Abstract
MicroRNAs are endogenous noncoding RNAs that play critical roles in biological processes and can be considered as molecular markers for early diagnosis and pathogenesis of diseases. The authors describe a highly sensitive electrochemical biosensor for microRNA that is based on the use of tetrahedral DNA nanostructure probes and guanine nanowire amplification. The DNA tetrahedral probe is self-assembled on a gold electrode and enhances reactivity, accessibility, and molecular recognition efficiency. Combined with the tetrahedral probe, the guanine nanowire amplifies the signal and improves the analytical performance of the biosensor. Operated best at a voltage of typically 150 mV (vs. Ag/AgCl), the sensor has a linear response to the logarithmic microRNA concentration in the 500 f. to 10 nM range, with a 176 f. detection limit. It is highly selective and can be applied to real samples. It is concluded that this strategy has a good potential with respect to the determination of microRNA in clinical diagnosis and in biological research.

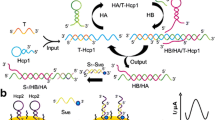
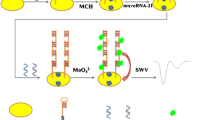
Schematic of a tetrahedral DNA nanostructure-based amperometric biosensor coupled to guanine nanowire amplification for analysis of microRNA-21. This strategy is highly selective and also performs well for the detection of microRNA levels of breast cancer patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) belong to small noncoding RNAs consisting of 19–24 nucleotides. They are involved in regulating gene expression through cell proliferation, differentiation, apoptosis and cancer development [1–3]. It has been reported that miRNAs levels are highly associated with cancers, viral infections and cardiovascular diseases [4, 5]. Therefore, miRNAs have been regarded as promising clinical biomarkers for cancer diagnosis, prognosis, and prediction [6, 7]. Simple, sensitive and selective methods for the detection of miRNAs become more attractive in disease diagnosis and biological research [8–11].
Many methods have been introduced for miRNAs analysis, such as northern blotting [12], in situ hybridization [13], microarray-based detection [14], surface Raman spectroscopy [15], surface plasmon resonance (SPR) [16], and reverse transcription polymerase chain reaction (TR-PCR) [17]. Northern blotting is considered as a good standard method for miRNA analysis, but it requires sophisticated equipment and has low detection sensitivity. Microarray-based detection has the advantages including high throughput and fast response. Nevertheless, the low sensitivity and poor specificity limit its applications. Real-time RT-PCR is used widely for highly sensitive and quantitative analysis of miRNA. However, it requires extraction and purification of miRNA from the real samples. The short length of miRNA also increases the sophistication to the experimental design. In addition, most of the techniques are time-consuming and require specialized reagents, sophisticated instruments and complicated decorating procedure. Therefore, a rapid, inexpensive and sensitive method is imperative for miRNA analysis [18, 19].
Electrochemical biosensors are attractive for bioanalysis owing to the inherent features such as low cost, simplicity, low detection limit, excellent selectivity and sensitivity [11, 20–22]. However, it is a challenge to control the orientation and density of probes on the electrode surface. Single-stranded DNA and hairpin DNA have been used widely as capture probes of the electrochemical biosensor [23, 24]. Generally, nonspecific interactions, the steric hindrance and the entanglement between the probes have a negative effect on the selectivity and reproducibility of the biosensor. To address these challenging problems, tetrahedral DNA nanostructure, which can use as a recognition probe, has attracted great attention [25–27]. Generally, there are three vertices of the tetrahedral structure which were modified on the gold electrode by Au-S bonds. And the fourth vertex appended with an extended DNA probe hybridizes with the target. Therefore, the rigid tetrahedral nanostructure can avoid the introduction of spacer molecules, and improves the reactivity and accessibility. Thus, the analytic performance of the electrochemical biosensor is enhanced significantly [28, 29].
There remains another challenge in the sensitivity for target detection and signal amplification of the electrochemical biosensor. Afterwards, guanine nanowire (G-wire) amplification has been reported with a potential use in the analysis by our group [30]. Guanine nanowire is considered as an efficient amplification method, which improves the sensing ability significantly in biosensor construction. Compared with the single G-quadruplex signal unit, the electrode signal was improved obviously via the signal amplification of guanine nanowire. However, research about the combination of DNA tetrahedral nanostructure and guanine nanowire amplification for miRNA detection has not been reported.
