Abstract
Knowledge of the molecular mechanisms of bone formation has been advanced by novel findings related to genetic control. Odd-skipped related 1 (Osr1) is known to play important roles in embryonic, heart, and urogenital development. To elucidate the in vivo function of Osr1 in bone formation, we generated transgenic mice overexpressing full-length Osr1 under control of its 2.8-kb promoter, which were smaller than their wild-type littermates. Notably, abnormalities in the skull of Osr1 transgenic mice were revealed by analysis of X-ray, skeletal preparation, and morphological findings, including round skull and cranial dysraphism. Furthermore, primary calvarial cells obtained from these mice showed increased proliferation and expression of chondrocyte markers, while expression of osteoblast markers was decreased. BMP2 reduced Osr1 expression and Osr1 knockdown by siRNA-induced alkaline phosphatase and osteocalcin expression in mesenchymal and osteoblastic cells. Together, our results suggest that Osr1 plays a coordinating role in appropriate skull closure and cranial bone formation by negative regulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The odd-skipped gene, initially identified as a Drosophila segmentation gene, contains four DNA-binding C2H2-type zinc fingers in the C-terminal half of the molecule as a pair-rule transcription factor [1]. The mammalian homologs Odd-skipped related 1 (Osr1) and Osr2 have been cloned from both mice and humans [2, 3]. Mouse Osr1 has a 65% homology with Osr2, whereas their tissue distribution differs. Human Osr1 has been detected in fetal lung, adult colon, small intestine, prostate, and testis tissues [3], whereas Osr2 is specifically expressed in limb, tooth, and kidney tissues [4].
Osr1 was shown to function in heart morphogenesis and urogenital development by targeted null mutation [5]. In that study, null mutant embryos did not succeed in forming an atrial septum, and exhibited dilated atria with hypoplastic venous valves and blood backflow from the heart into systemic veins. Expression of Osr1 is restricted to the central dorsal domain of the atrial myocardium during normal heart development. Furthermore, Osr1 null mutant embryos exhibit defects of adrenal glands, metanephric kidneys, gonads, and pericardium formation, while the key regulators of early intermediate mesoderm development are down-regulated. In addition, it was shown that nephrogenic mesenchyme underwent massive apoptosis, which caused a disruption of nephric duct elongation and failure of metanephric induction in Osr1 null mutant embryos [5].
We previously reported that Osr2 regulates osteoblast function, as dominant-negative Osr2 transgenic mice exhibited decreased osteoblast proliferation, and delayed mineralization in calvarial and tibial bone tissues [6]. By cloning the 5′-flanking 4.8-kb DNA region of the mouse Osr1 gene, our findings also revealed the regulation mechanism, and showed that Runx2 and Ikzf1 suppress Osr1 promoter activity [7]. Drosophila runt, hedgehog, and wingless belong to the group of segmentation genes, and their mammalian counterparts Runx2, Hh, and Wnt function in bone formation [8]. The Drosophila odd-skipped gene is involved in segmentation and it is expected that mammalian Osr1 functions in bone formation, though direct proof of the latter remains to be elucidated.
In the present study, we generated transgenic mice with full-length Osr1 under control of its own promoter. Round skull and cranial dysraphism were observed in these transgenic mice. Our findings are the first to show that Osr1 functions in adequate skull and cranial bone formation.
Materials and methods
Cell cultures and reagents
The mesenchymal cell line C3H10T1/2 (RCB0247), myoblastic cell line C2C12 (RCB0987), and osteoblastic cell line MC3T3-E1 (RCB1126) were purchased from Riken Bioresource Center (Tsukuba, Ibaragi, Japan). The HEK293A cell line was obtained from Life Technologies (Carlsbad, CA, USA). C3H10T1/2 and MC3T3-E1 cells were maintained in α-modified Eagle’s medium (α-MEM; Sigma-Aldrich, Saint Louis, MO, USA) supplemented with 10% fetal bovine serum (FBS; Life Technologies). HEK293A cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM, Sigma-Aldrich) supplemented with 10% FBS. Mouse Osr1 was amplified by polymerase chain reaction (PCR) using the mouse embryonic cDNA library and cloned into a pcDNA3 plasmid vector (Life Technologies). The sequences of these inserts were confirmed by DNA sequence analysis. Alkaline phosphatase (ALP) promoter-luciferase (ALP-Luc) and osteocalcin promoter (OG2)-luciferase (OCN-Luc) vectors were used as described in our previous report [6].
