Abstract
Salinity is the second most prevalent abiotic stress faced by plants, and rice is not an exception. Through this study, it has been tried upon, to study the relative salinity tolerance of eight local varieties of North East India. Preliminary screening was based on their dose- and time-dependent physiological responses to salinity stress. Among the cultivars, Tampha was found to be relatively more tolerant, whereas MSE9 the most sensitive. To further ascertain their tolerance capacity, MDA and H2O2 content was determined, which also confirmed the tolerance level of the two cultivars. Histochemical assays for root plasma membrane integrity and leaf and root H2O2 and O2 − content also showed more damage in Tampha in comparison to MSE9. Finally, gene expression analysis for Na+/K+ co-transporters, OsHKT2;1, OsHKT2;3 and OsHKT2;4, was performed to observe how the expression level of these transporters varies with the tolerance capacity of these two cultivars in leaves and roots under different time frames. The study reveals Tampha to be the most tolerant and MSE9 the most sensitive when compared to the other six screened cultivars for salinity stress.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L. spp. indica) is considered the second most important crop after wheat, taken as staple food by nearly half of the world’s population (Ghosh et al. 2016). Ninety percent of the world’s total rice produce is consumed by Asian population (IRRI 2013). Rice as a crop is hampered by various abiotic stresses especially salt stress (Mittal et al. 2016). Salt stress is the second most prevalent abiotic stress to plants next to drought, limiting production and productivity of crops worldwide. Saline soil is characterised by high electrical conductivity of above 4 dS/m (approximately 40 mM NaCl; Chinnusamy et al. 2005). High salinity has adversely affected an estimated area of around 800 million hectares of the total land area of the world (Munns and Tester 2008). So, finding relative salt tolerance capability of genotypes will enable to achieve higher productivity of rice for meeting the demands of ever-increasing world human population, and the selected salt-tolerant genotypes shall go a long way for “omic” studies to decipher salt stress tolerance mechanisms.
Soil salinity negatively impacts plants generally by two mechanisms, osmotic stress and ion toxicity (Vaid et al. 2015). Osmotic stress is the result of increased amount of salt in growth medium that hampers capacity of plant to retain and absorb water (Morales et al. 2012). Whereas, ion toxicity is caused by ionic imbalance due to higher accumulation of Na+ and Cl− ions at toxic level thereby lowering the availability of calcium (Ca2+) and potassium (K2+) (Hussain et al. 2013). Excessive Na+ may build up in apoplast thereby causing cell dehydration; they may also accumulate in cytosol thereby inhibiting enzymes responsible for metabolic processes including photosynthesis (Munns and Tester 2008). In case of rice, salinity induces biochemical and physiological alterations, causing growth inhibition and yield reduction (Ghosh et al. 2016). Growth reduction due to salinity differs greatly with species and cultivars within a species (Khan et al. 2002). Shoot growth is found to be more effected by stress than root. It may be explained as plants’ adaptive response to keep absorbing water by elongating roots even after the stress. As roots are in direct contact with saline growth medium, they need to elongate to access clean water. Whereas in case of shoots it is just the opposite, to adapt the osmotic stress due to salinity, they need to lower the transpiration pull for which there is reduction of foliage surface area. The significant biomass reduction is also due to impaired photosynthetic mechanisms and relatively decreased water content due to inability of plant for normal uptake of water. Severity and duration of the stress control the changes in stress-inducible parameters (Mishra et al. 2016).
Damage due to salinity is further accelerated due to overproduction of reactive oxygen species (ROS); like H2O2, O2 − drastically hampers metabolic homeostasis and cell membrane integrity (Hussain et al. 2013). Significant higher build-up of ROS leads to lipid peroxidation thereby interfering membrane stability (Chunthaburee et al. 2016). Sensitive rice plants show higher generation of H2O2 and lipid peroxidation molecules which are quantified as malondialdehyde (MDA). Tolerant cultivars thrive to survive by generating antioxidant enzymes that will catalyse the removal of ROS.
