Abstract
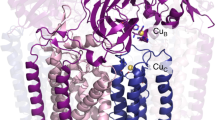
Nature’s primary methane-oxidizing enzyme, the membrane-bound particulate methane monooxygenase (pMMO), catalyses the oxidation of methane to methanol. Copper is required for pMMO activity, and decades of structural and spectroscopic studies have sought to identify the active site among three candidates: the CuB, CuC and CuD sites. Challenges associated with the isolation of active pMMO hindered identification of its catalytic centre; however, we have recently shown that reconstituting pMMO into native lipid nanodiscs stabilizes its structure and restores its activity. Here, such active samples were incubated with 2,2,2-trifluoroethanol, a product analogue that serves as a readily visualized active-site probe. Interactions between 2,2,2-trifluoroethanol and the CuD site were observed with pulsed electron nuclear double resonance spectroscopy and cryoelectron microscopy, implicating CuD and the surrounding hydrophobic pocket as the likely site of methane oxidation. Use of these orthogonal techniques on parallel samples is a powerful approach that can circumvent difficulties in interpreting metalloenzyme cryoelectron microscopy maps.






Similar content being viewed by others
Data availability
The models of pMMO in native lipid nanodiscs (7S4H), for KCN-treated (8SR5), KCN-treated and copper-reloaded (8SR4), 20× TFE (8OYI), cross-linked 20× TFE (8SQW), 20× TFB (8SR2) and cross-linked 20× TFB (8SR1) samples are available in the Protein Data Bank (PDB). Corresponding cryoEM maps are available at the Electron Microscopy Data Bank (EMDB). Other data are available in the main text, Supplementary Information or from the authors on reasonable request. Source Data are provided with this paper.
References
Arndtsen, B. A., Bergman, R. G., Mobley, T. A. & Peterson, T. H. Selective intermolecular carbon–hydrogen bond activation by synthetic metal complexes in homogeneous solution. Acc. Chem. Res. 28, 154–162 (1995).
Blanksby, S. J. & Ellison, G. B. Bond dissociation energies of organic molecules. Acc. Chem. Res. 36, 255–263 (2003).
Tang, P., Zhu, Q. J., Wu, Z. X. & Ma, D. Methane activation: the past and future. Energy Environ. Sci. 7, 2580–2591 (2014).
Haynes, C. A. & Gonzalez, R. Rethinking biological activation of methane and conversion to liquid fuels. Nat. Chem. Biol. 10, 331–339 (2014).
Ravi, M., Ranocchiari, M. & van Bokhoven, J. A. The direct catalytic oxidation of methane to methanol-a critical assessment. Angew. Chem. Int. Ed. 56, 16464–16483 (2017).
Dummer, N. F. et al. Methane oxidation to methanol. Chem. Rev. 123, 6359–6411 (2022).
Lawton, T. J. & Rosenzweig, A. C. Methane-oxidizing enzymes: an upstream problem in biological gas-to-liquids conversion. J. Am. Chem. Soc. 138, 9327–9340 (2016).
Hanson, R. S. & Hanson, T. E. Methanotrophic bacteria. Microbiol. Rev. 60, 439–471 (1996).
Ross, M. O. & Rosenzweig, A. C. A tale of two methane monooxygenases. J. Biol. Inorg. Chem. 22, 307–319 (2017).
Banerjee, R., Jones, J. C. & Lipscomb, J. D. Soluble methane monooxygenase. Annu. Rev. Biochem. 88, 409–431 (2019).
Koo, C. W. & Rosenzweig, A. C. Biochemistry of aerobic biological methane oxidation. Chem. Soc. Rev. 50, 3424–3436 (2021).
Prior, S. D. & Dalton, H. The effect of copper ions on membrane content and methane monooxygenase activity in methanol-grown cells of Methylococcus capsulatus (Bath). J. Gen. Microbiol. 131, 155–163 (1985).
Ross, M. O. et al. Particulate methane monooxygenase contains only mononuclear copper centers. Science 364, 566–570 (2019).
Jodts, R. J. et al. Coordination of the copper centers in particulate methane monooxygenase: comparison between methanotrophs and characterization of the Cuc site by EPR and ENDOR spectroscopies. J. Am. Chem. Soc. 143, 15358–15368 (2021).
Op den Camp, H. J. et al. Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ. Microbiol. Rep. 1, 293–306 (2009).
