Abstract
In Argentina, current procedures to ensure the safety of the blood supply for transfusion include the serologic detection of specific blood-borne infections. The aim of this study was to evaluate the prevalence and the genetic diversity of hepatitis B virus (HBV) and hepatitis D virus (HDV) in blood donor populations from two distantly located Argentine regions. Data from 56,983 blood donations from the Favaloro Foundation, in the city of Buenos Aires (Central Region), and the Central Blood Bank of Misiones Province (Northeast Region) were analyzed. Samples that were reactive for HBsAg were analyzed for HBV-DNA characterization and HDV serological and molecular analysis. The HBV prevalence was 0.12 % for HBsAg and 1.68 % for anti-HBc antibodies in Buenos Aires, and 0.73 % and 8.55 %, respectively, in Misiones. Seventy-seven HBsAg-reactive samples were analyzed by polymerase chain reaction for HBV-DNA. Subgenotypes A2, B2, C2, F1b and F4 (Buenos Aires) and F1b and D3 (Misiones) were detected. Several mutations within the major hydrophilic region of HBsAg, the reverse transcriptase, the basal core promoter, and the precore/core were detected. HDV genotype 1 was identified in Buenos Aires. This study confirms the circulation of several HBV subgenotypes, as well as known and newly identified variants, and the presence of HDV1 in this population. A thorough investigation has to be carried out to evaluate the clinical importance of some of the documented mutations as well as those detected in the HDV1 case.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatitis B virus (HBV) causes liver inflammation, frequently associated with cirrhosis and/or hepatocellular carcinoma (HCC) [1]. Hepatitis D virus (HDV) requires the helper function of HBV for its assembly and transmission [2]. At least nine major HBV genotypes (A-H, J), including multiple subgenotypes, and eight HDV genotypes (HDV 1-8) have been defined [3–5].
HBV infection is globally distributed, with high (>8 %), intermediate (between 2 and 8 %) and low (<2 %) endemic areas, while HDV is highly endemic in Mediterranean countries, the Middle East, Central Africa, and the northern areas of South America [6, 7]. Argentina shows a low prevalence of HBsAg, which ranges from 0.2 % to 0.3 % in blood donors (BDs) [8–10]. Genotypes A-F and H have been reported in different populations from Argentina, with the F genotype being the most common [11–16]. In addition, HDV infections were also observed in the 1990s, with a prevalence ranging from 0.7 % to 5.7 %, depending on the study group [17, 18]. Contrary to the recommended recruitment model proposed by the World Health Organization, the blood supply in Argentina is based mainly on repository or replacement donations. Current procedures to ensure the safety of blood transfusions include reviewing the records of BDs and the serological detection of specific blood-borne infections [19]. At present, only the detection of HBsAg and anti-HBc antibody markers for HBV is mandatory in Argentina.
The main objective of this study was to estimate the prevalence of HBV and HDV infections and to analyze the genetic diversity of the viruses detected in voluntary BDs from two geographically distant Argentine regions (approximately 1,600 km distance).
Materials and methods
Plasma samples
A retrospective analysis of serial cross-sectional data collected at two blood banks from the Central (Favaloro Foundation [FF], in the city of Buenos Aires) and Northeastern (Central Blood Bank of Misiones Province [CM]) regions of Argentina was carried out. Data from consecutive blood donations were obtained from a total of 56,983 donors: 50,445 from Favaloro Foundation (2003-2008) and 6,538 from Central Blood Bank of Misiones Province (2009). In both blood banks, serological tests were performed by enzyme immunoassays for HBsAg (Murex and Axsym, Abbott Diagnostics, USA) and total IgG anti-HBc (Hepanostika, Biomerieux, The Netherlands).
HBV-DNA amplification
Seventy-seven reactive samples for HBsAg were analyzed for HBV-DNA, which was extracted from 200 μl of the sample using a QIAmp DNA Mini Kit (QIAGEN AG, Basel, Switzerland). A polymerase chain reaction (PCR) for the S/Pol overlapping genomic region (S/P, nt 256-796) [20] and a nested-PCR (nPCR) for the preCore-Core region (preC/C, nt 1736-2471) [21] were performed. An amplification protocol for the full-length genotype C HBV-DNA was adapted from one described elsewhere [22]. Appropriate precautions were strictly followed to avoid cross-contamination [23]. Each HBV DNA amplified product was validated, since negative controls were added from the extraction step rendering the expected negative results. All positive PCR results were confirmed from the DNA extraction step in a second independent experiment. The potential Taq-dependent DNA misincorporation rate was investigated by bi-directionally sequencing PCR products derived from a GBV-C/HGV clone as previously described [24].
