Abstract
Bovine leukemia virus (BLV) infection in cattle causes persistent lymphocytosis, and a few percent of infected animals develop lymphoid tumors, namely enzootic bovine leukosis (EBL). In this study, a 440-bp fragment of the env gene was amplified from 204 tumor samples collected from different regions of Japan and analyzed by restriction fragment length polymorphism (RFLP) to determine the association of BLV with EBL. Of the seven RFLP types defined, types I, II, and III were dominant and found in 12.7, 75.0, and 8.3% of tumor samples, respectively. Cattle harboring type III virus were significantly older than other animals at the time of diagnosis of EBL. Type III viruses were found in approximately 33% and 5.5% of Japanese Black and Holstein cattle, respectively, with EBL. These findings indicate that genetically distinct BLV was associated with EBL in Japan and that the genetic profile may influence the leukemogenicity of the virus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
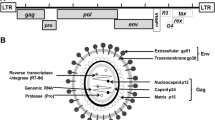
Bovine leukemia virus (BLV) is a member of the family Retroviridae, genus Deltaretrovirus, together with human T-lymphotropic virus (HTLV) types 1 and 2, and a causative agent of enzootic bovine leukosis (EBL). BLV generally infects B-lymphocytes [9], causing asymptomatic infection in cattle. About 33% of infected animals develop persistent lymphocytosis (PL), and 0.1-10% develop tumors when they are 3-5 years old or older [3, 8]. The virus is present in blood lymphocytes and tumor cells, and the provirus is integrated into the genome and found in the cellular fraction of various body fluids. Prenatal and milk transmission by colostrum or mature milk has been demonstrated in BLV infection. Contact with the affected animals may play a central role in transmission; however, calves did not show clinical signs of EBL when they were placed in contact with the virus at 6 weeks or older. Iatrogenic BLV infection by contaminated instruments is also possible [18].
The envelope glycoproteins of viruses play a crucial role in their life cycle. The BLV envelope contains two glycoproteins, gp30 and gp51, which are disulfide-linked and derived by posttranslational proteolytic cleavage of a precursor (gpr72) encoded by the env gene [7, 12]. The gp51 protein is responsible for virus attachment and entry into host cells, serving as a target for neutralizing antibodies [4, 5, 15]. The glycoprotein also induces a cell-mediated immune response, which may play a role in protective immunity against BLV infection, especially tumorigenesis [6].
Recently, we demonstrated through phylogenetic studies of the env gene that the majority of Japanese isolates were genotype 1, which is the common genotype found worldwide [13, 16]. Restriction fragment length polymorphism (RFLP) analysis of the env gene successfully categorized Japanese BLV isolated from cattle that developed PL into six different types [11], suggesting that some strains belonging to a particular RFLP type could not induce effective immune responses in virus-infected animals [1]. In the present study, to define the relationship between tumorigenicity and genetic background of BLV, we determined the RFLP types of the env genes obtained from 204 EBL tumor samples from different regions of Japan and compared the age of the animals affected by EBL.
Materials and methods
Tumor samples
Lymphocytic sarcomas were collected from 214 cattle, including 171 Holstein, 31 Japanese Black, 1 Jersey, and 11 crossbreeds, at various slaughterhouses around Japan in 2008-2010. The age of the host animals was traced from the Information for Individual Identification of Cattle provided by the National Livestock Breeding Center (Japan). The tissue samples were placed into cryotubes, the lids were sealed, and the samples were transported under frozen conditions to the Health Sciences University of Hokkaido and stored at −80°C until required.
DNA isolation and PCR
Provirus DNA was extracted from tumors using a QuickGene DNA tissue kit S (FUJIFILM, Tokyo, Japan) according to manufacturer’s instructions. PCR was carried out as described in the diagnostic manual for OIE (The World Organisation for Animal Health) [20]. Briefly, a 440-bp portion of the env gene corresponding to nucleotide positions 5029-5468 of the BLV genome [17] was amplified by using GoTaq Flexi DNA polymerase (Promega, Madison, WI) and a set of primers, OBLV1A (5′-CTTTGTGTGCCAAGTCTCCCAGATACA-3′) and OBLV6A (5′-CCAACATATAGCA CAGTCTGGGAAGGC-3′) [2]. The reaction mixtures contained 26.75 μl of distilled water, 10 μl of 5x GoTaq Flexi buffer (Promega, Madison, WI), 2 μl of 10 μM each primer, 5 μl of template DNA, 4 μl of a 25 mM MgCl2 solution and 0.25 μl of GoTaq Flexi DNA polymerase. The amplification was carried out for 5 cycles consisting of 94°C for 45 s, 60°C for 60 s and 72°C for 90 s, followed by 30 cycles of 94°C for 45 s, 55°C for 60 s and 72°C for 90 s. The thermal cycling protocol was concluded with a final step at 72°C for 7 min. The amplification products were analyzed on 1.5% agarose gels stained with ethidium bromide.
