Summary.
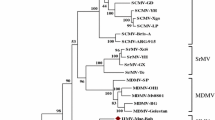
The 3′-terminal sequence of hop mosaic virus (HpMV) genomic RNA was determined. A cDNA of approximately 1.8 kbp was amplified from the HpMV genome by 3′ RACE using a degenerate primer, which was designed to anneal to the overlapping region of open reading frames (ORFs) 2 and 3 of eight carlavirus genomes. The sequence contained three ORFs, encoding proteins of 7-, 34-, and 11-kDa, which corresponded to ORFs 4, 5, and 6 of the carlavirus genome, respectively. The amino acid sequence of ORF 5, encoding the coat protein (CP) of HpMV, shows the highest identity (67%) to that of Hop latent virus (HpLV). The HpMV CP N-terminal sequence differs from that of HpLV, but the central and C-terminal sequences of the CP of both viruses are similar. The sequence similarity possibly causes the cross-reaction of heterologous antibodies of HpMV and HpLV. Phylogenetic analyses based on the CP amino acid and 3′ non-coding region sequences indicate close relationships among HpMV, HpLV, and Potato virus M. We report here the first molecular characterization of HpMV genomic RNA.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received November 30, 2000 Accepted April 11, 2001
Rights and permissions
About this article
Cite this article
Hataya, T., Arimoto, R., Suda, N. et al. Molecular characterization of Hop mosaic virus: its serological and molecular relationships to Hop latent virus . Arch. Virol. 146, 1935–1948 (2001). https://doi.org/10.1007/s007050170043
Issue Date:
DOI: https://doi.org/10.1007/s007050170043