Intracellular ion homeostasis is fundamental to the physiology of living cells, and K+ and Na+ homeostasis is more vital under salt stress (Mishra et al. 2016). Na+ competes for uptake with K+ into roots of rice plants after being exposed to stress. Low cytosolic Na+ and the low Na+/K ratio are required for osmotic and biochemical equilibrium in plant cells (Yao et al. 2010). Na+/K+ ratio is a key factor for salt tolerance in plants (Sun et al. 2014). Plants regulate the expression and activities of various membrane transporters to maintain this ratio. Among these, HKT (for high-affinity K+ transporters) are integral membrane proteins that facilitate cation transport across the membrane (Waters et al. 2013). They are subdivided into two subgroups HKT1 and HKT2 based on phylogenetic analysis (Platten et al. 2006). Class 1 HKT shows more Na+ transport activity, whereas the class 2 members show K+ permeability in addition to Na+. HKT1 are basically single-ion (Na+) transporters preventing overaccumulation of Na+ in the photosynthetic tissues, whereas HKT 2 transport both Na+ and K+ from the external medium depending on the concentration of each (Almeida et al. 2013). HKT2;1 is unique as it exhibits features of HKT class 1 transporters in having a serine residue instead of glycine in the “P loop”, responsible for binding ions (serine depicts more Na+ specificity and glycine more K+). HKT2;3 and HKT2;4 show around 93% sequence homology but differ in their function. Comparatively less studies have been done on HKT2;3, and its expression was found to have no effect on varying dosages of K+ and Na+ in growth medium. HKT2;4 shows Na+-independent K+ transport and is found to be dependent on a wide range of divalent cations like Ca2+ and Mg2+ (Horie et al. 2011). Expression analysis of HKT genes is presumed to give a better insight into ion transport under salinity stress.
This manuscript reports the relative salinity tolerance of eight indica rice genotypes, native to north eastern India using various physiological, biochemical and molecular strategies.
Materials and methods
Rice sample
Viable germplasms were collected from the Regional Agricultural Research Station, Karimganj, Assam and the Rice Research Centre, Thoubal, Manipur (Table 1).
Growth, treatment and relief
Seeds were surfaced-sterilised with 0.1% mercuric chloride and washed thrice thoroughly with distilled water. They were set for germination at 30 °C in dark. Uniformly germinated seeds were transferred to plastic pots containing Hoagland’s nutrient medium (Hoagland Arnon 1950), and growth conditions of plants were set at growth chamber with photon flux density of 52 μmol s−1 m−2 (PAR) and 16 h photoperiod. On the sixth day, plants were treated with 150 mM of salt. After 24 and 96 h duration, both control and stressed seedlings were excised for analysis (Fig. 1). After the completion of the 96 h stress period, Tampha and MSE9 were re-watered with normal nutrient solution and allowed to grow for 6 days (Castillo et al. 2007). Differences in revival potential of the two varieties have been documented.
Determination of plant growth
Growth of plants scored in the form of length of root, shoot and fresh weights was measured after 24 and 96 h of the treatment (Fig. 1).
Estimation of RWC
Relative water content of both shoot and root was determined (Barrs and Weatherley 1962). Plant tissue samples were soaked in double distilled water for 4 h to get turgid weight after taking respective fresh weights. Later, they were oven-dried at 60 °C for 72 h to get the dry weight.
Na+ and K+ content
A total of 0.1 g of dry sample were weighed and ashed at 550 °C for 3 h in a muffle furnace (dry ashing method). The ash was dissolved in 2 N HCl and extracted for Na+ and K+ (Chapman and Pratt 1962). The content was measured by flame photometer (Flame photometer 129, Systronics, Ahmedabad, India).