Nucleotide sequencing
The amplified products were purified from agarose gels using the Wizard® SV Gel and PCR Clean-Up System (Promega, Madison, WI). Purified amplicons were sequenced directly with the primers OBLV1A, OBLV6A, OBLV3 (5′-CTGTAAATGGCTATCCTAAGATCTACTGGC-3′) and OBLV5 (5′-GACAGAGGGAACCCAGTCACTGTTCAACTG-3′) [2] using a BigDye® Terminator ver1.1 Cycle Sequencing Kit (Applied Biosystems, Foster, CA) according to the manufacturer’s instructions. The sequences were analyzed with an ABI Prism 310 Genetic Analyzer (Applied Biosystems, Foster, CA).
RFLP analysis
The 440-bp amplicons were digested with 10 U of BamHI (Takara, Shiga, Japan), HaeIII (Takara, Shiga, Japan), MseI (New England Bio-Labs Inc., Ipswich, MA), MwoI (New England Bio-Labs Inc., Ipswich, MA) and TaqI (Takara, Shiga, Japan) restriction endonucleases according to the manufacturers’ instructions for 1 h and subsequently electrophoresed in 3% NuSieve 3:1 Agarose (Lonza, Rockland, ME) gels. The sizes of the resulting bands were compared with a 10-bp DNA ladder (Invtrogen, Carlsbad, CA) and a 100-bp DNA ladder (TOYOBO, Osaka, Japan).
Results
Detection of the BLV genome in tumor samples
Of the 214 tumor samples, the 440-bp fragments of interest were amplified from 204 samples (95.3%), and 62 were sequenced. The nucleotide sequences found in the fragments exhibited more than 96% identity with that of the λBLV-1 env gene [17] (accession no. AB598781-AB598805). Each fragment was also digested with restriction endonuclease BamHI and verified as the env gene, as denoted by two bands with sizes of 240 and 200 bp. Our findings indicate that the majority of lymphocytic sarcomas in Japanese cattle were diagnosed as EBL caused by BLV infection. As shown in Table 1, the mean age of cattle with EBL was approximately 75 months old. The mean age of cattle with tumor lacking BLV provirus was 45 months old.
Classification of BLV by RFLP analysis on the env gene
The 440-bp amplicons from the tumor samples were digested with HaeIII, MseI, MwoI, and TaqI. As shown in Fig. 1, HaeIII cleaved the majority of the amplicons into five bands, with sizes of 200, 90, 65, 35, and 25 bp; the remainder of the amplicons were cleaved into 290-, 65-, 35-, and 25-bp fragments. MseI digestion resulted in three different restriction patterns. A large number of the samples were cleaved into two bands, of 400 and 40 bp, with three samples that were subjected to enzyme treatment yielding 360- and 40-bp bands. Twenty-six of the amplicons were cleaved into three bands, of 260, 130, and 40 bp. MwoI digestion generated three bands, of 250, 160 and 30 bp, from the majority of samples. Only four samples were cleaved to yield two bands, of 280 and 160 bp. TaqI cleaved the 440-bp amplicon of one sample into three bands, of 230, 160, and 50 bp, and another into 250, 160, and 30 bp. The remaining samples yielded two bands, of 280 and 160 bp. Based on the restriction enzyme digestion patterns of the 440-bp env gene fragments, BLV proviruses collected in this study were classified into seven types (Table 2).
Relationship between RFLP type and age of host cattle for EBL diagnosis
As shown in Table 3, 75% of the BLV strains identified in this study were classified as RFLP type II, and their host animals were diagnosed with EBL at a mean age of 72 months. Twenty-six animals (12.7%) harbored a virus of RFLP type I and were also diagnosed at the same age. In contrast, for type III viruses, the host animals were significantly older (>100 months) than those infected with type I and II viruses when diagnosed with EBL. There was also a statistical difference between type III viruses and whole EBL samples.
Relationship between breeds of host cattle and RFLP type
Out of the 171 Holstein cattle tested, 163 (95.3%) were found to be infected with BLV. The breakdown of the RFLP type for BLV was as follows: type I, 14.1%; II, 76.7%; III, 5.5%; IV, 1.8%; V 1.2%, and VI, 0.6%. For Japanese Black cattle, all of the samples tested were also diagnosed with EBL, and the breakdown was as follows: type I, 6.5%; II, 58.1%; III, 29.0%; V, 3.2%, and VII, 3.2%. Nine of the 11 crossbreeding cattle tested were found to be infected with BLV of RFLP type II, one contained type I, and one animal harbored no virus at all. The affected crossbreeding cattle were significantly younger than the Holstein and Japanese Black breeds. There was also a statistical difference between Holstein and Japanese Black cattle (Table 4).
