Abstract
A fluorometric method is described for the determination of the tumor biomarker mucin 1 (MUC1). It is based on signal amplification of the hybridization chain reaction (HCR), and the interaction between a luminescent ruthenium(II) complex and CdZnTeS quantum dots (QDs). If MUC1 bind to the biotin-labeled aptamer, it will initiate HCR with hairpins H1 and H2 to form a long-range dsDNA. The long nucleic acid chains are then linked on the surface of streptavidin-modified magnetic microparticles (MMPs) through streptavidin-biotin interaction. The luminescent ruthenium(II) complex is then embedded in the long dsDNA linked to the MMPs. Hence, there is little Ru complex in the supernatant after magnetic separation, and the fluorescence of the CdZnTeS QDs (best measured at excitation/emission wavelengths of 350/530 nm) is only slightly quenched. In the absence of target, the fluorescence of the CdZnTeS QDs is strongly quenched. Fluorescence increases linearly in the 0.2–100 ng·mL−1 MUC1 concentration range, and the LOD is 0.13 ng·mL−1 (at S/N = 3). The method was applied to the determination of MUC1 in spiked human serum samples.

A fluorometric turn-on aptasensor for mucin 1 is described that is based on the interaction between a Ru(II) complex and quantum dots (QDs). The detection system includes biotin-labeled aptamer-H0, hairpins H1 and H2, streptavidin-modified magnetic microparticles (MMPs), Ru(bpy)2(dppx)2+ and CdZnTeS QDs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mucin 1 (MUC1) serves as an indicator in the early diagnosis of cancer [1,2,3,4]. However, it is still a challenge to detect MUC1 at the early stage, as the concentration of MUC1 is usually very low. Therefore, it is critical to develop a novel method to detect MUC1 sensitively and simply. Various methods for MUC1 detection have been developed, such as enzyme-linked immunosorbent assay (ELISA) [5], surface enhanced Raman scattering (SERS) [6], single molecular force spectroscopy (SMFS) [7], fluorescence (FL) [8, 9] and electrochemical techniques [10, 11]. Among the above methods, the fluorescent technique has attracted great attention due to the advantages of simpleness, efficiency and convenience.
Aptamers are oligonucleotides or peptide molecules and have special affinity with different targets. What’s more, aptamers have excellent advantages compared with antibodies, such as low cost, easy modification, small size, stability and lack of immunogenicity. Many successful aptasensors were reported for MUC1 detection [12,13,14,15,16]. These strategies were used for efficient and sensitive detection of MUC1 due to the specific binding of aptamers to MUC1. However, there were some details to be improved. For example, some of them need to immobilize DNA on the surface of the material, which had high spatial hindrance and low combined freedom, leading to low affinity efficiency between aptamers and proteins. In addition, most of them required labeling signal molecules that might result in high background signal. So it is very vital to establish a solution to solve these problems.
Quantum dots (QDs) are highlighted fluorescent probes, and they have attracted widespread attention in various research areas due to their unique advantages of wide excitation and narrow emission spectra, tunable size and high quantum yields [17,18,19,20,21]. Ru(bpy)2(dppx)2+(bpy = 2.2′- bipyridine, dppx = 7,8-dimethyldipyrido [3.2-a:2′,3′-c]-phenanthroline) is one of typical DNA molecular “light switches”, which can tightly bind toward DNA, especially dsDNA, and it can efficiently quench QDs via the photo induced electron transfer process. In the presence of dsDNA, Ru(bpy)2(dppx)2+ is bound by dsDNA and cannot quench QDs anymore [22, 23]. Magnetic microparticles (MMPs) have been extensively applied in the detection of DNA, small molecules, proteins, and cancer cells as a useful tool [16, 24,25,26], due to their high surface-to-volume ratio, easy surface modification and separation characteristics. Hybridization chain reaction (HCR) is an excellent signal amplification technique and can greatly amplify signal by DNA cascade hybridization without any nucleases involvement. H1 and H2 can self-assemble to form long-range dsDNA in the presence of an initiator [27]. In this experiment, Ru(bpy)2(dppx)2+ bound by dsDNA, independent of whether the HCR occurred or not. This reduces the quenching effect on QDs and leads to the high background signal. The streptavidin-modified MMPs can reduce the background signal significantly via the good separation performance.