Estimation of H2O2 and lipid peroxidation
Hydrogen peroxide contents of MSE9 and Tampha were estimated after 24 and 96 h of NaCl stress. For H2O2, 0.2 g of samples was extracted with 5% TCA and centrifuged at 12,500 rpm for 10 min (Sagisaka 1976). The reaction mixture contained 1.6 ml of supernatant, 0.4 ml of 50% TCA, 0.4 ml of 10 mM ferrous ammonium sulphate and 0.2 ml of 2.5 M KSCN. H2O2 content was measured at 480 nm against suitable blank. Lipid peroxidation was measured as MDA content (Khan and Panda 2008). A total of 0.1 g of sample were homogenised with 1 ml of 1% TCA and centrifuged at 15,000 rpm for 10 min. Then, 0.5 ml of supernatant was mixed with 1.5 ml of 0.5% TBA and incubated at 95 °C for 25 min. Absorbance was measured at 532 and 600 nm for non-specific turbidity.
Histochemical detection of H2O2, O2 − and plasma membrane integrity
Detection of hydrogen peroxide (H2O2) was done by 3,3-diaminobenzidine (DAB) staining according to the method of Ramel et al. (2009) and of superoxide radical (O2 −) by NBT staining following the modified method of Rao and Davis (1999) in leaf segments and roots under both stressed and unstressed conditions. The leaf segments and roots were immersed and infiltrated under vacuum with 1 mg/ml DAB staining solution, pH 7.8, dissolved in H2O for 6 h and 3 mg/ml nitro-blue tetrazolium (NBT) staining solution in 50 mM potassium phosphate buffer (pH 7.5) for 30 min at room temperature. Stained leaves were bleached in acetic acid:glycerol:ethanol (1:1:3 v/v) solution at 100 °C for 5 min and stored in glycerol:ethanol (1:4 v/v) solution until photographed.
The loss of plasma membrane integrity was evaluated using Evans blue staining method with slight modifications (Schutzendubel et al. 2001). Roots of intact seedlings were stained with 0.25% (w/v) Evans blue in 100 μM CaCl2 (pH 5.6) for 30 min; then, the stained roots were washed with 100 μM CaCl2 for 15 min. After rinsing with CaCl2, root tips were cut with sharp razor blade for stereoscopic microscope observation.
Gene expression analysis
For total RNA extraction, 0.2 g of fresh tissue was homogenised in liquid nitrogen. Then, extraction and isolation were done as per manufacturer’s instruction (Nucleopore RNA Sure Plant Kit, Genetix Biotech, New Delhi, India).
Total RNA extracted was processed for cDNA synthesis following manufacturer’s instruction (First strand cDNA synthesis kit, Thermo Scientific, USA). Amplification for gene used for expression analysis was done by polymerase chain reaction (Takara PCR Thermal cycler, Japan) for 30 cycles using respective primers (Supplementary 1).
Statistical analysis
For statistical analysis, 10–15 seedlings per replicate per experiment were taken into account. Statistical comparison between the variances was determined by ANOVA (analysis of variance), and significant differences between mean values (n = 3), where n is the number of times experiments repeated, were determined by LSD analysis. P ≤ 0.05 was deemed to show statistical significance.
Results
Changes in growth parameters
The effect of NaCl treatment on growth of plants in terms of root length and shoot length has been elucidated in Table 2. Salinity has caused reduction in the growth of roots of rice varieties except Krishna after 96 and 24 h of stress, respectively. Leima, Sana and Luit have been found to have root lengths significantly reduced from their respective controls. MSE9 has been observed to have the highest reduction in length of 27.83% over its control. Whereas root lengths of Krishna, Pankaj have not been significantly reduced showing tolerance to stress. The same trend has also been observed in Tampha after 24 h treatment and its decrease over control being 5.92 and 4.51% after 1st and 4th days of treatment (Fig. 1).
Shoot length of plants had demonstrated more susceptibility to salt stress as all the varieties have shown significantly reduced shoot lengths after 24 and 96 h of stress. Tampha had been least affected with an average percentage decrease of about 24% for both the treatment duration and MSE9 being the most affected as its treated shoot length has reduced by 57% in comparison to that of the control (Fig. 1). Krishna and Pankaj shoot lengths have been moderately effected (Table 2).