Kruse, T., Ratnadevi, C. M., Erikstad, H. A. & Birkeland, N. K. Complete genome sequence analysis of the thermoacidophilic verrucomicrobial methanotroph ‘Candidatus Methylacidiphilum kamchatkense’ strain Kam1 and comparison with its closest relatives. BMC Genomics 20, 642 (2019).
Hakemian, A. S. et al. The metal centers of particulate methane monooxygenase from Methylosinus trichosporium OB3b. Biochemistry 47, 6793–6801 (2008).
Sirajuddin, S. et al. Effects of zinc on particulate methane monooxygenase activity and structure. J. Biol. Chem. 289, 21782–21794 (2014).
Lieberman, R. L. & Rosenzweig, A. C. Crystal structure of a membrane-bound metalloenzyme that catalyses the biological oxidation of methane. Nature 434, 177–182 (2005).
Smith, S. M. et al. Crystal structure and characterization of particulate methane monooxygenase from Methylocystis species strain M. Biochemistry 50, 10231–10240 (2011).
Koo, C. W., Tucci, F. J., He, Y. & Rosenzweig, A. C. Recovery of particulate methane monooxygenase activity in a lipid bilayer. Science 375, 1287–1291 (2022).
Griese, J. J. & Högbom, M. Location-specific quantification of protein-bound metal ions by X-ray anomalous dispersion: Q-XAD. Acta Crystallogr. D 75, 764–771 (2019).
Ro, S. Y. et al. From micelles to bicelles: effect of the membrane on particulate methane monooxygenase activity. J. Biol. Chem. 293, 10457–10465 (2018).
Peisach, J. & Blumberg, W. E. Structural implications derived from the analysis of electron paramagnetic resonance spectra of natural and artificial copper proteins. Arch. Biochem. Biophys. 165, 691–708 (1974).
Pogni, R., Baratto, M. C., Diaz, A. & Basosi, R. EPR characterization of mono(thiosemicarbazones) copper(II) complexes. Note II. J. Inorg. Biochem. 79, 333–337 (2000).
Yu, S. S.-F. et al. Production of high-quality particulate methane monooxygenase in high yields from Methylococcus capsulatus (Bath) with a hollow-fiber membrane bioreactor. J. Bacteriol. 185, 5915–5924 (2003).
Davydov, R., Valentine, A. M., Komar-Panicucci, S., Hoffman, B. M. & Lippard, S. J. An EPR study of the dinuclear iron site in the soluble methane monooxygenase from Methylococcus capsulatus (Bath) reduced by one electron at 77 K: the effects of component interactions and the binding of small molecules to the diiron(III) center. Biochemistry 38, 4188–4197 (1999).
Smoukov, S. K. et al. Product binding to the diiron(III) and mixed-valence diiron centers of methane monooxygenase hydroxylase studied by 1,2H and 19F ENDOR spectroscopy. J. Am. Chem. Soc. 124, 2657–2663 (2002).
Smith, D. D. S. & Dalton, H. Solubilisation of methane monooxygenase from Methylococcus capsulatus (Bath). Eur. J. Biochem. 182, 667–671 (1989).
Furuto, T., Takeguchi, M. & Okura, I. Semicontinuous methanol biosynthesis by Methylosinus trichosporium OB3b. J. Mol. Catal. A: Chem. 144, 257–261 (1999).
Wieczorek, A. S., Drake, H. L. & Kolb, S. Organic acids and ethanol inhibit the oxidation of methane by mire methanotrophs. FEMS Microbiol. Ecol. 77, 28–39 (2011).
Mims, W. B. Pulsed endor experiments. Proc. Roy. Soc. Lond. 283, 452–457 (1965).
Grupp, A. & Mehring, M. in Modern Pulsed and Continuous-Wave Electron Spin Resonance (eds Kevan, L. & Bowman, M. K.) 195–229 (Wiley, 1990).
Doan, P. E., Lees, N. S., Shanmugam, M. & Hoffman, B. M. Simulating suppression effects in pulsed ENDOR, and the ‘hole in the middle’ of Mims and Davies ENDOR spectra. Appl. Magn. Reson. 37, 763–779 (2010).
Houseman, A. L. P. et al. 14,15N, 13C, 57Fe, and 1,2H Q-band ENDOR study of iron–sulfur proteins with clusters that have endogenous sulfur ligands. Biochemistry 31, 2073–2080 (1992).