We herein fabricate an electrochemical miRNA biosensor by the combination of DNA tetrahedral nanostructure and guanine nanowire amplification. The detection limit for miRNA is 176 fM. The specificity of the biosensor allows single-based mismatch to be distinguished. Furthermore, the biosensor exhibits an excellent analytical performance and can be applied in breast cancer serum samples. Therefore, this strategy might offer a potential use in clinical research and disease diagnosis.
Experimental
Materials and chemicals
The oligonucleotides were purchased and HPLC-purified by Sangon Biotech Co., Ltd. (Shanghai, China, www.sangon.com). The corresponding sequences are illustrated in Table S1 (Electronic Supplementary Material). Hemin was purchased from Sangon Biotech Co., Ltd. (Shanghai, China, www.sangon.com). Tris (2-carboxyethyl) phosphine hydrochloride (TCEP) was purchased from Sigma Chemical Co. (St. Louis, MO, USA, www.sigmaaldrich.com). 3,3′,5,5′-Tetramethyl benzidine dihydrochloride (TMB⋅2HCl) was purchased from Xiya Chemical Industry Co., Ltd. (Shandong, China, www.xiyashiji.com). Hydrogen peroxide (H2O2, 30%) and dimethylsulfoxide were purchased from Chengdu Kelong Chemical Reagents Factory (Chengdu, China, www.cdkelong.com). Human serum samples from breast cancer patients were supplied by the local hospital (Wuhan, China). Other chemicals mentioned were analytical reagent grade. Ultrapure water was used to prepare all solutions (18.2 MΩ cm resistivity).
Electrochemical measurements
Electrochemical experiments were measured with CHI660D electrochemical workstation (Shanghai CH Instruments Co., China, www.chinstruments.com). The traditional three electrode system was employed. The 2 mm diameter gold electrode acted as the working electrode. A platinum wire and Ag/AgCl (sat.KCl) acted as the auxiliary electrode and the reference electrode, respectively. The fabrication steps were recorded by cyclic voltammetry (CV) and electrochemical impedance spectroscopy (EIS). The CV curves were performed between −0.3 and 0.7 V. The scan rate was 100 mV s−1. EIS curves were measured in 0.1 M phosphate buffer (pH 7.4) containing 5 mM [Fe(CN)6]4−/3− with an amplitude of 5 mV. The frequency range was from 0.1 Hz to 10 kHz. Amperometric detection was measured at 150 mV and a steady state was recorded within 100 s.
Electrode preparation
Initially, the gold electrode was pretreated with piranha solution (30% H2O2: 98% H2SO4 = 1: 3) for 5 min (Caution: Piranha solution reacts violently with organic solvents and must be handled with care). Then the electrode was polished with 0.5 and 0.05 μm alumina powder to get a mirror-like surface. After that, the electrode was cleaned by sonication with ultrapure water, ethanol and ultrapure water for 5 min, respectively. The cleaned gold electrode was electrochemically cleaned in 0.5 M H2SO4 solution from a potential of −0.2 to 1.6 V with a scan rate of 100 mV s−1 until the stable cyclic peak was observed. Finally, the prepared electrode was dried with nitrogen and was ready for surface modification.
Self-assembly of DNA tetrahedron-structured probes on gold electrode surfaces
Buffer for dissolution of four DNA strands (tetrahedral A, B, C, and D) was 10 mM Tris-HCl (pH 8.0) containing 50 mM MgCl2 and 10 mM TCEP. The final concentration of each strand was 4 μM. Then, each strand with the volume of 25 μL was mixed. Afterward, the mixture was heated to 95 °C for 2 min and then cooled to 4 °C to form the tetrahedral DNA structure [31]. After that, the final concentration of tetrahedral structure probe was 1 μM. Then, the cleaned electrode was incubated with 10 μL of 1 μM tetrahedral structure probe and the reaction lasted for 8 h.
Hybridization with target miRNA
The tetrahedral DNA-modified electrode was carefully rinsed. Then, it was incubated with miRNA solution (10 mM phosphate buffer, 250 mM NaCl, pH 7.4) with different concentrations for 1.5 h, followed by thoroughly rinsing with washing buffer (10 mM phosphate buffer, 2.7 mM KCl, 140 mM NaCl, pH 7.4).