Relative water content
Relative water content (RWC) of eight rice varieties was found to be effected to different extent by salinity (Table 3), and RWC of each variety was significantly reduced due to stress compared to respective untreated condition. Among the varieties, Tampha showed least decrease having 80 and 74.12% RWC in roots and 84.92 and 74.52% RWC in shoots after 24 and 96 h of stress, respectively, while MSE9 variety showed highest decrease over control, i.e., 50% in case of both roots and shoots. Krishna and Pankaj gave better performance than the remaining varieties except Tampha. Punsi, Luit, Sana and Leima had higher value of RWC than that of MSE9 having around 70% RWC in roots and shoots after 24 h and 60% after 96 h. Time duration of the salt stress also negatively influenced RWC as we found that reduction after 96 h was more than that after 24 h stress in all varieties (Table 3).
Na+, K+ uptake and Na+/K+ ratio
All the varieties showed increased sodium uptake at the expense of decreased K+ uptake after NaCl treatment. Na+ content of all the plants significantly increased from their respective untreated condition at P ≤ 05 after stress (Tables 4 and 5).
After 24 h, Na+ content of rice shoot ranged from 12 to 26 μg/g while that of controls remained below 10 μg/g. MSE9 uptake was maximum, with a value of 26.65 μg/g of Na+ while for Tampha it was the least, with 12.68 μg/g (Table 4). After 96 h, Tampha variety had accumulated 17.42 μg/g of Na+ but MSE9 had 40.18 μg/g which was four times higher of its respective control. On the other hand, K+ uptake had been significantly reduced due to treatment, lowest being recorded for MSE9 at 35.77 μg/g after 96 h treatment. Rice varieties Krishna and Pankaj demonstrated moderate Na+ increase and K+ decrease, which attributed to their relatively low Na+/K+ ratios when compared to other varieties except Tampha. Tampha showed the highest tolerance with least Na+/K+ ratio, its increase over control being minimal.
Though Tampha showed the least decrease, significant increase of Na+/K+ ratio in all the varieties after treatment has been recorded with respect to control. Decreased K+ and increased Na+ uptake have been manifested most prominently in MSE9 showing its extreme sensitivity to salinity. MSE9 Na+/K+ ratio has been observed to be the highest, after 96 h treatment (Tables 4 and 5).
H2O2 and MDA content
H2O2 and MDA content were analysed in Tampha and MSE9 varieties to confirm their tolerance level. MDA content in both the varieties had significantly increased even though Tampha had lower MDA than MSE9. Tampha had 23.097 and 26.624 μM/gfwt in leaf and root, respectively, after 96 h stress, while MSE9 had 25.419 and 28.860 in leaf and roots, respectively. H2O2 had also been found to be in decreased quantity in Tampha when compared to MSE9 (Fig. 2).
Quantitative assay for H2O2 accumulation and lipid peroxidation (MDA) of shoot and root tissue samples at 24 and 96 h. Data represents mean values (n = 3) ± SE, where n is the number of times experiment repeated and SE denotes standard error. Statistically significant values at P < 0.05 using LSD analysis are indicated by star marks
Histochemical detection of H2O2, O2 − and plasma membrane integrity
H2O2, O2 − production and plasma membrane integrity in stressed and unstressed rice leaf segments and roots were investigated qualitatively using DAB, NBT and Evans blue histochemical staining, respectively (Fig. 3). Under normal physiological conditions, both Tampha (tolerant) and MSE9 (sensitive) showed low O2 − and H2O2 accumulation and almost intact plasma membrane. However, under salinity stress, rice leaf segments and roots of Tampha exhibited marked lower NBT, DAB and Evans blue staining than MSE9 which is an indication of less ROS production and less oxidative damage in Tampha (Fig. 3).