HBV serological markers
HBV-DNA-positive samples were tested for the detection of HBeAg (AxSYM, Abbott Diagnostics, USA), anti-HBe antibodies (AxSYM, Abbott Diagnostics, USA), and anti-HBs antibodies (AxSYM, Abbott Diagnostics, USA).
HDV serology and HDV-RNA amplification
The 77 HBV-reactive samples were tested in duplicate by ELISA for total anti-HDV antibodies (Murex, Abbott Diagnostics, USA). Concomitantly, RNA extraction was performed using 200 μl of plasma and TRIzol Reagent (Life Technologies, USA). The cDNA was prepared by reverse transcription (RT) with random hexamers and AMV reverse transcriptase (Promega, USA). Amplification of a partial region coding for HDAg was performed by nested PCR, in which one of the primers had been described elsewhere [26] and the remaining one was designed by the authors [27]. The first round produced a fragment of 415 bp (853-1267) [25], and the second round spanned a total of 353 bp (889-1241),
RFLP and cloning
To solve discrepancies in the assignment of S/P and preC/C genotypes, two different analyses were performed for some strains: a) RFLP of S amplicons according to a reference protocol [20] and b) cloning of the PCR products corresponding to different regions of the HBV genome (GR1: nt 1736-2471 and GR2: nt 1824-2654) into the pGEM-T Easy Vector system (Promega, Madison, WI) according to the manufacturer’s instructions.
Sequencing and phylogenetic analysis
PCR products and plasmid DNA were sequenced directly in both directions with the corresponding amplification primers using a BigDye Terminator Cycle Sequencing Ready Reaction Kit on an ABI Prism 3100 Genetic Analyzer (Applied Biosystems, Foster City, CA). Sequences were aligned using CLUSTALX v1.83 software [28]. Phylogenetic trees were obtained using the neighbor-joining algorithm with the Kimura two-parameter model of molecular evolution in MEGA v 4.0 [29]. The HBV genotype/subgenotype classification was assigned according to Shi et al. [30].
Analysis of the regulatory regions and ORFs of HBV and HDV
The nucleotide sequences representing the basal core promoter (BCP), the pC, C, and partial S (amino acids [aa] 55-210)/P (aa 64-220) regions for HBV and the HDAg-S (aa 110-195) and HDAg-L (aa 110-214) for HDV were translated into aa sequences according to their corresponding open reading frames (ORFs) and compared with the consensus sequences from the corresponding genotypes using BioEdit software.
Statistical analysis
Statistical tests were performed with the Student t-test and the chi-square test (Epidat v3.1 software) (PAHO/WHO, 2006) as appropriate; p-values <0.05 were considered significant.
Results
Prevalence of HBV/HDV infection
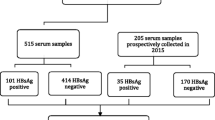
The seroprevalence was 0.12 % (61/50445; 95 % CI: 0.09-0.152) for HBsAg and 1.68 % (851/50445; 95 % CI: 1.57-1.8) for anti-HBc antibodies in BDs from the Favaloro Foundation and 0.73 % (48/6538; 95 % CI: 0.52-0.95) and 8.55 % (559/6538; 95 % CI: 7.86-9.23) in BDs from the Central Blood Bank of Misiones Province. These last values were significantly higher than those from the Favaloro Foundation (p<0.0000 and p<0.0000, respectively). Of the total HBsAg-reactive samples, 77 had enough volume for subsequent analysis (47 from Buenos Aires, and the remaining 30 from Misiones). Out of the 47 HBsAg-reactive samples from Buenos Aires, a single one (FF14) was found to be anti-HDV antibody reactive, reaching a final prevalence of 0.0000198 % (1/50,545) and 2.12 % in HBsAg-reactive samples. FF14 was reactive for both anti-HBc and anti-HBe antibodies. All HBsAg-reactive samples from Misiones were non-reactive for anti-HDV antibodies.
