of ycc and K72AK73AK79A, the E°′ values are approximately 0.4 V more negative than those of the corresponding native B1 state. A very similar negative shift has been determined for the state B2[6cLS], which exhibits the same haem pocket structure as B1 [38], and for cytochrome c in urea-containing solution [36, 41, 43, 44] and is consistent with a change of the ligand from S-Met to N-His [3, 35, 36, 47]. The difference in the reduction potential, ΔE°′, between these two states is 0.031 V more positive for ycc than for K72AK73AK79A (Table 2). The ΔE°′ values for ycc and K72AK73AK79A in solution, however, differ by only 0.014 V [44].
The B1 state displays negative enthalpy and entropy values for reduction [25, 26, 33–35, 44, 48–51]. The enthalpy term \( \Updelta H^{\circ \prime}_{\text{rc}} \) is considered to be the most important for the high reduction potentials of cytochromes c. It is mainly the result of stabilising the ferrous form owing to ligand binding interactions, the hydrophobicity in the haem pocket, and the limited solvent accessibility [2, 3, 35]. The electrostatic interactions of the charge of the redox centre with buried and surfaces charges, polar groups of the protein, solvent dipoles and the ionic environment are additional important factors constituting the enthalpic term [2, 3, 26, 35, 52–55]. Conversely, the entropic term \( \Updelta S^{\circ \prime}_{\text{rc}} \) disfavours protein reduction, yielding a negative contribution to the E°′ values. Here, solvent reorganisation effects, charge redistribution and changes in protein flexibility associated with the haem reduction play the major role in determining \( \Updelta S^{\circ \prime}_{\text{rc}} \) [25, 26, 35, 56–62]. At increasing urea concentration \( \Updelta S^{\circ \prime}_{\text{rc}} \) and \( \Updelta H^{\circ \prime}_{\text{rc}} \) of both proteins shift towards negative values (Table 1). This effect is larger for ycc than for K72AK73AK79A.
To sort out the underlying enthalpic and entropic contributions, we express the urea-induced changes of the reduction potential E°′ according to [45]
with
where \( E^{\circ \prime}_{\text{B1}} \) and \( E^{\circ \prime}_{\text{urea}} \) are the reduction potentials of the B1 state of the protein in its native form (without urea) and in the presence of different urea concentrations, respectively, \( \Updelta \Updelta H^{\circ \prime}_{\text{rc,solv}} \) is the change in \( \Updelta H^{\circ \prime}_{\text{rc}} \) due to solvent organisation induced by the unfolding of the protein, whereas \( \Updelta \Updelta H^{\circ \prime}_{\text{rc,int}} \) refers to internal protein structural changes such as the opening of the haem crevice. \( \Updelta \Updelta S^{\circ \prime}_{\text{rc,solv}} \) is the entropic contribution resulting from changes in the solvent organisation. Equation 3 assumes that the contribution of the intramolecular reaction entropy \( \Updelta S^{\circ \prime}_{\text{rc,int}} \) remains largely unchanged with increasing urea concentration such that \( \Updelta \Updelta S^{\circ \prime}_{\text{rc,int}} \) is zero [63–65]. Furthermore, the enthalpic and entropic contributions to the solvent reorganisation are assumed to compensate each other such that
and Eq. 2 simplifies to
Enthalpy/entropy compensation phenomena are well known for quite different processes of biopolymers [45, 51, 63, 66–69] and have been discussed on the basis of various models [64, 65, 70–74]. Such a compensation also refers to the present case of cytochrome c unfolding as shown in Fig. 4. Here, the standard enthalpy change \( (\Updelta H^{\circ \prime}_{\text{rc}} ) \) is plotted against the corresponding entropic terms \( (T\Updelta S^{\circ \prime}_{\text{rc}} ) \) at 293 K for adsorbed ycc and K72AK73AK79A at different urea concentrations.
Enthalpy–entropy compensation plots at different urea concentrations for the reduction thermodynamics of the B1 states of ycc (filled circles) and K72AK73AK79A (open circles) immobilised on an MUA/MU-modified gold electrode at 293 K. The solution contained 5 mM sodium perchlorate and 5 mM phosphate buffer (pH 7). The straight lines represent the least-squares fits to the data, yielding a slope of 0.94 and 0.51 for ycc and K72AK73AK79A, respectively
The plots in Fig. 4 clearly demonstrate that, within the error margins, enthalpy and entropy changes are linearly correlated, indicative of compensation effects. For ycc, the slope was determined to be 0.94 and thus close to 1, indicating a nearly fully compensatory effect. Conversely, a much smaller value of 0.51 was obtained for the immobilised K72AK73AK79A (Fig. 4) whereas K72AK73AK79A in solution exhibits an almost exact compensatory behaviour [45]. Evidently, the intramolecular contribution to the reaction enthalpy increases for the electrostatically bound mutant, underpinning the role of the three Lys residues in stabilising the protein structure of the adsorbed protein as discussed above. This conclusion is consistent with the higher \( \Updelta \Updelta H^{\circ \prime}_{\text{rc,int}} \) values for K72AK73AK79A as compared with ycc (Table 3).