Generation of transgenic mice
Osr1 transgenic mice have a fusion gene composed of a 2.8-kb fragment of the mouse Osr1 promoter [7] fused to the first intron of the rabbit beta-globin gene, Osr1 full-length cDNA, and 3′ untranslated and polyA signals of SV40. Transgenic founders were generated by pronuclear injection into mouse strain BDF1 (DBA2/C57BL6) using standard techniques [9]. The genotype of the transgenic animals was determined by PCR. The primer set for the mouse Osr1 gene was 5′-CCTGGACGTGACCAAGCTAT-3′ (forward) and 5′-TGTAGCGTCTTGTGGACAGC-3′ (reverse), and was not amplified Osr2 mRNA. All animal experiments were performed using protocols approved by the Osaka University Graduate School of Dentistry animal care and use committee (Permit number: Doushi-19-020-0).
Skeletal analysis
For skeletal preparations, newborn mice were dissected and fixed overnight in 95% ethanol. Staining with Alcian blue 8GX and Alizarin red S (Sigma-Aldrich) was performed using standard protocols [10]. For histological analysis, the mice were killed as newborns and calvaria were then obtained and fixed in 10% formalin and embedded in paraffin. Thereafter, 5-µm-thick sections were stained with hematoxylin and eosin (H&E, Sigma-Aldrich) using procedures described previously [11]. Gross appearance was imaged using 3-week-old male mice. Skeletons were analyzed with X-ray apparatus (Softex, Kanagawa, Japan) using 3-week-old male mice.
In vitro differentiation of osteoblasts
Primary cells were isolated from the calvarial bones of 1- to 3-day-old newborn mice, as previously described [12], then cultured in α-MEM with 10% FBS or mineralization medium (α-MEM with 10% FBS containing 50 μg/ml ascorbic acid, 10 mM β-glycerophosphate, and 100 ng/ml rhBMP2; R&D Systems, Minneapolis, MN, USA). Cells were maintained for 7 days to determine ALP and for 21 days to detect mineralized matrix (Alizarin red S), with standard protocols utilized [6]. Cell proliferation was determined using a CellTiter 96 AQueous One Solution Cell Proliferation Assay kit (Promega, Madison, WI, USA), according to the manufacturer’s instructions.
PCR assays
Total RNA from cells was prepared using TRIsure reagent (Bioline, London, UK) and reverse-transcribed with a PrimeScript RT reagent Kit (Takara Bio Inc., Shiga, Japan). PCR assays were performed using Taq PCR Master Mix (QIAGEN, Valencia, CA, USA). The primer set for the mouse Osr1 gene was 5′-CCTGGACGTGACCAAGCTAT-3′ (forward) and 5′-TGTAGCGTCTTGTGGACAGC-3′ (reverse), and for mouse GAPDH mRNA was 5′-CCGTAGACAAAATGGTGAAGGT-3′ (forward) and 5′-GTGAGTGGAGTCATACTGGAACAT-3′ (reverse). A quantitative real-time reverse transcription-PCR (qRT-PCR) assay was performed using the StepOnePlus System (Life Technologies), according to the manufacturer’s instructions, with the reaction carried out with KAPA SYBR FAST qRT-PCR Kit Master Mix (KAPA Biosystems, Woburn, MA, USA). The expression levels of mRNA are indicated as the relative expression normalized by GAPDH. Each procedure was performed in triplicate and repeated at least twice to assess reproducibility.
Transfection and luciferase activity assays
One day before transfection, the cells were plated in 24-well culture plates (Iwaki Glass, Chiba, Japan) at a density of 4 × 104 per well. An Osr1 expression vector and promoter firefly luciferase reporter vector, each 0.2 μg, were transiently transfected using FuGENE6 transfection reagent (1.2 μL/well; Roche Diagnostics, Indianapolis, IN, USA). A pRL-CMV vector (20 ng; Promega), in which the expression of Renilla luciferase was driven by the CMV promoter, was concomitantly transfected as an internal control. Approximately 48 h after transfection, the cells were lysed with passive-lysis buffer and the luciferase activities of both types were measured using a dual luciferase reporter assay system (Promega). Promoter firefly luciferase activity was normalized by Renilla luciferase activity. The experiments were performed in triplicate and repeated at least twice.
