Abstract
Nanopore-based sequencing platforms are transforming the field of genomic science. This review (containing 116 references) highlights some recent progress on nanopore-based nucleic acid analysis and sequencing. These studies are classified into three categories, biological, solid-state, and hybrid nanopores, according to their nanoporous materials. We begin with a brief description of the translocation-based detection mechanism of nanopores. Next, specific examples are given in nanopore-based nucleic acid analysis and sequencing, with an emphasis on identifying strategies that can improve the resolution of nanopores. This review concludes with a discussion of future research directions that will advance the practical applications of nanopore technology.

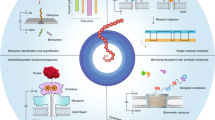
A review with 116 references that highlights the progress made in nanopore-based nucleic acid analysis and sequencing, and the mechanisms operative in analytical methods based on the use of nanopores
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gene sequencing is of central importance to both fundamental and applied aspects of genomic science, such as the emerging area of personalized genomic medicine. The first generation of gene sequencing is based on Sanger ladders consisting of short DNA fragments of unknown target sequences [1, 2]. These short DNA fragments are generated by selective incorporation of chain-terminating dideoxynucleotides during an in vitro DNA replication process. The dideoxynucleotides terminated at the 3′-end of DNA fragments are labeled and can be read by fluorescent or radioactive methods. The second generation of gene sequencing relies on the stochastic fragmentation of the target DNA sequences and subsequent amplification. Each fragment of the target sequence is amplified to millions of copies. The sequence information of these small copies is obtained separately and then assembled together to obtain the whole sequence of the target. For the first two generations of gene sequencing, both DNA amplification and labeling are necessary. Therefore, these sequencing techniques are costly and time-consuming. To overcome these disadvantages, many groups have been developing the third generation of gene sequencing techniques that can read single DNA chains in a label-free manner. Under the assumption of low error rates, only a small number of the target DNA molecules (or even a single DNA molecule) are needed to generate accurate sequencing results. In this review, we will focus on the third generation of sequencing techniques, viz. those based on nanopores.
In nanopore-based DNA sequencing, a nanometer-sized pore penetrating a lipid bilayer membrane or a solid-state membrane is used to link two fluid chambers (Fig. 1). The chambers are filled with saline solutions. When a voltage is applied between the two chambers, ions will be driven through the nanopore by the electric field, which thus generates a stable open-pore current. When a DNA strand translocates through a nanopore, it partially blocks the nanopore and decreases the ionic current. Therefore, DNA translocation events can be detected by the transient blockage of the ionic current. If each nucleotide on the DNA backbone can generate a separable current blockage, then the sequence of the DNA strand can be obtained through consecutive steps of the ionic current. This is the underlying principle of nanopore-based sequencing techniques.
There are generally two types of nanopores, namely biological and solid-state nanopores. A biological nanopore is usually generated by inserting a channel protein, such as α-hemolysin (α-HL) and MspA into a lipid bilayer. Engineered mutants of channel proteins have been developed to dramatically improve the resolution and sensitivity of biological nanopores [3, 4]. For a solid-state nanopore, a focused ion beam or electron beam is used to fabricate a nanometer-sized pore through an inorganic dielectric membrane [5, 6].
Although nanopore technologies offer promising opportunities for cheap and high-throughput gene sequencing, there are currently three challenges to be addressed before their practical applications in sequencing. The first challenge is the ultrafast translocation speed of DNA strands through nanopores [7]. Typical translocation speeds of DNA are from several to tens of microseconds per base through a wild-type biological nanopore, and it is even faster for solid-state nanopores. On the other hand, the bandwidth of a state-of-art current amplifier is at kHz level, which is thus too slow to record single nucleotide signals. Several methods have been developed to control the DNA translocation speeds, such as by the reduction of the pore sizes [8], the introduction of ion concentration gradients [9, 10], the introduction of pressure gradients [11, 12], and the use of optical and magnetic tweezers [13–15]. These methods will be individually discussed later in this review. The second challenge is to differentiate the current signals from the four nucleotides (A, T, G, and C). The difference of the volumetric blockade among the four nucleotides is usually too small to detect, especially in solid-state nanopores with large pore diameters. Furthermore, the interactions between DNA molecules and nanopores are weak. Genetic or chemical modification of biological nanopores can effectively increase the interactions between DNA molecules and nanopores, which has resulted in improved resolution of single nucleotides [3, 4]. DNA or other specially designed probing polymers can also be linked to solid-state nanopores for the differentiation of nucleotides [16–18]. The third challenge is related to the thickness of nanopores. The nanopores are usually several to tens of nanometers in depth and can thus accommodate more than one nucleotide. All the nucleotides in and adjacent to a nanopore determine the current blockade together, which greatly complicates the sequencing process. For example, the β-barrel of α-hemolysin (α-HL) [19] nanopore has a depth close to the length of 15 nucleotides. The depth of channel protein MspA [20, 21] is much shorter, in the range of 0.5 ~ 0.6 nm. Thus it can accommodate only one nucleotide at a time. However, the current blockade is still affected by 4 to 5 nucleotides adjacent to the MspA nanopore. We will discuss several promising methods that can be applied to address this problem in biological nanopores in Section 2. In addition, it is even more difficult to accurately control the depth of solid-state nanopores. Recently, ultrathin inorganic films have been applied as solid-state membranes, including monatomic thick graphene films, although single nucleotide resolution has yet to be achieved. Recent progress related with solid-state nanopores will be discussed in Section 3. In Section 4, we will discuss recent examples in developing hybrid nanopores that combine merits of biological and solid-state nanopores.