Discussion
The bovine leukosis complex is broken down into EBL, which is associated with BLV infection, and sporadic bovine leukosis (SBL) [14]. In the present study, of the 214 lymphocytic sarcoma samples, 204 (95.3%) tumors were found to possess the env gene of BLV. The mean age of 204 cattle affected by EBL was approximately 75 months; the mean age of the 10 remaining animals was 45 months. Because SBL, which includes calf lymphosarcoma, thymic lymphosarcoma, and skin lymphosarcoma, occurs most often in animals younger than 2 years, some of the 10 animals in which the env gene was not detected were possibly affected by EBL. A previous study has shown that about 25% of lymphoid tumors associated with EBL harbor BLV proviruses containing deletions [10].
RFLP analysis using HaeIII, MseI, MwoI, and TaqI endonucleases demonstrated seven different types of BLV. Three RFLP types (I, II, and III) were dominant and found in 12.7, 75.0, and 8.3% of tumors associated with EBL in Japan, respectively. A previous study using BclI, HaeIII, and PvuII on another 444-bp env fragment that overlapped the 440-bp fragment that was investigated in this study classified the virus into six types and revealed that one genotype accounted for 48.3% of nearly 400 samples collected from cattle with PL in Japan [1]. Based on the sequence data from the 62 tumor samples, the majority of type II and III viruses were predicted to be of this major genotype. Although some BLV strains (EF065659, EF065650, EF065662) found in PL in Japan were not classified into any of the RFLP types defined in this study, they showed close genetic relationships to type III, V, and VII viruses, which formed a mini-cluster exclusive of other types [13]. The phylogenetic analysis also demonstrated that all Japanese isolates, except for one type VI virus, were classified into the common genotype found worldwide. Type I viruses formed a unique cluster, while type II viruses were found to be disseminated within the genotype [13].
It is worthy of noting that host cattle harboring type III virus were significantly older than other animals. Although contact transmission of BLV is considered to be most common mode, none of the adult animals infected by contact demonstrated tumor formation [18]. Hence, the animals affected by EBL and associated with type III virus did not seem to be infected at an older age. It is likely that lymphocytes infected with certain BLV strains belonging to RFLP type III might be liable to become the target of cytotoxic T cells, as the envelope glycoprotein carries the CD8+ T cell epitope [6]. Alternatively, the viruses may possess reduced tumorigenic potency. The Tax protein, rather than the envelope glycoprotein, is thought to play a crucial role in leukemogenesis by BLV [19, 21].
Of the 31 viruses found in Japanese Black cattle, 11 (35.4%) were sorted into a type III-related group, whereas only 6.3% (11/174) of those in other breeding cattle were sorted into this group. Furthermore, 50% (11/22) of the type III-related viruses were found in Japanese Black cattle. It is possible that RFLP type-III-related viruses might be initially inherent in these breeding cattle. Crossbreeding cattle with EBL were significantly younger than the other cattle; these animals are usually slaughtered at less than 30 months of age; therefore, only young animals were examined.
We propose that the genetic profile of BLV may influence its leukemogenicity. To gain insight into the molecular mechanisms of leukemogenesis by deltaretroviruses, further investigations regarding the relationship between the age of onset of EBL and the genetics of BLV are required.
