Abstract
Moringa oleifera is a less used, drought-tolerant tropical plant, rich in nutritionally and nutraceutically important bioactive compounds. It is native to India and now under cultivation in many countries, but no data is available on genetic variability. Three DNA marker techniques, i.e., random amplified polymorphic DNA (RAPD), inter simple sequence repeat (ISSR) and cytochrome P450 gene-based markers were used for the detection of genetic variability in eight Indian cultivars of M. oleifera, collected from various states of India. A total of 17 RAPD, 6 ISSR and 7 pairs of cytochrome P450-based markers generated 48.68, 48.57 and 40.00 % polymorphisms, respectively. The marker index (MI) for each of these marker systems (3.25 for RAPD, 4.73 ISSR and 2.95 for Cyt P450-based markers) suggest that ISSR markers are the most effective for assessment of genetic diversity. Based on the three types of marker data, the eight cultivars of M. oleifera were grouped into four sub-clusters in a dendrogram, but without any distinct geographical pattern. This suggests spread of planting material and high rates of gene flow through cross pollination. High bootstrap values (94.4 and 82.3) were obtained at major nodes of the dendrogram using the winboot software. The dendrogram and PCA plots generated from the binary data matrices of the three marker systems were found highly concordant to each other. This study reveals a huge genetic diversity among the cultivars and this can be utilised for conservation and cultivar development in breeding programmes to produce high yielding, nutritionally superior cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Moringa oleifera is a tropical tree, commonly known as horseradish or drumstick tree, which is diploid with 28 chromosomes. The tree is indigenous to North-western India and is now distributed in Pakistan, the Philippines, Cambodia, Central, North and South America and the Caribbean Islands (Morton 1991). In India and other countries, immature pods, fresh leaves and flowers of this tree are used for culinary purposes (Oduro et al. 2008; D’souza and Kulkarni 1993), with medicinal uses of roots and seeds (Ruckmani et al. 1998; Lalas and Tsaknis 2002). The leaves and pods are known as potential sources of vitamins and minerals being used as raw, cooked and dried (Fuglie 2005). Anti-hypertensive, anti-pyretic and anti-microbial activities of the seeds of M. oleifera have been very well studied (The Wealth of India 1962; Oliveira et al. 1999; Lalas and Tsaknis 2002).
Molecular markers are very useful in taxonomic and genetic analysis. The most commonly used marker are restriction fragment length polymorphisms (Soller and Beckmann 1983), amplified fragment length polymorphisms (AFLPs) (Kim et al. 2002), random amplified polymorphic DNAs (RAPDs) (Jacobson and Hedren 2007), microsatellites or simple sequence repeats (SSRs) (Fang et al. 2012) and inter simple sequence repeats (ISSRs) (Zietkiewicz et al. 1994). These techniques have been extensively used for assessment of genetic diversity in a variety of annual and perennial plants like Zea mays (Garcia et al. 2004), finger millet (Panwar et al. 2010), artemisia (Badr et al. 2012). These markers have advantages and disadvantages, but produce reliable information on genetic diversity in genetically neutral regions (Karp 2002).
Cytochrome P450 genes are widely found in micro-organisms, animals and plants (Schalk et al. 1999) and play important roles in oxidative detoxification and the secondary metabolite production (Ohkawa et al. 1998). Cytochrome P450 gene-analogues have been successfully used as new genetic markers for genetic diversity studies in plants, which reflects both functional and genome-wide regions (Yamanaka et al. 2003b).
Previously, assessment of genetic diversity in M. oleifera has been done using AFLP (Muluvi et al. 1999; Muluvi et al. 2004) and RAPD (Mgendi et al. 2010) in accessions from Kenya and Tanzania, respectively; however, other methods have not been used on Indian Moringa oleifera cultivars. Quantitative variation in nutritionally important phytoconstituents has been found in leaves of M. oleifera cultivars (R. K. Saini et al., our unpublished data), indicating a considerable potential for improvement of this tree by breeding and selection for nutrient-rich food and wide range of adaptation to different climatic and soil conditions. In the present study, we therefore used RAPD, ISSR and cytochrome P450-based markers for detecting the genetic variability in eight cultivars of M. oleifera collected from various states of India.