Biological nanopores
The history of nanopore technology dated back to the year 1958 when the Coulter counter was firstly invented for the enumeration of blood cells [22]. However, this method didn’t attract much attention until 1996 when Kasianowicz et al. [23] used α-HL to detect the translocation of DNA molecules. Modern nanopore technologies have been enlightened by the substance transportation across the cell membranes through channel proteins. Therefore, the nanopores used at early times were mostly channel proteins with an intrinsic pore structure, such as α-HL [19], MspA [20], and the channel protein of bacteriophage phi29 [24] (Fig. 2). α-HL, the most commonly used biological nanopore, has a 10 nm-high mushroom-shaped pore with a ~10 nm-wide extramembranal cap and a ~2.6 nm-wide transmembrane β-barrel stem. The narrowest part of the pore is ~1.4 nm and matches with the diameter of DNA molecules, which thus determines both the current blockade amplitude and the duration of the nucleotide translocation events.
Sequencing with biological nanopores
Figure 3a illustrates the measurement setup of a biological nanopore and a typical ionic current signal when a DNA translocates through the nanopore [26]. The earliest demonstration of nanopore-based DNA analysis showed distinct current spikes related to single ssDNA translocation events [23] (Fig. 3b). Later, different RNA homopolymers [27] and DNA homopolymers [20] were distinguished by the amplitude of their current blockades during translocation (Fig. 3c). However, the significance of these two works should not be overestimated because a nucleic acid homopolymer can form distinctive secondary structures. Thus in these studies, the current blockades were mainly determined by the secondary structures of RNA or DNA homopolymers, instead of their nucleotide sequences.
DNA analysis with biological nanopores. a General schematic of a biological nanopore sensor (left) and a translocation current spike (right) [26]; b Real-time current spikes measured during the translocation of DNA homopolymers [23]; c Differentiation of DNA homopolymers with biological nanopores. Green, red, blue and black spikes denote poly (dT), poly (dC), poly (dG), poly (dA), respectively [20]
The interaction between ssDNA or ssRNA molecules with natural biological nanopores is not strong enough to detect the small difference among the four nucleotides. In addition, the translocation speed of ssDNA or ssRNA molecules through natural biological nanopores is beyond the temporal resolution of current state-of-the-art electronics. One way to increase the resolution of biological nanopores is by genetically engineering the channel proteins or modifying them with functional groups that can interact with translocating nucleotides. Stoddart et al. [3] developed an E111N/K147N mutant of the α-HL protein (Fig. 4a). There are three recognition positions in the β-barrel of the mutant which can improve the differentiation of transpassing nucleotides. This work represents a significant advance in nanopore technologies towards the aim of sequencing DNA strands. Rincon-Restrepo et al. [4] proposed the addition of positive charges within the α-HL by site-directed mutagenesis. The increased positive charges can slow down the translocation speed by interacting electrostatically with the negatively-charged nucleotides. In addition, chemical grafting of molecules onto biological nanopores has been shown to be an effective method to increase the specificity of nanopores. PEG linked α-HL has been experimentally demonstrated and characterized, which raises the possibility of using engineered nanopores in biosensing [28]. DNA oligomers have also been tethered to α-HL nanopores as identification probes [29, 30]. Complementary DNA strands can be temporally captured by hybridization with the tethered ssDNA when translocating through the nanopores, and the corresponding ionic current signals have been successfully detected.
Engineered nanopores that can differentiate the difference between the four nucleotides are sometimes too small to accommodate certain ssDNA sequences. To overcome this problem, enzymatic cleavage of targeted DNA strands was carried out within close proximity to the engineered nanopores, and the stripped nucleotides can then be sequentially detected by passing through the nanopores. Astier et al. [31] took the lead in realizing this idea. The signals of the four nucleotides can be clearly distinguished by an engineered α-HL nanopore. But the detection of the nucleotides in this work relies on a noncovalent cyclodextrin adapter which moves in and out of the nanopore during the process. When the adapter moves out of the nanopore, the signals from the passing nucleotides would be missed. Clarke et al. [32] overcame this problem by chemically linking the cyclodextrin adapter to the nanopore (Fig. 4b). However, sequential translocation of the stripped nucleotides was not realized in their work because the hydrolysis of DNA took place in the bulk solution, not near the nanopore. Therefore, accurate sequencing cannot be achieved. Meanwhile, the analysis of RNA sequences by a similar approach has also been reported recently [33].
MspA is another important channel protein with shorter depth than α-HL, and the structure of MspA is shown in Fig. 2b. As mentioned above, MspA provides a much better spatial resolution than α-HL due to its shorter nanopore channel. The differentiation of homopolymers by MspA was demonstrated in 2010 [20]. Similar to α-HL, MspA has been modified with positive charges on its side wall [34]. The channel protein of bacteriophage phi 29 has also been widely studied for nanopore analysis [24]. Their large pore diameter allows the translocation of dsDNA or DNA protein complex. DNA polymerase itself can also be used as a nanopore [35]. When a template DNA translocates through the DNA polymerase, the conductance of the polymerase varies as different nucleotides pass by the binding site of the polymerase. And a protein FET linked to a DNA polymerase could measure the conductance signals related with the polymerase. However, the stability of this system needs to be further improved. Furthermore, a ring-like protein SP1 with good stability and geometric symmetry was also applied in nanopore analysis in 2013 [36].