Based on this principle, we developed a novel aptasensor for the detection of MUC1. In the presence of MUC1, the initiated sequence of aptamer-H0 triggered the HCR. The HCR product, long-range dsDNA, had stronger interaction with Ru complex than single strand DNA or H1 and H2. Combined Ru complex, QDs and MMPs, which greatly improved the detection sensitivity, selectivity and practicability.
Experimental section
Chemicals and instruments
Trishydroxymethyl aminomethane (Tris), sodium phosphate dibasic (Na2HPO4), sodium phosphate monobasic (NaH2PO4), potassium chloride (KCl), sodium chloride (NaCl), mucin 1 (MUC1), Tween-20, N-acetyl-L-cysteine (NAC), 4,5-Dimeththyl-o-phenylene-diamine and 2,3-dimercaptopropane-2,3-dimercaptopropane-1-sulfonate (DMPS) were purchased from Sigma-Aldrich (USA.; https://www.sigmaaldrich.com). All above of them were analytical-reagent grade or better. CdCl2·2.5H2O, ZnCl2, Na2TeO3 and hydrochloric acid was purchased from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China; http://www.sinoreagent.com). Meso-2,3-dimercaptosuccinic acid (DMSA) was purchased from Aladdin Chemistry Co., Ltd. (China; http://www.aladdin-e.com). The streptavidin-modified magnetic microparticles (MMPs) (1 μm, 10 mg·mL−1) were obtained from Invitrogen (Norway; https://www.thermofisher.com).
Different sequences of the oligonucleotides were synthesized by Sangon Biotechnology Co. Ltd. (Shanghai, China; https://www.sangon.com) and purified by high performance liquid chromatography (HPLC). The sequences of the oligonucleotides in this experiment were as follows:
-
Aptamer-H0: 5′-TGA GGT AGT AGG TTG TAT AGT TGC AGT TGA TCC TTT GGA TAC CCT GGA ACT ATA CAA CCT ACT ACC TCA AAA AA-biotin-3′,
-
Hairpin H1: 5′-AGT AGG TTG TAT AGT TCA AAG TAA CTA TAC AAC CTA CTA CCT CA-3′,
-
Hairpin H2: 5’-ACT TTG AAC TAT ACA ACC TAC TTG AGG TAG TAG GTT GTA TAG TT-3′.
Ultrapure water (18.2 MΩ·cm) was produced by a Milli-Q Academic purification set (Millipore, USA). Fluorescence emission spectrum was measured with a RF-6000 PC spectrophotometer (Shimadzu, Japan). The Fluorescence intensity was measured by exciting the sample at 350 nm. The slits for excitation and emission were set at 10 and 5 nm, respectively. UV–vis absorption spectrum data was performed on a UV-2550 spectrophotometer (Shimadzu, Japan). Transmission electron microscopy (TEM) and high-resolution (HR) TEM images were obtained on a JEM-2100 electron microscope (Japan).
Synthesis of the Ru(II) complex and CdZnTeS QDs
Ru(bpy)2(dppx)2+ complex and CdZnTeS QDs were synthesized according to our previous reported method [21, 28, 29], respectively. The experimental procedures were described in supporting information (SI).
CdZnTeS QDs were dissolved in Tris buffer 1 (Tris 20 mM, pH 7.4) for further use.
MUC1 detection
Firstly, DNA sequence H1, H2 and biotin-labeled aptamer-H0 were heated at 90 °C for 10 min, respectively. Then the solution was cooled to room temperature for 1 h before use. Different concentration of MUC1 was mixed with 10 nM aptamer-H0 in PBS 1 (NaH2PO4 100 mM, Na2HPO4 100 mM, KCl 100 mM, pH 8.0) for 1 h at 37 °C, respectively. After that, 1 μM H1 and 1 μM H2 in 200 μL Tris buffer 2 (Tris 20 mM, NaCl 200 mM, pH 7.5) were added into above solution for 2 h at 37 °C to proceed the HCR.
