Abstract
A novel surface plasmon resonance (SPR) strategy is introduced for the specific determination of exosomes based on aptamer recognition and polydopamine-functionalized gold nanoparticle (Au@PDA NP)–assisted signal amplification. Exosomes derived from hepatic carcinoma SMMC-7721 were selected as the model target. SMMC-7721 exosomes can be specifically captured by the aptamer ZY-sls that was complementary to the DNA tetrahedron probes (DTPs), and then the CD63 aptamer–linked Au@PDA NPs recognized SMMC-7721 exosomes for signal amplification. The DTPs were modified on a Au film for preventing Au deposition on the surface during the introduction of HAuCl4, and PDA coated on the AuNPs was used to reduce HAuCl4 in situ without any reductant assistance. It results in a further enhanced SPR signal. The assay can clearly distinguish SMMC-7721 exosomes from others (HepG2 exosomes, Bel-7404 exosomes, L02 exosomes, MCF-7 exosomes, and SW480 exosomes, respectively). SMMC-7721 exosomes are specifically determined as low as 5.6 × 105 particles mL−1. The method has successfully achieved specific determination of SMMC-7721 exosomes even in 50% of human serum without any pretreatment.

A novel surface plasmon resonance (SPR) strategy was introduced for the specific determination of exosomes based on aptamer recognition and polydopamine functionalized gold nanoparticles (Au@PDA NPs). The SPR signal was improved using the Au@PDA NPs assisted amplification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Exosomes, nanoscale membrane-encapsulated extracellular vesicles with size ranging from 30 to 150 nm, are secreted by many mammalian cell [1, 2] and present in many different human biofluids [3]. In addition, exosomes carrying some biomolecules (such as RNA/DNA and proteins) can involve the exchange of information between cells and affect the physiological functions of cells [1, 4, 5]. They are well considered as potential and non-invasive biomarkers for significant disease diagnosis [1, 5]. Therefore, it is very necessary to develop an effective method for determining related cancerous exosomes.
There are some researches focusing on the determination of exosomes, such as electrochemistry [6, 7], microfluidics [8, 9], fluorescence [10], surface-enhanced Raman scattering (SERS) [11, 12], and surface plasmon resonance (SPR) [13,14,15,16]. Most of these methods had to depend on antibodies so that they have some downsides. For example, preparation of antibody is tedious, expensive, and time-consuming. Besides, given that antibody is easy to denature towards extreme chemical environment, it is difficult for most of these methods to standardize. Aptamer, in addition to having an antibody-like function, has several advantages, such as ease of modification, excellent specificity, and high affinity [17,18,19,20,21,22]. Hence, aptamer has high potential as a replacement for antibody in the specific determination of cancerous exosomes. Wang et al. developed a portable electrochemical aptasensor for specific determination of hepatocellular exosomes [18]. Huang et al. introduced an aptasensor based on rolling circle amplification–assisted fluorescence signal amplification for specific determination of leukemia-derived exosomes [19]. Zhang et al. also reported electrogenerated chemiluminescence biosensors for MCF-7 exosome–specific determination based on Ti3C2 MXene nanosheets and two aptamers (EpCAM aptamer and CD63 aptamer) [20]. To our knowledge, the application of aptamers on SPR strategy to determine exosomes has so far been relatively rare. Our group has developed gold nanoparticle (AuNP)–amplified SPR biosensor using CD63 aptamer as recognition element for sensitive determination of MCF-7 exosomes [21]. Since CD63 is usually present on most exosomes, the SPR biosensor has poor specific determination of MCF-7 exosomes. Therefore, it is necessary to develop a novel method for specific determination of cancerous exosomes.