Statistical analysis
Values are presented as the mean and standard deviation (SD) of results of separate experiments and compared using unpaired two-tailed Student’s t test. Values of P < 0.05 were considered to indicate a significant difference (*).
Results
Construction of Osr1 transgenic mice
To determine whether Osr1 affects bone formation in vivo, we generated transgenic mice expressing Osr1 under the control of the authentic 2.8-kb Osr1 promoter [7]. The transgene consisted of the Osr1 full-length coding sequence (Fig. 1a). Since we failed to measure the copy number by Southern blot technique, the expression level of transgene was estimated by qPCR. Among the transgenic founders, several lines, Tg12, Tg20, Tg25, Tg24, and Tg28, had a higher expression of the Osr1 transgene in qRT-PCR analysis of tail total RNA, and those showed a similar skeletal phenotype (Fig. 1b). The transgenic founder Tg28 showed the highest expression. Next, we performed qRT-PCR analysis to confirm transgene expression in calvaria, femur, tibia, heart, and kidney tissues of Tg28. The expression level of the transgene in the tibia was approximately ten-fold higher than the level of endogenous expression, and about seven-fold higher in femur and kidney tissues (Fig. 1c). A difference in Osr1 expression between Wt and Tg because of artificial 2.8-kb Osr1 promoter construct was observed. The body weight of Osr1 Tg28 mice was less and the weight difference became prominent after 11 days (Fig. 1d).
Generation of Osr1 transgenic mice. a Schematic representation of the Osr1 transgene. The transgene contained full length Osr1 under the control of a 2.8-kb Osr1 promoter fragment, and possessed the first intron of the rabbit beta-globin gene and polyA signals of SV40. b Expression level of Osr1 transgene in transgenic (Tg) founders. Total RNA samples were obtained from mouse tails and analyzed by qRT-PCR with transgene-specific primers. The transgenic founder Tg28 showed the highest expression. Data are shown as the mean and SD. Significant (P < 0.05) induction in comparison with the wild-type (Wt) is indicated by an asterisk (*). c The tissue distribution of Osr1 in Wt and Tg28 mice was determined by qRT-PCR assays. Data are shown as the mean and SD. *P < 0.05 (t test) versus Wt. The experiments were performed in triplicate and repeated at least twice. d Body weights of Wt and Tg28 male littermates (n = 3 in each group)
Osr1 transgenic mice exhibited shorter-bodied phenotype
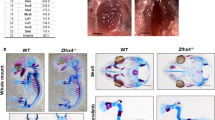
We analyzed 3-week-old male wild-type and transgenic littermates. Osr1 transgenic mice were smaller than the wild-type (Fig. 2a), which was confirmed by analysis of radiographic and skeletal preparation findings (Fig. 2b, c). The body length was 6.4 ± 0.84 and 5.8 ± 0.35 cm in Wt and Tg, respectively, meaning that Tg mice had an abnormality in the vertebrae. In contrast, long bone size was similar to that of the wild-type littermates (Fig. 2d, e). To determine whether Osr1 has a physiologic function in tooth development, we characterized the tooth phenotype of the transgenic mice. In mandible preparations, Osr1 transgenic mice displayed much shorter mandible and small condyle of mandible compared with their wild-type littermates (Fig. 2f). Osr1 transgenic mice displayed a white incisor compared with their wild-type littermates, suggesting a malfunction of ameloblasts (Fig. 2g). Furthermore, the present Osr1 transgenic mice did not demonstrate prenatal or postnatal lethality, while Osr1-deficient mice died between E11.5 and E12.5 [5]. There was no difference in the longevity and other tissues such as muscle between wild-type and Osr1 transgenic mice. Although Osr1 functions in heart morphogenesis and urogenital development [5], there was no abnormality in heart and kidney in Tg mice. Together, these results suggested that the small phenotype of Osr1 transgenic mice is caused by impaired bone formation and teeth abnormality from impaired teeth formation.