References
Asfaw Y, Tsuduku S, Konishi M, Murakami K, Tsuboi T, Wu D, Sentsui H (2005) Distribution and superinfection of bovine leukemia virus genotypes in Japan. Arch Virol 150:493–505
Ballagi-Pordany A, Klintevall K, Merza M, Klingeborn B, Belak S (1992) Direct detection of bovine leukaemia virus infection: practical applicability of a double polymerase chain reaction. J Vet Med [B] 39:69–77
Burny A, Cleuter Y, Kettman R, Mammerickx M, Marbaix G, Portelle D, Vanden Broeke A, Willems L, Thomas R (1998) Bovine leukaemia: facts and hypotheses derived from the study of an infectious cancer. Vet Microbiol 17:197–218
Bruck C, Portetelle D, Burny A, Zavada J (1982) Topographical analysis by monoclonal antibodies of BLV gp51 epitopes involved in viral functions. Virology 122:353–362
Callebaut I, Voneche V, Mager A, Fumiere O, Krchnak V, Merza M, Zavada J, Mammerickx M, Burny A, Portetelle D (1993) Mapping of B-neutralizing and T-helper epitopes on the bovine leukemia virus external glycoprotein gp51. J Virol 67:5321–5327
Gatei MH, Good MF, Daniel RC, Lavin MF (1993) T-cell responses to highly conserved CD4 and CD8 epitopes on the outer membrane protein of bovine leukemia virus: relevance to vaccine development. J Virol 67:1796–1802
Johnston ER, Radke K (2000) The SU and TM envelope protein subunits of bovine leukemia virus are linked by disulfide bonds, both in cells and in virions. J Virol 74:2930–2935
Kahrs RF (2001) Bovine leukemia virus and enzootic bovine leucosis. In: Viral diseases of cattle, 2nd edn. Iowa State University Press, Iowa, pp 103–112
Kenyon SJ, Piper CE (1977) Cellular basis of persistent lymphocytosis in cattle infected with bovine leukemia virus. Infect Immun 16:891–897
Kettmann R, Deschamps J, Cleuter Y, Couez D, Burny A, Marbaix G (1982) Leukemogenesis by bovine leukemia virus: proviral DNA integration and lack of RNA expression of viral long terminal repeat and 3’ proximate cellular sequences. Proc Natl Acad Sci USA 73:1014–1018
Licursi M, Inoshima Y, Wu D, Yokoyama T, González ET, Sentsui H (2003) Provirus variants of bovine leukemia virus in naturally infected cattle from Argentina and Japan. Vet Microbiol 96:17–23
Mamoun RZ, Astier T, Guillemain B, Duplan JF (1983) Bovine lymphosarcoma: processing of bovine leukemia virus-coded proteins. J Gen Virol 64:2791–2795
Matsumura K, Inoue E, Osawa Y, Okazaki K (2011) Molecular epidemiology of bovine leukemia virus associated with enzotic bovine leukosis in Japan. Virus Res 155:343–348
Parodi AL (1989) Pathology of enzootic bovine leucosis. Comparison with the sporadic form. In: Burny A, Mammerickx M (eds) Enzootic bovine leucosis and bovine leukemia virus. Martinus Nijhoff Publishing, Boston, pp 15–49
Portetelle D, Dandoy C, Burny A, Zavada J, Siakkou H, Gras-Masse H, Drobecq H, Tartar A (1989) Synthetic peptides approach to identification of epitopes on bovine leukemia virus envelope glycoprotein gp51. Virology 169:34–41
Rodriguez SM, Marcelo D, Golemba Campos RH, Trono K, Jones LR (2009) Bovine leukemia virus can be classified into seven genotypes: evidence for the existence of two novel clades. J Gen Virol 90:2788–2797
Sagata N, Yasunaga T, Tsuzuku-Kawamura J, Ohishi K, Ogawa Y, Ikawa Y (1985) Complete nucleotide sequence of the genome of bovine leukemia virus: its evolutionary relationship to other retroviruses. Proc Natl Acad Sci USA 82:677–681
Straub OC (1987) Natural and experimental transmission of bovine leukemia virus. In: Burny A, Mammerickx M (eds) Enzootic bovine leucosis and bovine leukemia virus. Martinus Nijhoff Publishing, Boston, pp 229–249
Szynal M, Cleuter Y, Beskorwayne T, Baganis C, Van Lint C, Kerkhofs P, Burny A, Martiat P, Griebel P, Van den Broke A (2003) Disruption of B-cell homeostatic mediated by the BLV-Tax oncoprotein: association with upregulation of Bcl-2 and signaling through NF-kB. Oncogene 22:4531–4542
The World Organisation for Animal Health (2004) Enzootic bovine leukosis. In: Manual of diagnostic tests and vaccines for terrestrial animals. http://www.oie.int/eng/normes/mmanual/A_00055.htm
Willems L, Heremans H, Chen G, Portetelle D, Billiau A, Burny A, Kettmann R (1990) Cooperation between bovine leukaemia virus transactivator protein and Ha-ras oncogene product in cellular transformation. EMBO J 9:1577–1581
Acknowledgments
This work was supported in part by a grant from the Ministry of Health, Labor and Welfare of Japan. The authors would like to thank the Hokkaido Obihiro, Hayakita, and Yakumo, Aomori Prefecture Towada, Tochigi Prefecture Kenkita, Ibaraki Prefecture Kenkita and Kennishi, and Shizuoka Prefecture Eastern District Meat Inspection Offices, Iwate, Gunma, Gifu, Okayama, Kagawa, and Tokushima Prefectural Meat Inspection Offices, Yokohama and Kumamoto City Meat Inspection Offices, as well as the Nara Prefectural Food Inspection Office for their extensive help in collecting samples.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Inoue, E., Matsumura, K., Maekawa, K. et al. Genetic heterogeneity among bovine leukemia viruses in Japan and their relationship to leukemogenicity. Arch Virol 156, 1137–1141 (2011). https://doi.org/10.1007/s00705-011-0955-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-011-0955-6