For the U[6cLS] state of both ycc and K72AK73AK79A, \( \Updelta S^{\circ \prime}_{\text{rc}} \) and \( \Updelta H^{\circ \prime}_{\text{rc}} \) are almost unaffected by the urea concentration, implying that once the Met → His ligand exchange and the coupled peptide rearrangements have taken place, no further structural changes are induced by the denaturant (Table 1). The view that the local structural change in the haem pocket including the ligand exchange is the main determinant for the large negative shift of the reduction potential is further supported by the quite similar reduction potentials of the state B2, in which the same haem pocket structural change is induced by electrostatic interactions rather than denaturants [38].
Most remarkable are the positive values for \( \Updelta H^{\circ \prime}_{\text{rc}} \) and \( \Updelta S^{\circ \prime}_{\text{rc}} \), which have not been observed for other states of cytochrome c [25, 26, 33–35, 44, 48–51].
Electron transfer kinetics of the adsorbed cytochrome c
The formal heterogeneous electron transfer rate constants were determined using Laviron’s method [75] (Table 4). In the absence of urea, the electron transfer rate constant for ycc is distinctly higher than that previously determined for that protein on an MUA/MU-coated silver electrode [32]. This discrepancy can be readily attributed to the higher electric field strength at the silver–SAM interface [76]. Thus, reorientation of the immobilised protein is slowed down and becomes the rate-limiting step of the interfacial redox process, unlike for the gold–SAM interface, where heterogeneous electron transfer is controlled by electron tunnelling. In fact, the rate constant of ycc at the gold–SAM interface is very similar to that of horse heart cytochrome c on an MUA-coated silver electrode where electron tunnelling is the rate-limiting step as well [77]. The respective rate constant for K72AK73AK79A is lower by a factor of 6.6 compared with ycc. This finding suggests that the three Lys residues are critical for prealignment of the protein prior to electron transfer.
Upon increasing the urea concentration, the rate constants steadily decrease, albeit more strongly for the mutant. The ratio of the rate constants for ycc and K72AK73AK79A increases from 6.6 (0 M urea) to 18.6 (4 M urea). On the other hand, the difference in the activation enthalpies remains largely unchanged (approximately 3 kJ mol−1) in this range (Table 4), pointing to significant and protein-specific entropic contributions of the reorganisation energy and/or changes in the tunnelling effects.
The heterogeneous electron transfer rate constant of U[6cLS] is substantially lower than that of state B1 even at high urea concentrations. This finding implies that the protein structural change associated with the B1 → U[6cLS] transition leads to a configuration that is highly unfavourable for the interfacial electron transfer. Again, this effect is more severe for the triple mutant than for the wild-type protein.
Conclusions
The combined use of electrochemical (CV) and spectroscopic (SERR spectroscopy) techniques has allowed characterisation of the interfacial redox process of immobilised cytochrome c in the presence of the denaturant urea. Increasing urea concentration leads to the formation of the conformational state U[6cLS] in which the Met80 ligand of the haem iron is substituted by a His (His33 or His26). The structural change associated with this transition causes an approximately 400 mV shift of the reduction potential to negative values. The potential is largely independent of the urea concentration, whereas the reduction potential of the native state B1 slightly shifts to negative values with increasing urea concentration. This effect is attributed to moderate urea-induced protein structural changes, including a gradually increasing opening of the haem crevice. Both the wild-type protein and the K72AK73AK79A variant reveal qualitatively the same response with increasing urea concentration such that the coordination of the haem iron by a Lys residue can safely be ruled out. However, the transition between states B1 and U[6cLS] occurs at a slightly lower urea concentration in the triple mutant than in the wild-type protein, pointing to the involvement of Lys72, Lys73 and Lys79 in stabilising the structure of the adsorbed protein. Furthermore, these Lys residues appear to be important for controlling the interfacial redox properties inasmuch as they stabilise the ferric form and also favour the appropriate prealignment of the protein for the heterogeneous electron transfer.