Gene expression analysis
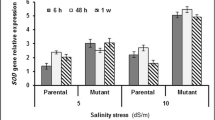
For OsHKT2;1, in leaves, first, there was increase in expression level with increase in stress duration then again decrease, but the intensity was more in the case of MSE9 in comparison to Tampha; whereas, in case of roots, there is gradual decrease throughout. OsHKT2;3 expression pattern did not show any deviation with increase in stress duration for both the varieties except in case of MSE9 leaves where there was a gradual increase. HKT2;3 expression was less in roots in comparison to leaves. No specific pattern was observed in case of OsHKT2;4. In some cases, there was gradual increase, whereas in others, there was increase and then decrease with increment of stress duration. All of these analyses were done after normalising the cDNA concentration with OsActin primers (Fig. 4).
Discussions
In this study, we have analysed physiological, biochemical and molecular characteristics in rice varieties of North East India under salt stress to show relative tolerance. Plant growth scores in the form of root length and shoot were found to be reduced implying that salt stress represses the growth of rice plants. Similar findings were earlier reported in rice (Jia et al. 2015; Yeo et al. 1990; Hussain et al. 2013). All throughout, shoot growth has been more susceptible compared to roots as it has been significantly reduced for all plants to different extent (Fig. 1, Table 2). It might be explained citing the fact that relative shoot length reduction compared to root would be helpful to plants for decreasing the water use, being already osmotically stressed (due to increased Na+ uptake) by salt (Munns and Tester 2008; Munns et al. 2006). Lower water potential in the cell causes stomatal closure and limits CO2 assimilation mounting single direct negative impact on photosynthesis which also causes growth reduction (Pattangul and Thitisaksakul 2008).
RWC is considered an appropriate measure of plant water status as well as osmotic adjustment under stress (Baisakh et al. 2012). It is known that osmotic stress due to salinity disturbs plant water status (Amirjani 2010) causing significant reduction in RWC of rice plants in both root and shoot (Qin et al. 2010; Rodriguez et al. 1997). Under such circumstances, plant responds by osmotic adjustment by increased uptake of Na+ and Cl− which is readily available in saline or treated growth condition. This, in turn, causes ion toxicity and inactivates various metabolic functions like photosynthesis and electron transport chain. But such adaptation is always not adequate enough, resulting in observation of decreased RWC in treated plants (Table 3; Pattangul and Thitisaksakul 2008).
Abiotic stress including salt stress leads to oxidative damage due to rapid and uncontrolled ROS production (Miller et al. 2010; Saha et al. 2016). The primary effects of salinity like membrane damage, ion toxicity and imbalance decrease assimilation of CO2 and reduce antioxidant enzyme activity leading to higher H2O2 production. Tampha variety after treatment showed lesser H2O2 accumulation compared to MSE9 even though both the varieties generate significantly higher H2O2 than their respective control (Fig. 2). This certainly concludes that Tampha is comparatively more tolerant and MSE9 is most sensitive. MDA, produced due to membrane lipid peroxidation, is often used as an indicator to differentiate between sensitive and tolerant cultivars (Dhanyalakshmi et al. 2013). In this study, Tampha contained less MDA than MSE9.
Low Na+/K+ ratio is an indicator of ionic homeostasis in plants (Rao et al. 2013). Under salt stress, Na+ ion competes with K+ due to its small size and abundant availability to be taken up by root through epidermal cells. The rice cultivars when arranged in decreasing order of Na+ content are as follows: MSE9, Leima, Sana, Luit, Punsi, Pankaj, Krishna and Tampha; the reverse is true for K+ uptake (Tables 4, 5). Lower accumulation of K+ and higher Na+ accumulation impart the higher Na+/K+ ratio in all varieties after treatment than the control. While comparing among the varieties, the lesser the ratio, the higher the salt tolerance adaptation (Kanawapee et al. 2013; Rao et al. 2013; Chunthaburee et al. 2016).
Accumulation of ROS (H2O2, O2 −) was histochemically observed in leaf segments and roots of both unstressed and stressed plants exposed to salinity stress (Fig. 3). It was observed that Tampha showed lesser accumulation of H2O2 and O2 − indicative of lesser oxidative damage and more tolerance to salinity stress in comparison to MSE9 (Saha et al. 2016). Also, Evans blue uptake exclusively at the root tip which was more in the case of MSE9 was observed. Evans blue uptake has largely been used as a marker for loss of plasma membrane integrity thus depicting excessive loss of integrity in MSE9 due to salinity stress (Fig. 3; Yamamoto et al. 2001; Zhang et al. 2016; Yang et al. 2016; Awasthi et al. 2017).