Nucleic acid analysis by biological nanopores
In addition to de novo sequencing, biological nanopores have also been intensively investigated for nucleic acid analysis. Many efforts have been made to distinguish wild and mutated DNA. Ashkenasy et al. [37] and Purnell et al. [38] used α-HL nanopores to investigate the variation of current blockade when a nucleotide mutation was introduced to a homopolymer DNA. The current signal characteristics were found to be dependent on the position of the mutation on the DNA strand. But in both works, the target DNA strands are homopolymers, which obviously limits the application of their techniques. Howorka et al. chemically tethered a probe DNA to a α-HL nanopore. When the target DNA translocated through the nanopore, the hybridization between the probe and the target DNA produced a prolonged ionic current signal. On the other hand, a mutation site on the target DNA would lead to a significantly faster translocation speeds [29]. The detection sensitivity of this method was further enhanced by concatenated hybridization of the target DNA and the probe DNA [39]. Very recently, the use of MspA to detect the methylation and hydromethylation of cytosine in nucleic acid was reviewed [40], highlighting the work by Laszlo et al. [41] and by Schreiber et al. [42].
The unzipping kinetics of DNA hairpins can also be investigated using biological nanopores. Since the nanopores of α-HL and MspA are too narrow for the translocation of dsDNA, DNA hairpins have to be unzipped by applying large electrophoretic forces before translocation. The unzipping kinetics of DNA hairpins has been measured with α-HL based nanopores [43]. In addition, the kinetics of DNA-nanopore interactions [44] and the helix-coil transition of DNA strands [45] have also been demonstrated using biological nanopores. Such analysis requires the target DNA to be stabilized inside the nanopore, which can be achieved by linking a biotinylated DNA to a streptavidin molecule [45] or forming a duplex at the end of the target DNA [44]. Both the streptavidin and the duplex at the end of the target DNA are too large to pass the β-barrel, thus the target DNA strand could be stabilized inside the nanopores.
By using biological nanopores, DNA-protein interactions have also been characterized at single-molecule level. For example, nanopore-based force spectroscopy has been successfully applied for real-time characterization of binding and unbinding between DNA and exonuclease [46]. The α-HL nanopore can also be used to study the stability of DNA-polymerase complex by measuring the dwell time of current spikes [47, 48]. An ssDNA was chemically linked to a DNA polymerase, and the complex was then electrophoretically driven to the orifice of α-HL nanopore. A blocker was linked to the binding site of ssDNA and polymerase so that replication process is inhibited in the bulk solution. When the transmembrane voltage exceeds a threshold, the blocker will be stripped, and the catalytic replication will be activated. Therefore, controlled DNA replication process can be characterized in real-time by recording the ionic current changes of the nanopore. Processive replication of ssDNA by bacteriophage T7 polymerase [49] and by phi29 DNA polymerase [50] was both realized based on similar mechanism. And the replication of ssDNA could even be observed with single nucleotide resolution by periodically adjusting the applied voltage and halting the DNA replication process [51]. Cherf et al. [52] achieved automated forward and reverse ratcheting of DNA using phi29 DNA polymerase, but the error rate was found to be as high as 10 to 24.5 % during replication.
Despite the ability of biological nanopores to read DNA sequences at single-nucleotide resolution, they are limited by the poor stability of either the pore itself or the lipid. In addition, the difficulty to integrate biological nanopores into large-scale arrays has been a long-standing hurdle in nanopore sequencing. Hence, solid-state nanopores have been developed to overcome the intrinsic limitations of biological nanopores.
Solid-state nanopores
Solid-state nanopores have emerged as a versatile alternative to biological nanopores. A solid-state nanopore can be fabricated by drilling a nanometer-sized pore through a solid-state membrane with an ion beam or electron beam. There are many choices of solid-state membrane materials such as ultra-thin silicon nitride (SiNx) membranes, silicon dioxide (SiO2) membranes, and atomic-thick graphene membranes. Next, we will discuss in details recent progress in solid-state nanopores.
Solid-state nanopore fabrication and DNA analysis
The fabrication process of solid-state nanopores has been extensively investigated since 2001 [5]. SiNx and SiO2 were chosen as dielectric membranes in early studies due to their thermal and chemical stability. Li et al. [5] fabricated the first solid-state nanopores by ion beam sculpting (Fig. 5a). Then translocation events of dsDNA [5, 53] and ssDNA [54] were demonstrated by using solid-state nanopores. The diameter of nanopores was found to depend strongly on the effective ion dosage, and a large variation of the nanopore geometry was observed in early studies [5]. Compared with ion beam, electron beam sculpting has been shown to provide sub-nanometer precision of nanopore diameters [6] (Fig. 5b). Feedback control mechanism has been further applied to improve the diameter control during both ion and electron beam sculpting [5, 55]. Another attempt used cold ion beam sculpting to fabricate nanopores through SiNx, which results in the precise control of the nanopore diameter [56].