Detection of HBV-DNA and HDV-RNA
Out of the 47 samples from the Favaloro Foundation (39 males; mean age = 33.5 ± 11.6 years), 12 allowed the amplification of HBV S/P region; and 8, the preC/C region. Out of 30 samples from Central Blood Bank of Misiones Province (27 males; mean age = 35.6 ± 10.6 years), 6 were positive for HBV-DNA (Table 1). FF14 was found to be HDV-RNA positive, although HBV-DNA could not be detected at either the S/P or preC/C region.
HBV and HDV phylogenetic analysis
The S/P region sequences from Buenos Aires were classified as subgenotype A2, B2, quasi-subgenotype C2, F1b and F4. Two of the four HBV sequences from Misiones were assigned to subgenotype F1b, while the remaining two clustered with “new D3” according to the nomenclature proposed by Shi et al. [30] (Fig. 1A). The preC/C region sequences (FF1, CM5, CM6 samples)–from which the S/P region could not be amplified–were classified as subgenotype “new D3” (Fig. 1B). Seven out of eight HBV C2 sequences clustered together, with a bootstrap value of 98 % (FF6, FF8-FF13). The complete genomes from one of these sequences (FF11) and from another C2 sequence (FF7; at a separate position in the phylogenetic tree) were analyzed. These two sequences were classified as quasi-subgenotype C2 (Fig. 2), and the identity matrix revealed 98 % identity between FF11 and C2 reference sequences obtained from GenBank.
Phylogenetic relationships of 31 HBV sequences from this study compared to representative sequences belonging to all known genotypes and subgenotypes, determined by neighbor joining. Bootstrap statistical analysis was performed using 1000 datasets, and the numbers at the nodes indicate the bootstrap values. A. Phylogenetic tree based on a portion of the S region (nt 369-769) of the HBV genome. B. Phylogenetic tree based on the the pC/C region (nt 1784-2391). All sequences obtained from GenBank are identified by accession number. The GenBank accession numbers of the sequences reported in this study are JX079922-JX079947. These sequences are indicated by the underlined initials FF (Favaloro Foundation, Buenos Aires) and CM (Central Blood Bank of Misiones). The scale bar indicates the number of nucleotide substitutions per site. The nomenclature of subgenotypes was proposed by Shi et al. [30]
Phylogenetic relationships of full-length C2 HBV genome sequences from this study (FF7 and FF11; Favaloro Foundation, Buenos Aires city) compared to representative sequences belonging to subgenotypes C1-C13, C15 and C16 using neighbor joining. Bootstrap statistical analysis was performed using 1000 datasets, and the numbers at the nodes indicate the bootstrap values. All sequences obtained from GenBank are identified by accession number. The GenBank accession numbers of the sequences reported in this study are KF485389 and KF485390. The scale bar indicates the number of nucleotide substitutions per site
The preC/C region of FF6 was assigned to the C2 quasi-subgenotype, and the S/P region was assigned to the A2 subgenotype (Table 1). To solve this discrepancy in the S/P and preC/C genotypes, two different analyses were performed. RFLP of the S region demonstrated the presence of only the C2 genotype, while phylogenetic analysis of the clones documented the presence of HBV A2 (n = 2) and C2 (n = 2) subgenotypes.
The FF14 HDV sequence was classified as genotype 1, clustering together with sequences from Asia and Argentine Amerindians from Misiones (Fig. 3).
Phylogenetic relationships of the sole HDV sequence obtained from Argentine blood donors to representative sequences belonging to all known genotypes using neighbor joining. Bootstrap statistical analysis was performed using 1000 datasets, and the numbers at the nodes indicate the bootstrap values. The tree represents the partial HDAg region (nt 912-1230) according to a HDV reference strain (GenBank accession number AB118849). All sequences obtained from GenBank are identified by accession number. The sequence reported in this study is indicated by adding the underlined initials FF14 (Favaloro Foundation, Buenos Aires; GenBank accession number JX011008)
Analysis of the HBV/HDV regulatory regions and ORFs
Six out of 12 preC/C sequences showed mutations in the BCP (A1762T and/or G1764A). Other mutations were detected in T cell core epitopes (Table 2). Several sequences showed different mutations within the major hydrophilic region (MHR) within the S protein. Regarding the RT domain (P protein), several mutations were observed in 7 of the 16 sequences; moreover, the rtY124H and rtV207I mutations were identified in the above-mentioned seven C2 sequences (Table 3).
