Abstract
Geminiviruses are a major threat to agriculture in tropical and subtropical regions of the world. Geminiviruses have small genome with limited coding capacity. Despite this limitation, these viruses have mastered hijacking the host cellular metabolism for their survival. To compensate for the small size of their genome, geminiviruses encode multifunctional proteins. In addition, geminiviruses associate themselves with satellite DNA molecules which also encode proteins that support the virus in establishing successful infection. Geminiviral proteins recruit multiple host factors, suppress the host defense, and manipulate host metabolism to establish infection. We have updated the knowledge accumulated about the proteins of geminiviruses and their satellites in the context of pathogenesis in a single review. We also discuss their interactions with host factors to provide a mechanistic understanding of the infection process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Geminiviruses are a group of non-enveloped plant viruses with small circular single-stranded DNA (ssDNA) with a size of 2.5-5.2 kb. The capsid is composed of coat protein subunits that form twinned incomplete icosahedral particles, giving rise to the name geminivirus [1]. Geminiviruses infect economically important crops worldwide, causing severe loss to agriculture [2, 3]. They are transmitted by insect vectors such as whiteflies, treehoppers, leafhoppers, and aphids [4]. The family Geminiviridae is divided into 14 genera, namely Becurtovirus, Begomovirus, Capulavirus, Citlodavirus, Curtovirus, Eragrovirus, Grablovirus, Maldovirus, Mastrevirus, Mulcrilevirus, Opunvirus Topilevirus, Topocuvirus, and Turncurtovirus, based on the insect vectors, genome organization, and the host range of their members [4,5,6]. At present, the family Geminiviridae includes more than 500 species [4]. The members of all of the genera of the family Geminiviridae except Begomovirus have a single genomic component, while the genus Begomovirus includes viruses with either a single genomic component, referred to as monopartite viruses, or two genomic components (DNA-A and DNA-B), referred to as bipartite viruses (Fig. 1) [2]. The genome of a monopartite begomovirus is equivalent to DNA-A of a bipartite begomovirus. Begomoviruses containing a monopartite genome are widespread in Old World countries in Africa, Asia, Australia, and Europe, whereas bipartite begomoviruses are mostly found in the New World, with some exceptions [7]. The genomic components DNA-A and DNA-B contain ORFs in a bidirectional orientation. The origin of replication (ori), a stem-loop region containing a nonanucleotide sequence (5’-TAATATTAC-3’), and the bidirectional promoter are located in the intergenic regions (IRs) of DNA-A and DNA-B [8]. The DNA-A component of the genome contains seven ORFs, two in the viral sense and five in the complementary sense. Genes oriented in the complementary sense (anticlockwise) such as AC1, AC2, AC3, AC4, and AC5 are responsible for replication and expression of the other viral genes. The AV1 (coding for the coat protein CP) and AV2 genes are in the viral sense (clockwise). The DNA-B component has two genes: BC1 (in the complementary sense, coding for a viral movement protein) and BV1 (in the viral sense, coding for a nuclear shuttle protein). Here, we review our current understanding of the role geminivirus proteins play in pathogenesis and how their interaction with host factors subverts the host defense, the overview of functions of the ORFs represented as in Fig. 2.
Genome organization of ssDNA plant viruses belonging to the family Geminiviridae. The yellow circle at the top represents the origin of replication. All the ORFs are labeled and color-coded. AC1/C1, replication-associated protein (Rep); AC2/C2, transcriptional activator protein (TrAP); AC3/C3, replication enhancer protein (REn); AV1/V1, coat protein; AV2/V2, pre-coat protein; BC1, movement protein (MP); BV1, nuclear shuttle protein (NSP). CR, SCR, and SIR refer to the common region, satellite conserved region, and short intergenic region, respectively. Recently, many small ORFs in the monopartite virus TYLCV and βV1 in a betasatellite were identified. ORFs of members of five recently created genera were identified using the ORF Finder programme (https://www.ncbi.nlm.nih.gov/orffinder/) and are appropriately positioned.
Functional summary of viral proteins associated with the geminivirus disease complexes. A pictorial representation of the components of geminivirus disease complexes and the effect of their encoded proteins on the host is shown. The core genomic components of geminivirus (helper virus or DNA-A, DNA-B) and the well-characterized satellite molecules DNA-α and DNA-β are indicated in different colors at the center. The important roles played by each viral protein and the corresponding genetic factors involved in host manipulation are indicated at the periphery with colors corresponding to their genetic origin.
DNA-A
AV1/V1
The AV1/V1 ORF encodes the only structural protein, the coat protein (CP), which is involved in the assembly and packaging of the geminivirus genome [9,10,11]. Although the CP is not required for geminivirus replication, deletion of the CP gene results in a reduction in viral ssDNA accumulation [12, 13]. In some instances, it has also been observed to elevate the expression of dsDNA [13]. The decrease in ssDNA could be related to the role of CP in packaging of ssDNA, with the lack of a CP causing the ssDNA to be exposed to nucleases, leading to its degradation. The increase in dsDNA hints at its role in the conversion of dsDNA to ssDNA [12, 13].
Since the monopartite viruses lack DNA-B, which encodes proteins that facilitate nucleocytoplasmic shuttling and inter- and intracellular movement for systemic spread, the other ORFs complement the movement function of DNA-B in monopartite viruses. The CP serves as a nucleocytoplasmic shuttling protein in the case of mastreviruses [14, 15] and monopartite begomoviruses [16, 17]. Interestingly, the property of nucleocytoplasmic shuttling is conserved in bipartite begomovirus as well [18, 19]. The CP is also involved in the nucleocytoplasmic shuttling of the viral DNA with the help of the nuclear localization signals (NLSs) present in its N-terminal, C-terminal, and central regions as well [10, 15, 20]. Analysis of the CP from tomato leaf curl Java virus (ToLCJV) revealed that arginine-rich stretches of amino acids at positions 16 to 20 (KVRRR) and 52 to 55 (RKPR) in the N-terminal region are essential for NLS activity and that a hydrophobic stretch in the C-terminal region from residues 245 to 250 (LKIRIY) is important for nuclear export signal (NES) activity [21]. Furthermore, CP interacts with the ssDNA and dsDNA forms of the geminivirus genome via its N-terminal domain [22]. The nucleocytoplasmic shuttling of the viral DNA is also facilitated by the interaction of CP with host proteins involved in nucleocytoplasmic trafficking, such as importin α and karyopherin α1 [16, 23]. In addition to the nuclear shuttling of the viral DNA, the CP of geminiviruses also helps in the systemic movement of viral DNA [12, 13, 15]. It is noteworthy that monopartite viruses require CP for systemic movement, while bipartite begomoviruses can exhibit systemic movement in the absence of the CP because this function is carried out by proteins encoded by DNA-B. However, it is dependent on the host in some cases. For example, in the case of tomato golden mosaic virus (TGMV), the virus without CP can still exhibit systemic movement in Nicotiana benthamiana, but not in Nicotiana tabacum or Datura stramonium, which suggests the possible involvement of host factors in the movement of the virus [24]. Studies have shown, however, that the coat proteins of bipartite begomoviruses are not required for cell-cell movement but can enhance the efficiency of cell-cell movement and systemic spread [18, 24].
The CP is required for vector-mediated transmission of geminiviruses, and it is known to interact with proteins of the vectors and their endosymbionts (Table 1). Mutational studies have demonstrated the importance of the CP and identified amino acids at positions 129, 130, 134 as essential for whitefly-mediated transmission and assembly of virion particles in the case of tomato yellow leaf curl Sardinia virus (TYLCSV) [25, 26]. Amino acids at positions 124, 149, and 174 of abutilon mosaic virus (AbMV) are important for whitefly transmission, and mutations at these positions can enhance vector-mediated transmission [27]. Residues in the CP also determine the specificity of the virus for insect vectors, and altering these residues affects host insect specificity. For example, a T147S substitution appears to be responsible for differences in whitefly transmissibility between the Asia-I and Asia-II-1 strains of squash leaf curl China virus (SLCCNV) [28]. Sometimes, viruses manipulate cellular events in the whitefly. For example, the CP of tomato yellow leaf curl virus (TYLCV) induces apoptosis in whiteflies. Interestingly, suppressing apoptosis reduces the viral titer in whiteflies, while its activation increases the viral titer. The mechanism by which this occurs is not known [29].
AV2/V2
The AV2/V2 gene encodes a pre-coat protein (also known as the AV2 protein). The 5’ end of the AV2/V2 ORF overlaps with the 3’ end of the CP ORF. The AV2 protein of many geminiviruses acts as a pathogenicity determinant. In East African cassava mosaic Cameroon virus (EACMCV), the pathogenicity is dependent on a conserved protein kinase C domain present in AV2 [40]. Mutation of the AV2 protein causes a reduction in the accumulation of ssDNA and dsDNA in plants, but not in protoplasts, indicating that mutation of AV2 impairs the movement of the virus. Furthermore, a localization study using an AV2-GFP fusion confirmed the role of the AV2 protein in geminivirus movement [13, 41]. In TYLCV, the V2 protein, together with exportin α, plays a role in the nuclear export of the coat protein (V1) and promotes the redistribution of V1 in the perinuclear region. Plants infected with TYLCV-V2 with a substitution mutation (V2C85S) that abolishes interaction with V1 exhibited delayed and mild symptoms compared to plants infected with wild-type virus [42]. The AV2 protein also inhibits the hypersensitive response (HR) by interacting with the host factor. TYLCV V2 inhibits HR by interacting with and inhibiting the activity of CYP1, a papain-like cysteine protease [43].
AV2 also acts as an RNAi silencing suppressor of both transcriptional gene silencing (TGS) and post-transcriptional gene silencing (PTGS). TYLCV-V2 suppresses PTGS by interacting with SlSGS3 (suppressor of gene silencing 3) from tomato. The Arabidopsis thaliana homolog, the SGS3 protein (AtSGS3), has already been reported to be involved in the RNA-silencing pathway [44, 45], and TYLCV V2 suppresses TGS by interacting with histone deacetylase 6 (HDA6). In this case, TYLCV V2 prevents the recruitment of the DNA methyltransferase MET1 by HDA6, resulting in hypomethylation of the viral DNA and the successful establishment of infection [46]. The V2 protein of the same virus also disrupts the de novo methylation of viral DNA in cajal bodies by interacting with AGO4 [47]. In another example, V2 from cotton leaf curl Multan virus (CLCuMuV) interacts with N. benthamiana AGO4 and suppresses RNA-dependent DNA methylation (RdDM)-mediated TGS [48]. The V2 from tomato yellow leaf curl China virus (TYLCCNV) suppresses silencing by binding to the 21-nt siRNA duplex and the 24-nt single-stranded siRNA [49]. TGS mediated by hypermethylation of DNA in the viral promoter, on the other hand, has been implicated in the ‘recovery’ of the host plant from geminivirus infection. Tomato leaf curl New Delhi virus (ToLCNDV)-AV2 blocks RDR1-mediated host recovery in tobacco plants. However, tomato leaf curl Gujarat virus (ToLCGV)-AV2 does not affect host recovery [50]. It would be interesting to study this recovery phenomenon in more detail.
AC1/C1
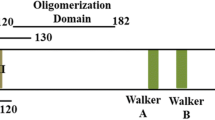
AC1/C1 encodes a replication-associated protein (Rep) that is highly conserved among geminiviruses. Rep is an early protein that is produced following entry into the host cell, and it plays an important role in the replication and transcription of the other viral genes. Rep contains three important domains: an N-terminal domain, a central domain, and a C-terminal domain. The N-terminal domain is essential for DNA binding and DNA nicking activity, the central domain, also referred to as the oligomerization domain, is involved in oligomerization of Rep, and the C-terminal domain is involved in ATPase activity and contains the Walker A and Walker B motifs essential for ATPase activity [51, 52].