Solid-state nanopores. a Solid-state nanopore fabricated by ion beam drilling of a SiNx membrane [53]. (i) TEM image of a SiNx nanopore; (ii) Schematic model of the DNA translocation measurement, and (iii) real-time recording of current spikes during DNA translocations through a SiNx nanopore; b Nanopore fabricated by electron beam drilling [55]. (i) TEM characterizations of the nanopores; (ii) Real-time current spikes recorded during DNA translocations
A key advantage of solid-state nanopore technology is that the geometry of solid-state nanopores can be easily manipulated during drilling, which facilitate the quantitative characterization of the relationship between the pore geometry and ionic conductance. Smeets et al. [57] investigated the dependence of the ionic conductance of SiNx nanopores on the concentration of the ions. The concentration of KCl in the bulk was decreased from 1 M to 1 μM. Under the assumption that the nanopore depth is long enough, the access resistance to the pore orifice can be ignored. Therefore, the relationship between the ionic conductance and ion concentrations was given as:
where nKCl are the number densities of K+ or Cl− in the solution; μK and μCl are the electrophoretic mobilities of K+ and Cl−; Lpore, dpore are the length and diameter of the nanopore, and σ refers to the surface charge. Equation (1) indicates that both the ion concentrations and surface charges of the nanopore contribute to the overall conductance of the solid-state nanopore. The experimental results confirmed that at high ion concentrations, the conductance of the nanopore was mainly determined by ion migration through the nanopore [57]. On the other hand, electroosmotic process played a vital role to the nanopore conductance at low ion concentrations [58].
When a DNA translocates through a nanopore, the nanopore is partially occupied by the DNA, which prevents certain amounts of K+ and Cl − from migrating through the nanopore and changes the ionic conductance of the nanopore. It has also been reported that DNA translocation can cause opposite effects on the nanopore signals at low and high ion concentration [57, 59]. The nanopore conductance signal caused by DNA translocation [57] was written by Smeets et al. as:
where Lpore is the length of the nanopore; dDNA is the diameter of DNA molecule; μK, μCl are the electrophoretic mobilities of the ions; nKCl is the number concentration of the salt; e is the elementary charge; and ql,DNA is the charge on DNA per unit length. The first term is related to the current blockade effects by the DNA molecules, and the second term is due to the charge shielding effects by counter ions. At low ion concentration, DNA translocation leaded to increased nanopore current signals due to the shielding effects of the negative charges on the DNA backbone by counter ions. On the other hand, nanopore current blockage was observed when the ion concentration was higher than 0.4 M, as confirmed by experimental results [58].
Nanopores of approximately 10 nm in diameter were drilled into freestanding 20 nm thick SiNx membranes by Skinner et al. in 2009 [60]. And different ss- and ds-RNA homopolymers were discriminated by these solid-state nanopores. However, high voltage was needed in their study to resolve the current signals from the translocating RNA strands due to the large pore diameter and depth. As indicated by Eqs. (1) and (2), decreasing the membrane thickness can increase the signal amplitudes during single-molecule translocations through the nanopore. Wanunu et al. developed a process to effectively reduce the thickness of SiNx nanopores [61]. By combining the methods of electron beam lithography and subsequent reactive ion etching, the thickness of SiNx nanopores was reduced from tens of nanometers to 6 nm. The current signal amplitude was dramatically increased during single-molecule translocation through as-prepared nanopores, and the use of 3 nm diameter pores in sub-10 nm membranes facilitated electronic discrimination among small RNA strands. Venta et al. [62] further optimized the fabrication process of solid-state nanopores, which results in nanopores of 0.8–2 nm in diameter and 5–8 nm in depth. The smaller diameter and depth of their nanopores lead to increased resolution and reduced noise for nucleic acid analysis.
Compared to biological nanopores, unmodified solid-state nanopores lack functional groups that can interact with the translocating nucleic acids. Therefore, the translocation speeds of nucleic acids through solid-state nanopores are even less controllable than biological nanopores. Surface modification has been used to improve the analysis capabilities of solid-state nanopores. It has been shown that a probe-DNA functionalized SiO2 nanopore can distinguish DNA with one base mismatch by hybridization [16]. Also the binding locations between RecA, a DNA-repair protein, with target DNA strands were also successfully mapped by using a RecA modified SiNx nanopore [63]. Nevertheless, there still lacks for facile and reliable methods for controllable surface modification of solid-state nanopores.