The partial amino acid sequence of HDAg (FF14) exhibited several non-synonymous changes: V149I, G151D, V188L, I198L (within the nuclear export signal), when compared with the HDV1 consensus sequence.
Discussion
Our study confirms a low HBV endemicity in BDs at two blood banks in central and northeastern Argentina and the presence of HDV in one BD in Buenos Aires. Although both HBsAg and anti-HBc antibodies from Buenos Aires donors were found at low prevalence, as reported in 2004 [12], these serological markers were significantly higher in BDs in Misiones. These data are consistent with the previously reported prevalence rates from Amerindians from Misiones [31], highlighting the need to improve health control policies in this region.
Concerning HBV genotypes, this study confirmed the circulation of F1b subgenotype in Misiones, as we have recently reported from Mbyá-guaraní natives living in this province [31]. Also, two sequences (CM1 and CM4) clustered together with D3, D6 and previously proposed “D3/D6 recombinant sequences” [32]. Further analysis of CM1 and CM4 sequences led to their final classification as subgenotype “new D3”, in agreement with a very recently proposed nomenclature [30, 33].
Regarding Buenos Aires, and in contrast to one published study in which genotype F was reported to be slightly predominant [12], the current survey demonstrates that quasi-subgenotype C2 was the most prevalent in this cohort. However, it should be taken into account that the time period during which the BDs visited the public bank(s) was not mentioned in the previous study, precluding proper temporal comparisons between both reports. A potential bias of the current study with respect to the high C2 prevalence in Buenos Aires might be associated with blood donations from related donors at the blood bank during the same year. These C2 sequences, which were confirmed by analysis of the full-length genome, were clustered with other C2 sequences recently detected in Sagua-Huarpes Argentine Amerindians (Central Western Region) [31], suggesting a recent internal migration event. An alternative hypothesis might involve recent migrations from Asia [34], as well as from other South American countries, since C2 has been recorded in Bolivia in both natives and Japanese immigrants [35]. The remaining HBV genotypes detected were A2, F1b and F4, confirming previous results [12], while a single B2 sequence was detected in BDs for the first time. Moreover, in one of the samples (FF6), the coexistence of variants assigned to C2 and A2 has been confirmed.
In Argentina, BCP and preC/C mutants have been detected in chronic hepatitis B patients, in HIV-1-coinfected patients, and in BDs [11, 12, 36]. In the present study, mutations in at least one nucleotide of these regions were found in 83.3 % of the samples. The A1762T/G1764A mutation, which was found in 33.3 % of the sequences, could also alter the X protein transactivator activity by causing amino acid changes (L130M and V131I) that are known to be associated with an increased risk of HCC [37]. On the other hand, G1896A preC mutations, which are responsible for the e-minus phenotype, and C1858T (16.66 % of our sequences) are not associated with the risk of HCC, regardless of HBeAg status or HBV genotype [38]. In our study, most of the HBsAg-positive samples proved to be HBeAg non-reactive (Table 1). Such a profile is known to exhibit a higher frequency of mutations at nucleotide positions 1846, 1896 and 1899 in preC/C than is found in HBeAg-positive subjects, as observed herein.
As reported for Amerindians from Misiones with occult HBV infection (OBI), the F179L (F1b) aa replacement [31] was detected in one non-OBI BD (CM3). The D3 sequences exhibited several mutations within the MHR, including the Y100S substitution, which has been reported to be associated with OBI [39]. The remaining D3 sequence exhibited the C107G replacement, which has been shown to be indispensable for efficient secretion. Interestingly, HBsAg was nevertheless detected in these isolates, suggesting that such mutations are not necessarily the sole cause of OBI cases. The M198I, W199L and Y200F mutations, which were present in seven C2 sequences, have been described recently in Sagua-Huarpes Amerindians [31]. Furthermore, the C107G and M198I mutations were shown to disrupt conformational region(s) of the HBsAg protein, affecting its antigenicity [40, 41]. Regarding the RT domain of the P protein, the D3 sequences exhibited rtS213T, rtQ215H and rtA194V mutations (Table 3). In this regard, rtS213T has been reported previously to be associated with resistance to adefovir in treated patients [42, 43]. Moreover, the rtQ215H mutation has been detected at baseline and after 6 months of lamivudine therapy [44], a true pretreatment mutation (also known as natural resistance). With regard to the rtA194V mutation, it is worth mentioning that residue rtA194 is located in a loop at the end of the B domain, which contains an alpha helix that interacts with the nucleic acid template. Since the rtA194T substitution, confers tenofovir resistance [42, 43], may affect polymerisation efficiency by causing allosteric changes that result in misalignment between the template and dNTP-binding site, the novel rtA194V replacement deserves to be studied further. It should be stressed that the clinical relevance of these substitutions has not yet been determined. Therefore, since rtA194V, rtS213T and rtQ215H, detected in this study, are not currently considered in genotypic drug resistance algorithms, our data warrant additional studies to analyze their possible associations with antiviral resistance.