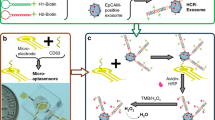
Herein, we first found that aptamer ZY-sls (Apt1), which was generated by the screening of human hepatic carcinoma (SMMC-7721) cells [23], can specifically bind SMMC-7721 exosomes using flow cytometry. Based on the finding, a novel SPR strategy was proposed for the specific determination of SMMC-7721 exosomes based on two aptamers (Apt1 and CD63 aptamer (Apt2)). The strategy is based on polydopamine-functionalized AuNP (Au@PDA NP)–assisted signal amplification, which consists of three main processes (Scheme 1). Firstly, the Apt1 that is complementary to DNA tetrahedron probes (DTPs) with efficient blocking ability and anti-gold deposition [24,25,26] can capture specifically SMMC-7721 exosomes. Then, Apt2-linked Au@PDA NPs (Au@PDA NP-Apt2) recognize SMMC-7721 exosomes captured by Apt1 to assist signal amplification. Subsequently, PDA on the surface of AuNPs can reduce HAuCl4 in situ without any reductant assistance when incubated with HAuCl4 [27, 28], resulting in further enhancement of the SPR signal. This work can also broaden the potential for specific determination of other cancerous exosomes by replacing other specific aptamers.
Experimental section
Preparation and characterization of Au@PDA NP-Apt2
Au@PDA NP-Apt2 was prepared as shown in Scheme S-1 in the Electronic Supporting Material (ESM). Typically, Au@PDA NPs were formed by in situ polymerizations of dopamine (DA) on the surface of AuNPs under alkaline conditions [29, 30]. Next, the preparation of Au@PDA NP-Apt2 was achieved by covalent bond cross-linking between the -NH2 on the PDA and the -COOH on Apt2 [27, 31]. The detail experimental procedures are listed in the ESM. Then, Au@PDA NPs and Au@PDA NP-Apt2 were characterized by UV-vis spectroscopy (Shimadzu, Japan), Zetasizer Nano ZS90 (Malvern Instruments, U.K.), Fourier transform-infrared (FT-IR) spectra (Brooke group), and transmission electron microscopy (TEM) (JEOL, Japan).
Exosome determination
SMMC-7721 exosomes were derived from human hepatic carcinoma (SMMC-7721) cell line that was purchased from Xiangya School of Medicine (Central South University, China). The specific determination of SMMC-7721 exosomes was conducted on a EC-SPR1010 device (Changchun Dingcheng Technology, China). The SPR chip was prepared following our previous work [21]. Firstly, DNA tetrahedral probes (DTPs) were assembled by referring to traditional simple denaturation and annealing processes [12, 13] (shown in ESM). Figure S-2 indicates that the DTPs are formed. Next, the DTPs (5.0 μM) were first incubated on Au film surface for 3 h at 25 °C. Figure S-3 suggests that DTPs are successfully modified on Au film. Then, the DTP-modified Au film surface was incubated with Apt1 (65 μL, 1.2 μM) for 40 min at 25 °C. Unbound Apt1 was repeatedly washed away with phosphate-buffered saline (10 mM KH2PO4/Na2HPO4, 137 mM NaCl, 2.7 mM KCl, pH 7.4), and the SPR spectrum was recorded. Next, SMMC-7721 exosomes were respectively introduced and incubated for 40 min at 25 °C, then rinsed thoroughly with phosphate-buffered saline and the SPR spectrum was recorded. Then, Au@PDA NP-Apt2 (65 μL, 4.2 nM) was introduced and incubated for 40 min at 25 °C, and the SPR spectrum was recorded after rinsing with phosphate-buffered saline thoroughly. Subsequently, HAuCl4 (65 μL, 0.24 mM) was injected and reacted for 20 min at 25 °C, and then the SPR spectrum was recorded after rinsing repeatedly with phosphate-buffered saline. Finally, the regeneration solution (65 μL, 0.1% SDS, 10 mM NaOH) was added for about 5 min to regenerate the SPR chip.
Analysis of real samples
SMMC-7721 exosome in normal human serum was also determined using this assay. The human serum was collected from healthy volunteers of the Hospital of Hunan University. The samples were first obtained through adding different concentrations of SMMC-7721 exosomes in 50% human serum. Then, the above samples were respectively injected and incubated with DTP-Au film for 40 min. After rinsing thoroughly with phosphate-buffered saline, SPR spectrum was recorded. Subsequently, Au@PDA NP-Apt2 (65 μL, 4.2 nM) were introduced and incubated for 40 min. SPR spectrum was recorded after being washed thoroughly with phosphate-buffered saline. Next, HAuCl4 (65 μL, 0.24 mM) was injected and reacted for 20 min. Afterward, SPR spectrum was recorded after rinsing repeatedly with phosphate-buffered saline. The concentration of the SMMC-7721 exosome in human serum was determined according to the change of resonance angle. The SPR chip was regenerated using 0.1% SDS/10 mM NaOH for the determination of other samples.