Materials and methods
Plant material
Commercially grown cultivars of M. oleifera were collected from different states of India. Cultivars PKM-1 and PKM-2 were collected from Kerala Agriculture University, Thrissur (Kerala). Cultivar CO-1 was collected from Tamil Nadu Agriculture University, Coimbatore (Tamil Nadu). Cultivars GKVK-1 and Dhanraj were collected from University of Agriculture Sciences, Bangalore (Karnataka). Cultivars Bhagya and Amar-32 were collected from University of Horticultural Sciences, Bagalkot (Karnataka) and the Amar Seeds Pvt. Ltd, Pune (Maharashtra), respectively. Cultivar PAVM-1 was collected from a progressive farmer in Tamil Nadu (India). The collected seeds were grown in plastic pots (10 l size) containing soil, sand and vermicompost (product of composting using earthworms) (3:1:1 v/v) in a greenhouse at ambient temperature (26–28 °C) (Saini et al. 2012). The soil type was red silt loam with fine-silty characteristics, which is the best for Moringa cultivation. At 2 months interval after germination, 10 g N–P–K (15:15:15) were applied to each pot. Newly opened (unfolded) leaves from 1-year-old M. oleifera plants (cultivars) were used for DNA extraction.
DNA extraction, purification and quantification
All the reagents used for DNA extraction, purification and polymerase chain reaction (PCR) amplification were of molecular biology grade and purchased from Sigma-Aldrich (Bangalore, India). Oligos were synthesised in BioServe Biotechnologies Pvt. Ltd, Hyderabad (India).
Genomic DNA was extracted from leaves of the different M. oleifera cultivars by the CTAB method, according to Murray and Thompson (1980) with some modifications to eliminate phenolics. From each cultivar, 1 g of leaves were ground in liquid nitrogen to a fine powder in a pre-chilled mortar and transferred into a 30 ml centrifuge tube containing 10 ml of DNA extraction buffer containing 0.1 M Tris chloride (pH 8.0), 0.02 M EDTA (pH 8.0), 1.4 M NaCl, 2 % CTAB (w/v), 2 % polyvinylpyrrolidone (w/v) and 0.2 % β-mercaptoethanol (v/v) and incubated in a water bath at 65 °C for 60 min. The tubes were cooled at room temperature and an equal amount of chloroform: isoamyl alcohol (24:1) was added, mixed thoroughly by gentle inversion and finally centrifuged at 10,000g for 15 min at 20 °C. The upper aqueous layer was transferred to a fresh sterile centrifuge tube and 0.7 vol chilled isopropanol was added and incubated overnight at −20 °C. The tubes were centrifuged at 10,000g for 15 min at 4 °C and the pellet was washed with 70 % ethanol, air-dried and dissolved in 200 μl TE buffer and stored at 4 °C. For DNA purification, 5 μl RNase (10 mg/ml) was added to total isolated DNA (200 μl) and incubated at 37 °C for 60 min. An equal volume of phenol:chloroform:isoamyl alcohol (25:24:1) was added and mixed gently. The tubes were centrifuged at 10,000g for 5 min and the aqueous layer was transferred to fresh Eppendorf tube and 1/10 volume of sodium acetate (3 M, pH 5.2) and a double volume of chilled absolute ethanol was added. After 30 min, the mixture was centrifuged at 10,000g for 5 min and finally, the pellet was washed with 70 % ethanol, dried and dissolved in 200 μl TE buffer. For quantification of genomic DNA, the absorbance of the DNA samples was measured at 260 nm in a Nano Drop 1000 (Thermo Scientific). After quantification, the quality of the purified DNA was analysed in a 0.7 % (w/v) agarose gel.
RAPD, ISSR and Cytochrome P450-based marker analyses
A total 25 RAPD primers, 10 ISSR primers and 11 Cytochrome P450-based primers were used for the polymorphism survey. The RAPD assay was carried out in 0.2 ml PCR vials containing 1× reaction buffer, 200 μM dNTPs mix, 0.4 μM primer, 50 ng DNA template, 1 U Taq DNA polymerase and sterile distilled water to a final volume of 25 μl. The content was gently mixed by spinning for a few seconds. The PCR amplification was performed with a thermocycler (Eppendorf, Germany). The standardised amplification was performed at an initial denaturation at 94 °C for 4 min, followed by 40 cycles of denaturation at 94 °C for 30 s; primer annealing based on Tm for 1 min; primer extension at 72 °C for 2 min and final primer extension at 72 °C for 10 min. For ISSR analysis, ISSR primers were used and other conditions was the same as for RAPD. For Cytochrome P450-based markers, PCR amplification was performed using 0.4 μM forward primer and 0.4 μM reverse primer in a volume of 25 μl with same PCR conditions as that of RAPD. The annealing temperature of each primer was calculated online by OligoAnalyzer 3.1 (http://www.idtdna.com/analyzer/Applications/OligoAnalyzer) according to the sequence of the oligos.