Role of Rep in replication initiation, cleavage, and ligation
Rep interacts with many host factors, including retinoblastoma-related protein (RBR), a regulator of cell division; proliferating cell nuclear antigen (PCNA), a DNA clamp and processivity factor of DNA polymerase; replication protein A (RPA), a single-stranded nucleic acid binding protein; replication factor C (RFC), a DNA clamp loader; radiation-activated DNA repair proteins (RAD) such as RAD51 and RAD54, which are indispensable for recombination; and minichromosome maintenance protein 2 (MCM2), a component of the pre-replication complex, all of which help in viral replication [53,54,55]. To initiate replication, Rep binds at two sites in the common region (CR) of the viral genome in a sequence-specific manner. One is in the iteron sequences, and another is in the nonanucleotide sequences. In the case of TGMV and squash leaf curl virus (SqLCV), the CR is a 200-nt region at the origin of replication (ori) that contains a 60-nt stem-loop structure that is essential for replication [56]. Iterons are iterative elements in the CR found between the transcription start site and the TATA promoter of AC1. They can be 8-12 nucleotides in length and can vary among different geminiviruses [57]. In the case of TGMV, Rep binds to a 13-bp region containing 5-bp repeat motifs separated by a 3-bp spacer to initiate replication. Interestingly, the repeat at the 3’ end is necessary for replication, while the repeat at the 5’ end might enhance the replication efficiency of geminiviruses [58]. Similar iterons were also identified in the TYLCV genome, and direct interaction with those regions has been demonstrated [59]. The Rep protein also binds to a hairpin region at the ori site to make a nick at the seventh/eighth residue of the conserved nonanucleotide sequence (5’TAATATTAC3’) in the sense strand to initiate replication. After completion of one round of rolling-circle amplification (RCA), Rep cleaves a phosphodiester bond, leading to the ligation of the end of newly formed nascent DNA. The N-terminus of TYLCV Rep catalyzes the cleavage and ligation of the viral DNA in vitro [60]. Site-directed mutation in the oligomerization domain of TGMV resulted in reduced levels of viral replication. Furthermore, a mutation in the oligomerization domain of TGMV Rep also inhibited viral replication by tenfold as compared to the wild type. This indicates that, in addition to the N-terminal DNA binding domain, the central oligomerization domain of Rep might play a role in viral DNA replication [61]. Geminiviruses replicate inside the nucleus, and Rep contains a nuclear localization signal (NLS). Removal of residues 1-120 of the Rep protein of tomato chlorotic mottle virus (TCMV) results in reduced nuclear accumulation of Rep. Similarly, mutation of the N-terminal residues of African cassava mosaic virus (ACMV) Rep significantly reduces its nuclear import. A recent study has suggested that lysine residues K67, K77, and K101 of TYLCV Rep are important for its nuclear localization [62]. These lysine residues have been shown to interact with E2 SUMO-conjugating enzyme 1 (SCE1), which mediates the sumoylation of lysine residues of the host cell cycle factors PCNA and RBR [63]. This study provides insight into how Rep controls the host cell machinery to facilitate viral replication.
The C-terminal portion of Rep possesses DNA helicase and ATPase activity, and based on domain homology, Rep has been classified as a member of superfamily 3 (SF3). However, Rep differs from other SF3 helicases by the absence of an arginine finger domain and its oligomerization properties [64]. Geminiviral Rep proteins can oligomerize to form complexes ranging from hexamers to dodecamers [65]. ToLCGV Rep oligomers have been shown to be stable even at high salt and low protein concentrations. The mutation K227A at the C-terminus of ToLCGV Rep abolishes its ability to bind to ATP and ssDNA [66], which is a further indication of the structural and functional similarity of geminiviral Rep proteins to SF3 helicases [66]. A recent study has identified conserved amino acids in the B’ motif in the C-terminus of ToLCNDV Rep that are necessary for replication of ToLCNDV. Mutation of these conserved residues negatively impacted the replication of the virus in planta. Interestingly, the mutant variants of the protein affected the helicase activity of Rep without affecting other activities, such as ATPase activity and ssDNA binding activity, that are associated with the C-terminal region of the protein, stressing the significance of helicase activity in geminiviral DNA replication [67].
Role of Rep in geminivirus transcription
Rep is a multifunctional protein that regulates the transcription of Rep and BC1 in addition to its principal role in replication [68]. In TGMV DNA-A, the regulatory rep-binding site essential for replication initiation is closely associated with a sequence that is important for its own transcription. The binding of Rep to this site negatively regulates Rep transcription, which could act as a switch that sets back the repression of early genes and further activation of late genes involved in the cell-to-cell movement. Although the BC1 promoters of TGMV DNA-B share homologous sequence due to sequence conservation, the BC1 transcription remains unaffected [68]. In contrast, Rep represses the active transcription of both Rep and BC1 in ACMV. However, the phenomenon of Rep autoregulation is not observed in members of the genus Curtovirus such as beet curly top virus (BCTV) and beet severe curly top virus (BSCTV) [69]. Interestingly, Rep enhances coat protein expression in the mastrevirus wheat dwarf virus (WDV) [70]. The binding site for autoregulation of its transcript is in a conserved iteron that lies between the transcription start site and the TATA box [71]. Interestingly, the CaMV 35S promoter, which contains the sequence of the Rep binding site is also repressed by Rep [72]. It has been observed that the ability of Rep to function in geminiviral replication and transcript autoregulation are independent events [72]. In the case of TYLCSV C1, a highly conserved RGG motif at position 124-126 in the N-terminal region has been reported to be associated with autoregulation [73]. The significance of autoregulation can be attributed to the expression of the AC2 and AC3 genes, since the transcription start site lies within the coding region of AC1 [69]. It is important to note that AC2 is vital for suppressing host defenses and expression of late genes, and its role will be discussed further in the following sections.
Role of Rep in stimulation of viral transcription
Chilli leaf curl virus (ChiLCV) forms a minichromosome-like structure by interacting with histone proteins, and to enhance viral gene transcription [74], ChiLCV Rep hijacks the host ubiquitin machinery. ChiLCV Rep interacts with ubiquitin-conjugating enzyme 2 (NbUBC2), and histone monoubiquitination1 (NbHUB1) in the nucleus of the host cell [74]. Furthermore, ChiLCV Rep re-localizes NbUBC2 from the cytoplasm to the nucleoplasm. This results in an increase in monoubiquitination of histone 2B (H2B) and trimethylation of histone 3 at lysine 4 (H3K4me3) and subsequently stimulates geminivirus transcription [74].
Role of Rep in pathogenesis
Rep also acts as a silencing suppressor. Rep represses the expression levels of plant DNA maintenance methyltransferases such as methyltransferase 1 (MET1) and chromomethylase 3 (CMT3), thereby interfering with the plant methylation cycle, resulting in a reduction in CG methylation of both the viral genome and host-defense-related genes [75]. Rep is also a target of the plant immune system, as ATG8h, which is important for autophagy, interacts with tomato leaf curl Yunnan virus (TLCYnV) C1 and translocates it to the cytoplasm from the nucleus via exportin 1 to induce autophagy [76]. ChiLCV Rep can also aid in pathogenesis by relocalizing the positive regulator of pathogenesis phosphatidylinositol 4-kinase (PI4K) into the nucleus [77]. In a recent study, a 7-amino-acid stretch was identified at the C-terminus of Sri Lankan cassava mosaic virus-Columbia (SLCMV-Col) that is essential for the accumulation of Rep and is a determinant of the higher virulence of SLCMV-Col when compared to the weaker SLCMV-HN7 strain. Interestingly, the same 7-amino-acid stretch also enhanced the triggering of salicylic acid signaling against SLCMV [78].
AC2/C2
AC2/C2 as transactivator
AC2/C2 encodes a protein referred to as transcriptional activator protein (TrAP), which acts as a central factor in the viral life cycle. TrAP regulates the promoter activity of the viral genes AV1, BV1, and BC1 [79,80,81]. It also interacts with several proteins of geminiviruses, including C3, C4, V2, and βC1 [82]. TrAP has an N-terminal region containing an NLS, a central DNA-binding domain with a zinc finger, and a C-terminal transactivation domain. The transactivation activity of AC2 has been mapped to 15 amino acids at its C-terminal end [83]. AC2 and C2 are examples of position homologs in geminiviruses, because C2, unlike AC2, lacks a transactivation domain and transactivation activity [84]. Computational analysis of 124 bipartite and 463 monopartite begomoviral AC2/C2 proteins has suggested that they have a C-terminal α-helix, like many transcriptional activator proteins (acidic activation domain) [85]. Deletion of TGMV AC2 resulted in reduced coat protein expression, indicating that AC2 is involved in the expression of the coat protein (AV1) [80, 86]. In this case, TrAP activates the TGMV CP promoter by interacting with the transcription factor PEAPOD [87]. In the case of mungbean yellow mosaic India virus (MYMIV), AC2 and AC1 synergistically activate the promoter in DNA-A, driving the expression of CP [88]. C2 of bhendi yellow vein mosaic virus (BYVMV) has an NLS at its N-terminal end comprising amino acids 17-31. The NLS of C2 interacts with karyopherin α, which is involved in the shuttling of molecules between the nucleus and the cytoplasm [89].
AC2/C2 as a silencing suppressor
AC2/C2 from a begomovirus was the first protein reported to have silencing suppressor activity [90]. Transgenic plants overexpressing AC2/C2 show reduced global cytosine methylation of the host genome [91]. BSCTV-C2 reduces the methylation level of the promoter of the genes studied in the plant by reducing the amount of siRNA corresponding to the locus of the methylated promoter [92]. The AC2 proteins of different geminiviruses exhibit varying strength of silencing [93] and employ different mechanisms to suppress the host silencing machinery (Table 2).
AC2/C2 as a pathogenicity determinant
AC2/C2 also plays a role in symptom development, suppression of HR, and inhibition of hormone-mediated defense. The 16-amino-acid hypervariable region in the C-terminus of C2 of tomato yellow leaf curl Sardinia virus (TYLCSV) induces HR [102]. Furthermore, the BYVMV C2 is essential for pathogenicity, whereas a virus with a termination codon in the C2 ORF caused less symptom development and grew to a lower titer. Symptoms were no longer observed when the virus was inoculated with a betasatellite [103]. The HR induced by proteins of geminiviruses is suppressed by the protein encoded by AC2. The HR induced by ToLCNDV NSP is inhibited by AC2 of the same virus [104]. Similarly, the HR induced by V2 of papaya leaf curl virus (PaLCuV) and cotton leaf curl Kokhran virus (CLCuKoV) is countered and suppressed by C2 of PaLCuV and cotton leaf curl Multan virus (CLCuMuV) [105].
Expression of ACMV AC2 in N. tabacum resulted in the upregulation of repressors of the jasmonic acid (JA) signaling pathway [106]. Tomato yellow leaf curl Sardinia virus (TYLCSV) C2 affects JA signaling by interfering with the ubiquitin pathway. TYLSCV C2 interacts with COP9 signalosome 5 (CSN5) and alters the derubylation activity of the CSN complex, which affects downstream signaling pathways such as those of auxin, gibberellic acid (GA), ethylene (ET), salicylic acid (SA), and JA [107]. Transcriptome analysis and challenge inoculation studies in transgenic A. thaliana plants expressing TYLSCV C2 have also suggested that TYLSCV C2 mediates suppression of JA-mediated defense [108]. TYLCV C2, like TYLSCV C2, compromises JA-mediated defense by interacting and altering the plant ubiquitin machinery, resulting in reduced degradation of jasmonate ZIM-domain 1 (JAZ1), a repressor of the JA signaling pathway, facilitating vector infestation in the infected plants [109].
AC3/C3
AC3/C3 encodes a replication enhancer protein (REn). Deletion of AC3 results in a reduction in viral DNA replication and symptom development [86, 110]. This observation is supported by recent evidence of TYLCV C3 recruiting DNA polymerase α and δ in N. benthamiana [111]. Some families of geminiviruses lack C3, and alternative mechanisms might be in place for such viruses. Furthermore, the AC3 from one virus can complement the function of AC3 in a related geminivirus [112]. REn interacts with geminivirus Rep and other replication-associated host factors to enhance viral DNA replication. In fact, REn enhances the ATPase activity of Rep in vitro. In addition, REn can form higher-order homo-oligomers through the hydrophobic domain in the central region of the protein [113, 114]. The hydrophobic region of REn is also essential for the interaction with PCNA, a DNA clamp essential for DNA replication. REn also interacts with RBR via polar residues in its N-terminal and C-terminal domains [114,115,116] and with the transcription factor NAC1 in tomato, leading to increased expression of NAC1. REn and NAC1 localize in the nucleus, and amino acids 1-70, comprising a putative α-helix of REn, are essential for this interaction. Overexpression of NAC1 enhances viral replication [117]. AC3 also enhances gene silencing in some geminiviruses [118].
AC4/C4
This ORF also encodes a multifunctional protein, and proteins from different viruses exhibit differences in their subcellular localization and functions, suggesting an incomplete functional overlap [84].
Role in symptom development and as an oncogenic protein
Deletion of tomato leaf curl virus (ToLCV) C4 leads to reduced symptom development in different hosts, but it does not affect the viral titer [119]. Transgenic N. benthamiana plants expressing BCTV C4 show tissue distortion and development of enations containing a large clustered mass of unorganized cellular material [120]. This can be suppressed by overexpression of pep receptor 2 (PEPR2), a receptor kinase associated with the danger peptide signaling pathway [121]. ACMV C4 also induces abnormal development in transgenic A. thaliana plants, which could partly be attributed to decreased miRNA accumulation [122]. There are many possible mechanisms for the induction of unorganized cell growth, which are discussed below. BSCTV C4 is S-acylated in planta, and S-acylated C4 interacts with CLAVATA1 (CLV1), a receptor kinase, which plays an important role in meristem maintenance. As a consequence, the expression level of WUSCHEL (WUS) is altered, resulting in abnormal development of the plant [123]. Cotyledons and hypocotyledons from transgenic A. thaliana expressing BSCTV C4 under the control of an inducible promoter show extensive cell division with no clear demarcation of vascular bundles [124, 125]. These observations indicate that C4 acts as a viral “oncogene” inhibiting the DNA damage checkpoint, resulting in cell cycle progression while also stimulating DNA replication by preventing programmed cell death [126]. BSCTV C4 expression in A. thaliana results in elevated levels of cell-cycle-associated proteins such as the cyclins cyc1, cyc2, and cyc2b, the cyclin-dependent kinases (CDK) cdc2a, cdc2b, and cdc25, and the cyclin-activated kinases (CAKs) cak1, cak2, and cak3. Some cell cycle inhibitors were also found to be suppressed in transgenic plants expressing C4, and upon BSCTV infection as well [127]. Furthermore, C4 stabilizes CDKs. BSCTV C4 induces the expression of RING finger E3 ligase, which is known to interact with the cell cycle inhibitors ICK and KRP and promote their degradation [128]. Recently, it has been demonstrated that TLCYnV C4 impairs phosphorylation-dependent degradation of CycD1;1 by Shaggy-like kinase (SKη) kinase by relocalizing SKη from the nucleus to the plasma membrane. Reduced phosphorylation-dependent degradation of CycD1;1 leads to the induction of cell division [129].