Results and discussion
Choice of materials
Given that SMMC-7721 exosome is one of the markers of human therapeutic carcinoma [32], it is selected as the model target. The SMMC-7721 exosomes were characterized using nanoparticle tracking analysis (NTA), dynamic light scattering (DLS), TEM, and scanning electron microscopy (SEM), respectively. Figure S-4 indicates that exosomes can be successfully obtained.
In our previous work [24], it was reported that DTP-Au film not only demonstrated superior antifouling property against protein and cell but also avoided Au deposition on the surface in the process of the catalytic growth of gold nanoparticles. Thus, DTP-Au film is used here for avoiding the Au deposition on the surface of the surface during the reduction of HAuCl4 by PDA.
Since the good optical properties of AuNPs can enhance the SPR signal and PDA can reduce HAuCl4 to form small AuNPs [27], Au@PDA NPs are selected here for signal amplification.
Characterization of Au@PDA NP-Apt2
The preparation of Au@PDA NP-Apt2 was confirmed by DLS, zeta potential, UV-vis absorption spectrum, and FT-IR spectra. As shown in Fig. 1(A), as the PDA and Apt2 are modified on the AuNPs respectively, the hydrodynamic size of the nanoparticles increases. As shown in Fig. 1(B), the zeta potential value decreases significantly, which is attributed to the shell of PDA and Apt2 attachment with a large amount of negative charge. Similarly, it also shows the maximum UV-vis absorption peak red shifts from 518 to 538 nm as PDA and Apt2 are modified on the AuNPs (Fig. S-5A). FT-IR profile is also used for the characterization of Au@PDA NP-Apt2. As shown in Fig. S-5B, the newly discovered absorption bands imply the presence of Au@PDA NP-Apt2 (detailed datums are showed in ESM). Simultaneously, the Au@PDA NPs also are confirmed by TEM images. Comparing with AuNPs (15.4 ± 4.5 nm) (Fig. 1(C)), the core-shell structure of Au@PDA NPs (24.2 ± 3.3 nm) is clearly visible under TEM image (Fig. 1(D)). In addition, the PDA layer (8.8 ± 1.2 nm) is clearly displayed on the periphery of the AuNPs. The results show that the Au@PDA NPs and Au@PDA NP-Apt2 can be obtained.
Investigation of the resistance of Au deposition using DTP-Au film
If HAuCl4 solution is incubated on the Au film, obvious background signal can be observed due to Au deposition on the Au film (Fig. S-6). Thus, the effect of DTP-Au film on the resistance of Au deposition was explored using the EC-SPR1010 device. After different concentrations of HAuCl4 solution were added and incubated for 20 min, the DTP-Au film was washed thoroughly with phosphate-buffered saline. Then the SPR response was recorded. Thus, the influence from the increase of the concentration of HAuCl4 solution can be ignored. As shown in Fig. 2, the resonance angle change (Δθ) increased with the increase of HAuCl4 concentration. Δθ states a background signal which is caused by the Au deposition on the chip as different concentrations of HAuCl4 solution are added. Given that the Δθ (> 0.0015°) is acted as a signal [29], 0.24 mM HAuCl4 is selected for the following experiment.