Secondly, a certain amount of streptavidin-modified MMPs were washed three times with 100 μL PBS 2 (NaH2PO4 10 mM, Na2HPO4 10 mM, NaCl 15 mM, pH 7.4), then they were mixed with the HCR products, respectively. After incubating at 37 °C with gentle shaking for 30 min, the conjugated MMPs were washed three times with 400 μL washing buffer (Tris 10 mM, NaCl 100 mM, Tween-20 0.01% (w/v), pH 7.5).
After the redundant DNA that absorbed on the surface of the MMPs were removed, 20 μL of 0.1 mM Ru(bpy)2(dppx)2+ was mixed with the MMPs in 360 μL Tris buffer 1 for 30 min with gentle shaking. Finally, 7.5 nM CdZnTeS QDs were added into the supernatant which had been separated with a magnet. The fluorescence intensity of the supernatant was implemented by RF-6000 PC spectrophotometer (measured at excitation/emission wavelengths of 340/530 nm).
Results and discussion
Working principle
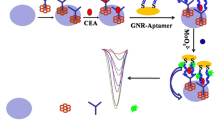
The principle of MUC1 detection is illustrated in Scheme 1. The detection system includes biotin-labeled aptamer-H0, hairpins H1 and H2, streptavidin-modified MMPs, Ru(bpy)2(dppx)2+ and CdZnTeS QDs. The sequence of DNA in this experiment referred the method of Ma C et al. [8]. The structure of H0 includes stem and loop, and the sequence of aptamer is in the loop of H0. In the absence of MUC1, the biotin-labeled aptamer-H0 kept stable hairpin structure from proceeding HCR with H1 and H2. When streptavidin-modified MMPs were mixed with above solution, only biotin-labeled aptamer-H0 were linked on the surface of streptavidin-modified MMPs. Subsequently, Ru(bpy)2(dppx)2+ was mixed with the above MMPs under stirring. There were a large amount of Ru(bpy)2(dppx)2+ in the supernatant after magnetic separation due to the failure of embedding in dsDNA. After CdZnTeS QDs were mixed with the supernatant, the fluorescence intensity of CdZnTeS QDs was at a low level, as Ru complex highly quenched CdZnTeS QDs. When MUC1 is present, it binds with aptamer specifically, whose structure will be changed [30]. The initiated sequence of aptamer-H0 will proceed HCR with H1 and H2 to form long-range dsDNA structures. After streptavidin-modified MMPs were mixed with the above solution, the long nucleic acid chains were linked on the surface of streptavidin-modified MMPs through streptavidin-biotin interaction. Subsequently, Ru(bpy)2(dppx)2+ was mixed with the above MMPs under stirring, then MMPs were separated with a magnet. There was little Ru(bpy)2(dppx)2+ in the supernatant, as Ru(bpy)2(dppx)2+ was embedded in dsDNA. So the fluorescence intensity of the supernatant was very high after CdZnTeS QDs were mixed with the supernatant.
Schematic representation of the MUC1 detection based on fluorescent aptasensor turn-on strategy with interaction between Ru complex and QDs. The detection system includes biotin-labeled aptamer-H0, hairpins H1 and H2, streptavidin-modified magnetic microparticles (MMPs), Ru(bpy)2(dppx)2+ and CdZnTeS QDs
Characterization of the CdZnTeS quantum dots (QDs)
The synthesized QDs were characterized by TEM, UV-vis absorption and fluorescence spectrum. As shown in Fig. 1, TEM image shows that the QDs have good mono-dispersion with an average size of 3.40 ± 0.40 nm. The UV-vis absorption and fluorescence spectra of CdZnTeS QDs are shown in Fig. 2, respectively. The absorption peak is at 500 nm and the emission peak is at 530 nm. According to the absorption peak intensity, the concentration of QDs is obtained [31].
Feasibility of the design strategy
The feasibility of this strategy is shown in Fig. 3. In the absence of MUC1, only biotin-labeled aptamer-H0 was linked on the streptavidin-modified MMPs surface, leading a lot of Ru(bpy)2(dppx)2+ dispersed in the supernatant. When CdZnTeS QDs were added into the supernatant, their fluorescence of CdZnTeS QDs was quenched by Ru(bpy)2(dppx)2+ that resulted in low fluorescence signal (Fig. 3b). When MUC1 was present without H1 and H2, only small amount of Ru complex bound to the MMPs, leading to a slight peak (Fig. 3a). Only had H1 or H2 in the system, the fluorescence intensity of QDs was still at low level as HCR product was formed unsuccessfully (Fig. 3c and d). When MUC1 and HCR products presented simultaneously, the supernatant with QDs showed high fluorescence signal, as a large amount of Ru(bpy)2(dppx)2+ were embedded in dsDNA (Fig. 3e). The fluorescence of QDs at different conditions is shown in the inset of Fig. 3.