Direct growth of guanine nanowire
The oligonucleotide, c-myc, was heated at 95 °C for 5 min and cooled to room temperature. The above electrode was immersed in 50 μL of 10 mM Tris-HCl buffer (pH 7.4) containing 0.5 M K+ for 1 h to form G-quadruplex structure. Then, the electrode was incubated in 1.5 μM c-myc solution (10 mM Tris-HCl buffer, 0.12 M Mg2+, 0.5 M KCl, pH 7.4) to form guanine nanowire structure. After the reaction, the electrode was incubated in 25 μL of 0.2 mM hemin solution (10 mM HEPES buffer, 1% dimethylsulfoxide, 50 mM KCl, pH 8.0) for 1 h, and then rinsed before the electrochemical measurements in a 2 mM TMB – 5 mM H2O2 system (pH 6.5).
Results and discussion
Sensing mechanism
The DNA tetrahedral structure is designed by four single strand oligonucleotides with a pendant hairpin. Tetrahedral A, B, C and D, four ssDNAs, are self-assembled to constitute the six edges of the DNA tetrahedral nanostructure [32, 33]. Tetrahedral B, C and D are modified with thiols, which can be fixed on the electrode via thiols [34]. As shown in Scheme 1, a hairpin structure of the tetrahedron is designed to recognize target miRNA. The presence of target opens the hairpin of tetrahedral A. Then, the released stem sequence from the hairpin, which consists of c-myc sequence at 3′ termini, can self-assemble with K+ to constitute the parallel G-quadruplex. In the presence of c-myc, the parallel G-quadruplex induces the construction of guanine nanowire within 10 min with the help of Mg2+ [30, 35]. After the addition of hemin, the G-quadruplex-hemin repeat units can catalyze the oxidation of TMB. And the increased electrochemical signals reflect target miRNA concentration.
Electrochemical characterization of electrode modification
CV was used to characterize the stepwise modification process of the Au electrode. As shown in Fig. 1a, the cyclic voltammogram showed the behavior of [Fe(CN)6]3−/4- over the gold electrode surface before and after the process of modification. The bare gold electrode exhibited a pair of redox peaks (curve a), indicating the redox behavior in [Fe(CN)6]3−/4-. After the modification of DNA tetrahedral nanostructure, the peak current decreased (curve b). This was because the successful immobilization of tetrahedral DNA nanostructure occurred on the electrode surface, which hindered interfacial electron transfer between solution and electrode surface. In the presence of miRNA, the peak current decreased slightly compared to that of the tetrahedral DNA nanostructure modified electrode (curve c). After the treatment with c-myc (curve d), the peak current decreased observably, demonstrating that the high-order guanine nanowire superstructure formed on the electrode surface.
To confirm the stepwise process of modification on the electrode surface, EIS was performed. The Randles circuit (inset, Fig. 1b) was used to fit the EIS data, including the solution resistance (R s), the charge transfer resistance (R ct), the double-layer capacitance (C dl), and the Warburg impedance (Z w). As depicted in Fig. 1b, the bare gold electrode showed a very small semicircle domain (curve a, R ct = 189.9 Ω), which can be attributed to the rapid electron transfer of the redox probe [Fe(CN)6]3−/4-. After the modification of the thiolated tetrahedral DNA structure onto the electrode surface, the electronic transfer resistance R ct increased obviously (curve b, R ct = 10,870 Ω). The reason was that the negatively charged tetrahedral DNA structure effectively repelled the negatively charged redox probe [Fe(CN)6]3−/4-, and the charge-transfer resistance was enhanced. After the incubation with miRNA, the R ct increased substantially (curve c, R ct = 14,770 Ω). And a further increased R ct was observed (curve d, R ct = 21,700 Ω) when the c-myc was added, indicating the formation of guanine nanowire on the electrode surface.