Membrane materials for solid-state nanopores
Solid-state nanopores can be made with a wide variety of membrane materials. For example, Al2O3 based membranes have attracted much attention to fabricate solid-state nanopores [64, 65]. During nanopore drilling through an Al2O3 membrane by electron beam, partial dissociation of Al2O3 occurred near the nanopore. The local stoichiometry of Al2O3 was changed with the ratio of oxygen to aluminum atoms decreasing from 1.5 to 0.6. This metallization process provides a method for electrode insertion near the nanopore, which can thus be applied for transverse current measurements [64]. Besides, the phase transformation of Al2O3 induced by the electron beam could be utilized to modulate the surface charges of the nanopores, since different phases of Al2O3 have different isoelectric points in aqueous solutions. In addition, all the phases of Al2O3 have an isoelectric point higher than 7, thus the surface charging of Al2O3 is always positive in neutral solutions. Therefore, electrostatic interactions between the positively charged nanopores and negative charged DNA molecules can be used to reduce the translocation speeds of DNA molecules. Glass capillaries have also been investigated for nanopore fabrication. Laser pipette puller was used to stretch a glass capillary and narrow its diameter. However, the pore diameter of the glass pipette cannot be reduce below 45 nm [66, 67], thus they can only be applied to study large analytes. Owing to the superior chemical and mechanical stability of HfO2, they are also being intensively studied as membrane materials for solid-state nanopores [68–71]. For example, ultra-stable nanopores fabricated with HfO2 membranes have been demonstrated by Larkin et al. [68]. The thickness of the HfO2 nanopores is only 2–7 nm. The mean DNA velocities through the HfO2 nanopores were found to be slower than velocities through similar SiNx pores due to favorable physicochemical interactions of the HfO2 nanopores with nucleic acids [68].
To further reduce the nanopore thickness for single nucleotide resolution, two dimensional nanomaterials have been investigated as membrane materials, such as graphene [72–74], single-layer BN [75], and MoS2 [76] (Fig. 6). Among them, graphene is especially attractive due to its superb electrical and mechanical properties. DNA translocation events have been recorded by using graphene-based nanopores [72–74]. Due to pinholes and cracks of these ultrathin membranes, their insulating properties are poor, which leads to large noise level of the ionic currents. To solve this problem, a thin layer of TiO2 has been deposited on a graphene membrane by atomic layer deposition [73], which was found to effectively reduce the noise of the ionic currents. Furthermore, the hydrophobicity nature of graphene leads to strong interactions between graphene and translocating DNA molecules, which can result in clogging of the graphene nanopores. Therefore, hydrophilicity treatment of the graphene nanopore has also been reported [77].
To combine the merits of different solid-state membrane materials, layer-by-layer stacking of inorganic materials has been investigated to fabricate hybridized solid-state nanopores. A metal-insulator-metal sandwiched structure was designed to achieve base-by-base ratcheting [78]. Stacked graphene-HfO2 nanopore has been shown to possess enhanced mechanical strength than pure HfO2 membranes [79]. Until now, single-nucleotide discrimination has not been achieved using ionic current through solid-state nanopores due to their large pore diameters and depths. Graphene-based nanopores offer subnanometer membrane thickness. However, the pore diameters of current graphene nanopores are still too large to resolve the difference between the four nucleotides by ionic currents. In addition, the translocation speed of ds-DNA through graphene nanopores is still about 10–100 nucleotides per microsecond, which is too fast for the state-of-art electronics.
Transverse current measurements of DNA translocation
In addition to ionic currents through nanopores, transverse current can also be used to analyze translocating DNA molecules. There are currently two types of transverse currents based on nanopores, including tunneling currents and semiconductor currents. For transverse tunneling current measurements, a transverse voltage is applied between a pair of electrodes near a nanopore. The distance between the two electrodes is within nanometer scale. During DNA translocation through the nanopore, the potential barrier between the two electrodes is changed by the DNA, which results in a tunneling current signal related to the translocating DNA. The idea of tunneling current measurements was firstly proposed theoretically by Lagerqvist et al. [80]. According to Lagerqvist et al., the tunneling current of the nanopore can be written as:
where f t (E) and f b (E) are the Fermi-Dirac function of top electrode and bottom electrode, respectively. The tunneling coefficient T(E) depends on the retarded Green function ζ DNA which reflects the effects of DNA translocation.
Ivanov et al. [81] developed the first device for nanopore-based tunneling current measurements. Two platinum electrodes were integrated near a SiNx nanopore by electron beam deposition (Fig. 7a). The ionic current and transverse tunneling current were simultaneously measured during the translocation of λ-DNA. More recently, heat-induced diffusion [82] and electron beam lithography [83] have also been used to fabricate metallic nanogaps within solid-state nanopores, and transverse tunneling current signals were recorded during DNA translocation. The first sequencing attempt using transverse current signal was reported by Tsutsui et al. [84]. The signals of different nucleotides were experimentally detected and identified (Fig. 7b). Furthermore, functionalized groups linked to the electrodes were shown to interact with passing nucleotides, which resulted in increased tunneling current signals [85–87].
DNA analysis based on transverse current measurements. a (i) Schematics of solid-state nanopores embedded with two Pt electrodes; (ii) simultaneously recording of the ionic and transverse current spikes during DNA translocations [81]; b Schematics of single nucleotide detection by two gold electrodes in a nanopore (i) and the differentiation of different nucleotides with transverse tunneling current recorded (ii) [84]
The second type of transverse current measurements depends on the conductivity signals of a semiconductor near a nanopore. According to the model developed by Xie et al. [88], direct gating effects of the semiconductor by the negative charges on the translocating DNA can be ignored. On the other hand, DNA translocation changes the electric field near the nanopore, which then changes the effective gate voltage and conductance of the semiconductor. The gate voltage change can be written as:
where V is the applied voltage; A is area of DNA cross section; l, d are the thickness and diameter of the nanopore, respectively; C cis and C trans are ionic concentration of cis and trans side of nanopore, respectively; r is the distance of the semiconductor to the nanopore opening.