Abbreviations
- 6cLS:
-
Six-coordinated low spin
- CV:
-
Cyclic voltammetry
- MU:
-
11-Mercapto-1-undecanol
- MUA:
-
11-Mercapto-1-undecanoic acid
- SAM:
-
Self-assembled monolayer
- SCE:
-
Saturated calomel electrode
- SERR:
-
Surface-enhanced resonance Raman
- ycc:
-
Recombinant non-trimethylated Saccharomyces cerevisiae iso-1-cytochrome c
References
Messerschmidt A, Huber R, Poulos T, Wieghardt K (eds) (2001) Handbook of metalloproteins, vol 1. Wiley, Chichester
Scott RA, Mauk GA (eds) (1996) Cytochrome c: a multidisciplinary approach. University Science Books, Sausalito
Moore GR, Pettigrew GW (1990) Cytochromes c: evolutionary, structural, and physicochemical aspects. Springer, Berlin
Kagan VE, Bayr HA, Belikova NA, Kapralov O, Tyurina YY, Jang J, Stoyanovsky DA, Wipf P, Kochanek PM, Greenberger JS, Pitt B, Shvedova AA, Borisenko G (2009) Free Radic Biol Med 46:1439–1453
Bond AM (1994) Inorg Chim Acta 226:293–340
Hill HAO, Hunt NI (1993) Methods Enzymol 227:501–522
Armstrong FA (1990) Struct Bonding 72:137–222
Armstrong FA, Hill HAO, Walton NJ (1986) Q Rev Biophys 18:261–322
Murgida DH, Hildebrandt P (2008) Chem Soc Rev 37:937–945
Bryngelson JD, Onuchic JN, Socci ND, Wolynes PG (1995) Proteins 21:167–195
Dobson CM, Sali A, Karplus M (1998) Angew Chem Int Ed 37:868–893
Dobson CM, Karplus M (1999) Curr Opin Struct Biol 9:92–101
Yeh SR, Rousseau DL (1998) Nat Struct Biol 5:222–228
Xu Y, Mayne L, Englander SW (1998) Nat Struct Biol 5:774–778
Russell BS, Melenkivitz R, Bren KL (2000) Proc Natl Acad Sci USA 97:8312–8317
Myer YP, MacDonald LH, Verma BC, Pande A (1980) Biochemistry 19:199–207
Yeh SR, Han SW, Rousseau DL (1998) Acc Chem Res 31:727–736
Zhou J, Zheng J, Jiang S (2004) J Phys Chem B 108:17418–17424
Xu J, Bowden EF (2006) J Am Chem Soc 128:6813–6822
Battistuzzi G, Borsari M, De Rienzo F, Di Rocco G, Ranieri A, Sola M (2007) Biochemistry 46:1694–1702
Rosell FI, Ferrer JC, Mauk AG (1998) J Am Chem Soc 120:11234–11245
Pollock WBR, Rosell FI, Twitchett MB, Dumont ME, Mauk AG (1998) Biochemistry 37:6124–6131
Cutler RJ, Pielak GJ, Mauk AG, Smith M (1987) Protein Eng 1:95–99
Liang N, Mauk AG, Pielak GJ, Johnson JA, Smith M, Hoffmann B (1988) Science 240:311–313
Battistuzzi G, Borsari M, Sola M, Francia F (1997) Biochemistry 36:16247–16258
Battistuzzi G, Borsari M, Bortolotti CA, Di Rocco G, Ranieri A, Sola M (2007) J Phys Chem B 111:10281–10287
Yee EL, Cave RJ, Guyer KL, Tyma PD, Weaver MJ (1979) J Am Chem Soc 101:1131–1137
Yee EL, Weaver MJ (1980) Inorg Chem 19:1077–1079
Song S, Clark RA, Bowden EF, Tarlov MJ (1993) J Phys Chem 97:6564–6572
Weaver MJ (1979) J Phys Chem 13:1748–1757
Bonifacio A, Millo D, Gooijer C, Boegschoten R, van der Zwan G (2004) Anal Chem 76:1529–1531
Feng JJ, Murgida DH, Utesch T, Mroginski MA, Hildebrandt P, Weidinger I (2008) J Phys Chem B 112:15202–15211
Millo D, Bonifacio A, Ranieri A, Borsari M, Gooijer C, van der Zwan G (2007) Langmuir 23:4340–4345
Millo D, Bonifacio A, Ranieri A, Borsari M, Gooijer C, van der Zwan G (2007) Langmuir 23:9898–9904
Battistuzzi G, Borsari M, Sola M (2001) Eur J Inorg Chem 2989–3004
Fedurco M, Augustynski J, Indiani C, Smulevich G, Antalík M, Bánó M, Sedlák E, Galscock MC, Dawson JH (2005) J Am Chem Soc 127:7638–7646
Oellerich S, Wackerbarth H, Hildebrandt P (2002) J Phys Chem B 106:6566–6580
Wackerbarth H, Hildebrandt P (2003) ChemPhysChem 4:714–724
Murgida DH, Hildebrandt P (2001) J Phys Chem B 105:1578–1586
Hildebrandt P (1991) J Mol Struct 242:379–395
Fedurco M, Augustynski J, Indiani C, Smulevich G, Antalík M, Bánó M, Sedlák E, Galscock MC, Dawson JH (2004) Biochim Biophys Acta 1703:31–41