Feasibility for miRNA detection
To investigate the feasibility of this method for miRNA analysis, amperometric measurements were carried out on the tetrahedral DNA nanostructure modified electrode in the absence/presence of miRNA with/without guanine nanowire amplification. As depicted in Fig. 2, in the absence of miRNA, a tiny signal (recorded as I 0) was observed on the tetrahedral DNA nanostructure modified electrode (curve a). However, in the presence of 10 nM miRNA without the incubation of c-myc solution (curve b), the current increment was 12.2 nA. This was attributed to the formation of G-quadruplex at 3′ terminus of tetrahedral A, which can strongly bind hemin to form the catalytic DNAzyme. When the tetrahedral DNA nanostructure modified electrode incubated with 10 nM miRNA and c-myc solution (curve c), there was an evident increase of the response current (recorded as I) and the current increment was 24.9 nA. Such change was basically due to the construction of guanine nanowire, which significantly improved the electrochemical signal. CV was utilized to verify the feasibility of designed amplification process (Fig. S1). With the formation of G-wire, a pair of asymmetric redox peaks was the characteristic of the occurrence of electrocatalysis. These results indicate that this strategy can detect miRNA sensitively.
Sensitivity for miRNA analysis
To achieve the high hybridization efficiency and maximize the sensitivity, the following parameters were optimized: (a) miRNA hybridization time; (b) incubation time of hemin. Respective data and figures are given in Figs. S2 and S3. We found the following experiment conditions to give best results: (a) the hybridization time of 90 min was chosen as optimum hybridization time between miRNA and the tetrahedral DNA nanostructure modified electrode; (b) the incubation time of hemin was set as 60 min in the detection assay. Moreover, miRNA hybridization time of the DNA tetrahedron decorated electrode was much shorter than that of the hairpin probe modified electrode (Fig. S4).
The analytical performance of this method was investigated under the optimal conditions. Amperometry, a simple and classic electrochemical method, was used to characterize the enzyme-catalyzed electrochemical process. As depicted in Fig. 3a, it demonstrates that the amperometric current increases with increasing miRNA concentration. Fig. 3b reveals the linear relationship between the current variation (ΔI = I - I 0, where I 0 and I denote the amperometric current with and without miRNAs, respectively) and the logarithm of miRNA concentration ranging from 500 f. to 10 nM. The detection limit is 176 f. (S/N = 3). The fitting equation is ΔI = 66.4201 + 5.2803 log C (R = 0.9935), where ΔI is the amperometric current variation and C denotes the miRNA concentration. As shown in Table 1, compared with other reported electrochemical biosensors for miRNA analysis, this biosensor demonstrates a lower detection limit than other methods.
a Amperometric responses in the TMB – H2O2 system of the tetrahedral DNA decorated electrode for miRNA detection with the concentrations of 0, 500 fM, 1 pM, 10 pM, 100 pM, 1 nM, and 10 nM. b The linear relationship between the current variation and logarithm of miRNA concentration. The voltage was held at 150 mV (vs. Ag/AgCl) and the reduction current was recorded at 100 s. Error bars represent the standard deviation of three independent experiments
Selectivity, repeatability and stability of the electrochemical biosensor
To investigate the selectivity of the method, a comparison study was performed on single-base mismatch miRNAs and the complementary target miRNA under the same and optimum conditions. As shown in Fig. 4, the amperometric current variation (ΔI) of the complementary target miRNA was much higher than those of the single-base mismatched sequences at different concentrations. At the same concentration of 100 pM, ΔI of the complementary target was 12.0 nA, which is much higher than that of other mismatch sequences with ΔI of 2.6 nA (mismatch 1), 1.3 nA (mismatch 2) and 1.7 nA (mismatch 3). Even if the concentration of the single-base mismatch (10 nM) is 100 times higher than the complementary target (100 pM), the values of single-base mismatches were significantly lower than that of the complementary target miRNA. These results indicated that this sensor is selective for the target miRNA from its analogous sequences.
The repeatability of this method is also of great importance in practical application. Five different electrodes were fabricated under the same conditions and used for the detection target miRNA (10 nM). The above five biosensors exhibited similar amperometric currents, the relative standard deviation (RSD) was 2.82%. The results demonstrated that this electrochemical biosensor has satisfactory repeatability for the detection of target miRNA. Furthermore, the stability of the biosensor was investigated by three independent experiments. After stored the DNA tetrahedral decorated gold electrode in the refrigerator at 4 °C for 5 days, the response showed no apparent change and can retain about 94.9% of its initial values. The results indicated that the electrochemical biosensor has satisfactory stability.