Role as silencing suppressor
ACMV AC4 synergistically suppresses host PTGS along with EACMCV AC2 and enhances EACMCV DNA accumulation by approximately eightfold [130]. ACMV AC4 interacts with single-stranded siRNA and miRNA, but not with any double-stranded RNA forms, indicating that ACMV AC4 blocks PTGS at the mature stage of small RNA biogenesis [122]. Specific localization of AC4 seems to be essential for its silencing suppressor activity. EACMCV AC4 is localized in the plasma membrane, perinucleus, and cytoplasm. It has a consensus N-myristoylation site, and mutations such as G2A and C3A abolish its plasma membrane localization as well as its silencing suppressor activity [131]. Furthermore, C4 interacts with BAM1 (barely any meristem 1), which is a receptor-like kinase (RLK) as well as a positive regulator of cell-to-cell movement of silencing, and it blocks systemic silencing in the host [132, 133]. MYMV AC4 binds to 21- to 25-nucleotide siRNAs and targets them to the plasma membrane via S-palmitoylation [134]. Together, this indicates that AC4/C4 exhibits silencing suppressor activity by blocking the cell-cell movement of siRNAs. ToLCV-C4 interacts with SKη from tomato in the nucleus region and this interaction has been mapped to 12 amino acids in the C-terminal portion of C4. Deletion of these 12 amino acids abolished this interaction as well as the silencing suppressor activity of C4 [135]. C4 proteins from different viruses show different affinity towards SKη, and interestingly, the symptom-induction property of C4 appears to correlate with its affinity for SKη [136]. Phosphorylation of C4 by SKη and its subsequent myristoylation are also essential for the nucleocytoplasmic shuttling and pathogenicity of C4 in the case of TLCYnV [137]. Like other geminiviral suppressors of RNA silencing (VSRs), CLCuMuV C4 interacts with and inhibits the activity of the essential methyl cycle enzyme S-adenosyl methionine synthetase (SAMS), resulting in suppression of both TGS and PTGS [138]. Using another strategy, TLCYnV C4 suppresses TGS by disrupting self-interaction of N. benthamiana domains rearranged methylase 2 (NbDRM2), a major methyltransferase, which catalyzes the addition of methyl groups on cytosine of viral DNA. As a result, NbDRM2 no longer binds to DNA, resulting in suppression of TGS. Plants infected with TLCYnV C4 harboring an S43A substitution mutation (which has impaired ability to disrupt NbDRM2 interaction) showed higher recovery [139].
Role in systemic movement
There is evidence that C4 complements the function of viral movement. For example, in the case of TYLCV, the systemic movement of TYLCV in tomato was completely blocked as a result of mutation in C4. However, in N. benthamiana, a mutation in TYLCV C4 did not impair the systemic movement of the virus but did result in a reduction in the severity of the symptoms. Similar observations were also made in the case of BSCTV, where a virus with two termination codons in the C4 ORF could replicate in protoplasts and leaf discs but could not establish symptoms in A. thaliana and N. benthamiana plants. The newly emerged leaves from the mutant-virus-infected plants showed no viral DNA accumulation, while the exogenous application of wild-type BSCTV C4 could complement the movement function of mutated BSCTV-C4 [140]. Interestingly, TYLCV C4 can provide the function of movement in the bipartite begomovirus ToLCNDV [141].
Other functions
In addition to the functions of AC4/C4 discussed above, recent reports have demonstrated its ability to suppress HR [142] and SA-mediated defense [143] as well as confer drought stress tolerance independent of abscisic acid in plants [144]. The mechanism of HR suppression involves preventing self-association of the hypersensitive-induced reaction 1 (HIR1) protein and elevating the level of the leucine-rich-repeat (LRR) protein, which promotes degradation of HIR1 [142]. Suppression of SA-mediated plant defense occurs by a mechanism that is conserved among different phytopathogens. C4 translocates from the membrane to the chloroplast, where it suppresses calcium-sensing signal (CAS)-mediated activation of SA signaling. This is supported by the observations that knockout lines of CAS or depletion of SA enhances virus accumulation and can complement C4-null mutations in the virus [143]. The precise mechanism by which C4 confers drought stress tolerance is not known. AC4 of ToLCNDV is also an avirulent gene in the case of the resistant tomato cultivar H-88-78-1, which harbors the resistance gene SlSw5a. SlSw5a interacts with ToLCNDV AC4 to elicit HR and the production of reactive oxygen species to limit the spread of the virus. The amino acid motif “RTSK” present in the C-terminal portion of AC4 is essential for this interaction [145]. The interaction of AC4/C4 with kinases and their significance in silencing suppression is discussed above. A recent finding suggests that TYLCV C4 can also interact with kinases, including NSP-interacting kinase 1 (NIK1), which plays various roles in the pathogenesis of bipartite begomoviruses. The precise mechanism of pathogenesis is described in a later section. The interaction of TYLCV C4 with NIK1 raises the question whether C4 perform the function of the DNA-B-encoded nuclear shuttle protein in monopartite begomovirus, and this will require elaborate experimental studies [146].
Despite its multifunctional nature, this ORF is the least conserved ORF of geminiviruses. The diversity of AC4/C4 provides an adaptive advantage to the virus, since it does not perform any obligatory function in basic processes such as replication, transcription, or encapsidation, suggesting that it is dispensable. However, AC4/C4 supports virus growth by silencing suppression of RNAi, and in viruses without AC4/C4, other proteins such as TrAP, AV2, or Rep might take over this role.
AC5/C5
The AC5/C5 ORF is located downstream of AC3/C3 and overlaps with ORF-AC1 [147,148,149]. The function of the AC5/C5 protein in geminiviruses has not been studied as well like other geminivirus ORFs. AC5 was found to be essential for DNA replication in the case of MYMIV [150]. Deletion of tomato leaf deformation virus (ToLDeV) C5 has been shown to reduce symptom severity [149]. Furthermore, MYMIV AC5 also acts as a silencing suppressor and a pathogenicity determinant. Mutations in MYMIV AC5 have been shown to result in a decrease in infectivity and viral DNA accumulation. AC5 can induce a hypersensitive response when carried by a potato virus X (PVX) vector, but this phenotype has not been observed in MYMIV-infected plants. AC5 also suppresses PTGS, and its N-terminal region is essential for PTGS suppressor activity. Interestingly, AC5 can also suppress TGS by suppressing the expression of domain-rearranged methyltransferase 2 (DRM2), a methyltransferase responsible for de novo CHH methylation and its maintenance. Thirty-three amino acids at the C-terminus of the protein have been shown to be essential for TGS suppressor activity as well as symptom development [176].
DNA-B
After replication, viral DNA must be transported to neighboring cells for successful infection. To achieve this, it must cross the nucleus through a nuclear pore to reach the cytoplasm, and from the cytoplasm it has to move intracellularly to the plasmodesmata to reach uninfected cells. In the case of bipartite viruses, these functions are mediated by proteins encoded by DNA-B. DNA-B contains two ORFs: one on the virion strand, designated as BV1, and one on the complementary strand, designated as BC1.
BV1
The BV1 ORF of DNA-B of bipartite viruses encodes a nuclear shuttle protein (NSP), which is responsible for nucleocytoplasmic shuttling of the viral DNA (vDNA). The NSP has a strong affinity for nucleic acids [151,152,153,154]. The mechanism of nuclear export involves the interaction and cooperation of several host factors. For example, NSP interacts with histone 3 (H3) to form an H3-NSP-viral DNA complex that allows the viral DNA to be exported from the nucleus [155]. The NSP possesses a nuclear export signal (NES), but the exportin has not been identified. The region including amino acids 177-198 in the NSP of SqLCV is leucine-rich. It has been shown that the TFIIIA protein of Xenopus could complement the NES of NSP [156]. The NSP, along with viral DNA, once it reaches the nuclear envelope, is released into the cytoplasm with the help of NIG (NSP-interacting GTPase), a cytosolic GTPase that accumulates around the nuclear envelope in the cytosol. It acts as a cofactor in mediating the intracellular movement of the viral genome. NIG interacts with NSP and shuttles from the nucleus to the cytoplasm [157, 158]. In the cytoplasm, yet another cofactor, NISP (NSP-interacting syntaxin domain-containing protein), a plant-specific syntaxin-6 protein, interacts with the NIG-NSP-vDNA complex and facilitates the intracellular movement of the complex from the cytosol to endosomes [159].
In addition to its role in viral export, BV1 also acts as a pathogenicity determinant. Cabbage leaf curl virus (CaLCuV) NSP suppresses host immunity by inducing expression of the asymmetric leaves 2 (AS2) protein and translocating it out of the nucleus to the cytoplasm. AS2 activates the decapping enzyme DCP2 in the cytoplasm, compromising host defense against the virus [160]. Furthermore, CaLCuV NSP mimics the transcription factor MYC2 and suppresses terpene biosynthesis, making the host more attractive to vectors, thereby increasing the chances that the virus will be transmitted [161]. A 38-amino-acid region close to the NES in CaLCuV NSP interacts with the A. thaliana nuclear shuttle protein interactor (NSI), which plays a role in histone acetylation. The NSP-AtNSI interaction results in acetylation of coat protein bound to viral DNA. Then, the NSP-AtNSI complex replaces CP in the CP-viral DNA complex and exports the viral DNA from the nucleus. Overexpression of AtNSI results in a higher viral titer, while the mutation in NSP that abolishes NSI-AtNSP interaction results in reduced symptom severity and delayed systemic spread of the virus [151, 162, 163]. NSP also interacts with a group of kinases referred to as NIKs. NIK1-3 are leucine-rich repeat receptor-like kinases (LRR-RLK) that localize to membranes. Interaction between NSP and NIK leads to suppression of the kinase activity of NIKs. The loss of function of NIKs supports the replication of CaLCuV. NIK has an 80-amino-acid stretch that contains the putative serine-threonine kinase active site and activation loop, which are the targets of NSP. Loss of NIK1-3 enhances the susceptibility of the host [164]. It has been observed that NIK1 activation leads to phosphorylation and translocation of ribosomal protein 10a (RPL10a) from the cytoplasm to the nucleus [165]. In the nucleus, RLP10a represses global translation by interacting with L10-interacting MYB domain-containing protein (LIMYB). The RPL10-LIMYB interaction leads to suppression of transcription of ribosomal protein genes, resulting in reduced translation of viral proteins [166]. Yet another host factor that interacts with CaLCuV NSP is NsAK (NSP-associated kinase), which is a proline-rich-extension-like receptor protein kinase (PERK). NsAK phosphorylates NSP, and the loss of NsAK results in a decrease in viral titer [167]. ToLCNDV NSP induces a hypersensitive response in N. tabacum and tomato through the N-terminal region of the protein, whereas in N. benthamiana, the same protein induces leaf curling symptoms similar to those associated with viral infection [168].
BC1
BC1 codes for the movement protein (MP), which is responsible for the cell-cell movement of viral DNA. MP increases the size exclusion limit of plasmodesmata of mesophyll cells, thereby facilitating the movement of the virus [152]. MP localizes as small punctuate bodies in the cell periphery and around the nucleus. In the sink leaves of the host, it forms a disc-like structure in the periphery. When BC1 is inoculated with its cognate DNA-A and DNA-B, it forms needle-like structures in sink leaves without altering its subcellular localization [169,170,171,172]. MP interacts with synaptotagmin (SYTA), a regulator of recycling of endosomes to promote transport of the viral genome to plasmodesmata for cell-cell transport through the SYTA-mediated endosomal recycling pathway. The role of SYTA in endosome recycling is evident by the depletion of plasma-membrane-derived endosomes in dominant negative mutant lines [173]. As discussed in the earlier section pertaining to NSP, NISP recruits NSP-NIG-vDNA to endosomes, and from the endosomes, the MP facilitates the transport of the vDNA complex to uninfected neighboring cells through plasmodesmata [174].