Polymerase chain reaction amplified products were analysed by agarose gel electrophoresis using 1.5 % agarose in 1× TAE buffer. The amplicon sizes were measured with a 100–5,000 bp DNA ladder. Respective gels were stained with 10 ppm ethidium bromide followed by image capturing using a gel documentation system (Herolab, Germany). The procedure was repeated twice for each DNA set and reproducible primers were subjected for diversity analysis.
Data analysis
The various sizes of amplified products were scored for presence (1) or absence (0) in the eight cultivars to generate a binary matrix. Binary matrix data were analysed by the software NTSYS-pc, version 2.11w, to calculate the Jaccard’s similarity coefficient (using the simqual programme) and generate a dendrogram based on the unweighted pair-group method using the arithmetic average (UPGMA) method (Rohlf 2001). Genetic similarity between cultivars was calculated according to Jaccard’s similarity coefficient (Jaccard 1980). The computer programme WINBOOT was used to determine the robustness of the dendrogram, with 2,000 replications along with Jaccard’s coefficient (Yap and Nelson 1996).
Percent polymorphisms were calculated for each primer combination according to the formula: % Polymorphism = p/(m + p), where p is total number of polymorphic bands and m is the total number of monomorphic bands of the primer combination used. Multiplex ratio (MR) for each marker was calculated by using the formula: MR = (m + p)/n, where p is the total number of polymorphic bands, m is the total number of monomorphic bands and n is the total number of primer combinations used (Powell et al. 1996). Polymorphic information content (PIC) was calculated using the formula: PIC = 1 − ∑pi2, where ‘pi’ is the frequency of the ith allele. Average heterozygosity (Hav) was obtained by taking the average of PIC values obtained for all the markers. Marker index (MI) was obtained by multiplying the average heterozygosity (Hav) with the multiplex ratio (MR).
Results
RAPD analysis
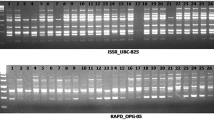
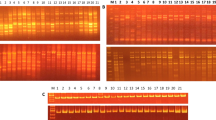
Randomly amplified polymorphic DNA assays were performed with 25 random primers. Of these, 17 produced polymorphic and reproducible bands and were selected for further screening. In an assay of the 17 RAPD primers on the eight cultivars, 76 bands were scored in the size range 0.3–3.0 kb, with an average of 4.47 bands per primer. Of the 76 bands, 37 were polymorphic (Table 1), with 48.68 % polymorphisms. RAPD-15 was found most efficient with 87.50 % polymorphism out of eight bands. A representative RAPD profile obtained by primer RAPD-10 and 11 in the 8 cultivars of M. oleifera is shown in Fig. 1a and b. The PIC values, a measure of informativeness of a marker, were found in the range from 0.49 (RAPD-17) to 0.817 (RAPD-15) with a mean of 0.726 (average heterozygosity).
ISSR analysis
Inter-Simple Sequence Repeat (ISSR) assays were performed with 10 ISSR primers. Of these, six gave polymorphic and reproducible bands and were selected for further screening. In an assay of the 6 ISSR primers on the 8 cultivars, 35 bands were scored in the size range 0.2–2.5 kb, with an average of 5.83 bands per primer. Of the 35 bands, 17 were polymorphic (Table 2), with 48.57 % polymorphism. ISSR-6 was found most efficient with four polymorphic bands out of six bands (66.66 %). A representative ISSR profile obtained by primer ISSR-3 and six in the eight cultivars of M. oleifera is shown in Fig. 1c and d. The PIC values were found in the range from 0.744 (ISSR-4) to 0.871 (ISSR-12) with a mean of 0.810.