Effect of HAuCl4 on Au@PDA NP-Apt2
After DTP-Au film had good anti-gold deposition effect on HAuCl4 (0.24 mM), the effect of HAuCl4 (0.24 mM) on Au@PDA NP-Apt2 was also investigated. Figure 3 shows that the maximum UV-vis absorption peak moves from 538 to 545 nm as Au@PDA NP-Apt2 was incubated with 0.24 mM HAuCl4. Meanwhile, the color of the solution that incubated with 0.24 mM HAuCl4 changes from dark purple to deep blue. For the AuNP-Apt2 without PDA functionalization (Fig. S-1), the shift in the UV-vis spectrum of AuNP-Apt2 is observed to be negligible after incubation with 0.24 mM HAuCl4 (Fig. 3(B)). Meantime, the color of the solution hardly changes after incubation with 0.24 mM HAuCl4. It is supposed that the size of Au@PDA NP-Apt2 is changed by the reduction of HAuCl4 (0.24 mM) by PDA. Besides, the results are also further confirmed by TEM images. It is clearly observed that many small nanoparticles were coated on the Au@PDA NPs after incubating with HAuCl4 (Fig. 3(C)), while almost no changed is observed for AuNP-Apt2 (Fig. 3(D)). It also suggests that HAuCl4 is reduced by PDA on the Au@PDA NP surface. As shown in Fig. 3, the weight content of Au is increased from 13.5 to 28.07% by an energy dispersive spectrometer (EDS) profile (JEOL, Japan) (Fig. 3(E)), while a negligible change in the weight content of Au was also obtained by EDS profile (Fig. 3(F)). The results also demonstrate that the small AuNPs are deposited on the Au@PDA NP-Apt2 due to the reduction of HAuCl4 by PDA. Therefore, it suggests that Au@PDA NP-Apt2 can provide a feasible platform for assisted SPR signal amplification.
Effect of HAuCl4 (0.24 mM) on Au@PDA NP-Apt2. The change of UV-vis absorption spectra of Au@PDA NP-Apt2 (A) and AuNP-Apt2 (B) after incubating with HAuCl4. Inset: Photographs of nanoparticles before (left) and after (right) incubated with HAuCl4. TEM images of Au@PDA NP-Apt2 (C) and AuNP-Apt2 (D) after incubating with HAuCl4. EDS profiles of Au@PDA NP-Apt2 (E) and AuNP-Apt2 (F) after incubating with HAuCl4. Wt% represents the weight content of Au
Investigation of the feasibility
The specific recognition of the SMMC-7721 exosomes by the aptamer ZY-sls (Apt1) is a prerequisite in the successful construction of the SPR assay. This was first investigated using flow cytometry (Scheme S-2). As shown in Fig. 4(A), the comparison between the fluorescence intensities shows that the experiment group is approximately 80-fold higher than the control group. It provides solid evidence of the specific recognition between the Apt1 and SMMC-7721 exosomes. Simultaneously, the specific recognition between the Apt1 and SMMC-7721 exosomes was further investigated using EC-SPR1010 device. As shown in Fig. 4(B), in experiment group I, SMMC-7721 exosomes are incubated with Apt1 assembled on DTP-Au film to obtain a significant Δθ (0.0492° ± 0.0032). However, control group II is also performed using random sequence1 instead of Apt1, and a low Δθ is observed in comparison with the experiment group I. In addition, HepG2 exosomes (III), Bel-7404 exosomes (IV), and L02 exosomes (V) were used for the control exosomes. The results show that Δθ induced by these control exosomes is negligible. Therefore, the results suggest that the SPR assay for specific determination of SMMC-7721 exosomes can be constructed using Apt1 as the recognition probe.