BC1 proteins from different viruses differ in their affinity for different forms of DNA. BC1 from SqLCV and bean dwarf mosaic virus (BDMV) has weak affinity for single-stranded DNA, whereas BC1 from MYMIV has strong affinity for single-stranded DNA [153, 154, 175].
Transgenic plants expressing tomato mottle virus (ToMoV) MP and BDMV MP exhibit an abnormal phenotype, with symptoms resembling those induced by viral infection [176, 177]. In one study, plants expressing a mutated form of ToMoV BC1 did not produce viral-like symptoms and showed resistance to ToMoV and CaLCuV infection, possibly because of a transdominant negative effect [177].
Satellite molecules
Monopartite begomoviruses are often associated with satellite DNA molecules referred to as alphasatellites, betasatellites, or, more recently, deltasatellites. Satellite molecules are dependent on helper viruses for their replication and movement and are nearly half the size of the helper virus genome [178, 179].
Alphasatellites
Alphasatellites are approximately 1375 nt in size, with an A-rich region ranging from 150 to 200 nt, a hairpin loop containing the conserved nonanucleotide for replication, and an ORF encoding a Rep protein, referred to as alpha-rep, of approximately 37 kDa in size. Since alphasatellites encode their own Rep, they are not strictly considered satellites. Alphasatellites have not been shown to play a role in symptom development or pathogenicity [179], but they negatively affect the transmission of some helper viruses by whiteflies and affect the DNA level of some betasatellites [180, 181]. Some Rep proteins encoded by alphasatellites such as those of Gossypium darwinii symptomless alphasatellite, Gossypium mustelinum symptomless alphasatellite, and cotton leaf curl Multan alphasatellite, have been reported to exhibit silencing suppressor activity as well [182, 183].
Betasatellites
Unlike alphasatellites, betasatellites are essential for their helper viruses, as they function as pathogenicity determinants. They are known to induce symptoms such as leaf curling, enations, and yellowing [184, 185]. Betasatellites are approximately 1350 nt in size with no sequence similarity to the helper virus other than the conserved nonanucleotide. Betasatellites have a satellite conserved region (SCR), which is highly conserved among betasatellites, an A-rich region, and a single ORF on the complementary strand, encoding a protein referred to as βC1. βC1 is a multitasking protein that plays an essential role during pathogenesis by suppressing host TGS and PTGS, suppressing host defense, and promoting symptom development (Fig. 2) [179, 184, 186]. Covering the work on βC1s merits a separate review, but considering the space limitations, this review will only touch upon its key roles with examples for illustration.
βC1 as a pathogenicity determinant and silencing suppressor
βC1 is a pathogenicity determinant that plays a vital role in viral pathogenesis. It has evolved multiple strategies to mitigate the host defense. It interacts with several host factors to evade host defense mechanisms and support helper viruses in establishing disease [184]. βC1 induces characteristic symptoms in the leaves of the infected host, which is mainly due to the interaction of βC1 with the host factors. βC1 from TYLCCNV interacts with AS1, replacing AS2, and interferes with the normal leaf development. The AS1-AS2 interaction is essential for downregulation of miR165/166 as well as upregulation of the transcription factor HD-ZIP III, which are essential for normal leaf development [187]. βC1 also induces vein clearing or yellowing in veins. βC1 from radish leaf curl betasatellite (RaLCB) has been shown to be localized in chloroplasts, where it alters their ultrastructure as well the expression of chloroplast-encoded proteins, resulting in reduced photosynthesis. Furthermore, RaLCB βC1 interacts with oxygen-evolving enhancer protein 2 (encoded by PsbP) and interferes with its binding to RaLCB DNA, which is essential for plant immunity to the virus, as silencing PsbP transiently increases the viral load [188, 189]. A recent study has suggested that βC1 from tomato leaf curl Patna betasatellite (ToLCPaB) regulates the titer of the helper virus as well as the betasatellite and the transcript accumulation of Rep and βC1 through its ATPase activity. Interestingly, ATPase activity is conserved in βC1 from diverse betasatellites. Unlike the Rep protein, βC1 lacks the canonical Walker A and Walker B motifs that are essential for ATPase activity, and they have a non-canonical ATPase domain that overlaps with the DNA-binding domain [190]. βC1 also interacts with the host ubiquitin system, resulting in symptom development. βC1 interacts with the ubiquitin-conjugating enzyme SlUBC3 in tomato, resulting in reduced polyubiquitination of many target proteins. The myristoylation-like motif GMDVNE at the C-terminus of βC1 is essential for its interaction with SlUBC3, and mutation in this site results in reduced symptom severity [191]. In another example, tomato yellow leaf curl China betasatellite (TYLCCNB) βC1 interacts with a RING-finger protein from tobacco (NtRFP1), which is an E3 ubiquitin ligase that is involved in polyubiquitination of βC1 to promote degradation of βC1 via the 26S proteasomal pathway [192]. It is not clear whether the NtRFP1-βC1 interaction is a survival mechanism of the plant or the virus, but experimental evidence shows that the plant gains immunity against the virus by promoting the proteasomal degradation pathway through NtRFP1 [192]. A recent report also suggested that βC1 undergoes SUMOylation through its SUMO-interacting motif to escape from degradation in the host [193].
βC1 nullifies the antiviral defense of plants by suppressing both TGS and PTGS. βC1 of tomato yellow leaf curl China betasatellite (TYLCCNB) can complement BCTV C2 and reverse TGS by directly interacting with S-adenosylhomocysteine hydroxylase (SAHH), which is an essential component of the methyl cycle, replenishing the methyltransferase cofactor S-adenosyl methionine (SAM). The βC1-SAHH interaction leads to a decrease in SAM [194]. Tyrosine residues at positions 5 and 110 are essential for the reversal of TGS activity [195]. Transgenic plants expressing cotton leaf curl Multan betasatellite (CLCuMuB) βC1 were shown to have elevated levels of both AGO1 and DCL1 gene expression, and the CLCuMuB βC1 protein was found to physically interact with AGO1 [196]. In another mechanism, βC1 suppresses host silencing by upregulating a silencing suppressor. One such silencing suppressor is the calmodulin-like protein rgs-Cam, which is an endogenous suppressor of the gene silencing pathway that suppresses the expression of RDR6. Overexpression of Nbrgs-Cam has been shown to produce a phenotype similar to that caused by βC1 in plants [197].
The TYLCCNB βC1 protein interacts with sucrose non-fermenting-1-related kinase from Solanum lycopersicum (SlSnRK1). SlSnRK1 inactivates βC1-mediated pathogenicity by phosphorylating βC1 at serine 33 and threonine 78. Expression of βC1 with phosphomimic mutants and overexpression of SlSnRK1 result in a delay in viral symptoms as well as a decrease in the viral titer [198]. SlSnRK1 also phosphorylates tyrosine at positions 5 and 110 resulting in milder disease symptoms, a weakened reversal of TGS, and a lack of interaction with AS1 [195].
TYLCCNV, together with TYLCCNB βC1, induces upregulation of NbCaM and NbCaM, which in turn interact with SGS3, resulting in degradation of SGS3, mediated by the phosphatidylinositol 3-kinase complex. This suggests a role in the βC1-dependent autophagy in geminivirus infection [199]. ATG8, an autophagy-related protein, interacts with CLCuMB βC1 in a manner involving a valine residue at position 32. A V32A mutation in βC1 was found to enhance symptom severity and accumulation of viral DNA. Interestingly, silencing of other autophagy-related proteins, ATG5 and ATG7, made plants susceptible to different viruses [200]. A recent study suggested a novel role of TYLCNNB βC1 in which it interferes with the mitogen-activated protein kinase (MAPK or MPK) pathway to mitigate the host defense. It selectively inhibits the activity of MAPK pathway members such as MPK4 and MAPK kinase 2 (MKK2) to suppress the host defense [201].
βC1 in systemic movement
In infections with monopartite begomoviruses, βC1 can also complement functions associated with DNA-B of bipartite viruses. ToLCNDV-A alone can induce local movement but is impaired in the systemic movement of the virus. However, ToLCNDV-A inoculated with CLCuMuB betasatellite can induce both local and systemic movement. Interestingly, ToLCNDV-A, when inoculated with a CLCuMuB betasatellite with a disrupted βC1 gene failed to exhibit systemic movement. Similar observations were made in the case of ageratum yellow vein virus [141, 202]. βC1 possesses an NES or NLS and interacts with the CP of BYVMV and the host nuclear import protein karyopherin α. The failure of βC1 to localize in the nucleus results in a lack of symptom development [203, 204].
Host-vector-virus tripartite interaction
The βC1 protein promotes host-vector-virus tripartite interaction [184] by suppressing JA signaling and promoting emission of the volatile compound linalool [205]. The βC1 protein interferes with AS1/AS2 complex formation [187] as well as interfering with dimerization of the transcription factor MYC2 [161]. Both of these interactions suppress JA-mediated synthesis of terpenes and other metabolites (which repel insects), resulting in the attraction of more insects to infected plants. Apart from increasing the vector activity on infected plants, βC1 can also deter infestation of infected plants by non-vector insects. Such observations have been reported in the case of CLCuMuV and its associated betasatellite, in which βC1 binds to the transcription factor WRKY20 in the phloem to modulate chemical immunity in the host to support propagation of its vector (i.e., whitefly) and deter non-vectors such as cotton bollworms and aphids [206].
Satellite- and DNA-B-encoded proteins share functional homology in several aspects. For instance, both βC1 and DNA-B-encoded-proteins assist in the movement of the viral genome, and βC1 and BV1 mimic other proteins to suppress JA acid signaling to attract whiteflies for transmission. This clearly demonstrates that although they diverged in the process of evolution, these proteins still retain properties that are essential for pathogenesis.
βV1 and other small ORFs
βV1 is a novel ORF discovered very recently encoded on the viral-sense strand of the betasatellite genome. This ORF is conserved in position and sequence in nearly 40% of betasatellite sequences. Intriguingly, it positively affects viral infection by triggering HR in leaves of N. benthamiana [207]. Exploration of genome sequences for small ORFs encoding microproteins has led to the discovery of possible functional roles for small ORFs. An important and interesting study by Gong et al. revealed the presence of six small ORFs in the TYLCV genome. One of these, ORF6, also designated as V3, is conserved among begomoviruses. Interestingly, a TYLCV variant with a mutated V3 exhibited an impaired ability to induce symptoms in the host and was also found to silence both TGS and PTGS suppressors [208].
Conclusions and future directions
Over the past few years, studies have established the multifarious nature of geminivirus proteins and the complexity of their interaction with host factors. These studies have identified interactions that are essential for infection and provided insight into how geminiviruses redirect plant processes and counteract host defense responses [209, 210]. However, structural studies of these viral proteins could shed more light on the mechanism involved in their interaction with host factors. It would also help to identify the protein domains required for the various functions performed by geminivirus proteins. With the recent identification of positional homologs with incomplete functional overlap in proteins such as AC2/C2 and AC4/C4 suggesting that novel functions of these proteins remain to be discovered, one needs to be cautious when predicting the function of an ORF based on its position [84]. Furthermore, the discovery of novel ORFs encoding small proteins such as βV1 and V3 might suggest additional complexity in host-virus interactions. Application of new and emerging techniques in the fields of cell biology, molecular biology, and biochemistry, especially systems biology and CRISPR-Cas9-related techniques, will provide a new dimension and improve our understanding of one of the largest families of plant viruses.