Detection of miRNA in human serum samples
Serum samples can be conveniently utilized for the analysis of different disease biomarkers [46]. To investigate the practical application of this strategy in serum sample analysis, we performed the miRNA assay in serum samples from breast cancer patients. Initially, serum samples were detected directly. Then, two different concentrations (1 pM and 10 pM) of target miRNA were added to the samples, and then the content of target miRNA was tested by this electrochemical biosensor. The detected results and the recoveries are listed in Table 2. Accordingly, the electrochemical biosensor can be applied directly for miRNA detection in serum samples without further pretreatment. In addition, this strategy might be a potential tool for miRNA assay and possessed excellent practical application in biological fluids.
Conclusions
In conclusion, we have fabricated an electrochemical biosensor for miRNA analysis through guanine nanowire amplification on a DNA tetrahedron decorated electrode. Compared to traditional amplification methods, guanine nanowire can significantly amplify the electrochemical signal and reduce the reaction time from hours to minutes. This sensing system is reliable, simple, easy to operate, and exhibits good analytical characteristics like low LOD. The sensor also displays fine selectivity, stability, and practical utility in real serum samples from breast cancer patients, which demonstrates its great potential application in clinical diagnostics and bioanalysis. This biosensor cannot be regenerated, and we will do more research on widening the linear range of target miRNA and promoting the degree of automation in the future. We also expect that our strategy would be extended in combination with other detection methods like enzyme-assisted target recycling and nanomaterials in sensing platform.
References
Png KJ, Halberg N, Yoshida M, Tavazoie SF (2012) A microRNA regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 481:190–194
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM (2013) Bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113:25–36
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebet BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838
Ha TY (2011) MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw 11:135–154
Cortez MA, Calin GA (2009) MicroRNA identification in plasma and serum: a new tool to diagnose and monitor diseases. Exp Opin Biol Ther 9:703–711
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, Zhang Y, Wang J, Jiang X, Xiang Y, Xu C, Zheng P, Zhang J, Li R, Zhang H, Shang X, Gong T, Ning G, Wang J, Zen K, Zhang J, Zhang C (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006
Teo AKL, Lim CL, Gao Z (2014) The development of electrochemical assays for microRNAs. Electrochim Acta 126:19–30
Miao P, Tang Y, Yin J (2015) MicroRNA detection based on analyte triggered nanoparticle localization on a tetrahedral DNA modified electrode followed by hybridization chain reaction dual amplification. Chem Commun 51:15629–15632
Bi S, Chen M, Jia X, Dong Y (2015) A hot-spot-active magnetic graphene oxide substrate for microRNA detection based on cascaded chemiluminescence resonance energy transfer. Nano 7:3745–3753
Yang J, Tang M, Diao W, Cheng W, Zhang Y, Yan Y (2016) Electrochemical strategy for ultrasensitive detection of microRNA based on MNAzyme-mediated rolling circle amplification on a gold electrode. Microchim Acta 183:3061–3067
Zhou Y, Yin H, Li J, Li B, Li X, Ai S, Zhang X (2016) Electrochemical biosensor for microRNA detection based on poly (U) polymerase mediated isothermal signal amplification. Biosens Bioelectron 79:79–85
Kim SW, Li Z, Moore PS, Monaghan AP, Chang Y, Nichols M, John B (2010) A sensitive non-radioactive northern blot method to detect small RNAs. Nucleic Acids Res 38:e98