On the other hand, the rtY124H and rtV207I mutations were identified in C2 sequences from Buenos Aires and have recently been reported in Sagua-Huarpes Amerindians [31]. The former mutation has been described in untreated Chinese patients, and the latter is associated with resistance to lamivudine [45]. It is worth emphasizing that, in our study, the above-mentioned mutations occurred in untreated (naïve) subjects, as observed previously in randomly selected patients with chronic hepatitis B [46]. These data suggest the circulation of HBV natural resistance variants among BDs. Moreover, other new variants described in Table 3 have not been reported elsewhere.
Concerning HDV, a report published in 1987 showed 1.4 % anti-HDV antibody prevalence among HBsAg-positive Argentine BDs [17], similar to the value reported in this study. One HDV1 sequence (FF14) was detected recently in Amerindians from Misiones [27], confirming the circulation of this genotype in at least two distantly located regions of Argentina. Interestingly, the single HDV case described in this study exhibited several non-synonymous changes when compared with HDV1 reference sequences. With the exception of I198L (involved in the nuclear export signal), there is no published data regarding the functional meaning of the remaining mutations.
In summary, several HBV genotypes were observed in BDs from two Argentine geographic regions. Several sequences exhibited mutations within the BCP, the preC/C region, the HBsAg MHR, and the reverse transcriptase, some of them associated with natural resistance. A thorough investigation has to be carried out to evaluate the clinical impact of some of the documented mutations as well as those detected in the HDV1 case.
References
El-Serag H, Ruldolph K (2007) Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology 132:2557–2576. doi:10.1053/j.gastro.2007.04.061
Shih HH, Jeng KS, Syu WJ, Huang YH, Su CW, Peng WL, Sheen IJ, Wu JC (2008) Hepatitis B surface antigen levels and sequences of natural hepatitis B virus variants influence the assembly and secretion of hepatitis D virus. J Virol 82:2250–2264. doi:10.1128/JVI.02155-07
McMahon BJ (2009) The natural history of chronic hepatitis B virus infection. Hepatology 49:45–55. doi:10.1002/hep.22898
Tatematsu K, Tanaka Y, Kurbanov F, Sugauchi F, Mano S, Maeshiro T, Nakayoshi T, Wakuta M, Miyakawa Y, Mizokami M (2009) A genetic variant of hepatitis B virus divergent from known human and ape genotypes isolated from a Japanese patient and provisionally assigned to new genotype J. J Virol 83:10538–10547. doi:10.1128/JVI.00462-09
Hughes SA, Wedemeyer H, Harrison PM (2011) Hepatitis delta virus. Lancet 378:73–85. doi:10.1016/S0140-6736(10)61931-9
Radjef N, Gordien E, Ivaniushina V, Gault E, Anaïs P, Drugan T, Trinchet JC, Roulot D, Tamby M, Milinkovitch MC, Dény P (2004) Molecular phylogenetic analyses indicate a wide and ancient radiation of African hepatitis delta virus, suggesting a delta virus genus of at least seven major clades. J Virol 78:2537–2544. doi:10.1128/JVI.78.5.2537-2544.2004
Monsalve-Castillo F, Echevarría JM, Atencio R, Suárez A, Estévez J, Costa-León L, Montiel P, Molero T, Zambrano M (2008) Amerindians in Japreira, Zulia State, Venezuela. Cad Saude Pública 24:1183–1186
World Health Organization (1998) Blood bank situation in Latin America, 1996: serological markers for communicable diseases in blood donors. Epidemiol Bull 19: 12–14. Fact sheet N°204. http://www.who.int. and Pan American Health Organization (Revised August 2008)