Cytochrome P450-based markers analysis
Cytochrome P450-based marker assays were performed with 11 set of Cyt P450-based (forward and reverse) primers. Of these, seven produced polymorphic and reproducible bands and were selected for further screening. Tests of 7 sets of Cyt P450-based primers on the 8 cultivars gave 30 bands in the size range 0.1–1.2 kb, with an average of 4.28 bands per primer. Of the 30 bands, 12 were polymorphic (Table 3), with 40.00 % polymorphisms. CYP1A1 F with CYP1A1 R was found to be most efficient with 2 polymorphic bands out of 2 bands (100.0 %). A representative profile obtained by primer Cyt 02 F with Cyt 02 R and Cyt 03 F with Cyt 03 R in the eight cultivars of M. oleifera is shown in Fig. 1e and f. The PIC values were recorded in the range from 0.444 (CYP1A1F with CYP1A1R) to 0.838 (Cyt 02 F with Cyt 02 R) with a mean of 0.689.
Statistical analysis
Jaccard’s similarity coefficient, which reflects the genetic similarity among the cultivars, was calculated using binary data based on the RAPD, ISSR and cytochrome P450-based markers and is shown in Table 4. The Jaccard’s similarity coefficient ranged from 0.689 (between PKM-1 and PAVM-1) to 0.875 (between Dhanraj and Amar-32). The dendrogram, constructed according to the UPGMA method and based on the RAPD, ISSR and Cyt P450-based markers data, grouped the eight cultivars into two separate clusters (Fig. 2). The principal component analysis of RAPD, ISSR and Cyt P450-based markers revealed that the genotypes belonging to a particular sub-cluster were grouped together in the PCA plots (Fig. 3). The coefficient of correlation (Pearson’s correlation coefficients) was determined by a mantel test using matrices generated by these markers. The correlation coefficient between the ISSR and Cyt P450-based markers, between the RAPD and Cyt P450-based markers and between the ISSR and RAPD markers was 0.115, 0.097 and 0.0013, respectively. The correlation coefficient between Cyt P450-based markers and the combined RAPD, ISSR and Cyt P450-based markers, between the ISSR and the combined RAPD and ISSR and between the Cyt P450-based marker, the RAPD markers and the combined RAPD, ISSR and Cyt P450-based markers was 0.557, 0.237 and 0.580, respectively. The three molecular markers, ISSR, RAPD and Cyt P450, were compared on the basis of Percentage polymorphism, Multiplex ratio (MR), Average heterozygosity (Hav) and Marker index (Table 5). In relation to per cent polymorphisms, ISSR markers scored higher (48.57 %) than RAPD (48.68 %) and cytochrome P450-based markers (40 %). The Marker index (MI), which is a measure to evaluate the overall usefulness of a marker system considering all parameters, was highest for ISSR (4.726), compared to RAPD (3.246) and Cyt P450-based markers (2.954), making the former a highly efficient marker for the genetic diversity studies in M. oleifera.
The eight cultivars of M. oleifera were grouped into two major clusters using a Jaccard similarity coefficient of 0.75. Cluster 1 comprises 5 cultivars; PKM-1, PKM-2, GKVK-1, Dhanraj and Amar-32. Cluster 2 contains 3 cultivars; CO-1, Bhagya and PAVM-1. Bootstrap analysis from the binary data was used to determine the robustness of the dendrogram and confidence values as percentages at each node. Very high bootstrap values (94.4 and 82.3) were obtained from the major nodes (Fig. 2), indicating the robustness of these groups.
Discussion
Information on genetic variability of wild and cultivated plants is important for identification, conservation and cultivar development (Demir et al. 2010) and DNA markers are the most appropriate to study the genetic diversity among a set of genotypes or cultivars. In the present study, the combination of non-functional (RAPD and ISSR) and functional (Cyt P450 based) marker systems were used to provide wider genome coverage and, therefore, will be a better indicator of the genetic relationships among the eight cultivars.
Assessment of genetic diversity in M. oleifera accessions has been reported using AFLP (Muluvi et al. 1999) and RAPD markers (Muluvi et al. 2004; Mgendi et al. 2010). Muluvi et al. (1999) analysed the genetic diversity by AFLP among 140 accessions of M. oleifera, collected from south India, Southern Malawi and Kenya and found the highest levels of genetic diversity within the Indian genotypes. They used only one cultivar (PKM-1) from India whereas the other genotypes were from natural populations. In our study, we used eight commercially used Indian cultivars, among these 40.00 % (by ISSR markers) to 48.68 % (by RAPD markers) polymorphisms were recorded. This is the first report on genetic diversity among commercially grown Indian cultivars. Similarly, Mgendi et al. (2010) investigated the genetic diversity by RAPD markers between and within cultivated and non-cultivated provenances of M. oleifera from coastal regions of Tanzania and found that wild provenances were more diverse then cultivated. In cultivated plants, genetic changes are slowed down because of reduced competition and natural selection, due to optimum environmental conditions (Manoko et al. 2008). However, there is lack of information for Indian cultivated genotypes since India is the centre of origin for M. oleifera. Similarly, there is a need to evaluate the efficiency of other molecular markers for proper assessment of genetic diversity in M. oleifera, particularly markers based on Cyt P450 genes to be exploited for assessing the intra-species diversity.