Expression analysis through semi-quantitative PCR was performed for HKT group of Na+ and K+ transporters, viz. HKT2;1, HKT2;3 and HKT2;4, in order to ascertain the implications of salinity on these transporters. HKT2;1 showed gradual increase in expression at 24 h in comparison to control, but at 96 h, it decreases. Tampha showed lesser expression when compared to MSE9 (Fig. 4). Depending upon exterior Na+ and K+ concentrations, HKT2;1 acts as symporter or uniporter (Jabnoune et al. 2009). Since in case of salinity, stress exterior Na+ concentration is excessive, HKT2;1 acts as Na+ uniporter leading to its increased accumulation in cells. Tampha showed lesser expression of HKT2;1 which might had led to lesser accumulation of Na+ hence greater tolerance. No change in expression pattern of HKT2;3 was observed with stress and increase in time frame which indicated of its being independent of stress. Wu et al. (2009) reported similar results. HKT2;4 showed no pattern in expression; so, it remains inconclusive.
Conclusions
In this present study, we demonstrated relative salt-tolerant capacity of eight rice varieties of North East India based on physiological parameters. Tampha and MSE9 were found to be most tolerant and most sensitive, respectively, among the others. The two varieties were further analysed based on lipid peroxidation, H2O2 content and HKT transporter gene expression. Based on the findings, we conclude that differences in salinity tolerance mechanisms might be partially due to differences in regulation of gene expression of HKT2 transporter proteins and plants’ metabolomic adaptations to resist damage caused by oxidative stress.
References
Almeida P, Katschnig D, de Boer AH (2013) HKT transporters—state of the art. Int J Mol Sci 14(10):20359–20385
Amirjani MR (2010) Effect of NaCl on some physiological parameters of rice. Eur J Biol Sci 3(1):6–16
Awasthi JP, Saha B, Regon P, Sahoo S, Chowra U, Pradhan A et al (2017) Morpho-physiological analysis of tolerance to aluminum toxicity in rice varieties of North East India. PLoS One 12(4):e0176357
Baisakh N, Ramana Rao MV, Rajasekaran K, Subudhi P, Janda J, Galbraith D, Pereira A (2012) Enhanced salt stress tolerance of rice plants expressing a vacuolar H+-ATPase subunit c1 (SaVHAc1) gene from the halophyte grass Spartina alterniflora Löisel. Plant Biotechnol J 10(4):453–464
Barrs HD, Weatherley PE (1962) A re-examination of the relative turgidity technique for estimating water deficits in leaves. Aust J Biol Sci 15(3):413–428
Castillo EG, Tuong TP, Ismail AM, Inubushi K (2007) Response to salinity in rice: comparative effects of osmotic and ionic stresses. Plant Prod Sci 10(2):159–170
Chapman HD, Pratt PF (1962) Methods of analysis for soils, plants and waters. Soil Sci 93(1):68
Chinnusamy V, Jagendorf A, Zhu JK (2005) Understanding and improving salt tolerance in plants. Crop Sci 45(2):437–448
Chunthaburee S, Dongsansuk A, Sanitchon J, Pattanagul W, Theerakulpisut P (2016) Physiological and biochemical parameters for evaluation and clustering of rice cultivars differing in salt tolerance at seedling stage. Saudi J Biol Sci 23(4):467–477
Dhanyalakshmi KH, Vijayalakshmi C, Boominathan P (2013) Evaluation of physiological and biochemical responses of rice (Oryza sativa L.) varieties to salt stress. Indian J Agric Res 47(2):91–99
García Morales S, Trejo-Téllez LI, Gómez Merino FC, Caldana C, Espinosa-Victoria D, Herrera Cabrera BE (2012) Growth, photosynthetic activity, and potassium and sodium concentration in rice plants under salt stress. Acta Sci Agron 34(3):317–324