References
Zerbini FM, Briddon RW, Idris A et al (2017) ICTV virus taxonomy profile: geminiviridae. J Gen Virol 98:131–133. https://doi.org/10.1099/jgv.0.000738
Rojas MR, Macedo MA, Maliano MR et al (2018) World management of geminiviruses. Annu Rev Phytopathol 56:637–677. https://doi.org/10.1146/annurev-phyto-080615-100327
Devendran R, Kumar M, Ghosh D et al (2021) Capsicum-infecting begomoviruses as global pathogens: host-virus interplay, pathogenesis, and management. Trends Microbiol. https://doi.org/10.1016/j.tim.2021.05.007
Fiallo-Olivé E, Lett J-M, Martin DP et al (2021) ICTV virus taxonomy profile: geminiviridae 2021. J Gen Virol 102:1–2. https://doi.org/10.1099/jgv.0.001696
Varsani A, Navas-Castillo J, Moriones E et al (2014) Establishment of three new genera in the family Geminiviridae: Becurtovirus, Eragrovirus and Turncurtovirus. Arch Virol 159:2193–2203. https://doi.org/10.1007/s00705-014-2050-2
Varsani A, Roumagnac P, Fuchs M et al (2017) Capulavirus and Grablovirus: two new genera in the family Geminiviridae. Arch Virol 162:1819–1831. https://doi.org/10.1007/s00705-017-3268-6
Brown JK, Zerbini FM, Navas-Castillo J et al (2015) Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch Virol 160:1593–1619. https://doi.org/10.1007/s00705-015-2398-y
Roshan P, Kulshreshtha A, Hallan V (2017) Genome organization of begomoviruses. In: Saxena S, Tiwari AK (eds) Begomoviruses: occurrence and management in Asia and Africa. Springer Singapore, Singapore, pp 11–32
Boulton MI, Pallaghy CK, Chatani M et al (1993) Replication of maize streak virus mutants in maize protoplasts: evidence for a movement protein. Virology 192:85–93. https://doi.org/10.1006/viro.1993.1010
Liu H, Boulton MI, Thomas CL et al (1999) Maize streak virus coat protein is karyophyllic and facilitates nuclear transport of viral DNA. Mol Plant Microbe Interact 12:894–900. https://doi.org/10.1094/MPMI.1999.12.10.894
Briddon RW, Pinner MS, Stanley J, Markham PG (1990) Geminivirus coat protein gene replacement alters insect specificity. Virology 177:85–94. https://doi.org/10.1016/0042-6822(90)90462-Z
Briddon RW, Watts J, Markham PG, Stanley J (1989) The coat protein of beet curly top virus is essential for infectivity. Virology 172:628–633. https://doi.org/10.1016/0042-6822(89)90205-5
Padidam M, Beachy RN, Fauquet CM (1996) The role of AV2 ('precoat’) and coat protein in viral replication and movement in tomato leaf curl geminivirus. Virology 224:390–404. https://doi.org/10.1006/viro.1996.0546
Liu L, Saunders K, Thomas CL et al (1999) Bean yellow dwarf virus RepA, but not rep, binds to maize retinoblastoma protein, and the virus tolerates mutations in the consensus binding motif. Virology 256:270–279. https://doi.org/10.1006/viro.1999.9616
Liu H, Lucy AP, Davies JW, Boulton MI (2001) A single amino acid change in the coat protein of Maize streak virus abolishes systemic infection, but not interaction with viral DNA or movement protein. Mol Plant Pathol 2:223–228. https://doi.org/10.1046/j.1464-6722.2001.00068.x
Kunik T, Mizrachy L, Citovsky V, Gafni Y (1999) Characterization of a tomato karyopherin α that interacts with the Tomato Yellow Leaf Curl Virus (TYLCV) capsid protein1. J Exp Bot 50:731–732. https://doi.org/10.1093/jxb/50.334.731
Rojas MR, Jiang H, Salati R et al (2001) Functional analysis of proteins involved in movement of the monopartite begomovirus, Tomato yellow leaf curl virus. Virology 291:110–125. https://doi.org/10.1006/viro.2001.1194
Ingham DJ, Pascal E, Lazarowitz SG (1995) Both bipartite geminivirus movement proteins define viral host range, but only BL1 determines viral pathogenicity. Virology 207:191–204. https://doi.org/10.1006/viro.1995.1066
Qin S, Ward BM, Lazarowitz SG (1998) The bipartite geminivirus coat protein aids BR1 function in viral movement by affecting the accumulation of viral single-stranded DNA. J Virol 72:9247–9256. https://doi.org/10.1128/JVI.72.11.9247-9256.1998
Unseld S, Höhnle M, Ringel M, Frischmuth T (2001) Subcellular targeting of the coat protein of African cassava mosaic geminivirus. Virology 286:373–383. https://doi.org/10.1006/viro.2001.1003
Sharma P, Ikegami M (2009) Characterization of signals that dictate nuclear/nucleolar and cytoplasmic shuttling of the capsid protein of Tomato leaf curl Java virus associated with DNA beta satellite. Virus Res 144:145–153. https://doi.org/10.1016/j.virusres.2009.04.019
Liu H, Boulton MI, Davies JW (1997) Maize streak virus coat protein binds single- and double-stranded DNA in vitro. J Gen Virol 78(Pt 6):1265–1270. https://doi.org/10.1099/0022-1317-78-6-1265
Guerra-Peraza O, Kirk D, Seltzer V et al (2005) Coat proteins of Rice tungro bacilliform virus and Mungbean yellow mosaic virus contain multiple nuclear-localization signals and interact with importin alpha. J Gen Virol 86:1815–1826. https://doi.org/10.1099/vir.0.80920-0
Pooma W, Petty IT (1996) Tomato golden mosaic virus open reading frame AL4 is genetically distinct from its C4 analogue in monopartite geminiviruses. J Gen Virol 77(Pt 8):1947–1951. https://doi.org/10.1099/0022-1317-77-8-1947
Noris E, Vaira AM, Caciagli P et al (1998) Amino acids in the capsid protein of tomato yellow leaf curl virus that are crucial for systemic infection, particle formation, and insect transmission. J Virol 72:10050–10057. https://doi.org/10.1128/JVI.72.12.10050-10057.1998
Caciagli P, Medina Piles V, Marian D et al (2009) Virion stability is important for the circulative transmission of tomato yellow leaf curl sardinia virus by Bemisia tabaci but virion access to salivary glands does not guarantee transmissibility. J Virol 83:5784–5795. https://doi.org/10.1128/JVI.02267-08
Höhnle M, Höfer P, Bedford ID et al (2001) Exchange of three amino acids in the coat protein results in efficient whitefly transmission of a nontransmissible abutilon mosaic virus isolate. Virology 290:164–171. https://doi.org/10.1006/viro.2001.1140
Pan L-L, Chi Y, Liu C et al (2020) Mutations in the coat protein of a begomovirus result in altered transmission by different species of whitefly vectors. Virus Evol. https://doi.org/10.1093/ve/veaa014
Wang X-R, Wang C, Ban F-X et al (2020) Apoptosis in a whitefly vector activated by a begomovirus enhances viral transmission. mSystems 5:e0043320. https://doi.org/10.1128/mSystems.00433-20
Saurav GK, Rana VS, Popli S et al (2019) A thioredoxin-like protein of Bemisia tabaci interacts with coat protein of begomoviruses. Virus Genes 55:356–367. https://doi.org/10.1007/s11262-019-01657-z
Götz M, Popovski S, Kollenberg M et al (2012) Implication of Bemisia tabaci heat shock protein 70 in begomovirus-whitefly interactions. J Virol 86:13241–13252. https://doi.org/10.1128/JVI.00880-12
Ohnesorge S, Bejarano ER (2009) Begomovirus coat protein interacts with a small heat-shock protein of its transmission vector (Bemisia tabaci). Insect Mol Biol 18:693–703. https://doi.org/10.1111/j.1365-2583.2009.00906.x
Rana VS, Popli S, Saurav GK et al (2016) A Bemisia tabaci midgut protein interacts with begomoviruses and plays a role in virus transmission. Cell Microbiol 18:663–678. https://doi.org/10.1111/cmi.12538
Kanakala S, Ghanim M (2016) Implication of the whitefly Bemisia tabaci cyclophilin b protein in the transmission of tomato yellow leaf curl virus. Front Plant Sci 7:1702. https://doi.org/10.3389/fpls.2016.01702
Gottlieb Y, Zchori-Fein E, Mozes-Daube N et al (2010) The transmission efficiency of tomato yellow leaf curl virus by the whitefly Bemisia tabaci is correlated with the presence of a specific symbiotic bacterium species. J Virol 84:9310–9317. https://doi.org/10.1128/JVI.00423-10
Zhao J, Guo T, Lei T et al (2020) Proteomic analyses of whitefly-begomovirus interactions reveal the inhibitory role of tumorous imaginal discs in viral retention. Front Immunol 11:1596. https://doi.org/10.3389/fimmu.2020.01596
Chi Y, Pan L-L, Liu S-S et al (2021) Implication of the whitefly protein vps twenty associated 1 (vta1) in the transmission of cotton leaf curl multan virus. Microorganisms 9:304
Pan L-L, Chen Q-F, Zhao J-J et al (2017) Clathrin-mediated endocytosis is involved in Tomato yellow leaf curl virus transport across the midgut barrier of its whitefly vector. Virology 502:152–159. https://doi.org/10.1016/j.virol.2016.12.029
Fan Y-Y, Zhong Y-W, Zhao J et al (2021) Bemisia tabaci vesicle-associated membrane protein 2 interacts with begomoviruses and plays a role in virus acquisition. Cells 10:1700
Chowda-Reddy RV, Achenjang F, Felton C et al (2008) Role of a geminivirus AV2 protein putative protein kinase C motif on subcellular localization and pathogenicity. Virus Res 135:115–124. https://doi.org/10.1016/j.virusres.2008.02.014
Rothenstein D, Krenz B, Selchow O, Jeske H (2007) Tissue and cell tropism of Indian cassava mosaic virus (ICMV) and its AV2 (precoat) gene product. Virology 359:137–145. https://doi.org/10.1016/j.virol.2006.09.014
Zhao W, Wu S, Barton E et al (2020) Tomato yellow leaf curl virus v2 protein plays a critical role in the nuclear export of v1 protein and viral systemic infection. Front Microbiol 11:1243. https://doi.org/10.3389/fmicb.2020.01243
Bar-Ziv A, Levy Y, Citovsky V, Gafni Y (2015) The Tomato yellow leaf curl virus (TYLCV) V2 protein inhibits enzymatic activity of the host papain-like cysteine protease CYP1. Biochem Biophys Res Commun 460:525–529. https://doi.org/10.1016/j.bbrc.2015.03.063
Muangsan N, Beclin C, Vaucheret H, Robertson D (2004) Geminivirus VIGS of endogenous genes requires SGS2/SDE1 and SGS3 and defines a new branch in the genetic pathway for silencing in plants. Plant J 38:1004–1014. https://doi.org/10.1111/j.1365-313X.2004.02103.x
Glick E, Zrachya A, Levy Y et al (2008) Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc Natl Acad Sci USA 105:157–161. https://doi.org/10.1073/pnas.0709036105
Wang B, Yang X, Wang Y et al (2018) Tomato yellow leaf curl virus v2 interacts with host histone deacetylase 6 to suppress methylation-mediated transcriptional gene silencing in plants. J Virol 92:e00036-18. https://doi.org/10.1128/jvi.00036-18
Wang L, Ding Y, He L et al (2020) A virus-encoded protein suppresses methylation of the viral genome through its interaction with AGO4 in the Cajal body. Elife 9:e55542. https://doi.org/10.7554/eLife.55542
Wang Y, Wu Y, Gong Q et al (2019) Geminiviral V2 protein suppresses transcriptional gene silencing through interaction with AGO4. J Virol. https://doi.org/10.1128/JVI.01675-18
Zhang J, Dong J, Xu Y, Wu J (2012) V2 protein encoded by Tomato yellow leaf curl China virus is an RNA silencing suppressor. Virus Res 163:51–58. https://doi.org/10.1016/j.virusres.2011.08.009
Basu S, Kumar Kushwaha N, Kumar Singh A et al (2018) Dynamics of a geminivirus-encoded pre-coat protein and host RNA-dependent RNA polymerase 1 in regulating symptom recovery in tobacco. J Exp Bot 69:2085–2102. https://doi.org/10.1093/jxb/ery043
Desbiez C, David C, Mettouchi A et al (1995) Rep protein of tomato yellow leaf curl geminivirus has an ATPase activity required for viral DNA replication. Proc Natl Acad Sci USA 92:5640–5644. https://doi.org/10.1073/pnas.92.12.5640
Orozco BM, Miller AB, Settlage SB, Hanley-Bowdoin L (1997) Functional domains of a geminivirus replication protein. J Biol Chem 272:9840–9846. https://doi.org/10.1074/jbc.272.15.9840
Hanley-Bowdoin L, Settlage SB, Orozco BM et al (2000) Geminiviruses: models for plant DNA replication, transcription, and cell cycle regulation. Crit Rev Biochem Mol Biol 35:105–140
Ruhel R, Chakraborty S (2019) Multifunctional roles of geminivirus encoded replication initiator protein. VirusDisease 30:66–73. https://doi.org/10.1007/s13337-018-0458-0