In the present study, we used RAPD, ISSR and Cyt P450-based markers to analyse the genetic diversity between eight Indian cultivars of M. oleifera”. The three molecular markers used detected an average of 46.80 % polymorphisms (genetic diversity) among the cultivars. RAPD and ISSR revealed an average of 48.68 and 48.57 % polymorphism across all cultivars. The Jaccard’s similarity coefficient was calculated based on RAPD, ISSR and cytochrome P450 markers and it was found in the range of 0.689 to 0.875. Similar values of this coefficient were obtained between cultivars of close relatives of M. oleifera, i.e., papaya and Phaseolus vulgaris (Stiles et al. 1993; Maciel et al. 2003).
In the cluster analysis, constructed according to the UPGMA method and based on the three sets of marker data, the cultivars were not clustered according to their geographical origin. This may be due to spread of planting materials and high rates of gene flow through cross-pollination since this plant is highly cross-pollinated and different cultivars were collected from different states of India, i.e., Karnataka, Tamil Nadu, Maharashtra and Kerala. These states are located in close proximity of each other. Several studies have reported similar observations in other plants, e.g., Andrographis paniculata and Olea europaea (Padmesh et al. 1999; Bronzini de Caraffa et al. 2002), whereas some other studies reported the grouping of genotypes in dendrogram on the basis of their geographical collection centres (Das et al. 2007; Panwar et al. 2010). The cluster (dendrogram) was bootstrapped by the WINBOOT program (Yap and Nelson 1996) with 2,000 replications as suggested by Hedges (1992) and the values obtained in terms of percentages were high (94.4 and 82.3) at major nodes, suggesting the robustness of the dendrogram.
The comparative analysis of RAPD, ISSR and Cyt P450-based marker systems revealed ISSR to be the best marker as it generated highest percentage of polymorphisms, marker index (MI), average heterogeneity (Hav) and multiplex ration (MR). In the present study, Cytochrome P450-based markers were not found very efficient, due to low value of PIC compared to RAPD and ISSR markers, but these new markers have been found very efficient in evaluation of diverse species such as rice and various endemic wild plants (Tanaka et al. 2001; Yamanaka et al. 2003a). Similarity Pearson’s correlation coefficients was found very low (R 2 = 0.115, between ISSR and Cyt P450-based markers), showing a very poor fit of similarity matrix between two markers. This indicates that the two sets of markers target different regions of the genome. Similar results of low levels of correlation between RAPD and ISSR markers were observed by Lalhruaitluanga and Prasad (2009) in Melocanna baccifera. However, Panwar et al. (2010) observed the high value of coefficient of correlation between RAPD and Cyt P450 gene-based markers.The dendrogram and PCA plots generated from the binary data matrices of the three marker systems was found highly concordant to each other. In dendrogram, all the eight cultivars were grouped in four major sub-clusters, similar pattern of grouping was recorded in PCA plot. The similar type of agreements between dendrogram and PCA plot was reported by Panwar et al. (2010).
The high genetic diversity (variability) found in the present study is directly linked with higher levels of biodiversity, which is useful for food security, productivity and ecological sustainability. Significant variability can be utilised in breeding programmes to produce high-yielding nutritionally superior cultivars with better adaptations to different climatic conditions. In India, major breeding programmes are on-going for the development of fast growing, disease resistant and high pod yielding cultivars of M. oleifera. Thus, cv. Bhagya (KDM-1) has recently been developed by University of Horticultural Sciences, Bagalkot (Karnataka) and it is gaining popularity among farmers, due to its high yield of pods. Similarly, in future, attention is required to develop cultivars for high foliage yield, since Moringa leaves are a rich and affordable source of major nutritional and nutraceutical phytoconstituents (Oduro et al. 2008).