Erlandsen, H., Flatmark, T., Stevens, R. C. & Hough, E. Crystallographic analysis of the human phenylalanine hydroxylase catalytic domain with bound catechol inhibitors at 2.0 A resolution. Biochemistry 37, 15638–15646 (1998).
Goldfeder, M., Kanteev, M., Isaschar-Ovdat, S., Adir, N. & Fishman, A. Determination of tyrosinase substrate-binding modes reveals mechanistic differences between type-3 copper proteins. Nat. Commun. 5, 4505 (2014).
Burrows, K. J., Cornish, A., Scott, D. & Higgins, I. J. Substrate specificities of the soluble and particulate methane monooxygenases of Methylosinus trichosporium OB3b. J. Gen. Microbiol. 130, 327–3333 (1984).
Carosati, E., Sciabola, S. & Cruciani, G. Hydrogen bonding interactions of covalently bonded fluorine atoms: from crystallographic data to a new angular function in the GRID force field. J. Med. Chem. 47, 5114–5125 (2004).
Bartesaghi, A., Matthies, D., Banerjee, S., Merk, A. & Subramaniam, S. Structure of β-galactosidase at 3.2-Å resolution obtained by cryo-electron microscopy. Proc. Natl Acad. Sci. USA 111, 11709–11714 (2014).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Fang, J. S. & Findlay, R. H. The use of a classic lipid extraction method for simultaneous recovery of organic pollutants and microbial lipids from sediments. J. Microbiol. Methods 27, 63–71 (1996).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Ro, S. Y. et al. Native top-down mass spectrometry provides insights into the copper centers of membrane-bound methane monooxygenase. Nat. Commun. 10, 2675 (2019).
Davoust, C. E., Doan, P. E. & Hoffman, B. M. Q-band pulsed electron spin-echo spectrometer and its application to ENDOR and ESEEM. J. Magn. Reson. 119, 38–44 (1996).
Doan, P. E. Combining steady-state and dynamic methods for determining absolute signs of hyperfine interactions: pulsed ENDOR saturation and recovery (PESTRE). J. Magn. Reson. 208, 76–86 (2011).
Schweiger, A. & Jeschke, G. Principles of Pulse Electron Paramagnetic Resonance (Oxford Univ. Press, 2001).
Hoffman, B. M., Martinsen, J. & Venters, R.A. General theory of polycrystalline ENDOR patterns. g and hyperfine tensors of arbitrary symmetry and relative orientation. J. Magn. Reson. 59, 110–123 (1984).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Cryst. D66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: atructure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Farabella, I. et al. TEMPy: a Python library for assessment of three-dimensional electron microscopy density fits. J. Appl. Crystallogr. 48, 1314–1323 (2015).
Acknowledgements
This work was supported by NIH grants R35GM118035 (A.C.R.), R01GM111097 (B.M.H.), T32GM105538 (F.J.T.), F31ES034283 (F.J.T.), and T32GM008382 (R.J.J.), as well as the NSF MCB-1908587 (B.M.H) This work used resources of the Northwestern Structural Biology Facility and the Northwestern Keck Biophysics Facility, which are supported by the NCI CCSG P30 CA060553 grant awarded to the Robert H. Lurie Comprehensive Cancer Center. Metal analysis was performed at the Northwestern University Quantitative Bio-element Imaging Center generously supported by NASA Ames Research Center NNA06CB93G. Some of this work was performed at the National Center for CryoEM Access and Training (NCCAT) and the Simons Electron Microscopy Center located at the New York Structural Biology Center, supported by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy program (U24 GM129539) and by grants from the Simons Foundation (SF349247) and NY State Assembly. We thank L. Mazhar for experimental assistance, Y. He for guidance in cryoEM, P. Doan for helpful discussions, J. Pattie for computer-related support and M. Ho for assistance during revisions.
Author information
Authors and Affiliations
Contributions
F.J.T., R.J.J., B.M.H. and A.C.R. conceptualized the work and designed experiments. F.J.T. and R.J.J. performed experiments, analysed data and prepared figures. F.J.T., R.J.J., B.M.H. and A.C.R. contributed to writing and editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Kallol Ray, Uhn-Soo Cho and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–24 and Table 1.
Source data
Source Data Fig. 2
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tucci, F.J., Jodts, R.J., Hoffman, B.M. et al. Product analogue binding identifies the copper active site of particulate methane monooxygenase. Nat Catal 6, 1194–1204 (2023). https://doi.org/10.1038/s41929-023-01051-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-023-01051-x
- Springer Nature Limited