Rizvi I, Choudhury NR, Tuteja N (2015) Insights into the functional characteristics of geminivirus rolling-circle replication initiator protein and its interaction with host factors affecting viral DNA replication. Arch Virol 160:375–387. https://doi.org/10.1007/s00705-014-2297-7
Lazarowitz SG, Wu LC, Rogers SG, Elmer JS (1992) Sequence-specific interaction with the viral AL1 protein identifies a geminivirus DNA replication origin. Plant Cell 4:799–809. https://doi.org/10.1105/tpc.4.7.799
Argüello-Astorga GR, Guevara-González RG, Herrera-Estrella LR, Rivera-Bustamante RF (1994) Geminivirus replication origins have a group-specific organization of iterative elements: a model for replication. Virology 203:90–100. https://doi.org/10.1006/viro.1994.1458
Fontes EP, Eagle PA, Sipe PS et al (1994) Interaction between a geminivirus replication protein and origin DNA is essential for viral replication. J Biol Chem 269:8459–8465
Akbar Behjatnia SA, Dry IB, Ali Rezaian M (1998) Identification of the replication-associated protein binding domain within the intergenic region of tomato leaf curl geminivirus. Nucleic Acids Res 26:925–931. https://doi.org/10.1093/nar/26.4.925
Laufs J, Traut W, Heyraud F et al (1995) In vitro cleavage and joining at the viral origin of replication by the replication initiator protein of tomato yellow leaf curl virus. Proc Natl Acad Sci USA 92:3879–3883. https://doi.org/10.1073/pnas.92.9.3879
Orozco BM, Kong LJ, Batts LA et al (2000) The multifunctional character of a geminivirus replication protein is reflected by its complex oligomerization properties. J Biol Chem 275:6114–6122. https://doi.org/10.1074/jbc.275.9.6114
Maio F, Arroyo-Mateos M, Bobay BG et al (2019) A lysine residue essential for geminivirus replication also controls nuclear localization of the tomato yellow leaf curl virus rep protein. J Virol. https://doi.org/10.1128/JVI.01910-18
Arroyo-Mateos M, Sabarit B, Maio F et al (2018) Geminivirus replication protein impairs SUMO conjugation of proliferating cellular nuclear antigen at two acceptor sites. J Virol. https://doi.org/10.1128/jvi.00611-18
Clérot D, Bernardi F (2006) DNA helicase activity is associated with the replication initiator protein rep of tomato yellow leaf curl geminivirus. J Virol 80:11322–11330. https://doi.org/10.1128/JVI.00924-06
Choudhury NR, Malik PS, Singh DK et al (2006) The oligomeric Rep protein of Mungbean yellow mosaic India virus (MYMIV) is a likely replicative helicase. Nucleic Acids Res 34:6362–6377. https://doi.org/10.1093/nar/gkl903
George B, Ruhel R, Mazumder M et al (2014) Mutational analysis of the helicase domain of a replication initiator protein reveals critical roles of Lys 272 of the B’ motif and Lys 289 of the β-hairpin loop in geminivirus replication. J Gen Virol 95:1591–1602. https://doi.org/10.1099/vir.0.064923-0
Ruhel R, Mazumder M, Gnanasekaran P et al (2021) Functional implications of residues of the B′ motif of geminivirus replication initiator protein in its helicase activity. FEBS J. https://doi.org/10.1111/febs.16053
Sunter G, Hartitz MD, Bisaro DM (1993) Tomato golden mosaic virus leftward gene expression: autoregulation of geminivirus replication protein. Virology 195:275–280. https://doi.org/10.1006/viro.1993.1374
Hur J, Buckley K, Lee S, Davis K (2007) Transcriptional activator elements for curtovirus C1 expression reside in the 3’ coding region of ORF C1. Mol Cells 23:80–87
Hofer JM, Dekker EL, Reynolds HV et al (1992) Coordinate regulation of replication and virion sense gene expression in wheat dwarf virus. Plant Cell 4:213–223. https://doi.org/10.1105/tpc.4.2.213
Eagle PA, Orozco BM, Hanley-Bowdoin L (1994) A DNA sequence required for geminivirus replication also mediates transcriptional regulation. Plant Cell 6:1157–1170. https://doi.org/10.1105/tpc.6.8.1157
Shung CY, Sunter G (2007) AL1-dependent repression of transcription enhances expression of Tomato golden mosaic virus AL2 and AL3. Virology 364:112–122. https://doi.org/10.1016/j.virol.2007.03.006
Sardo L, Lucioli A, Tavazza M et al (2011) An RGG sequence in the replication-associated protein (Rep) of Tomato yellow leaf curl Sardinia virus is involved in transcriptional repression and severely impacts resistance in Rep-expressing plants. J Gen Virol 92:204–209. https://doi.org/10.1099/vir.0.025817-0
Kushwaha NK, Bhardwaj M, Chakraborty S (2017) The replication initiator protein of a geminivirus interacts with host monoubiquitination machinery and stimulates transcription of the viral genome. PLoS Pathog 13:e1006587. https://doi.org/10.1371/journal.ppat.1006587
Rodríguez-Negrete E, Lozano-Durán R, Piedra-Aguilera A et al (2013) Geminivirus Rep protein interferes with the plant DNA methylation machinery and suppresses transcriptional gene silencing. New Phytol 199:464–475. https://doi.org/10.1111/nph.12286
Li F, Zhang M, Zhang C, Zhou X (2020) Nuclear autophagy degrades a geminivirus nuclear protein to restrict viral infection in solanaceous plants. New Phytol 225:1746–1761. https://doi.org/10.1111/nph.16268
Kushwaha NK, Singh AK et al (2019) Nicotiana benthamiana phosphatidylinositol 4-kinase type II regulates chilli leaf curl virus pathogenesis. Mol Plant Pathol 20:1408–1424. https://doi.org/10.1111/mpp.12846
Wang D, Zhang X, Yao X et al (2020) A 7-amino-acid motif of rep protein essential for virulence is critical for triggering host defense against Sri Lankan cassava mosaic virus. Mol Plant Microbe Interact 33:78–86. https://doi.org/10.1094/MPMI-06-19-0163-FI
Haley A, Zhan X, Richardson K et al (1992) Regulation of the activities of African cassava mosaic virus promoters by the AC1, AC2, and AC3 gene products. Virology 188:905–909. https://doi.org/10.1016/0042-6822(92)90551-y
Sunter G, Bisaro DM (1991) Transactivation in a geminivirus: AL2 gene product is needed for coat protein expression. Virology 180:416–419. https://doi.org/10.1016/0042-6822(91)90049-H
Sunter G, Bisaro DM (1992) Transactivation of geminivirus AR1 and BR1 gene expression by the viral AL2 gene product occurs at the level of transcription. Plant Cell 4:1321–1331. https://doi.org/10.1105/tpc.4.10.1321
Li H, Li F, Zhang M et al (2020) Dynamic subcellular localization, accumulation, and interactions of proteins from tomato yellow leaf curl china virus and its associated betasatellite. Front Plant Sci 11:840. https://doi.org/10.3389/fpls.2020.00840
Hartitz MD, Sunter G, Bisaro DM (1999) The tomato golden mosaic virus transactivator (TrAP) is a single- stranded DNA and zinc-binding phosphoprotein with an acidic activation domain. Virology 263:1–14. https://doi.org/10.1006/viro.1999.9925
Luna AP, Lozano-Durán R (2020) Geminivirus-encoded proteins: not all positional homologs are made equal. Front Microbiol. https://doi.org/10.3389/fmicb.2020.00878
Babu KSD, Manoharan P, Pandi G (2018) Computational studies on Begomoviral AC2/C2 proteins. Bioinformation 14:294–303. https://doi.org/10.6026/97320630014294
Sunter G, Hartitz MD, Hormuzdi SG et al (1990) Genetic analysis of tomato golden mosaic virus: ORF AL2 is required for coat protein accumulation while ORF AL3 is necessary for efficient DNA replication. Virology 179:69–77. https://doi.org/10.1016/0042-6822(90)90275-V
Lacatus G, Sunter G (2009) The Arabidopsis PEAPOD2 transcription factor interacts with geminivirus AL2 protein and the coat protein promoter. Virology 392:196–202. https://doi.org/10.1016/j.virol.2009.07.004
Shivaprasad PV, Akbergenov R, Trinks D et al (2005) Promoters, transcripts, and regulatory proteins of mungbean yellow mosaic geminivirus. J Virol 79:8149–8163. https://doi.org/10.1128/JVI.79.13.8149-8163.2005
Chandran SA, Levy Y, Mett A et al (2012) Mapping of functional region conferring nuclear localization and karyopherin α-binding activity of the C2 protein of bhendi yellow vein mosaic virus. J Gen Virol 93:1367–1374. https://doi.org/10.1099/vir.0.038943-0
Voinnet O, Pinto YM, Baulcombe DC (1999) Suppression of gene silencing: a general strategy used by diverse DNA and RNA viruses of plants. Proc Natl Acad Sci 96:14147–14152. https://doi.org/10.1073/pnas.96.24.14147
Buchmann RC, Asad S, Wolf JN et al (2009) Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. J Virol 83:5005–5013. https://doi.org/10.1128/JVI.01771-08
Yang L-P, Fang Y-Y, An C-P et al (2013) C2-mediated decrease in DNA methylation, accumulation of siRNAs, and increase in expression for genes involved in defense pathways in plants infected with beet severe curly top virus. Plant J 73:910–917. https://doi.org/10.1111/tpj.12081
Sundaresan G, Das SS, Tripathi A et al (2020) Evaluating the strength of RNA silencing suppressor proteins encoded by two geminiviruses using assay based on reversal of GFP silencing. Australas Plant Pathol 49:95–106. https://doi.org/10.1007/s13313-019-00678-4
Wang H, Hao L, Shung C-Y et al (2003) Adenosine kinase is inactivated by geminivirus al2 and l2 proteins. Plant Cell 15:3020–3032. https://doi.org/10.1105/tpc.015180
Wang H, Buckley KJ, Yang X et al (2005) Adenosine kinase inhibition and suppression of RNA silencing by geminivirus AL2 and L2 proteins. J Virol 79:7410–7418. https://doi.org/10.1128/jvi.79.12.7410-7418.2005
Hao L, Wang H, Sunter G, Bisaro DM (2003) Geminivirus AL2 and L2 Proteins Interact with and Inactivate SNF1 Kinase. Plant Cell 15:1034–1048. https://doi.org/10.1105/tpc.009530
Trinks D, Rajeswaran R, Shivaprasad PV et al (2005) Suppression of RNA silencing by a geminivirus nuclear protein, AC2, correlates with transactivation of host genes. J Virol 79:2517–2527. https://doi.org/10.1128/JVI.79.4.2517-2527.2005
Tu Y-C, Tsai W-S, Wei J-Y et al (2017) The C2 protein of tomato leaf curl Taiwan virus is a pathogenicity determinant that interferes with expression of host genes encoding chromomethylases. Physiol Plant 161:515–531. https://doi.org/10.1111/ppl.12615
Castillo-González C, Liu X, Huang C et al (2015) Geminivirus-encoded TrAP suppressor inhibits the histone methyltransferase SUVH4/KYP to counter host defense. Elife 4:e06671. https://doi.org/10.7554/eLife.06671
Kumar V, Mishra SK, Rahman J et al (2015) Mungbean yellow mosaic Indian virus encoded AC2 protein suppresses RNA silencing by inhibiting Arabidopsis RDR6 and AGO1 activities. Virology 486:158–172. https://doi.org/10.1016/j.virol.2015.08.015
Zhang Z, Chen H, Huang X et al (2011) BSCTV C2 attenuates the degradation of SAMDC1 to suppress DNA methylation-mediated gene silencing in Arabidopsis. Plant Cell 23:273–288. https://doi.org/10.1105/tpc.110.081695
Matić S, Pegoraro M, Noris E (2016) The C2 protein of tomato yellow leaf curl Sardinia virus acts as a pathogenicity determinant and a 16-amino acid domain is responsible for inducing a hypersensitive response in plants. Virus Res 215:12–19. https://doi.org/10.1016/j.virusres.2016.01.014
Chandran SA, Jeyabharathy C, Usha R (2014) The C2 protein of Bhendi yellow vein mosaic virus plays an important role in symptom determination and virus replication. Virus Genes 48:203–207. https://doi.org/10.1007/s11262-013-0992-1
Hussain M, Mansoor S, Iram S et al (2007) The hypersensitive response to tomato leaf curl New Delhi virus nuclear shuttle protein is inhibited by transcriptional activator protein. Mol Plant Microbe Interact 20:1581–1588. https://doi.org/10.1094/MPMI-20-12-1581
Mubin M, Amin I, Amrao L et al (2010) The hypersensitive response induced by the V2 protein of a monopartite begomovirus is countered by the C2 protein. Mol Plant Pathol 11:245–254. https://doi.org/10.1111/j.1364-3703.2009.00601.x
Soitamo AJ, Jada B, Lehto K (2012) Expression of geminiviral AC2 RNA silencing suppressor changes sugar and jasmonate responsive gene expression in transgenic tobacco plants. BMC Plant Biol 12:204. https://doi.org/10.1186/1471-2229-12-204