Proyecto Programa Nacional de Control de Hepatitis Virales. Epidemiología Informe N°6-9. Instituto Nacional de Enfermedades Infecciosas (INEI). Administración Nacional de Laboratorios e Institutos de Salud (ANLIS) “Dr. Carlos Gregorio Malbrán”. http://www.hepatitisviral.com.ar/ppnhv.htm
Mathet V, Cuestas ML, Trinks J, Minassian ML, Ruiz V, Rivero CW, Andreetta AM, Weissenbacher MC, Oubiña JR (2007) Chapter X: genetic diversity and variability of Hepatitis B virus (HBV) in Latin America and the Caribbean region: implications in epidemiological, clinical, diagnostic, prophylactic and therapeutic approaches. In: Denyer DV (ed) Progress in hepatitis B. Nova Science Publishers Inc., NY, pp 277–351
Trinks J, Cuestas ML, Tanaka Y, Mathet VL, Minassian ML, Rivero CW, Benetucci JA, Gímenez ED, Segura M, Bobillo MC, Corach D, Ghiringhelli PD, Sánchez DO, Avila MM, Peralta LA, Kurbanov F, Weissenbacher MC, Simmonds P, Mizokami M, Oubiña JR (2008) Two simultaneous hepatitis B virus epidemics among injecting drug users and men who have sex with men in Buenos Aires, Argentina: characterization of the first D/A recombinant from the American continent. J Viral Hepat 15:827–838. doi:10.1111/j.1365-2893.2008.00997.x
França PH, González JE, Munné MS, Brandão LH, Gouvea VS, Sablon E, Vanderborght BO (2004) Strong association between genotype F and hepatitis B virus (HBV) e antigen-negative variants among HBV-infected Argentinean blood donors. J Clin Microbiol 42:5015–5021. doi:10.1128/JCM.42.11.5015-5021.2004
Piñeiro Y, Leone FG, Pezzano SC, Torres C, Rodríguez CE, Eugenia Garay M, Fainboim HA, Remondegui C, Sorrentino AP, Mbayed VA, Campos RH (2008) Hepatitis B virus genetic diversity in Argentina: dissimilar genotype distribution in two different geographical regions; description of hepatitis B surface antigen variants. J Clin Virol 42:381–388. doi:10.1016/j.jcv.2008.01.018
Mathet VL, Cuestas ML, Ruiz V, Minassian ML, Rivero C, Trinks J, Daleoso G, León LM, Sala A, Libellara B, Corach D, Oubiña JR (2006) Detection of hepatitis B virus (HBV) genotype E carried -even in the presence of high titers of anti-HBs antibodies- by an Argentinean patient of African descent who had received vaccination against HBV. J Clin Microbiol 44:3435–3439. doi:10.1128/JCM.00866-06
Flichman D, Galdame O, Livellara B, Viaut M, Gadano A, Campos R (2009) Full-length genome characterization of hepatitis B virus genotype H strain isolated from serum samples collected from two chronically infected patients in Argentina. J Clin Microbiol 47:4191–4193. doi:10.1128/JCM.01337-09
Quarleri J, Moretti F, Bouzas MB, Laufer N, Carrillo MG, Giuliano SF, Pérez H, Cahn P, Salomon H (2007) Hepatitis B virus genotype distribution and its lamivudine-resistant mutants in HIV-coinfected patients with chronic and occult hepatitis B. AIDS Res Hum Retroviruses 23:525–531. doi:10.1089/aid.2006.0172
Fay O, Tanno H, Gatti H, Basualdo JA, Ciocca M, Fainboin H, Fainboin L, Jorge A, Motta A, Naval M et al (1987) Anti-delta antibody in various HBsAg positive Argentine populations. J Med Virol 22:257–262
Fainboim H, González J, Fassio E, Martínez A, Otegui L, Eposto M, Cahn P, Marino R, Landeira G, Suaya G, Gancedo E, Castro R, Brajterman L, Laplumé H (1999) Prevalence of hepatitis viruses in an anti-human immunodeficiency virus-positive population from Argentina. A multicentre study. J Viral Hepat 6:53–57. doi:10.1046/j.1365-2893.1999.t01-1-6120135.x
Schmunis GA, Dias JC (2000) Health care reform, decentralization, prevention and control of vector-borne diseases. Cad Saude Publica 16:117–123
Lindh M, Andersson AS, Gusdal A (1997) Genotypes, nt 1858 variants, and geographic origin of hepatitis B virus-large-scale analysis using a new genotyping method. J Infect Dis 175:1285–1293