Xie et al. [88] combined silicon nanowire FETs with solid-state nanopores (Fig. 8a). The strong correlation of ionic current spikes of the nanopore and transverse current spikes of the FET has been successfully demonstrated experimentally. Graphene nanoribbons were also a promising candidate in FET sensors to record transverse current during DNA translocation [89] (Fig. 8b). However, the signal-to-noise ratio of graphene electrodes still needs to be improved for single nucleotide resolution. To reduce the noise of semiconducting graphene, Saha et al. [90] proposed the application of metallic graphene nanoribbons (GNRs) near a nanopore. When DNA translocates through a nanopore, the energy level of charge carriers in a metallic GNR varies, which can result in conductance change of the nanoribbon. The noise level can be dramatically reduced due to the indirect interference between the metallic GNR and the translocating DNA. Min et al. [91] proposed a device schematic in which a trench is fabricated on the bulk SiNx as a nanochannel, and a graphene nanoribbon is placed on the channel as a sensing probe. When a DNA strand is electrophoretically driven through the nanochannel, the π-π stacking between nucleotides of the DNA and GNRs can lead to conductance changes of GNR. The key advantage of transverse currents is that the transverse electrodes or transistors are highly localized near the nanopore, which thus facilitates the integration of high-density nanopore arrays. However, nanopore technology based on transverse current measurement is still in its infancy.
Nanopore-FET based DNA analysis. a Nanopore-nanowire FETs for DNA analysis. Schematics of a nanopore-nanowire FET (left) and the simultaneously recording of ionic current and semiconductor current during DNA translocations [88]; b A graphene nanoribbon based nanopore-FET device for the recording of DNA translocations [89]
The control of translocation speeds through solid-state nanopores
DNA translocation processes through nanopores in early reports were fast and lack of control. Molecular dynamics simulation [59, 92] has shown that the translocation events depend upon the ion concentration, applied voltage, and nanopore geometry. The effects by the applied voltage have been noted since the earliest time of the nanopore analysis [23, 93]. The dependence of translocation on nanopore geometry has also been investigated in details [8]. To control the translocation of DNA, ion concentration gradients have been applied to increase the DNA capture rates by accumulating positive charges near the nanopore entrance [10]. It was shown that ion concentration gradients also leaded to cationic electro-osmotic flow, which reduced the DNA translocation speeds [9]. The substitution of K+ with Li+ [94] or Mg2+ [95] was also investigated to reduce the DNA translocation speeds. The translocation speed reduction was due to an increased cation adsorption on the DNA backbone by Li+ or Mg2+, which decreased the effective charges on the DNA molecule.
By lowering the temperature to 2 °C, it was found that DNA translocation speeds can be dramatically reduced [93]. However, we note that the effects of temperature on DNA translocation are complicated. For example, local heating was shown to cause DNA stretching near a nanopore, which results in reduced conformational fluctuation and current noise [96]. By applying a pressure gradient through a nanopore, both the reduction of translocation speed and the translocation of charge-neutral molecules have been demonstrated [11]. And a comprehensive analysis of the pressure gradient effects on molecule translocation was also reported [12]. Chemical modification of solid-state nanopores has also been widely applied for both translocation control and noise reduction [17, 18].
A promising way to overcome the ultrafast DNA translocation is based on the optical detection of converted DNA strands. Each of the four nucleotides on the target DNA was converted to a predefined sequence of oligonucleotides, and then the converted DNA was hybridized with fluorophore-labeled molecular beacons. As the converted DNA translocates through a solid-state nanopore, the molecular beacons were stripped sequentially from the DNA strand, which resulted in specific bursts in fluorescence signals. In this way, each nucleotide on the target DNA was elongated to an oligonucleotide of 100 nt in the converted DNA, which results in longer translocation time for each nucleotide on the target DNA and improves their differentiation [97]. In addition, a waveguide has also been introduced to increase the fluorescence signal-to-noise-ratio of this technique [98].
Optical tweezers were also used to control the translocation of molecules through nanopores. Keyser et al. [13] applied optical tweezers to control the translocation of DNA through SiNx nanopores in 2006. A polystyrene bead was linked to a DNA molecule via streptavidin-biotin interaction. An infrared laser was used to preciously control the position of the bead. The tension force on the bead can be calculated based on Hooke’s law: F = −ktrap∆Z, where ∆Z is the linear deformation of the polystyrene bead. To trap the translocating DNA inside the nanopore, the tension force was tuned to balance the electric force on the DNA. Therefore, the tension by the bead can be used to both reduce the speed of DNA translocation and even pull the DNA molecule out of a nanopore [14]. In recent years, optical tweezers have become a routine tool for controlling the translocation process through nanopores [68].
Magnetic tweezers provide another way to control the translocating DNA by tension force. A magnetic bead was linked to a DNA molecule through streptavidin-biotin interaction. Then a magnetic field was applied to control the magnetic bead and thus the position of the DNA molecule. Reduced translocation speeds and even reverse electrophoresis have been observed experimentally [15]. In addition, Hyun et al. reported the control of molecule translocation by using the tip of a scanning-probe microscope [99] or a probe tip linked to a tuning fork [100].