The feasibility analysis of the method. Flow cytometry (A) and SPR (B) investigation of the specific recognition between Apt1 and SMMC-7721 exosomes. SPR response (Δθ) of Apt1 recognizing SMMC-7721 exosomes (I); random sequence1 recognizing SMMC-7721 exosomes (II); Apt1 recognizing HepG2 exosomes (III); Apt1 recognizing Bel-7404 exosomes (IV); and Apt1 recognizing L02 exosomes (V). (C) SPR spectra of DTP-Au film (1); hybridization with Apt1 (2); recognizing SMMC-7721 exosomes (3); reaction with Au@PDA NP-Apt2 (4); reaction with HAuCl4 (0.24 mM) (5); and regeneration (6). (D) Determination of target exosome using Au@PDA NPs (a), AuNP-Apt2 (b), and Au@PDA NP-Apt2 (c) as signal amplification. The final concentration of all exosomes was 6.2 × 106 particles mL−1
The feasibility of the assay was then studied. As shown in Fig. 4(C), when Apt1 was incubated with DTPs modified on the Au film, a tiny Δθ (0.0063°) is observed (curve 1 to curve 2). It demonstrates that the Apt1 is assembled on DTP-Au film through hybridization. Next, when SMMC-7721 exosomes were reacted with the Apt1 assembled on DTP-Au film, the SPR spectrum shifts from curve 2 to curve 3 and a clear Δθ (0.0492°) is obtained. It indicates that the SMMC-7721 exosomes are specifically captured by Apt1 assembled on DTP-Au film. Then, Au@PDA NP-Apt2 was injected; the SPR spectrum moves from curve 3 to curve 4 and a Δθ (0.0790°) is observed. It suggested that Au@PDA NP-Apt2 can enhance the SPR signal, implying that the Au@PDA NP-Apt2 are attached on the surface of SMMC-7721 exosomes through the recognition between Apt2 and SMMC-7721 exosomes. Furthermore, the binding between Au@PDA NP-Apt2 and exosomes is characterized using TEM. As shown in Fig. S-7 of the ESM, Au@PDA NPs can attach on the surface of exosomes via Apt2 recognition. Subsequently, HAuCl4 (0.24 mM) were injected; the SPR spectrum moves from curve 4 to curve 5 and a Δθ (0.0644°) is observed. It implies that Au@PDA NP-Apt2 can reduce HAuCl4 in situ to further significantly enhance the SPR signals. The results are also shown in Fig. 4(D); compared with Au@PDA NPs (curve a) and AuNP-Apt2 (curve b), a significant Δθ is obtained for Au@PDA NP-Apt2 after treatment with HAuCl4 (curve c). Finally, after regeneration solution was added, the SPR spectrum moves from curve 5 to curve 6, and then Δθ substantially coincides with the curve 6 to curve 1, indicating that the chip can achieve regeneration. Hence, it is concluded that the assay is feasible for the specific determination of SMMC-7721 exosomes under the Au@PDA NP-Apt2-assisted amplification of the SPR signal.
Exosome determination
The sensitivity of the method was significantly affected by the concentration of Apt1 and the reaction time of SMMC-7721 exosomes. Thus, the effects of the concentration of Apt1 and the reaction time of SMMC-7721 exosomes were tested. The results indicate that the optimal conditions for the SPR assay are 1.2 μM for Apt1 (Fig. S-8A) and 40 min for SMMC-7721 exosome reaction (Fig. S-8B) Next, the SPR biosensor is used to determine SMMC-7721 exosomes under optimal conditions. As shown in Fig. 5(A), the SPR resonance angle gradually increases as the concentration of SMMC-7721 exosomes increased (from 0 to 7.9 × 107 particles mL−1). Figure 5 (B) illustrates the comparison between the Δθ response and the concentration of SMMC-7721 exosomes for the three processes (direct determination of SMMC-7721 exosomes (curve a); Au@PDA NP-Apt2-amplified SPR assay (curve b); Au@PDA NP-Apt2- and HAuCl4-amplified SPR assay (curve c)). The results show that the determination limit of direct assay, Au@PDA NP-Apt2-amplified assay, and Au@PDA NP-Apt2- and HAuCl4-amplified assay is 7.0 × 107 particles mL−1, 6.2 × 106 particles mL−1, and 5.6 × 105 particles mL−1, respectively. It further indicates that the method significantly improves the sensitivity of SMMC-7721 exosomes with the assistance of Au@PDA NP-Apt2 and HAuCl4. Compared with the existing method, the SPR assay based on aptamer and Au@PDA NPs is sensitive and specific for determining target exosomes in complex samples without any pretreatment (shown in Table S-2). Although many methods can sensitively determine the exosomes, most of these methods use antibody as the recognition probe, which was easy to denature and difficult to standardize. Here, the method for the determination of exosomes based on economical and stable aptamers probe has the same performance as the existing methods. This work broadens the application of aptamer based SPR technology.