Lozano-Durán R, Rosas-Díaz T, Gusmaroli G et al (2011) Geminiviruses subvert ubiquitination by altering CSN-mediated derubylation of SCF E3 ligase complexes and inhibit jasmonate signaling in Arabidopsis thaliana. Plant Cell 23:1014–1032. https://doi.org/10.1105/tpc.110.080267
Rosas-Díaz T, Macho AP, Beuzón CR et al (2016) The C2 protein from the geminivirus tomato yellow leaf curl sardinia virus decreases sensitivity to jasmonates and suppresses jasmonate-mediated defences. Plants (Basel, Switzerland) 5:8. https://doi.org/10.3390/plants5010008
Li P, Liu C, Deng W-H et al (2019) Plant begomoviruses subvert ubiquitination to suppress plant defenses against insect vectors. PLoS Pathog 15:e1007607. https://doi.org/10.1371/journal.ppat.1007607
Etessami P, Saunders K, Watts J, Stanley J (1991) Mutational analysis of complementary-sense genes of African cassava mosaic virus DNA A. J Gen Virol 72(Pt 5):1005–1012. https://doi.org/10.1099/0022-1317-72-5-1005
Wu M, Wei H, Tan H et al (2021) Plant DNA polymerases α and δ mediate replication of geminiviruses. Nat Commun 12:2780. https://doi.org/10.1038/s41467-021-23013-2
Sunter G, Stenger DC, Bisaro DM (1994) Heterologous complementation by geminivirus AL2 and AL3 genes. Virology 203:203–210. https://doi.org/10.1006/viro.1994.1477
Pasumarthy KK, Choudhury NR, Mukherjee SK (2010) Tomato leaf curl Kerala virus (ToLCKeV) AC3 protein forms a higher order oligomer and enhances ATPase activity of replication initiator protein (Rep/AC1). Virol J. https://doi.org/10.1186/1743-422X-7-128
Settlage SB, Miller AB, Hanley-Bowdoin L (1996) Interactions between geminivirus replication proteins. J Virol 70:6790–6795. https://doi.org/10.1128/JVI.70.10.6790-6795.1996
Settlage SB, Miller AB, Gruissem W, Hanley-Bowdoin L (2001) Dual interaction of a geminivirus replication accessory factor with a viral replication protein and a plant cell cycle regulator. Virology 279:570–576. https://doi.org/10.1006/viro.2000.0719
Settlage SB, See RG, Hanley-Bowdoin L (2005) Geminivirus C3 protein: replication enhancement and protein interactions. J Virol 79:9885–9895. https://doi.org/10.1128/JVI.79.15.9885-9895.2005
Selth LA, Dogra SC, Saif Rasheed M et al (2005) A NAC domain protein interacts with tomato leaf curl virus replication accessory protein and enhances viral replication. Plant Cell 17:311–325. https://doi.org/10.1105/tpc.104.027235
Pasumarthy KK, Mukherjee SK, Choudhury NR (2011) The presence of tomato leaf curl Kerala virus AC3 protein enhances viral DNA replication and modulates virus induced gene-silencing mechanism in tomato plants. Virol J. https://doi.org/10.1186/1743-422X-8-178
Rigden JE, Krake LR, Rezaian MA, Dry IB (1994) ORF C4 of tomato leaf curl geminivirus is a determinant of symptom severity. Virology 204:847–850. https://doi.org/10.1006/viro.1994.1606
Latham JR, Saunders K, Pinner MS, Stanley J (1997) Induction of plant cell division by beet curly top virus gene C4. Plant J 11:1273–1283. https://doi.org/10.1046/j.1365-313X.1997.11061273.x
Zeng R, Liu X, Li H et al (2020) Danger peptide signaling enhances internalization of a geminivirus symptom determinant in plant cells during infection. J Exp Bot 71:2817–2827. https://doi.org/10.1093/jxb/eraa053
Chellappan P, Vanitharani R, Fauquet CM (2005) MicroRNA-binding viral protein interferes with Arabidopsis development. Proc Natl Acad Sci U S A 102:10381–10386. https://doi.org/10.1073/pnas.0504439102
Li H, Zeng R, Chen Z et al (2018) S-acylation of a geminivirus C4 protein is essential for regulating the CLAVATA pathway in symptom determination. J Exp Bot 69:4459–4468. https://doi.org/10.1093/jxb/ery228
Mills-Lujan K, Andrews DL, Chou C, Deom CM (2015) The Roles of phosphorylation and shaggy-like protein kinases in geminivirus c4 protein induced hyperplasia. PLoS One 10:e0122356. https://doi.org/10.1371/journal.pone.0122356
Mills-Lujan K, Deom CM (2010) Geminivirus C4 protein alters Arabidopsis development. Protoplasma 239:95–110. https://doi.org/10.1007/s00709-009-0086-z
Nikitin PA, Luftig MA (2012) The DNA damage response in viral-induced cellular transformation. Br J Cancer 106:429–435. https://doi.org/10.1038/bjc.2011.612
Park J, Hwang H-S, Buckley KJ et al (2010) C4 protein of Beet severe curly top virus is a pathomorphogenetic factor in Arabidopsis. Plant Cell Rep 29:1377–1389. https://doi.org/10.1007/s00299-010-0923-8
Lai J, Chen H, Teng K et al (2009) RKP, a RING finger E3 ligase induced by BSCTV C4 protein, affects geminivirus infection by regulation of the plant cell cycle. Plant J 57:905–917. https://doi.org/10.1111/j.1365-313X.2008.03737.x
Mei Y, Yang X, Huang C et al (2018) Tomato leaf curl Yunnan virus-encoded C4 induces cell division through enhancing stability of Cyclin D 1.1 via impairing NbSKη -mediated phosphorylation in Nicotiana benthamiana. PLOS Pathog 14:e1006789
Vanitharani R, Chellappan P, Pita JS, Fauquet CM (2004) Differential roles of AC2 and AC4 of cassava geminiviruses in mediating synergism and suppression of posttranscriptional gene silencing. J Virol 78:9487–9498. https://doi.org/10.1128/jvi.78.17.9487-9498.2004
Fondong VN, Reddy RV, Lu C et al (2007) The consensus N-myristoylation motif of a geminivirus AC4 protein is required for membrane binding and pathogenicity. Mol Plant Microbe Interact 20:380–391. https://doi.org/10.1094/mpmi-20-4-0380
Rosas-Diaz T, Zhang D, Fan P et al (2018) A virus-targeted plant receptor-like kinase promotes cell-to-cell spread of RNAi. Proc Natl Acad Sci 115:1388–1393. https://doi.org/10.1073/pnas.1715556115
Li Z, Du Z, Tang Y et al (2020) C4, the pathogenic determinant of tomato leaf curl guangdong virus, may suppress post-transcriptional gene silencing by interacting with BAM1 protein. Front Microbiol 11:851. https://doi.org/10.3389/fmicb.2020.00851
Carluccio AV, Prigigallo MI, Rosas-Diaz T et al (2018) S-acylation mediates Mungbean yellow mosaic virus AC4 localization to the plasma membrane and in turns gene silencing suppression. PLoS Pathog 14:e1007207. https://doi.org/10.1371/journal.ppat.1007207
Dogra SC, Eini O, Rezaian MA, Randles JW (2009) A novel shaggy-like kinase interacts with the Tomato leaf curl virus pathogenicity determinant C4 protein. Plant Mol Biol 71:25–38. https://doi.org/10.1007/s11103-009-9506-x
Mei Y, Zhang F, Wang M et al (2020) Divergent symptoms caused by geminivirus-encoded C4 proteins correlate with their ability to bind NbSKη. J Virol. https://doi.org/10.1128/jvi.01307-20
Mei Y, Wang Y, Hu T et al (2018) Nucleocytoplasmic shuttling of geminivirus C4 protein mediated by phosphorylation and myristoylation is critical for viral pathogenicity. Mol Plant 11:1466–1481. https://doi.org/10.1016/j.molp.2018.10.004
Ismayil A, Haxim Y, Wang Y et al (2018) Cotton Leaf Curl Multan virus C4 protein suppresses both transcriptional and post-transcriptional gene silencing by interacting with SAM synthetase. PLoS Pathog 14:e1007282. https://doi.org/10.1371/journal.ppat.1007282
Mei Y, Wang Y, Li F, Zhou X (2020) The C4 protein encoded by tomato leaf curl Yunnan virus reverses transcriptional gene silencing by interacting with NbDRM2 and impairing its DNA-binding ability. PLoS Pathog 16:e1008829. https://doi.org/10.1371/journal.ppat.1008829
Teng K, Chen H, Lai J et al (2010) Involvement of C4 protein of beet severe curly top virus (family Geminiviridae) in virus movement. PLoS One 5:e11280. https://doi.org/10.1371/journal.pone.0011280
Saeed M, Zafar Y, Randles JW, Rezaian MA (2007) A monopartite begomovirus-associated DNA beta satellite substitutes for the DNA B of a bipartite begomovirus to permit systemic infection. J Gen Virol 88:2881–2889. https://doi.org/10.1099/vir.0.83049-0
Mei Y, Ma Z (2020) Geminivirus C4 antagonizes the HIR1-mediated hypersensitive response by inhibiting the HIR1 self-interaction and promoting degradation of the protein. New Phytol 225:1311–1326. https://doi.org/10.1111/nph.16208
Medina-Puche L, Tan H, Dogra V et al (2020) A defense pathway linking plasma membrane and chloroplasts and co-opted by pathogens. Cell 182:1109-1124.e25. https://doi.org/10.1016/j.cell.2020.07.020
Corrales-Gutierrez M, Medina-Puche L, Yu Y et al (2020) The C4 protein from the geminivirus Tomato yellow leaf curl virus confers drought tolerance in Arabidopsis through an ABA-independent mechanism. Plant Biotechnol J 18:1121–1123. https://doi.org/10.1111/pbi.13280
Sharma N, Sahu PP, Prasad A et al (2021) The Sw5a gene confers resistance to ToLCNDV and triggers an HR response after direct AC4 effector recognition. Proc Natl Acad Sci 118:e2101833118. https://doi.org/10.1073/pnas.2101833118
Garnelo Gómez B, Zhang D, Rosas-Díaz T et al (2019) The C4 protein from tomato yellow leaf curl virus can broadly interact with plant receptor-like kinases. Viruses 11:1009
Fontenelle MR, Luz DF, Gomes APS et al (2007) Functional analysis of the naturally recombinant DNA-A of the bipartite begomovirus Tomato chlorotic mottle virus. Virus Res 126:262–267. https://doi.org/10.1016/j.virusres.2007.02.009
Kheyr-Pour A, Bananej K, Dafalla GA et al (2000) Watermelon chlorotic stunt virus from the Sudan and Iran: sequence comparisons and identification of a whitefly-transmission determinant. Phytopathology® 90:629–635. https://doi.org/10.1094/PHYTO.2000.90.6.629
Melgarejo TA, Kon T, Rojas MR et al (2013) Characterization of a new world monopartite begomovirus causing leaf curl disease of tomato in Ecuador and Peru reveals a new direction in geminivirus evolution. J Virol 87:5397–5413. https://doi.org/10.1128/JVI.00234-13
Raghavan V, Malik PS, Choudhury NR, Mukherjee SK (2004) The DNA-A component of a plant geminivirus (Indian mung bean yellow mosaic virus) replicates in budding yeast cells. J Virol 78:2405–2413. https://doi.org/10.1128/jvi.78.5.2405-2413.2004
McGarry RC, Barron YD, Carvalho MF et al (2003) A novel Arabidopsis acetyltransferase interacts with the geminivirus movement protein NSP. Plant Cell 15:1605–1618. https://doi.org/10.1105/tpc.012120
Noueiry AO, Lucas WJ, Gilbertson RL (1994) Two proteins of a plant DNA virus coordinate nuclear and plasmodesmal transport. Cell 76:925–932. https://doi.org/10.1016/0092-8674(94)90366-2
Rojas MR, Noueiry AO, Lucas WJ, Gilbertson RL (1998) Bean dwarf mosaic geminivirus movement proteins recognize DNA in a form- and size-specific manner. Cell 95:105–113. https://doi.org/10.1016/s0092-8674(00)81786-9
Pascal E, Sanderfoot AA, Ward BM et al (1994) The geminivirus BR1 movement protein binds single-stranded DNA and localizes to the cell nucleus. Plant Cell 6:995–1006. https://doi.org/10.1105/tpc.6.7.995
Zhou Y, Rojas MR, Park M-R et al (2011) Histone H3 interacts and colocalizes with the nuclear shuttle protein and the movement protein of a geminivirus. J Virol 85:11821–11832. https://doi.org/10.1128/JVI.00082-11
Ward BM, Lazarowitz SG (1999) Nuclear export in plants. Use of geminivirus movement proteins for a cell-based export assay. Plant Cell 11:1267–1276. https://doi.org/10.1105/tpc.11.7.1267
Carvalho CM, Machado JPB, Zerbini FM, Fontes EPB (2008) NSP-Interacting GTPase. Plant Signal Behav 3:752–754. https://doi.org/10.4161/psb.3.9.6641
Carvalho CM, Fontenelle MR, Florentino LH et al (2008) A novel nucleocytoplasmic traffic GTPase identified as a functional target of the bipartite geminivirus nuclear shuttle protein. Plant J 55:869–880. https://doi.org/10.1111/j.1365-313X.2008.03556.x
Gouveia-Mageste BC, Martins LGC, Dal-Bianco M et al (2021) A plant-specific syntaxin-6 protein contributes to the intracytoplasmic route for the begomovirus CabLCV. Plant Physiol 187:158–173. https://doi.org/10.1093/plphys/kiab252
Ye J, Yang J, Sun Y et al (2015) Geminivirus activates ASYMMETRIC LEAVES 2 to accelerate cytoplasmic DCP2-mediated mRNA turnover and weakens RNA silencing in Arabidopsis. PLoS Pathog 11:e1005196. https://doi.org/10.1371/journal.ppat.1005196