Ghosh B, Nasim A Md, Gantait S (2016) Response of rice under salinity stress: a review update. Rice research: open access, 2016
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Circ Calif Agric Exp Station 347(2nd edn)
Horie T, Brodsky DE, Costa A, Kaneko T, Schiavo FL, Katsuhara M, Schroeder JI (2011) K+ transport by the OsHKT2;4 transporter from rice with atypical Na+ transport properties and competition in permeation of K+ over Mg2+ and Ca2+ ions. Plant Physiol 156(3):1493–1507
Hussain M, Park HW, Farooq M, Jabran K, Lee DJ (2013) Morphological and physiological basis of salt resistance in different rice genotypes. Int J Agric Biol 15:113–118
IRRI (International Rice Research Institute) (2013) IRRI World rice Statistics (WRS). online at. http://ricestat.irri.org:8080/wrsv3/entrypoint.htm
Jabnoune M, Espeout S, Mieulet D, Fizames C, Verdeil JL, Conéjéro G, Véry AA (2009) Diversity in expression patterns and functional properties in the rice HKT transporter family. Plant Physiol 150(4):1955–1971
Jia H, Shao M, He Y, Guan R, Chu P, Jiang H (2015) Proteome dynamics and physiological responses to short-term salt stress in Brassica napus leaves. PLoS One 10(12):e0144808
Kanawapee N, Sanitchon J, Srihaban P, Theerakulpisut P (2013) Physiological changes during development of rice (Oryza sativa L.) varieties differing in salt tolerance under saline field condition. Plant Soil 370(1–2):89–101
Khan MH, Panda SK (2008) Alterations in root lipid peroxidation and antioxidative responses in two rice cultivars under NaCl-salinity stress. Acta Physiol Plant 30(1):81
Khan MH, Singha KL, Panda SK (2002) Changes in antioxidant levels in Oryza sativa L. roots subjected to NaCl-salinity stress. Acta Physiol Plant 24(2):145–148
Miller GAD, Suzuki N, CIFTCI-YILMAZ SULTAN, Mittler RON (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33(4):453–467
Mishra S, Kumar S, Saha B, Awasthi J, Dey M, Panda S K, Sahoo L (2016) Crosstalk between Salt, Drought, and Cold Stress in Plants: Toward Genetic Engineering for Stress Tolerance. In: Tuteja N, Gill S S (eds) Abiotic Stress Response in Plants. Wiley, New York, pp 57-88.
Mittal D, Sharma N, Sharma V, Sopory SK, Sanan-Mishra N (2016) Role of microRNAs in rice plant under salt stress. Ann Appl Biol 168(1):2–18
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Läuchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exp Bot 57(5):1025–1043
Pattanagul W, Thitisaksakul M (2008) Effect of salinity stress on growth and carbohydrate metabolism in three rice (Oryza sativa L.) cultivars differing in salinity tolerance. Indian J Exp Bot 46:736–742
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Mäser P (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11(8):372–374
Qin J, Dong WY, He KN, Yu Y, Tan GD, Han L et al (2010) NaCl salinity-induced changes in water status, ion contents and photosynthetic properties of Shepherdia argentea (Pursh) Nutt. seedlings. Plant Soil Environ 56(7):325–332
Ramel F, Sulmon C, Bogard M, Couée I, Gouesbet G (2009) Differential patterns of reactive oxygen species and antioxidative mechanisms during atrazine injury and sucrose-induced tolerance in Arabidopsis thaliana plantlets. BMC Plant Biol 9(1):28
Rao MV, Davis KR (1999) Ozone-induced cell death occurs via two distinct mechanisms in Arabidopsis: the role of salicylic acid. Plant J 17(6):603–614
Rao PS, Mishra B, Gupta SR (2013) Zinc/phosphorous ratio in shoot as an index of evaluating rice salt tolerance. Afr J Agric Res 8(1):70–76
Rodriguez HG, Roberts JK, Jordan WR, Drew MC (1997) Growth, water relations, and accumulation of organic and inorganic solutes in roots of maize seedlings during salt stress. Plant Physiol 113(3):881–893