Exosome determination using the SPR assay. (A) SPR spectra of different concentrations of SMMC-7721 exosomes. (B) The relationship between the Δθ and the concentration of SMMC-7721 exosomes by (a) directing determination, (b) Au@PDA NP-Apt2-amplified SPR assay, (c) Au@PDA NP-Apt2- and HAuCl4-amplified SPR assay. (C) The selective investigation of the SPR assay. The final concentration of all exosomes was 6.2 × 106 particles mL−1. (D) Reproducibility investigation of the SPR assay
Next, the specificity of the SPR assay was investigated by using HepG2 exosomes, Bel-7404 exosomes, L02 exosomes, MCF-7 exosomes, and SW480 exosomes under the same condition. As shown in Fig. 5(C), the Δθ exhibits negligible response for HepG2 exosomes, Bel-7404 exosomes, L02 exosomes, MCF-7 exosomes, and SW480 exosomes. However, in the case of SMMC-7721 exosomes or SMMC-7721 exosomes with a mix of other exosomes, it causes a significant change of Δθ (0.1204° ± 0.078). The selectivity of the SPR assay was further investigated to specifically determine SMMC-7721 exosomes in a mixture of SMMC-7721 exosomes and other exosomes (HepG2 exosomes, Bel-7404 exosomes, L02 exosomes, MCF-7 exosomes, and SW480 exosomes, respectively). As shown in Fig. S-9, the Δθ response caused by SMMC-7721 exosomes is similar to that aroused using the mixing of SMMC-7721 exosomes with other exosomes. The results demonstrate that SPR assay show excellent selectivity for SMMC-7721 exosomes.
The regeneration of the SPR assay was studied. As shown in Fig. 5(D), as the SPR chip is dealt with regeneration solution after each determination of target exosomes (6.2 × 106 particles mL−1), and 6 cycles of regeneration are observed without significant signal attenuation. The relative standard deviation (RSD) of 4.3% was also obtained. The results suggest that the assay has good reproducibility.
Determination of SMMC-7721 exosomes in serum
Given that the assay has excellent specificity for SMMC-7721 exosomes (Fig. 5(C)), it can avoid the concern of high levels of biological noise in the absence of a series of human serum pretreatments [19]. Here, for investigating the application of this assay, SMMC-7721 exosomes in 50% human serum are determined without any pretreatment. Figure S-10 illustrates the comparison of the Δθ obtained with determining in 50% human serum and in PBS buffer. It shows that Δθ caused by the same concentration of exosomes in 50% human serum and PBS buffer is almost the same. Besides, the SMMC-7721 exosomes in 50% human serum samples were also determined using the standard addition method. As shown in Table 1, the recoveries of SMMC-7721 exosomes in 50% human serum range from 95.3 to 103.6%, and the RSD is less than 5%. The results indicate that the method has the potential for the specific determination of SMMC-7721 exosomes in liquid biopsy.
Conclusions
A novel SPR assay based on aptamers and Au@PDA NPs was developed for the specific determination of SMMC-7721 exosomes. This assay shows excellent specificity and stability for the determination of the SMMC-7721 exosome and can be used in 50% human serum analysis. It has the potential for the determination of SMMC-7721 exosomes in liquid biopsy. Theoretically, this assay can potentially be used for determining the other target exosome, as long as an appropriate aptamer can be replaced. However, it has complex operation steps and is limited to use in the laboratory. Therefore, it is required to simplify the operation steps in the future.