Abbreviations
- PCR:
-
Polymerase chain reaction
- RAPD:
-
Random amplified polymorphic DNA
- ISSR:
-
Inter simple sequence repeats
- PIC:
-
Polymorphism information content
- UPGMA:
-
Unweighted pair group method with arithmetic averages
- PCA:
-
Principal component analysis
- MI:
-
Marker index
- MR:
-
Multiplex ratio
References
Badr A, El-Shazly HH, Helail NS, El-Ghanim W (2012) Genetic diversity of Artemisia populations in central and north Saudi Arabia based on morphological variation and RAPD polymorphism. Plant Syst Evol 298(5):871–886
Bronzini de Caraffa V, Giannettini J, Gambotti C, Maury J (2002) Genetic relationships between cultivated and wild olives of corsica and sardinia using RAPD markers. Euphytica 123:263–271
D’souza J, Kulkarni AR (1993) Comparative studies on nutritive values of tender foliage of seedlings and mature plants of Moringa oleifera Lam. J Econ Taxon Bot 17:479–485
Das S, Mishra RC, Rout GR, Aparajita S (2007) Genetic variability and relationships among thirty genotypes of finger millet Eleusine coracana L. Gaertn. using RAPD markers. Z Naturforsch 62c:116–122
Demir K, Bakir M, Sarikamiş G, Acunalp S (2010) Genetic diversity of eggplant (Solanum melongena) germplasm from Turkey assessed by SSR and RAPD markers. Genet Mol Res 9(3):1568–1576
Fang W, Cheng H, Duan Y, Jiang X, Li X (2012) Genetic diversity and relationship of clonal tea (Camellia sinensis) cultivars in China as revealed by SSR markers. Plant Syst Evol 298(2):469–483
Fuglie LJ (2005) The Moringa Tree: a local solution to malnutrition? Church World Service in Senegal
GarciaI AAF, Benchimo LL, Barbosa AMM, Geraldi IO, Souza CL Jr, De Souza AP (2004) Comparison of RAPD, RFLP, AFLP and SSR markers for diversity studies in tropical maize inbred lines. Genet Mol Biol 27:579–588
Hedges SB (1992) The number of replications needed for accurate estimation of the bootstrap value in phylogenetic studies. Mol Biol Evol 9:366–369
Jaccard P (1980) New Researchers on the distribution florale. Waldensian Soc Nat Sci 44:22–270
Jacobson A, Hedren M (2007) Phylogenetic relationships in Alisma (Alismataceae) based on RAPDs, and sequence data from ITS and trnL. Plant Syst Evol 265:27–44
Karp A (2002) The new genetic era: will it help us in managing genetic diversity? In: Engels JMM, Rao VR, Brown AHD, Jackson MT (eds) Managing plant genetic diversity. CAB Publishing, Wallingford, pp 43–56
Kim MS, Moore PH, Zee F, Fitch MMM, Steiger DL, Manshardt RE et al (2002) Genetic diversity of Carica papaya as revealed by AFLP markers. Genome 45(3):503–512
Lalas S, Tsaknis J (2002) Extraction and identification of natural antioxidants from the seeds of Moringa oleifera tree variety of Malavi. J Am Oil Chem Soc 79:677–683
Lalhruaitluanga H, Prasad MNV (2009) Comparative results of RAPD and ISSR markers for genetic diversity assessment in Melocanna baccifera Roxb. growing in Mizoram State of India. Afr J Biotechnol 8(22):6053–6062
Maciel FL, Echeverrigaray S, Gerald LTS, Grazziotin FG (2003) Genetic relationships and diversity among Brazilian cultivars and landraces of common beans (Phaseolus vulgaris L.) revealed by AFLP markers. Genet Resour Crop Evol 50(8):887–893
Manoko MLK, van den Berg RG, Feron RMC, Van der Weerden GM, Mariani C (2008) Genetic diversity of the African hexaploid species Solanum scrabrum Mill. and Solanum nigrum L. (Solanaceae). Genet Resour Crop Evol 55(3):409–418
Mgendi MG, Manoko MK, Nyomora AM (2010) Genetic diversity between cultivated and non-cultivated Moringa oleifera Lam. provenances assessed by RAPD markers. J Cell Mol Biol 8(2):95–102
Morton JF (1991) The horse radish tree: M. pterigosperma (Moringacea). A boon to arid lands. Econ Bot 45:318–333
Muluvi GM, Sprent JI, Soranzo N, Provan J, Odee D, Folkard G, McNicol JW, Powell W (1999) Amplified fragment length polymorphism (AFLP) analysis of genetic variation in Moringa oleifera Lam. Mol Ecol 8:463–470