Yamamichi N, Shimomura R, Inada K, Sakurai K, Haraguchi T, Ozaki Y, Fujita S, Mizutani T, Furukawa C, Fujishiro M, Ichinose M, Shiogama K, Tsutsumi Y, Omata M, Iba H (2009) Locked nucleic acid in situ hybridization analysis of miR-21 expression during colorectal cancer development. Clin Cancer Res 15:4009–4016
Roy S, Soh JH, Ying JY (2016) A microarray platform for detecting disease-specific circulating miRNA in human serum. Biosens Bioelectron 75:238–246
Ye LP, Hu J, Liang L, Zhang C (2014) Surface-enhanced Raman spectroscopy for simultaneous sensitive detection of multiple microRNAs in lung cancer cells. Chem Commun 50:11883–11886
Zhang D, Yan Y, Cheng W, Zhang W, Li Y, Ju H, Ding S (2013) Streptavidin-enhanced surface plasmon resonance biosensor for highly sensitive and specific detection of microRNA. Microchim Acta 180:397–403
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ (2005) Real-time quantification of microRNAs by stem–loop RT–PCR. Nucleic Acids Res 33:e179
Lee H, Shapiro SJ, Chapin SC, Doyle PS (2016) Encoded hydrogel microparticles for sensitive and multiplex microRNA detection directly from raw cell lysates. Anal Chem 88:3075–3081
Chen A, Gui GF, Zhuo Y, Chai YQ, Xiang Y, Yuan R (2015) Signal-off electrochemiluminescence biosensor based on Phi29 DNA polymerase mediated strand displacement amplification for microRNA detection. Anal Chem 87:6328–6334
Zhou Y, Wang M, Yang Z, Lu L (2016) Electrochemical biosensor for microRNA detection based on hybridization protection against nuclease S1 digestion. J Solid State Electrochem 20:413–419
Miao P, Tang Y, Wang B, Jiang C, Gao L, Bo B, Wang J (2016) Nuclease assisted target recycling and spherical nucleic acids gold nanoparticles recruitment for ultrasensitive detection of microRNA. Electrochim Acta 190:396–401
Liu H, Bei X, Xia Q, Fu Y, Zhang S, Liu M, Fan K, Zhang M, Yang Y (2016) Enzyme-free electrochemical detection of microRNA-21 using immobilized hairpin probes and a target-triggered hybridization chain reaction amplification strategy. Microchim Acta 183:297–304
Wang M, Li B, Zhou Q, Yin H, Zhou Y, Ai S (2015) Label-free, ultrasensitive and electrochemical immunosensing platform for microRNA detection using anti-DNA: RNA hybrid antibody and enzymatic signal amplification. Electrochim Acta 165:130–135
Li X, Guo J, Zhai Q, Xia J, Yi G (2016) Ultrasensitive electrochemical biosensor for specific detection of DNA based on molecular beacon mediated circular strand displacement polymerization and hyperbranched rolling circle amplification. Anal Chim Acta 934:52–58
Ge Z, Lin M, Wang P, Pei H, Yan J, Shi J, Huang Q, He D, Fan C, Zuo X (2014) Hybridization chain reaction amplification of microRNA detection with a tetrahedral DNA nanostructure-based electrochemical biosensor. Anal Chem 86:2124–2130
Zeng D, Zhang H, Zhu D, Li J, San L, Wang Z, Wang C, Wang Y, Wang L, Zuo X, Mi X (2015) A novel ultrasensitive electrochemical DNA sensor based on double tetrahedral nanostructures. Biosens Bioelectron 71:434–438
Lu N, Pei H, Ge Z, Simmons C, Yan H, Fan C (2012) Charge transport within a three-dimensional DNA nanostructure framework. J Am Chem Soc 134:13148–13151
Miao P, Wang B, Chen X, Li X, Tang Y (2015) Tetrahedral DNA nanostructure-based microRNA biosensor coupled with catalytic recycling of the analyte. ACS Appl Mater Interfaces 7:6238–6243
Pei H, Wan Y, Li J, Hu HY, Su Y, Huang Q, Fan CH (2011) Regenerable electrochemical immunological sensing at DNA nanostructure-decorated gold surfaces. Chem Comum 47:6254–6256
Gao ZF, Huang YL, Ren W, Luo HQ, Li NB (2016) Guanine nanowire based amplification strategy: enzyme-free biosensing of nucleic acids and proteins. Biosens Bioelectron 78:351–357
Miao P, Wang B, Meng F, Yin J, Tang Y (2015) Ultrasensitive detection of microRNA through rolling circle amplification on a DNA tetrahedron decorated electrode. Bioconjug Chem 26:602–607