Fluorescence spectra at different conditions: (a) taget only; (b) H1 + H2; (c) target + H2; (d) target + H1; (e) target and H1 + H2; inset: fluorescence intensity of CdZnTeS QDs at different condition, respectively. The concentration of MUC1, H1, H2, QDs and Ru(II) complex was 100 ng·mL−1, 1 μM, 1 μM, 7.5 nM and 5 μM, respectively
Optimization of reaction conditions
The effect of Ru(bpy)2(dppx)2+ concentration on fluorescence of QDs is explored in Fig. S1. Different concentration of Ru(bpy)2(dppx)2+ (0, 1.25, 2.50, 3.75, 5, 6.25, 7.50 μM) was mixed with 7.5 nM QDs in Tris buffer 1, respectively. It reveals that the fluorescence intensity of QDs was decreased with the increasing concentration of Ru(bpy)2(dppx)2+, and the QDs showed fairly low fluorescence intensity with 5 μM Ru(bpy)2(dppx)2+. We have obtained the quenching type according to the Stern-Volmer plot (Fig. S2). The curve equation is “y = 0.020 x2 – 0.083 x + 1.3”, which is consistent with the modified Stern-Volmer equation “Io/If = (1 + Kd[Q]) (1 + Ks[Q])=1 + (Kd + Ks)[Q] + KdKs[Q]2”. So it is indicating both static and dynamic quenching. The amount of MMPs used in the detection system was investigated. Fig. S3 shows that the fluorescence intensity of supernatant had positive correlation with the amount of MMPs from 2 to 10 μL, and the fluorescence intensity was almost stable when the adding volume of MMPs was 8 μL. The concentration of aptamer was optimized, as shown in Fig. S4. The more aptamer was, the higher efficiency of binding with MUC1 was, and the fluorescence intensity of QDs reached to a plateau when the concentration of aptamer was 10 nM. The concentrations of H1 and H2 were also optimized (Fig. S5). The fluorescent signal increased with the increasing concentration from 0 ~ 1 μM. However, when the concentration continued to increase, the binding efficiency between MMP and aptamer-H0 decreased, as a large amount of DNA in the solution increased the difficulty of recognition between streptavidin-modified MMPs and biotin-labeled aptamer-H0. So the optimal concentration of H1 and H2 were both 1 μM.
MUC1 detection
In order to investigate the sensitivity of the detection strategy, different concentration of MUC1 mixed with aptamer-H0 in optimal conditions was measured, respectively, as shown in Fig. 4. The fluorescence intensity of QDs was enhanced with the increasing concentration of MUC1, and the relative fluorescence intensity showed linear relationship with the concentration of MUC1 from 0.2–100 ng·mL−1 (LOD = 0.13 ng·mL−1, S/N = 3). Compared with the previous method (Table S1), the sensitive of this fluorescent aptasensor is better than the most of them. The result shows that this method can be used to detect MUC1 sensitively.
Specificity of MUC 1 detection
For proving the specificity, MUC1 and other proteins, such as AFP, CEA and ALP were tested. As shown in Fig. 5, the fluorescence intensity of QDs was effectively quenched by introducing other proteins, in spite of their concentrations were ten times as that of MUC1. So this detection method can be used for specific detection of MUC1. The reason is attributed to the unique recognition capability of the aptamer of MUC1.
MUC 1 detection in human serum
For evaluating the function of this method in complex biological environment, the fluorescent turn-on aptasensor was performed to detect MUC1 in the 20-fold-diluted human serum samples. The recovery was carried on four different levels of MUC1 samples (12.5, 25, 50 and 100 ng·mL−1) with the proposed aptasensor. The recovery is in the range of 92.9–108%, and the relative standard deviation (RSD) varies from 3.00% to 6.25%, which is shown in Table 1. Therefore, the aptasensor is applicable for the determination of MUC1 in the real sample with satisfied results.