Muluvi GM, Sprent JI, Odee D, Powell W (2004) Estimates of outcrossing rates in Moringa oleifera using Amplified fragment length polymorphism (AFLP). Afr J Biotechnol 3(2):146–151
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Oduro I, Ellis WO, Owusu D (2008) Nutritional potential of two leafy vegetables: Moringa oleifera and Ipomoea batatas leaves. Sci Res Essay 3(2):57–60
Ohkawa H, Imaishi H, Shiota N, Yamada T, Inui H, Ohkawa Y (1998) Molecular mechanisms of herbicide resistance with special emphasis on cytochrome P450 mono-oxigenases. Plant Biotech 15:173–176
Oliveira JTA, Silveira SB, Vasconcelos IM, Cavada BS, Moreira RA (1999) Compositional and nutritional attributes of seeds from the multipurpose tree Moringa oleifera Lamarck. J Sci Food Agric 79:815–820
Padmesh P, Sabu KK, Seeni S, Pushpangadan P (1999) The use of RAPD in assessing genetic variability in Andrographis paniculata Nees, a hepatoprotective drug. Curr Sci 76:833–835
Panwar P, Saini RK, Sharma N, Yadav D, Kumar A (2010) Efficiency of RAPD, SSR and Cytochrome P450 gene based markers in accessing genetic variability amongst finger millet (Eleusine coracana) accessions. Mol Bio Rep 37:4075–4082
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) marker for germplasm analysis. Mol Breed 2(3):225–238
Rohlf FJ (2001) NTSYS-pc numerical taxonomy and multivariate analysis system.Version 5.1. Exeter Publishing Ltd, Setauket
Ruckmani K, Kavimani S, Anandan R, Jaykar B (1998) Effect of Moringa oleifera Lam on paracetamol-induced hepatoxicity. Indian J Pharm Sci 60:33–35
Saini RK, Shetty NP, Giridhar P, Ravishankar GA (2012) Rapid in vitro regeneration method for Moringa oleifera and performance evaluation of field grown nutritionally enriched tissue cultured plants. 3 Biotech 2(3):187–192
Schalk M, Nedelkina S, Schoch G, Batard Y, Werck-Reichhart D (1999) Role of unusual amino-acid residues in the proximal and distal heme regions of a plant P450, CYP73A1. Biochem 38:6093–6103
Soller M, Beckmann JS (1983) Genetic polymorphism in varietal identification and genetic improvement. Theor Appl Genet 67:25–33
Stiles JI, Lemme C, Sondur S, Morshidi MB, Manshardt R (1993) Using randomly amplified polymorphic DNA for evaluating genetic relationships among papaya cultivar. Theor Appl Genet 85:697–701
Tanaka M, Watanabe JA, Watanabe KN (2001) Presence of genetic variation in the endangered semi-cultivated species Suita Kuwai (Sagittaria triforia L. var. suitensis). Breed Res 3:162
The Wealth of India (1962) A dictionary of Indian raw materials and industrial products, raw materials, vol. VI: L–M; Council of Scientific and Industrial Research, New Delhi
Yamanaka S, Nakamura I, Sato YI, Watanabe KN (2003a) Evaluation on genetic resources based on the diversity of functional regions 1 Genetic diversity of wild rice estimated by variations on xenobiotics-metabolizing cytochrome P450 gene-analogues. Breeding Res 5:249
Yamanaka S, Suzuki E, Tanaka M, Takeda Y, Watanabe JA, Watanabe KN (2003b) Assessment of cytochrome P450 sequences offers a useful tool for determining genetic diversity in higher plant species. Theor Appl Genet 108:1–9
Yap IV, Nelson RJ (1996) In WinBoot: A program for performing bootstrap analysis for binary data to determine the confidence limits of UPGMA-based dendrograms. Discussion paper series number 14, International Rice Research Institute, Manila
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saini, R.K., Saad, K.R., Ravishankar, G.A. et al. Genetic diversity of commercially grown Moringa oleifera Lam. cultivars from India by RAPD, ISSR and cytochrome P450-based markers. Plant Syst Evol 299, 1205–1213 (2013). https://doi.org/10.1007/s00606-013-0789-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-013-0789-7