Bhuyan AK, Udgaonkar JB (2001) J Mol Biol 312:1135–1160
Pilard R, Haladjian J, Bianco P, Serre P-A, Brabec V (1983) Biophys Chem 17:131–137
Monari S, Ranieri A, Di Rocco G, van der Zwan G, Peressini S, Tavagnacco C, Millo D, Borsari M (2009) J Appl Electrochem 39:2181–2190
Battistuzzi G, Borsari M, Di Rocco G, Ranieri A, Sola M (2004) J Biol Inorg Chem 9:23–26
Paggi DA, Martín DF, Kranich A, Hildebrandt P, Martí M, Murgida DH (2009) Electrochim Acta 54:4963–4970
Battistuzzi G, Borsari M, Cowan JA, Ranieri A, Sola M (2002) J Am Chem Soc 124:5315–5324
Bortolotti CA, Battistuzzi G, Borsari M, Facci P, Ranieri A, Sola M (2006) J Am Chem Soc 128:5444–5451
Grealis C, Magner E (2003) Langmuir 19:1282–1286
Battistuzzi G, Borsari M, Canters GW, De Waal E, Loschi L, Warmerdam G, Sola M (2001) Biochemistry 40:6707–6712
Bertrand P, Mbarki O, Asso M, Blanchard L, Guerlesquin F, Tegoni M (1995) Biochemistry 34:11071–11079
Gunner MR, Alexov E, Torres E, Lipovaca S (1997) J Biol Inorg Chem 2:126–134
Mauk AG, Moore GR (1997) J Biol Inorg Chem 2:119–125
Tezcan FA, Winkler JR, Gray HB (1998) J Am Chem Soc 120:13383–13388
Warshel A, Papazyan A, Muegge I (1997) J Biol Inorg Chem 2:143–152
Banci L, Bertini I, Rosato A, Varani G (1999) J Biol Inorg Chem 4:824–837
Battistuzzi G, Loschi L, Borsari M, Sola M (1999) J Biol Inorg Chem 4:601–607
Borsari M, Bellei M, Tavagnacco C, Peressini S, Millo D, Costa G (2003) Inorg Chim Acta 349:182–188
Banci L, Bertini I, Gray HB, Luchinat C, Redding T, Rosato A, Turano P (1997) Biochemistry 36:9867–9877
Furlan S, La Penna G, Banci L, Mealli C (2007) J Phys Chem B 111:1157–1164
La Penna G, Furlan S, Banci L (2007) J Biol Inorg Chem 12:180–193
Mao J, Hauser K, Gunner MR (2003) Biochemistry 42:9829–9840
Liu L, Guo Q-X (2001) Chem Rev 101:673–695
Grünwald E, Steel C (1995) J Am Chem Soc 117:5687–5692
Grünwald E (1986) J Am Chem Soc 108:5726–5731
Searle MS, Weatwell MS, Williams DH (1995) J Chem Soc Perkin Trans 2 141–151
Rekharsky M, Inoue Y (2000) J Am Chem Soc 122:4418–4435
Liu L, Yang C, Guo Q-X (2000) Biophys Chem 84:239–251
Strazewski P (2002) J Am Chem Soc 124:3546–3554
Blokzijl W, Engberts JBNF (1993) Angew Chem Int Ed Engl 32:1545–1579
Lumry R, Rajender S (1970) Biopolymers 9:1125–1227
Krug RR, Hunter WG, Grieger RA (1976) J Phys Chem 80:2335–2351
Ben-Naim A (1975) Biopolymers 14:1337–1355
Lee B, Graziano G (1996) J Am Chem Soc 118:5163–5168
Laviron E (1979) J Electroanal Chem 101:19–28
Millo D, Ranieri A, Gross P, Ly HK, Borsari M, Hildebrandt P, Wuite GJL, Gooijer C, van der Zwan G (2009) J Phys Chem C 113:2861–2866
Murgida DH, Hildebrandt P (2001) J Am Chem Soc 123:4062–4068
Acknowledgments
We gratefully acknowledge Murat Sezer for supporting the SERR spectroscopy measurements in Berlin. This work was performed with financial support from MIUR (COFIN 2007, protocollo 20079Y9578_002, Bioelettrochimica: trasferimento di carica in sistemi di rilevanza biologica), the University of Modena and Reggio Emilia, the Deutsche Forschungsgemeinschaft (Sfb498), the Alexander von Humboldt Foundation (D.M.) and the European Community Access to Research Infrastructures Action of The Improving Human Potential (contract no. HPRI-CT-1999-00064) (A.R.).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Monari, S., Millo, D., Ranieri, A. et al. The impact of urea-induced unfolding on the redox process of immobilised cytochrome c . J Biol Inorg Chem 15, 1233–1242 (2010). https://doi.org/10.1007/s00775-010-0681-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-010-0681-7