References
van Niel G, D’Angelo G, Raposo G (2018) Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19:213–228
Yang BW, Chen Y, Shi JL (2018) Exosome biochemistry and advanced nanotechnology for next-generation theranostic platforms. Adv Mater:31. https://doi.org/10.1002/adma.201802896
Brown L, Wolf JM, Prados-Rosales R, Casadevall A (2015) Through the wall: extracellular vesicles in gram-positive bacteria, mycobacteria and fungi. Nat Rev Microbiol 13:620–630
Shao H, Im H, Castro CM, Breakefield X, Weissleder R, Lee H (2018) New technologies for analysis of extracellular vesicles. Chem Rev 118:1917–1950
Pick H, Alves AC, Vogel H (2018) Single-vesicle assays using liposomes and cell-derived vesicles: from modeling complex membrane processes to synthetic biology and biomedical applications. Chem Rev 118:8598–8654
Boriachek K, Masud MK, Palma C, Phan HP, Yamauchi Y, Hossain MSA, Nguyen NT, Salomon C, Shiddiky MJA (2019) Avoiding pre-isolation step in exosome analysis: direct isolation and sensitive detection of exosomes using gold-loaded nanoporous ferric oxide nanozymes. Anal Chem 91:3827–3834
Boriachek K, Islam MN, Gopalan V, Lam AK, Nguyen NT, Shiddiky MJA (2017) Quantum dot-based sensitive detection of disease specific exosome in serum. Analyst 142:2211–2220
Chen Z, Cheng SB, Cao P, Qiu QF, Chen Y, Xie M, Xu Y, Huang WH (2018) Detection of exosomes by ZnO nanowires coated three-dimensional scaffold chip device. Biosens Bioelectron 122:211–217
Kang YT, Kim YJ, Bu J, Cho YH, Han SW, Moon BI (2017) High-purity capture and release of circulating exosomes using an exosome-specific dual-patterned immunofiltration (ExoDIF) device. Nanoscale 9:13495–13505
Lewis JM, Vyas AD, Qiu Y, Messer KS, White R, Heller MJ (2018) Integrated analysis of exosomal protein biomarkers on alternating current electrokinetic chips enables rapid detection of pancreatic cancer in patient blood. ACS Nano 12:3311–3320
Li T, Zhang R, Chen H, Huang Z, Wang XH, Deng A, Kong J (2018) An ultrasensitive polydopamine bi-functionalized SERS immunoassay for exosome-based diagnosis and classification of pancreatic cancer. Chem Sci 9:5372–5382
Zong SF, Wang L, Chen C, Lu J, Zhu D, Zhang YZ, Wang ZY, Cui YP (2016) Facile detection of tumor-derived exosomes using magnetic nanobeads and SERS nanoprobes. Anal Methods 8:5001–5008
Liu C, Zeng X, An Z, Yang Y, Eisenbaum M, Gu X, Jornet JM, Dy GK, Reid ME, Gan Q, Wu Y (2018) Sensitive detection of exosomal proteins via a compact surface plasmon resonance biosensor for cancer diagnosis. ACS Sens 3:1471–1479
Qiu GY, Thakur A, Xu C, Ng SP, Lee Y, Wu CML (2019) Detection of glioma-derived exosomes with the biotinylated antibody-functionalized titanium nitride plasmonic biosensor. Adv Funct Mater 29(9). https://doi.org/10.1002/adfm.201806761
Liang K, Liu F, Fan J, Sun D, Liu C, Lyon CJ, Bernard DW, Li Y, Yokoi K, Katz MH, Koay EJ, Zhao Z, Hu Y (2017) Nanoplasmonic quantification of tumour-derived extracellular vesicles in plasma microsamples for diagnosis and treatment monitoring. Nat Bio Eng 1(4):1–11. https://doi.org/10.1038/s41551-016-0021
Picciolini S, Gualerzi A, Vanna R, Sguassero A, Gramatica F, Bedoni M, Masserini M, Morasso C (2018) Detection and characterization of different brain-derived subpopulations of plasma exosomes by surface plasmon resonance imaging. Anal Chem 90:8873–8880
Wang YM, Liu JW, Adkins GB, Shen W, Trinh MP, Duan LY, Jiang JH, Zhong WW (2017) Enhancement of the intrinsic peroxidase-like activity of graphitic carbon nitride nanosheets by ssDNAs and its application for detection of exosomes. Anal Chem 89:12327–12333
Wang S, Zhang L, Wan S, Cansiz S, Cui C, Liu Y, Cai R, Hong C, Teng I, Shi M, Wu Y, Dong Y, Tan W (2017) Aptasensor with expanded nucleotide using DNA nanotetrahedra for electrochemical detection of cancerous exosomes. ACS Nano 11:3943–3949