Phenotype of Osr1 transgenic mice. a Gross appearance of 3-week-old male wild-type (Wt) and Osr1-expressing (Tg) mice. b Radiographic analyses of skeletons from 3-week-old male Wt and Tg mice. c Skeletal preparations of newborn Wt and Tg mice. The skeletons were stained first with Alcian blue (cartilage), then Alizarin red (bone). d, e Radiographic images of tibia (d) and femur (e) specimens. f Surgical extraction of mandibular bone from 3-week-old male mice. g Enlarged mandibular image (incisor) from f. The experiments were performed in the Tg28 line (n = 3 in each group) (color figure online)
Skull formation abnormality in Osr1 transgenic mice
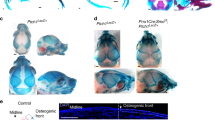
Next, we performed detailed analysis of the skull phenotype and found a calvaria abnormality in the transgenic mice indicating that intramembranous ossification was affected by Osr1 overexpression. The calvaria of the transgenic mice was shorter and more round than that of their wild-type littermates (Fig. 3a). The length of the anteroposterior axis of the cranium in Tg mice was shorter than in Wt mice (Fig. 3b). The facial bones, especially both the maxilla and mandible, were much shorter in Tg mice (Fig. 3c). Delayed cranial closure was also observed in the transgenic mice, indicating that Osr1 overexpression caused cranial dysraphism. Notably, the spaces of both the lambdoid and sagittal sutures were wider in the transgenic mice (Fig. 3d). No difference was observed in Alcian blue staining, meaning that there was no difference in chondrocalvarium (cartilage appeared transiently in calvaria of fetal and newborn mice, Fig. 3d). Histological analysis of calvaria specimens from Osr1 transgenic mice showed an abnormality in cranial closure (Fig. 3e). We then performed ALP and Alizarin red staining of calvaria from wild-type and Osr1 transgenic mice, and weaker staining in the Osr1 transgenic calvaria was detected in the area of cranial closure (Fig. 3f, g). These observations indicated that Osr1 is involved in cranial closure and skull formation.
Skull of Osr1 transgenic mice. a Surgical extraction of skulls from 3-week-old male mice. b Quantification for cranium length. Data are shown as the mean and SD. *P < 0.05 (t test) versus Wt. c Radiographic analyses of skulls from 3-week-old male mice. d Skeletal preparations of newborn Wt and Tg mice with alcian blue (cartilage) and Alizarin red (bone). Arrows indicate delayed cranial closure. e Histological analysis of calvaria from newborn mice. Coronal sections of parietal bones were stained using H&E staining. Arrows indicate parietal bones. f, g Calvaria of newborn mice. The calvaria specimens were stained with ALP (f) and Alizarin red (AR, g). Arrows indicate delayed cranial closure. The experiments were performed in the Tg28 line (n = 3 in each group) (color figure online)
Proliferation and differentiation of primary calvarial cells
We analyzed the activities of primary calvarial osteoblasts obtained from wild-type and Osr1 transgenic mice to confirm that the phenotype of the latter was caused by an intrinsic defect of osteoblasts. Proliferation of osteoblasts was evaluated using an MTS assay and found to be increased in the transgenic mice (Fig. 4a). We considered that the enhanced osteoblastic proliferation in those might be caused by an aberrant function of osteoblasts. Thus, we performed RT-PCR expression analysis to compare osteoblast differentiation between the wild-type and transgenic mice. RNA was prepared from 7-day cultures of primary calvarial cells, in which we observed overexpression of Osr2 as well as Osr1, whereas the expression of Msx1 and Msx2 was suppressed (Fig. 4b). The expression of osteoblastic genes was also affected, as osteocalcin and Osterix were decreased, ALP, type I α2 collagen, and Runx2 were unchanged, and type I α1 collagen was increased in calvaria cells from the Osr1 transgenic mice (Fig. 4c). Chondrogenic genes including type II α1 and type X α1 collagen, and Sox9 were increased, whereas aggrecan was decreased (Fig. 4d). Together, these findings suggest that Osr1 negatively regulates osteoblast differentiation, while it positively regulates cell proliferation and chondrocyte differentiation.