Conclusion
In summary, we establish a new method for MUC1 detection, based on signal amplification via a hybridization chain reaction and the interaction between a luminescent ruthenium(II) complex and CdZnTeS quantum dots. This MUC1 detection method doesn’t require any nucleases or fluorescent labels and can be used to selectively detect MUC1 in real sample. Furthermore, this sensitive fluorescent MUC1 detection biosensor has great potential in the area of cancer diagnosis.
References
Donald WK (2009) Mucins in cancer: function, prognosis and therapy. Nat Rev Cancer 9:874–885
Singh R, Bandyopadhyay D (2007) MUC1: a target molecule for cancer therapy. Cancer Biol Ther 6:481–486
Chinen AB, Guan CM, Ferrer JR, Barnaby SN, Merkel TJ, Mirkin CA (2015) Nanoparticle probes for the detection of cancer biomarkers, cells, and tissues by fluorescence. Chem Rev 115:10530–10574
Chikkaveeraiah BV, Bhirde A, Morgan NY, Eden HS, Chen XY (2012) Electrochemical immunosensors for detection of cancer protein biomarkers. ACS Nano 6:6546–6561
Falahat R, Wiranowska M, Gallant ND, Toomey R, Hill R, Alcantar N (2015) A cell ELISA for the quantification of MUC1 mucin (CD227) expressed by cancer cells of epithelial and neuroectodermal origin. Cell Immunol 298:96–103
Feng JJ, Wu XL, Ma W, Kuang H, Xu LG, Xu CL (2015) A SERS active bimetallic core–satellite nanostructure for the ultrasensitive detection of Mucin-1. Chem Commun 51:14761–14763
Wang N, Zhang MM, Chen XJ, Ma XX, Li C, Zhang Z, Tang JL (2017) Mapping the interaction sites of mucin 1 and DNA aptamer by atomic force microscopy. Analyst 142:3800–3804
Ma C, Liu HY, Tian T, Song XR, Yu JH, Yan M (2016) A simple and rapid detection assay for peptides based on the specific recognition of aptamer and signal amplification of hybridization chain reaction. Biosens Bioelectron 83:15–18
He Y, Lin Y, Hong HW, Pang DW (2012) A graphene oxide-based fluorescent aptasensor for the turn-on detection of epithelial tumor marker mucin 1. Nanoscale 4:2054–2059
Hu R, Wen W, Wang QL, Xiong HY, Zhang XH, Gu HS, Wang SF (2014) Novel electrochemical aptamer biosensor based on an enzyme–gold nanoparticle dual label for the ultrasensitive detection of epithelial tumour marker MUC1. Biosens Bioelectron 53:384–389
Liu CY, Liu X, Qin Y, Deng CY, Xiang J (2016) A simple regenerable electrochemical aptasensor for the parallel and continuous detection of biomarkers. RSC Adv 6:58569–58476
Deng CY, Pi XM, Qian P, Chen XQ, Wu WM, Xiang J (2017) High-performance ratiometric electrochemical method based on the combination of signal probe and inner reference probe in one hairpin-structured DNA. Anal Chem 89:966–973
Jiang XY, Wang HJ, Wang HJ, Zhuo Y, Yuan R, Chai YQ (2017) Electrochemiluminescence biosensor based on 3-D DNA nanomachine signal probe powered by protein-aptamer binding com- plex for ultrasensitive mucin 1 detection. Anal Chem 89:4280–4286
Guo QJ, Li XZ, Shen CC, Zhang SB, Qi HZ, Li T, Yang MH (2015) Electrochemical immunoassay for the protein biomarker mucin 1 and for MCF-7 cancer cells based on signal enhancement by silver nanoclusters. Microchim Acta 182:1483–1489
Yazdanparast S, Benvidi A, Banaei M, Nikukar H, Tezerjani MD, Azimzadeh M (2018) Dual-aptamer based electrochemical sandwich biosensor for MCF-7 human breast cancer cells using silver nanoparticle labels and a poly(glutamic acid)/MWNT nanocomposite. Microchim Acta 185:405
Ma N, Jiang WT, Li T, Zhang ZQ, Qi HZ, Yang MH (2015) Fluorescence aggregation assay for the protein biomarker mucin 1 using carbon dot-labeled antibodies and aptamers. Microchim Acta 182:443–447