Hybrid biological/solid-state nanopores
Biological nanopores possess atomically precise structures that can be genetically engineered to achieve single-nucleotide discrimination, while they are traditionally limited by the fragile nature of the supporting lipid substrate and the difficulty to achieve high-density nanopore arrays. On the other hand, solid-state nanopores offer the advantages of durability and are suitable for large-scale integration into high-density arrays, but they cannot yet differentiate the small difference among the four nucleotides due to their large pore sizes. To overcome these limitations in biological and solid-state nanopore technologies, a hybrid biological/solid-state nanopore architecture that combines the merits of biological and solid-state nanopores was developed by Hall et al. [101]. A long dsDNA tail was chemically tethered to a genetically engineered α-HL. When a voltage is applied between the two chambers, the electrophoretic force acts on the negatively charged DNA, and the α-HL was inserted into a SiNx nanopore. Translocation events of poly(dA)100 strands through the hybridized nanopore were then successfully measured by the ionic current signals, confirming the functionality of these hybridized nanopores. However, large leakage currents have been observed in the biological/solid-state nanopore, which was attributed to some leakage path at the interface between the protein and the membrane or protein deformation induced by the insertion. Nevertheless, hybrid biological/solid-state nanopores are becoming an attractive research direction in nanopore technology for high-throughput sequencing.
Nanopores formed by the self-assembly of DNA origami represent another attractive platform for single-molecule analysis [102–104]. The flexibility and versatility in the design of DNA origami makes them ideally suited for construction of nanopores with a broad variety of geometry and surface functionality. Recently, DNA origami has been applied to form hybrid architectures with solid-state nanopores [105, 106], and a variety of applications in DNA analysis were demonstrated to highlight the versatility of these DNA origami nanopores. For example, by tuning the pore size of the DNA origami, controlled folding of dsDNA molecules was demonstrated during their translocation. In addition, by specifically introducing binding sites in the DNA origami nanopores, selective detection of ssDNA strands was further demonstrated [106]. A key challenge related with current DNA origami nanopores is their high leakage currents, which thus results in poor signal-to-noise ratio of single-molecule analysis. The origin of the high leakage currents may be related to the presence of conductive paths both at the interface between DNA and membrane and through the body of the DNA origami. Future studies are needed to reduce the leakage currents and thus improve the signal-to-noise ratio of DNA origami nanopores. Furthermore, the interface at biological and solid-state nanopores needs to be characterized as a function of channel geometry, temperature, and electric field.
Prospects
Since the first publication of modern nanopore technology for DNA analysis in 1996 [23], tremendous advances have been made towards reliable DNA sequencing with nanopores. However, numerous research efforts are still needed for the practical applications of nanopore-based technologies. For example, noise reduction and parallel sequencing are among the key challenges for accurate, cheap, and high-throughput nanopore based sequencing. Noise reduction is still not satisfactory in nanopore analysis, which can result in high error rates. Both low and high frequency noise of nanopore conductance have been investigated by many studies. Low frequency noise was found to be in the form of flicker noise, which can be described with Hooge’s formula [107, 108]. Former studies have shown that the low frequency noise of nanopores can be decreased by reducing local fluctuations, such as geometrical inhomogeneity of nanopores and turbulence or bubbles in the solution [107, 109]. High frequency noise is mostly related to the capacitance of the substrates [109]. It has been shown that the high frequency noise can be decreased by reducing the permeability of the substrates [73]. Another significant challenge in nanopore technologies lies in the need for parallel sequencing. To reduce the cost and increase the throughput of nanopores, it is necessary to simultaneously monitor a large number of nanopores. The main challenges to achieve parallel sequencing are due to both the difficulty of fabricating biological nanopore arrays and also the cross talking between nanopores during current measurements. The arrays of solid-state nanopores are straightforward to fabricate with modern microfabrication techniques [55, 110], although their resolution has not been high enough for direct DNA sequencing. On the other hand, biological nanopores are difficult to be multiplexed and individually addressed. An inorganic template is needed for the fabrication of a biological nanopore array [111, 112]. For both biological and solid-state nanopore arrays, multi-channel, high-speed recording of ionic current has not been demonstrated yet.
In 2012, Oxford Nanopore Technologies claimed the invention of the first two commercial sequencers, MinION and GridION, based on nanopore technology [113]. The first sequencing results by MinION were presented at the Advances in Genome Biology and Technology meeting in 2014. DNA fragments up to 10 kilobases long have been sequenced by the device, which highlights an exciting advance for practical applications of nanopore-based sequencing [114]. Integrated systems have also been paving the way to the commercialization of nanopore analysis [115, 116]. Nevertheless, numerous challenges in nanopore technologies are still to be resolved towards low error rates, rapid and highly parallel recording, and long read length up to 100 kilobases. Thus, we expect that nanopore technologies will continue to be an active and engaging research area that incorporates new ideas and innovative approaches from a wide range of science and engineering disciplines.