Primary cultures of calvarial cells from newborn wild-type and transgenic mice. a Proliferation of primary calvarial cells was analyzed with an MTS assay. b–d RT-PCR analysis of expression in primary cultures of cells from newborn mice after 7 days of differentiation. Primary cultures prepared from two separate calvaria were analyzed. Abbreviations; OCN (osteocalcin), Osx (Osterix), Col1a1 (type I alpha 1 collagen), Col1a2 (type I alpha 2 collagen), Agc (aggrecan), Col2a1 (type II alpha 1 collagen), Col10a2 (type X alpha 1 collagen). The experiments were performed in triplicate and repeated at least twice. *Significant difference from Wt (P < 0.05)
BMP2 down-regulates Osr1 expression
BMP2 induces ALP production in osteoblastic cells [13]. Primary cells from calvaria were cultured for 7 days in BMP2-treated or mineralization medium and then subjected to ALP staining. The level of ALP staining of cells from the transgenic mice was reduced compared to those from the wild-type cultured in both types of media (Fig. 5a). The staining intensity of ALP of the transgenic mice was reduced compared to the wild-type cultured in mineralization medium (Fig. 5b). Primary cells from calvaria were then cultured for another 21 days and subjected to Alizarin red S staining for detecting calcium deposits. Staining intensity was weakly mineralized in Osr1 calvaria cells cultured in mineralization medium, whereas primary cell cultures treated with BMP2 displayed less staining (Fig. 5a). Next, we analyzed Osr1 expression after BMP2 treatment in mesenchymal C3H10T1/2 and osteoblastic MC3T3-E1 cells (Fig. 5c), and the results of those assays indicated that BMP2 negatively regulated Osr1 expression. To further analyze whether Osr1 affects osteoblast differentiation, Osr1 was knocked down in the cell lines. siRNA for Osr1 and the luciferase reporter were transfected into mesenchymal C3H10T1/2 or osteoblastic MC3T3-E1 cells, which were then cultured for 2 days, after which luciferase activity was measured. That activity was enhanced by Osr1 knockdown. In addition, a reporter assay for the ALP promoter indicated that Osr1 siRNA induced ALP promoter and osteocalcin promoter activities (Fig. 5d–g). Together, these results show that Osr1 negatively regulates the expression of ALP and osteocalcin.
Involvement of Osr1 in osteogenic gene expression. a Primary cultures of calvarial cells from newborn mice were treated with BMP2 (lane B) or cultured in mineralization medium alone (lane M), followed by ALP staining after 7 days and Alizarin red staining (AR) after 21 days of culture. C control. b Quantification for ALP staining in mineralization medium (M). Data are shown as the mean and SD. *P < 0.05 (t test) versus Wt. c Effect of Osr1 expression by BMP2 in C3H10T1/2 and MC3T3-E1 cells. Cells were treated with 100 ng/mL of BMP2 for 2 days, and the expression levels of Osr1, ALP, and osteocalcin (OCN) were analyzed by qRT-PCR. (d–g) Reporter assay for ALP (ALP-Luc) and the osteocalcin promoter (OCN-Luc) in C3H10T1/2 and MC3T3-E1cells. Cells were transfected by siRNA for Osr1 (siOsr1) and reporter genes for 2 days. The experiments were performed in triplicate and repeated at least twice. *Significant difference from siCtrl (P < 0.05) (color figure online)
Discussion
In this study, the zinc-finger transcription factor Osr1 was found to function in both skull and cranial bone formation. Our genetic analyses showed that transgenic mice overexpressing Osr1 had round skulls and delayed skull closure in the calvaria. We also noted essential functions of Osr1 in the processes of proliferation and differentiation of cranial osteoblasts along with alternation of expression of osteoblast marker genes. These results strongly suggest that Osr1 plays a regulatory role in critical skull and cranial bone formation.
Osr1 transgenic mice display an abnormal cranial suture. Several genes, such as Gsk3a and Gsk3b [14], Runx2 [15], Msx2 [16], and ameloblastin [17], have also been demonstrated to play crucial roles in precise skull formation. For example, Runx2 expression was found in cranial suture samples [18] and Runx2 deficiency was seen in cleidocranial dysplasia [19], while we previously reported that Runx2 decreased Osr1 expression [7].