Jr MB, Moronne M, Gin P, Weiss S, Alivisatos AP (1998) Semiconductor nanocrystals as fluorescent biological labels. Science 281:2013–2016
Peng ZA, Peng XG (2001) Formation of high-quality CdTe, CdSe, and CdS nanocrystals using CdO as precursor. J Am Chem Soc 123:183–184
Liu ZY, Tang A, Wang M, Yang CH, Teng F (2015) Heating-up synthesis of cadimum-free and color-tunable quaternary and five-component Cu–In–Zn–S-based semiconductor nanocrystals. J Mater Chem C 3:10114–10120
Mao GB, Cai Q, Wang FB, Luo CL, Ji XH, He ZK (2017) One-step synthesis of Rox-DNA functionalized CdZnTeS quantum dots for the visual detection of hydrogen peroxide and blood Gl- ucose. Anal Chem 89:11628–11635
Mao GB, Liu C, Du MY, Zhang YW, Ji XH, He ZK (2018) One-pot synthesis of the stable CdZnTeS quantum dots for the rapid and sensitive detection of copper-activated enzyme. Talanta 185:123–131
Zhao D, Chan WH, He ZK, Qiu T (2009) Quantum dot-ruthenium complex dyads: recognition of double-Strand DNA through dual-color fluorescence detection. Anal Chem 81:3537–3543
Xiang X, Chen L, Zhuang QG, Ji XH, He ZK (2012) Real-time luminescence-based colorimetric determination of double-strand DNA in droplet on demand. Biosens Bioelectron 32:43–49
Liu YF, Luo M, Yan J, Xiang X, Ji XH, Zhou GH, He ZK (2013) An ultrasensitive biosensor for DNA detection based on hybridization chain reaction coupled with the efficient quenching of a ruthenium complex to CdTe quantum dots. Chem Commun 49:7424–7426
Zhang Z, Xiang X, Huang FH, Zheng MM, Xia XY, Han L (2018) Mercury ion-mediated “molecular beacon” integrating with hybridization chain reaction: application to fluorescence turn-on detection of glutathione by using quantum dots and Ru complex. Sensors Actuators B Chem 273:159–166
Liu SW, Xu NH, Tan CY, Fang W, Tan Y (2018) A sensitive colorimetric aptasensor based on trivalent peroxidasemimic DNAzyme and magnetic nanoparticles. Anal Chim Acta 1018:86–93
Huang J, Wu YR, Chen Y, Zhu Z, Yang XH, Yang CYJ (2011) Pyrene-excimer probes based on the hybridization chain reaction for the detection of nucleic acids in complex biological fluids. Angew Chem Int Ed 50:401–404
Ling LS, He ZK, Song GW, Yuan D, Zeng YE (2000) A novel method for determination of DNA by use of molecular ‘light switch’ complex of Ru(bipy)2(dppx)2+. Anal Chim Acta 403:209–217
Zhao D, Fang Y, Wang HY, He ZK (2011) Synthesis and characterization of high-quality water-soluble CdTe: Zn2+ quantum dots capped by N-acetyl-L-cysteine via hydrothermal method. J Mater Chem 21:13365–13370
Ferreira CSM, Matthews CS, Missailidis S (2006) DNA aptamers that bind to MUC1 tumor marker: design and characterization of MUC1-binding single-stranded DNA aptamers. Tumour Biol 27:289–301
Yu WW, Qu LH, Guo WZ, Peng XG (2003) Experimental determination of the extinction coefficient of CdTe, CdSe, and CdS nanocrystals. Chem Mater 15:2854–2860
Acknowledgements
This work was supported by the National Natural Science Foundation of China (21675119) and National Major Science and Technology Projects (2018ZX10301405).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The author(s) declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 122 kb)
Rights and permissions
About this article
Cite this article
Li, Z., Mao, G., Du, M. et al. A fluorometric turn-on aptasensor for mucin 1 based on signal amplification via a hybridization chain reaction and the interaction between a luminescent ruthenium(II) complex and CdZnTeS quantum dots. Microchim Acta 186, 233 (2019). https://doi.org/10.1007/s00604-019-3347-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-3347-3