Pei H, Lu N, Wen YL, Song SP, Liu Y, Yan H, Fan CH (2010) A DNA nanostructure-based biomolecular probe carrier platform for electrochemical biosensing. Adv Mater 22:4754–4758
Liang L, Li J, Li Q, Huang Q, Shi JY, Yan H, Fan CH (2014) Single-particle tracking and modulation of cell entry pathways of a tetrahedral DNA nanostructure in live cells. Angew. Chem. Int Ed 53:7745–7750
Hsu MH, Chuang H, Cheng FY, Huang YP, Han CC, Chen JY, Huang SC, Chen JK, Wu DS, Chu HL, Chang CC (2014) Directly thiolated modification onto the surface of detonation nanodiamonds. ACS Appl Mater Interfaces 6:7198–7203
Shi Y, Luo HQ, Li NB (2013) A highly sensitive resonance rayleigh scattering method to discriminate a parallel-stranded G-Quadruplex from DNA with other topologies and structures. Chem Commun 49:6209–6211
Zhou Y, Wang M, Meng X, Yin H, Ai S (2012) Amplified electrochemical microRNA biosensor using a hemin-G-quadruplex complex as the sensing element. RSC Adv 2:7140–7145
Zhang Y, Yan Y, Chen W, Cheng W, Li S, Ding X, Li D, Wang H, Ju H, Ding S (2015) A simple electrochemical biosensor for highly sensitive and specific detection of microRNA based on mismatched catalytic hairpin assembly. Biosens Bioelectron 68:343–349
Zhou W, Liang W, Li X, Chai Y, Yuan R, Xiang Y (2015) MicroRNA-triggered, cascaded and catalytic self-assembly of functional “DNAzyme ferris wheel” nanostructures for highly sensitive colorimetric detection of cancer cells. Nano 7:9055–9061
Zheng J, Bai J, Zhou Q, Li J, Li Y, Yang J, Yang R (2015) DNA-templated in situ growth of AgNPs on SWNTs: a new approach for highly sensitive SERS assay of microRNA. Chem Commun 51:6552–6555
Xu F, Shi H, He X, Wang K, He D, Guo Q, Qing Z, Yan L, Ye X, Li D, Tang J (2014) Concatemeric dsDNA-templated copper nanoparticles strategy with improved sensitivity and stability based on rolling circle replication and its application in microRNA detection. Anal Chem 86:6976–6982
Cui L, Lin X, Lin N, Song Y, Zhu Z, Chen X, Yang CJ (2012) Graphene oxide-protected DNA probes for multiplex microRNA analysis in complex biological samples based on a cyclic enzymatic amplification method. 2012. Chem Commun 48:194–196
Wang W, Kong T, Zhang D, Zhang J, Cheng G (2015) Label-free MicroRNA detection based on fluorescence quenching of gold nanoparticles with a competitive hybridization. Anal Chem 87:10822–10829
Xi Q, Zhou DM, Kan YY, Ge J, Wu ZK, Yu RQ, Jiang JH (2014) Highly sensitive and selective strategy for microRNA detection based on WS2 nanosheet mediated fluorescence quenching and duplex-specific nuclease signal amplification. Anal Chem 86:1361–1365
Zhao H, Qu Y, Yuan F, Quan X (2016) A visible and label-free colorimetric sensor for miRNA-21 detection based on peroxidase-like activity of graphene/gold-nanoparticle hybrids. Anal Methods 8:2005–2012
Liu S, Su W, Li Z, Ding X (2015) Electrochemical detection of lung cancer specific microRNAs using 3D DNA origami nanostructures. Biosens Bioelectron 71:57–61
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, Magri E, Pedriali M, Fabbri M, Campiglio M, Ménard S, Palazzo JP, Rosenberg A, Musiani P, Volinia S, Nenci I, Calin GA, Querzoli P, Negrini M, Croce CM (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (No. 21675131, 21273174) and the Municipal Natural Science Foundation of Chongqing City (No. CSTC-2013jjB00002, CSTC-2015jcyjB50001) and the Innovation Foundation of Chongqing City for Postgraduate (No. CYB14052).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The author(s) declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOC 1.11 mb)
Rights and permissions
About this article
Cite this article
Huang, Y.L., Mo, S., Gao, Z.F. et al. Amperometric biosensor for microRNA based on the use of tetrahedral DNA nanostructure probes and guanine nanowire amplification. Microchim Acta 184, 2597–2604 (2017). https://doi.org/10.1007/s00604-017-2246-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-017-2246-8