Regulation of Osr1 expression by Runx2 seems to be critical for skull formation. Nevertheless, the interaction between Osr1 and Runx2, and modulation of Runx2 expression by Osr1 remain to be elucidated. Msx2 was reported to be involved in premature suture closure [16]. In the present study, we found that Osr1 overexpression resulted in delayed closure and Msx2 expression was down-regulated in Osr1 transgenic mice. These findings suggest that adequate adjustment of these genes is important for proper suture closure. Although the relationships between Osr1 and Gsk3a, Gsk3b, and ameloblastin should be further analyzed, the present results demonstrated that Osr1 is an important transcriptional factor involved in skull shape formation and suture fusion.
It is also important to identify the molecular mechanisms regulating Osr1 expression during bone formation. Recently, James et al. reported that a low concentration of BMP activated Osr1 gene expression, whereas a high concentration repressed it [20], suggesting that the function of Osr1 in bone formation is regulated by the BMP signaling pathway. In the present study, we analyzed Osr1 expression by undifferentiated C3H10T1/2 and differentiated MC3T3-E1 cells cultured with a high dose of BMP2. Consistent with previously reported results, we found that Osr1 expression was decreased by BMP2 treatment. This control of expression suggests that Osr1 functions during osteoblast differentiation induced by BMP2. siRNA knockdown of Osr1 induced ALP (an early osteogenic marker) and osteocalcin (a late marker), which also suggests that Osr1 functions in commitment to osteoblast lineage. Based on our previous studies regarding the Osr2 binding sequence [6], a possible explanation is regulation of the expression of osteoblast marker genes via the Osr binding sequence by Osr1. We also found induction of chondrocyte genes, suggesting a function of Osr1 in chondrogenesis. These molecular activities in chondrogenesis require further elucidation in a future study, although our present results indicate a genetic function of Osr1 in osteoblasts and chondrocytes during formation of skeletal tissue. The expression of Osr1 in osteoclasts was 40 times lower than in osteoblasts (unpublished observation). As the relationship between Osr1 and osteoclastogenesis remains unclear, further studies are needed to elucidate this association.
The kidneys regulate homeostasis of calcium and phosphate, which is related to bone metabolism, while 1,25-dihydroxy vitamin D, parathyroid hormone, and FGF23 regulate the serum levels of calcium and phosphate. James et al. reported that the Osr1 gene functions in generation of kidney precursor cells and their differentiation into nephrons [21], and it was also reported that the OSR1 rs12329305(T) allele in Caucasians alters the activity of the exon 2 splice enhancer, and reduces spliceosome-binding affinity and stability of OSR1 mRNA. Another study found that the OSR1 rs12329305(T) allele had a relationship with small kidney volume and high cystatin C levels in cord blood [22]. An investigation of Osr1 knockout mice also showed that Osr1 functions in heart morphogenesis and urogenital development [5]. However, Osr1 transgenic mice did not show obvious abnormalities in those tissues, even though high expression levels were found in the heart and kidneys. Elucidation of the detailed mechanisms of Osr1 in heart and kidney tissues of Osr1 transgenic mice requires additional investigation.
In summary, our findings are the first to show the underlying functions of Osr1 in skull and bone formation. As Osr1 transgenic mice exhibited decreased osteoblast activity as well as delayed skull closure in the calvaria, we speculate that Osr1 is a regulator of osteoblast function. Our results also indicate the existence of a novel alternative cascade involved in skull formation. Thus, regulation of Osr1 function by pharmacological compounds is a potential strategy for abnormal skull development.