Huang L, Wang DB, Singh N, Yang F, Gu N, Zhang XE (2018) A dual-signal amplification platform for sensitive fluorescence biosensing of leukemia-derived exosomes. Nanoscale 10:20289–20295
Zhang HX, Wang ZH, Zhang QX, Wang F, Liu Y (2019) Ti3C2 MXenes nanosheets catalyzed highly efficient electrogenerated chemiluminescence biosensor for the detection of exosomes. Biosens Bioelectron 124-125:184–190
Wang Q, Zou L, Yang X, Liu X, Nie W, Zheng Y, Cheng Q, Wang K (2019) Direct quantification of cancerous exosomes via surface plasmon resonance with dual gold nanoparticle-assisted signal amplification. Biosens Bioelectron 135:129–136
Huang R, He L, Xia Y, Xu H, Liu C, Xie H, Wang S, Peng L, Liu Y, Liu Y, He N, Li Z (2019) A sensitive aptasensor based on a hemin/G-quadruplex assisted signal amplification strategy for electrochemical detection of gastric cancer exosomes. Small 15. https://doi.org/10.1002/smll.201900735
Tan Y, Guo Q, Xie Q, Wang K, Yuan B, Zhou Y, Liu J, Huang J, He X, Yang X, He C, Zhao X (2014) Single-walled carbon nanotubes (SWCNTs)-assisted cell-systematic evolution of ligands by exponential enrichment (cell-SELEX) for improving screening efficiency. Anal Chem 86:9466–9472
Nie W, Wang Q, Zou L, Zheng Y, Liu X, Yang X, Wang K (2018) Low-fouling surface plasmon resonance sensor for highly sensitive detection of microRNA in a complex matrix based on the DNA tetrahedron. Anal Chem 90:12584–12591
Ye DK, Zuo XL, Fan CH (2018) DNA nanotechnology-enabled interfacial engineering for biosensor development. Annu Rev Anal Chem 11:171–195
Huang YL, Mo S, Gao ZF, Chen JR, Lei JL, Luo HQ, Li NB (2017) Amperometric biosensor for microRNA based on the use of tetrahedral DNA nanostructure probes and guanine nanowire amplification. Microchim Acta 184:2597–2604
Hu W, He G, Zhang H, Wu X, Li J, Zhao Z, Qiao Y, Lu Z, Liu Y, Li CM (2014) Polydopamine-functionalization of graphene oxide to enable dual signal amplification for sensitive surface plasmon resonance imaging detection of biomarker. Anal Chem 86:4488–4493
Scarano S, Pascale E, Palladino P, Fratini E, Minunni M (2018) Determination of fermentable sugars in beer wort by gold nanoparticles@polydopamine: a layer-by-layer approach for localized surface plasmon resonance measurements at fixed wavelength. Talanta 183:24–32
Choi CK, Li J, Wei K, Xu YJ, Ho LW, Zhu M, To KK, Choi CH, Bian L (2015) A gold@polydopamine core-shell nanoprobe for long-term intracellular detection of microRNAs in differentiating stem cells. J Am Chem Soc 137:7337–7346
Zhang JJ, Mou L, Jiang XY (2018) Hydrogels incorporating Au@polydopamine nanoparticles: robust performance for optical sensing. Anal Chem 90:11423–11430
Ye Q, Zhou F, Liu WM (2011) Bioinspired catecholic chemistry for surface modification. Chem Soc Rev 40:4244–4258
Fang T, Lv HW, Lv GS, Li T, Wang CZ, Han Q, Yu LX, Su B, Guo LN, Huang SN, Cao D, Tang L, Tang SH, Wu MC, Yang W, Wang HY (2018) Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat Commun 9:191. https://doi.org/10.1038/s41467-017-02583-0
Funding
This work was carried out within the framework of the National Natural Science Foundation of China (21675047, 21375034, and 21735002) and the National Science Foundation for Distinguished Young Scholars of Hunan Province (2016JJ1008).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 992 kb)
Rights and permissions
About this article
Cite this article
Liao, G., Liu, X., Yang, X. et al. Surface plasmon resonance assay for exosomes based on aptamer recognition and polydopamine-functionalized gold nanoparticles for signal amplification. Microchim Acta 187, 251 (2020). https://doi.org/10.1007/s00604-020-4183-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-020-4183-1