Birkenmeyer L, Mushahwar I (1994) Detection of hepatitis A, B and D virus by the polymerase chain reaction. J Virol Methods 49:101–112
Sugauchi F, Mizokami M, Orito E, Ohno T, Kato H, Suzuki S, Kimura Y, Ueda R, Butterworth LA, Cooksley WG (2001) A novel variant genotype C of hepatitis B virus identified in isolates from Australian Aborigines: complete genome sequence and phylogenetic relatedness. J Gen Virol 82:883–892
Kwok S, Higuchi R (1989) Avoiding false positives with PCR. Nature 339(6221):237–238
Mathet VL, Feld M, Espinola L, Sanchez DO, Ruiz V, Mando O, Carballal G, Quarleri JF, D’Mello F, Howard CR, Oubina JR (2003) Hepatitis B virus S gene mutants in a patient with chronic active hepatitis with circulating anti-HBs antibodies. J Med Virol 69:18–26
Nakano T, Shapiro CN, Hadler SC, Hadler SC, Casey JL, Mizokami M, Orito E, Robertson BH (2001) Characterization of hepatitis D virus genotype III among Yucpa Indians in Venezuela. J Gen Virol 82:2183–2189
Wang KS, Choo QL, Weiner AJ, Ou JH, Najarian RC, Thayer RM, Mullenbach GT, Denniston KJ, Gerin JL, Houghton M (1986) Structure, sequence and expression of the hepatitis delta (delta) viral genome. Nature 323:508–514
Delfino CM, Eirin ME, Berini C, Malan R, Gentile E, Castillo A, Pedrozo W, Krupp R, Blejer J, Oubiña JR, Mathet VL, Biglione MM (2012) HDAg-L variants in covert hepatitis D and HBV occult infection among Amerindians of Argentina: new insights. J Clin Virol 54:223–228. doi:10.1016/j.jcv.2012.04.014
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTALX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Shi W, Zhang Z, Ling C, Zheng W, Zhu C, Carr MJ, Higgins DG (2013) Hepatitis B virus subgenotyping: History, effects of recombination, misclassifications, and corrections. Infect Genet Evol 16:355–361. doi:10.1016/j.meegid.2013.03.021
Delfino CM, Berini C, Eirin ME, Malan R, Pedrozo W, Krupp R, Blejer J, Espejo R, Fierro L, Puca A, Oubiña JR, Mathet VL, Biglione MM (2012) New natural variants of hepatitis B virus among Amerindians from Argentina with mainly occult infections. J Clin Virol 54:174–179. doi:10.1016/j.jcv.2012.02.023
Reis LM, Soares MA, França PH, Soares EA, Bonvicino CR (2011) Clonal analysis of hepatitis B viruses among blood donors from Joinville, Brazil: evidence of dual infections, intragenotype recombination and markers of risk for hepatocellular carcinoma. J Med Virol 83:2103–2112. doi:10.1002/jmv.22246
Yousif M, Kramvis A (2012) Genotype D of hepatitis B virus and its subgenotypes: An update. Hepatol Res 43(4):355–364. doi:10.1111/j.1872-034X.2012.01090.x
Pezzano SC, Torres C, Fainboim HA, Bouzas MB, Schroder T, Giuliano SF, Paz S, Alvarez E, Campos RH, Mbayed VA (2011) Hepatitis B virus in Buenos Aires, Argentina: genotypes, virological characteristics and clinical outcomes. Clin Microbiol Infect 17:223–231. doi:10.1111/j.1469-0691.2010.03283.x
Khan A, Tanaka Y, Saito H, Ebinuma H, Sekiguchi H, Iwama H, Wakabayashi G, Kamiya T, Kurbanov F, Elkady A, Mizokami M (2008) Transmission of hepatitis B virus (HBV) genotypes among Japanese immigrants and natives in Bolivia. Virus Res 132:174–180. doi:10.1016/j.virusres.2007.12.005
López JL, Mbayed VA, Telenta PF, González JE, Campos RH (2002) ‘Hbe minus’ mutants of hepatitis B virus. Molecular characterization and its relation to viral genotypes. Virus Res 87:41–49. doi:10.1016/S0168-1702(02)00078-3
Tan YJ (2011) Hepatitis B virus infection and the risk of hepatocellular carcinoma. World J Gastroenterol 17:4853–4857. doi:10.3748/wjg.v17.i44.4853