Li R, Weldegergis BT, Li J et al (2014) Virulence factors of geminivirus interact with myc2 to subvert plant resistance and promote vector performance. Plant Cell 26:4991–5008. https://doi.org/10.1105/tpc.114.133181
Carvalho MF, Lazarowitz SG (2004) Interaction of the movement protein NSP and the ArabidopsisAcetyltransferase AtNSI Is necessary for cabbage leaf curl geminivirus infection and pathogenicity. J Virol 78:11161–11171. https://doi.org/10.1128/jvi.78.20.11161-11171.2004
Carvalho MF, Turgeon R, Lazarowitz SG (2006) The geminivirus nuclear shuttle protein NSP inhibits the activity of AtNSI, a vascular-expressed arabidopsis acetyltransferase regulated with the sink-to-source transition. Plant Physiol 140:1317–1330. https://doi.org/10.1104/pp.105.075556
Fontes EPB, Santos AA, Luz DF et al (2004) The geminivirus nuclear shuttle protein is a virulence factor that suppresses transmembrane receptor kinase activity. Genes Dev 18:2545–2556. https://doi.org/10.1101/gad.1245904
Carvalho CM, Santos AA, Pires SR et al (2008) Regulated nuclear trafficking of rpL10A mediated by NIK1 represents a defense strategy of plant cells against virus. PLoS Pathog 4:e1000247. https://doi.org/10.1371/journal.ppat.1000247
Zorzatto C, Machado JP, Lopes KV et al (2015) NIK1-mediated translation suppression functions as a plant antiviral immunity mechanism. Nature 520:679–682. https://doi.org/10.1038/nature14171
Florentino LH, Santos AA, Fontenelle MR et al (2006) A PERK-like receptor kinase interacts with the geminivirus nuclear shuttle protein and potentiates viral infection. J Virol 80:6648–6656. https://doi.org/10.1128/JVI.00173-06
Hussain M, Mansoor S, Iram S et al (2005) The nuclear shuttle protein of Tomato leaf curl New Delhi virus is a pathogenicity determinant. J Virol 79:4434–4439. https://doi.org/10.1128/JVI.79.7.4434-4439.2005
Zhang SC, Wege C, Jeske H (2001) Movement proteins (BC1 and BV1) of Abutilon mosaic geminivirus are cotransported in and between cells of sink but not of source leaves as detected by green fluorescent protein tagging. Virology 290:249–260. https://doi.org/10.1006/viro.2001.1185
Frischmuth S, Wege C, Hülser D, Jeske H (2007) The movement protein BC1 promotes redirection of the nuclear shuttle protein BV1 of Abutilon mosaic geminivirus to the plasma membrane in fission yeast. Protoplasma 230:117–123. https://doi.org/10.1007/s00709-006-0223-x
Zhan B, Zhao W, Li S et al (2018) Functional scanning of apple geminivirus proteins as symptom determinants and suppressors of posttranscriptional gene silencing. Viruses 10:488. https://doi.org/10.3390/v10090488
Zhang SC, Ghosh R, Jeske H (2002) Subcellular targeting domains of Abutilon mosaic geminivirus movement protein BC1. Arch Virol 147:2349–2363. https://doi.org/10.1007/s00705-002-0880-9
Lewis JD, Lazarowitz SG (2010) Arabidopsis synaptotagmin SYTA regulates endocytosis and virus movement protein cell-to-cell transport. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.0909080107
Martins LGC, Raimundo GAS, Ribeiro NGA et al (2020) A begomovirus nuclear shuttle protein-interacting immune Hub: Hijacking host transport activities and suppressing incompatible functions. Front Plant Sci 11:398
Radhakrishnan GK, Splitter GA, Usha R (2008) DNA recognition properties of the cell-to-cell movement protein (MP) of soybean isolate of Mungbean yellow mosaic India virus (MYMIV-Sb). Virus Res 131:152–159. https://doi.org/10.1016/j.virusres.2007.09.002
Hou YM, Sanders R, Ursin VM, Gilbertson RL (2000) Transgenic plants expressing geminivirus movement proteins: abnormal phenotypes and delayed infection by Tomato mottle virus in transgenic tomatoes expressing the Bean dwarf mosaic virus BV1 or BC1 proteins. Mol Plant Microbe Interact 13:297–308. https://doi.org/10.1094/MPMI.2000.13.3.297
Duan Y-P, Powell CA, Webb SE et al (1997) geminivirus resistance in transgenic tobacco expressing mutated BC1 protein. Mol Plant-Microbe Interact 10:617–623. https://doi.org/10.1094/MPMI.1997.10.5.617
Fiallo-Olivé E, Martínez-Zubiaur Y, Moriones E, Navas-Castillo J (2012) A novel class of DNA satellites associated with New World begomoviruses. Virology 426:1–6. https://doi.org/10.1016/j.virol.2012.01.024
Zhou X (2013) Advances in understanding begomovirus satellites. Annu Rev Phytopathol 51:357–381. https://doi.org/10.1146/annurev-phyto-082712-102234
Vinoth Kumar R, Singh D, Singh AK, Chakraborty S (2017) Molecular diversity, recombination and population structure of alphasatellites associated with begomovirus disease complexes. Infect Genet Evol 49:39–47. https://doi.org/10.1016/j.meegid.2017.01.001
Mar TB, Mendes IR, Lau D et al (2017) Interaction between the New World begomovirus Euphorbia yellow mosaic virus and its associated alphasatellite: effects on infection and transmission by the whitefly Bemisia tabaci. J Gen Virol 98:1552–1562. https://doi.org/10.1099/jgv.0.000814
Abbas Q, Amin I, Mansoor S et al (2019) The Rep proteins encoded by alphasatellites restore expression of a transcriptionally silenced green fluorescent protein transgene in Nicotiana benthamiana. VirusDisease 30:101–105. https://doi.org/10.1007/s13337-017-0413-5
Nawaz-Ul-Rehman MS, Nahid N, Mansoor S et al (2010) Post-transcriptional gene silencing suppressor activity of two non-pathogenic alphasatellites associated with a begomovirus. Virology 405:300–308. https://doi.org/10.1016/j.virol.2010.06.024
Gnanasekaran P, KishoreKumar R, Bhattacharyya D et al (2019) Multifaceted role of geminivirus associated betasatellite in pathogenesis. Mol Plant Pathol 20:1019–1033. https://doi.org/10.1111/mpp.12800
Reddy K, Bhattacharyya D, Chakraborty S (2020) Mutational study of radish leaf curl betasatellite to understand the role of the non-coding region in begomovirus pathogenesis. Physiol Mol Plant Pathol 112:101549. https://doi.org/10.1016/j.pmpp.2020.101549
Li F, Yang X, Bisaro DM, Zhou X (2018) The βC1 protein of geminivirus-betasatellite complexes: a target and repressor of host defenses. Mol Plant 11:1424–1426. https://doi.org/10.1016/j.molp.2018.10.007
Yang JY, Iwasaki M, Machida C et al (2008) betaC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev 22:2564–2577. https://doi.org/10.1101/gad.1682208
Bhattacharyya D, Gnanasekaran P, Kumar RK et al (2015) A geminivirus betasatellite damages the structural and functional integrity of chloroplasts leading to symptom formation and inhibition of photosynthesis. J Exp Bot 66:5881–5895. https://doi.org/10.1093/jxb/erv299
Gnanasekaran P, Ponnusamy K, Chakraborty S (2019) A geminivirus betasatellite encoded βC1 protein interacts with PsbP and subverts PsbP-mediated antiviral defence in plants. Mol Plant Pathol 20:943–960. https://doi.org/10.1111/mpp.12804
Prabu G, Neha G, Kalaiarasan P et al (2021) Geminivirus betasatellite-encoded βC1 protein exhibits novel ATP hydrolysis activity that influences its DNA-binding activity and viral pathogenesis. J Virol 95:e00475-21. https://doi.org/10.1128/JVI.00475-21
Eini O, Dogra S, Selth LA et al (2009) Interaction with a host ubiquitin-conjugating enzyme is required for the pathogenicity of a geminiviral DNA beta satellite. Mol Plant Microbe Interact 22:737–746. https://doi.org/10.1094/MPMI-22-6-0737
Shen Q, Hu T, Bao M et al (2016) Tobacco RING E3 ligase NtRFP1 mediates ubiquitination and proteasomal degradation of a geminivirus-encoded βC1. Mol Plant 9:911–925. https://doi.org/10.1016/j.molp.2016.03.008
Nair A, Chatterjee KS, Jha V et al (2020) Stability of Begomoviral pathogenicity determinant βC1 is modulated by mutually antagonistic SUMOylation and SIM interactions. BMC Biol 18:110. https://doi.org/10.1186/s12915-020-00843-y
Yang X, Xie Y, Raja P et al (2011) Suppression of methylation-mediated transcriptional gene silencing by βC1-SAHH protein interaction during geminivirus-betasatellite infection. PLoS Pathog 7:e1002329. https://doi.org/10.1371/journal.ppat.1002329
Zhong X, Wang ZQ, Xiao R et al (2017) Mimic phosphorylation of a βC1 protein encoded by TYLCCNB impairs its functions as a viral suppressor of RNA silencing and a symptom determinant. J Virol 91:e00300–e00317. https://doi.org/10.1128/jvi.00300-17
Eini O (2017) A betasatellite-encoded protein regulates key components of gene silencing system in plants. Mol Biol 51:656–663. https://doi.org/10.7868/s002689841703003x
Li F, Huang C, Li Z, Zhou X (2014) Suppression of RNA silencing by a plant DNA virus satellite requires a host calmodulin-like protein to repress RDR6 expression. PLOS Pathog 10:e1003921
Shen Q, Liu Z, Song F et al (2011) Tomato SlSnRK1 protein interacts with and phosphorylates βC1, a pathogenesis protein encoded by a geminivirus β-satellite. Plant Physiol 157:1394–1406. https://doi.org/10.1104/pp.111.184648
Li F, Zhao N, Li Z et al (2017) A calmodulin-like protein suppresses RNA silencing and promotes geminivirus infection by degrading SGS3 via the autophagy pathway in Nicotiana benthamiana. PLOS Pathog 13:e1006213
Haxim Y, Ismayil A, Jia Q et al (2017) Autophagy functions as an antiviral mechanism against geminiviruses in plants. Elife. https://doi.org/10.7554/eLife.23897
Hu T, Huang C, He Y et al (2019) βC1 protein encoded in geminivirus satellite concertedly targets MKK2 and MPK4 to counter host defense. PLOS Pathog 15:e1007728
Saunders K, Bedford ID, Briddon RW et al (2000) A unique virus complex causes Ageratum yellow vein disease. Proc Natl Acad Sci USA 97:6890–6895. https://doi.org/10.1073/pnas.97.12.6890
Cui X, Li G, Wang D et al (2005) A Begomovirus DNAbeta-encoded protein binds DNA, functions as a suppressor of RNA silencing, and targets the cell nucleus. J Virol 79:10764–10775. https://doi.org/10.1128/JVI.79.16.10764-10775.2005
Kumar PP, Usha R, Zrachya A et al (2006) Protein–protein interactions and nuclear trafficking of coat protein and βC1 protein associated with Bhendi yellow vein mosaic disease. Virus Res 122:127–136. https://doi.org/10.1016/j.virusres.2006.07.007
Salvaudon L, De Moraes CM, Yang J-Y et al (2013) Effects of the virus satellite gene βC1 on host plant defense signaling and volatile emission. Plant Signal Behav 8:e23317. https://doi.org/10.4161/psb.23317
Zhao P, Yao X, Cai C et al (2019) Viruses mobilize plant immunity to deter nonvector insect herbivores. Sci Adv 5:eaav9801. https://doi.org/10.1126/sciadv.aav9801
Hu T, Song Y, Wang Y, Zhou X (2020) Functional analysis of a novel βV1 gene identified in a geminivirus betasatellite. Sci China Life Sci 63:688–696. https://doi.org/10.1007/s11427-020-1654-x
Gong P, Tan H, Zhao S et al (2021) Geminiviruses encode additional small proteins with specific subcellular localizations and virulence function. Nat Commun 12:4278. https://doi.org/10.1038/s41467-021-24617-4
Gupta N, Reddy K, Bhattacharyya D, Chakraborty S (2021) Plant responses to geminivirus infection: guardians of the plant immunity. Virol J 18:143. https://doi.org/10.1186/s12985-021-01612-1
Ghosh D, Chakraborty S (2021) Molecular interplay between phytohormones and geminiviruses: a saga of a never-ending arms race. J Exp Bot 72:2903–2917. https://doi.org/10.1093/jxb/erab061
Acknowledgements
We gratefully acknowledge the financial support from the Department of Biotechnology, Govt of India, DBT BUILDER (SLS/BUILDER/SC/2021) and DST-FIST II (SLS/FIST-II/SC/2021). We also thank Malavika M for critically reading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing financial interests.
Additional information
Handling Editor: Jesús Navas-Castillo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Devendran, R., Namgial, T., Reddy, K.K. et al. Insights into the multifunctional roles of geminivirus-encoded proteins in pathogenesis. Arch Virol 167, 307–326 (2022). https://doi.org/10.1007/s00705-021-05338-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05338-x