References
Coulter DE, Swaykus EA, Beran-Koehn MA, Goldberg D, Wieschaus E et al (1990) Molecular analysis of odd-skipped, a zinc finger encoding segmentation gene with a novel pair-rule expression pattern. EMBO J 9:3795–3804
So PL, Danielian PS (1999) Cloning and expression analysis of a mouse gene related to Drosophila odd-skipped. Mech Dev 84:157–160
Katoh M (2002) Molecular cloning and characterization of OSR1 on human chromosome 2p24. Int J Mol Med 10:221–225
Kawai S, Michikami I, Kitagaki J, Hashino E, Amano A (2013) Expression pattern of zinc-finger transcription factor Odd-skipped related 2 in murine development and neonatal stage. Gene Expr Patterns 13:372–376
Wang Q, Lan Y, Cho ES, Maltby KM, Jiang R (2005) Odd-skipped related 1 (Odd 1) is an essential regulator of heart and urogenital development. Dev Biol 288:582–594
Kawai S, Yamauchi M, Wakisaka S, Ooshima T, Amano A (2007) Zinc-finger transcription factor odd-skipped related 2 is one of the regulators in osteoblast proliferation and bone formation. J Bone Miner Res 22:1362–1372
Yamauchi M, Kawai S, Kato T, Ooshima T, Amano A (2008) Odd-skipped related 1 gene expression is regulated by Runx2 and Ikzf1 transcription factors. Gene 426:81–90
Deng ZL, Sharff KA, Tang N, Song WX, Luo J et al (2008) Regulation of osteogenic differentiation during skeletal development. Front Biosci 13:2001–2021
Gordon JW, Scangos GA, Plotkin DJ, Barbosa JA, Ruddle FH (1980) Genetic transformation of mouse embryos by microinjection of purified DNA. Proc Natl Acad Sci USA 77:7380–7384
McLeod MJ (1980) Differential staining of cartilage and bone in whole mouse fetuses by alcian blue and alizarin red S. Teratology 22:299–301
Lyle HM (1947) An improved tissue technique with hematoxylin-eosin stain. Am J Med Technol 13:178–181
Farley JR, Tarbaux NM, Hall SL, Linkhart TA, Baylink DJ (1988) The anti-bone-resorptive agent calcitonin also acts in vitro to directly increase bone formation and bone cell proliferation. Endocrinology 123:159–167
Rawadi G, Vayssiere B, Dunn F, Baron R, Roman-Roman S (2003) BMP-2 controls alkaline phosphatase expression and osteoblast mineralization by a Wnt autocrine loop. J Bone Miner Res 18:1842–1853
Barrell WB, Szabo-Rogers HL, Liu KJ (2012) Novel reporter alleles of GSK-3alpha and GSK-3beta. PLoS One 7:e50422
Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K et al (1997) Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89:755–764
Liu YH, Kundu R, Wu L, Luo W, Ignelzi MA Jr et al (1995) Premature suture closure and ectopic cranial bone in mice expressing Msx2 transgenes in the developing skull. Proc Natl Acad Sci USA 92:6137–6141
Atsawasuwan P, Lu X, Ito Y, Zhang Y, Evans CA et al (2013) Ameloblastin inhibits cranial suture closure by modulating MSX2 expression and proliferation. PLoS One 8:e52800
Park MH, Shin HI, Choi JY, Nam SH, Kim YJ et al (2001) Differential expression patterns of Runx2 isoforms in cranial suture morphogenesis. J Bone Miner Res 16:885–892
Otto F, Kanegane H, Mundlos S (2002) Mutations in the RUNX2 gene in patients with cleidocranial dysplasia. Hum Mutat 19:209–216
James RG, Schultheiss TM (2005) Bmp signaling promotes intermediate mesoderm gene expression in a dose-dependent, cell-autonomous and translation-dependent manner. Dev Biol 288:113–125
James RG, Kamei CN, Wang Q, Jiang R, Schultheiss TM (2006) Odd-skipped related 1 is required for development of the metanephric kidney and regulates formation and differentiation of kidney precursor cells. Development 133:2995–3004
Zhang Z, Iglesias D, Eliopoulos N, El Kares R, Chu L et al (2011) A variant OSR1 allele which disturbs OSR1 mRNA expression in renal progenitor cells is associated with reduction of newborn kidney size and function. Hum Mol Genet 20:4167–4174
Acknowledgements
We thank all the members of the Challenge to Intractable Oral Diseases and Center for Frontier Oral Science for their assistance and encouragement, as well as Mark Benton for their comments regarding our manuscript. This work was supported by Grants-in-Aid for Scientific Research from the Japanese Society for the Promotion of Science (KAKENHI no. C21592356, C24592796), and by special funds for Challenge to Intractable Oral Diseases.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All authors declare that they have no competing interests.
About this article
Cite this article
Kawai, S., Yamauchi, M. & Amano, A. Zinc-finger transcription factor Odd-skipped related 1 regulates cranial bone formation. J Bone Miner Metab 36, 640–647 (2018). https://doi.org/10.1007/s00774-017-0885-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00774-017-0885-9