Sagisaka S (1976) The occurrence of peroxide in a perennial plant, Populusgelrica. Plant Physiol 57(2):308–309
Saha B, Mishra S, Awasthi JP, Sahoo L, Panda SK (2016) Enhanced drought and salinity tolerance in transgenic mustard [Brassica juncea (L.) Czern & Coss.] overexpressing Arabidopsis group 4 late embryogenesis abundant gene (AtLEA4-1). Environ Exp Bot 128:99–111
Schützendübel A, Schwanz P, Teichmann T, Gross K, Langenfeld-Heyser R, Godbold DL, Polle A (2001) Cadmium-induced changes in antioxidative systems, hydrogen peroxide content, and differentiation in Scots pine roots. Plant Physiol 127(3):887–898
Sun J, Zou DT, Luan FS, Zhao HW, Wang JG, Liu HL, Liu ZL (2014) Dynamic QTL analysis of the Na+ content, K+ content, and Na+/K+ ratio in rice roots during the field growth under salt stress. Biol Plant 58(4):689–696
Vaid N, Pandey P, Srivastava VK, Tuteja N (2015) Pea lectin receptor-like kinase functions in salinity adaptation without yield penalty, by alleviating osmotic and ionic stresses and upregulating stress-responsive genes. Plant Mol Biol 88(1–2):193–206
Waters S, Gilliham M, Hrmova M (2013) Plant high-affinity potassium (HKT) transporters involved in salinity tolerance: structural insights to probe differences in ion selectivity. Int J Mol Sci 14(4):7660–7680
Wu Y, Hu Y, Xu G (2009) Interactive effects of potassium and sodium on root growth and expression of K/Na transporter genes in rice. Plant Growth Regul 57(3):271–280
Yamamoto Y, Kobayashi Y, Matsumoto H (2001) Lipid peroxidation is an early symptom triggered by aluminum, but not the primary cause of elongation inhibition in pea roots. Plant Physiol 125(1):199–208
Yang G, Wang C, Wang Y, Guo Y, Zhao Y, Yang C, Gao C (2016) Overexpression of ThVHAc1 and its potential upstream regulator, ThWRKY7, improved plant tolerance of cadmium stress. Sci Rep 6:18752
Yao X, Horie T, Xue S, Leung HY, Katsuhara M, Brodsky DE et al (2010) Differential sodium and potassium transport selectivities of the rice OsHKT2; 1 and OsHKT2; 2 transporters in plant cells. Plant Physiol 152(1):341–355
Yeo AR, Yeo ME, Flowers SA, Flowers TJ (1990) Screening of rice (Oryza sativa L.) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. TAG Theor Appl Genet 79(3):377–384
Zhang M, Deng X, Yin L, Qi L, Wang X, Wang S, Li H (2016) Regulation of galactolipid biosynthesis by overexpression of the rice MGD gene contributes to enhanced aluminum tolerance in tobacco. Front Plant Sci 7:337
Acknowledgements
TO thanks the Regional Agricultural Research Station, Karimganj, Assam and the Rice Research Centre, Thoubal Manipur for providing rice seeds throughout the experimental work and also to the UGC for financial support (UGC Non-NET fellowship).
Author information
Authors and Affiliations
Contributions
TO and SKP conceived and designed the experiment. TO and SS procured seeds for the experimental work. TO and BS performed all the experimental works. TO, BS and SKP contributed to writing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest in the present investigation.
Additional information
Handling Editor: Peter Nick
Electronic supplementary material
ESM 1
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Omisun, T., Sahoo, S., Saha, B. et al. Relative salinity tolerance of rice cultivars native to North East India: a physiological, biochemical and molecular perspective. Protoplasma 255, 193–202 (2018). https://doi.org/10.1007/s00709-017-1142-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-017-1142-8