Cao GW (2009) Clinical relevance and public health significance of hepatitis B virus genomic variations. World J Gastroenterol 15:5761–5769. doi:10.3748/wjg.15.5761
Svicher V, Cento V, Bernassola M, Neumann-Fraune M, Van Hemert F, Chen M, Salpini R, Liu C, Longo R, Visca M, Romano S, Micheli V, Bertoli A, Gori C, Ceccherini-Silberstein F, Sarrecchia C, Andreoni M, Angelico M, Ursitti A, Spanò A, Zhang JM, Verheyen J, Cappiello G, Perno CF (2012) Novel HBsAg markers tightly correlate with occult HBV infection and strongly affect HBsAg detection. Antiviral Res 93:86–93. doi:10.1016/j.antiviral.2011.10.022
Mangold CM, Unckell F, Werr M, Streeck RE (1995) Secretion and antigenicity of hepatitis B virus small envelope proteins lacking cysteines in the major antigenic region. Virology 211:535–543. doi:10.1006/viro.1995.1435
Torresi J, Earnest-Silveira L, Deliyannis G, Edgtton K, Zhuang H, Locarnini SA, Fyfe J, Sozzi T, Jackson DC (2002) Reduced antigenicity of the hepatitis B virus HBsAg protein arising as a consequence of sequence changes in the overlapping polymerase genes that are selected by lamivudine therapy. Virology 293:305–313. doi:10.1006/viro.2001.1246
Shaw T, Bartholomeusz A, Locarnini S (2006) HBV drug resistance: mechanisms, detection and interpretation. J Hepatol 44(3):593–606
Sheldon J, Ramos B, Garcia-Samaniego J, Rios P, Bartholomeusz A, Romero M, Locarnini S, Zoulim F, Soriano V (2007) Selection of hepatitis B virus (HBV) vaccine escape mutants in HBV-infected and HBV/HIV-coinfected patients failing antiretroviral drugs with anti-HBV activity. J Acquir Immune Defic Syndr 46(3):279–282
Kim do Y, Chang HY, Lim SM, Kim SU, Park JY, Kim JK, Lee KS, Han KH, Chon CY, Ahn SH (2013) Quasispecies and pre-existing drug-resistant mutations of hepatitis B virus in patients with chronic hepatitis B. Gut Liver 7(3):329–334. doi:10.5009/gnl.2013.7.3.329
Liu BM, Li T, Xu J, Li XG, Dong JP, Yan P, Yang JX, Yan L, Gao ZY, Li WP, Sun XW, Wang YH, Jiao XJ, Hou CS, Zhuang H (2010) Characterization of potential antiviral resistance mutations in hepatitis B virus reverse transcriptase sequences in treatment-naïve Chinese patients. Antiviral Res 85:512–519. doi:10.1016/j.antiviral.2009.12.006
Cuestas ML, Rivero CW, Minassian ML, Castillo AI, Gentile EA, Trinks J, León L, Daleoso G, Frider B, Lezama C, Galoppo M, Giacove G, Mathet VL, Oubiña JR (2010) Naturally occurring hepatitis B virus (HBV) variants with primary resistance to antiviral therapy and S-mutants with potential primary resistance to adefovir in Argentina. Antiviral Res 87:74–77. doi:10.1016/j.antiviral.2010.04.005
Acknowledgments
The authors are grateful for the invaluable assistance provided by Horacio Salamone and Ramón Krupp during manuscript preparation. This work was supported by funds from B.I.D. 1728/OC-AR PICT 2007 01639 (Scientific and Technologic Research Projects), Fundación Florencio Fiorini; UBACyT 20020100101063 and 20020090200189, (University of Buenos Aires), Buenos Aires, Argentina, and CONICET PIP 112-200801-01773 and from Département de Virologie, Unité d’épidémiologie et physiopathologie des virus oncogènes, Institut Pasteur, Paris, France.
Conflict of interest
No conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. M. Biglione and V. L. Mathet contributed equally.
Rights and permissions
About this article
Cite this article
Delfino, C.M., Gentile, E.A., Castillo, A.I. et al. Hepatitis B virus and hepatitis D virus in blood donors from Argentina: circulation of HBsAg and reverse transcriptase mutants. Arch Virol 159, 1109–1117 (2014). https://doi.org/10.1007/s00705-013-1917-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1917-y