References
McGinn S, Gut IG (2013) DNA sequencing – spanning the generations. New Biotechnol 30(4):366–372
Ku C-S, Roukos DH (2013) From next-generation sequencing to nanopore sequencing technology: paving the way to personalized genomic medicine. Expert Rev Med Devices 10(1):1–6
Stoddart D, Heron AJ, Mikhailova E, Maglia G, Bayley H (2009) Single-nucleotide discrimination in immobilized DNA oligonucleotides with a biological nanopore. Proc Natl Acad Sci U S A 106(19):7702–7707
Rincon-Restrepo M, Mikhailova E, Bayley H, Maglia G (2011) Controlled translocation of individual DNA molecules through protein nanopores with engineered molecular brakes. Nano Lett 11(2):746–750
Li J, Stein D, McMullan C, Branton D, Aziz MJ, Golovchenko JA (2001) Ion-beam sculpting at nanometre length scales. Nature 412(6843):166–169
Storm A, Chen J, Ling X, Zandbergen H, Dekker C (2003) Fabrication of solid-state nanopores with single-nanometre precision. Nat Mater 2(8):537–540
Ying YL, Zhang J, Gao R, Long YT (2013) Nanopore‐based sequencing and detection of nucleic acids. Angew Chem Int Ed Engl 52(50):13154–13161
van Dorp S, Keyser UF, Dekker NH, Dekker C, Lemay SG (2009) Origin of the electrophoretic force on DNA in solid-state nanopores. Nat Phys 5(5):347–351
Wanunu M, Morrison W, Rabin Y, Grosberg AY, Meller A (2010) Electrostatic focusing of unlabelled DNA into nanoscale pores using a salt gradient. Nat Nanotechnol 5(2):160–165
He Y, Tsutsui M, Scheicher RH, Fan C, Taniguchi M, Kawai T (2013) Mechanism of how salt-gradient-induced charges affect the translocation of DNA molecules through a nanopore. Biophys J 105(3):776–782
Lu B, Hoogerheide DP, Zhao Q, Zhang H, Tang Z, Yu D, Golovchenko JA (2013) Pressure-controlled motion of single polymers through solid-state nanopores. Nano Lett 13(7):3048–3052
Zhang H, Zhao Q, Tang Z, Liu S, Li Q, Fan Z, Yang F, You L, Li X, Zhang J (2013) Slowing down DNA translocation through solid‐state nanopores by pressure. Small 9(24):4112–4117
Keyser UF, Koeleman BN, Van Dorp S, Krapf D, Smeets RM, Lemay SG, Dekker NH, Dekker C (2006) Direct force measurements on DNA in a solid-state nanopore. Nat Phys 2(7):473–477
Trepagnier EH, Radenovic A, Sivak D, Geissler P, Liphardt J (2007) Controlling DNA capture and propagation through artificial nanopores. Nano Lett 7(9):2824–2830
Peng H, Ling XS (2009) Reverse DNA translocation through a solid-state nanopore by magnetic tweezers. Nanotechnology 20(18):185101
Iqbal SM, Akin D, Bashir R (2007) Solid-state nanopore channels with DNA selectivity. Nat Nanotechnol 2(4):243–248
Krishnakumar P, Gyarfas B, Song W, Sen S, Zhang P, Krstic P, Lindsay S (2013) Slowing DNA translocation through a nanopore using a functionalized electrode. ACS Nano 7(11):10319–10326
Squires AH, Hersey JS, Grinstaff MW, Meller A (2013) A nanopore-nanofiber mesh biosensor to control DNA translocation. J Am Chem Soc 135(44):16304–16307
Song L, Hobaugh MR, Shustak C, Cheley S, Bayley H, Gouaux JE (1996) Structure of staphylococcal α-hemolysin, a heptameric transmembrane pore. Science 274(5294):1859–1865
Derrington IM, Butler TZ, Collins MD, Manrao E, Pavlenok M, Niederweis M, Gundlach JH (2010) Nanopore DNA sequencing with MspA. Proc Natl Acad Sci U S A 107(37):16060–16065
Faller M, Niederweis M, Schulz GE (2004) The structure of a mycobacterial outer-membrane channel. Science 303(5661):1189–1192
Kubitschek HE (1958) Electronic counting and sizing of bacteria. Nature 182(4630):234–235
Kasianowicz JJ, Brandin E, Branton D, Deamer DW (1996) Characterization of individual polynucleotide molecules using a membrane channel. Proc Natl Acad Sci U S A 93(24):13770–13773
Wendell D, Jing P, Geng J, Subramaniam V, Lee TJ, Montemagno C, Guo P (2009) Translocation of double-stranded DNA through membrane-adapted phi29 motor protein nanopores. Nat Nanotechnol 4(11):765–772
Haque F, Li J, Wu H-C, Liang X-J, Guo P (2013) Solid-state and biological nanopore for real-time sensing of single chemical and sequencing of DNA. Nano Today 8(1):56–74
Keyser UF (2011) Controlling molecular transport through nanopores. J R Soc Interface:rsif20110222
Akeson M, Branton D, Kasianowicz JJ, Brandin E, Deamer DW (1999) Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid, and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys J 77(6):3227–3233
Howorka S, Movileanu L, Lu X, Magnon M, Cheley S, Braha O, Bayley H (2000) A protein pore with a single polymer chain tethered within the lumen. J Am Chem Soc 122(11):2411–2416
Howorka S, Cheley S, Bayley H (2001) Sequence-specific detection of individual DNA strands using engineered nanopores. Nat Biotechnol 19(7):636–639
Howorka S, Bayley H (2002) Probing distance and electrical potential within a protein pore with tethered DNA. Biophys J 83(6):3202–3210
Astier Y, Braha O, Bayley H (2006) Toward single molecule DNA sequencing: direct identification of ribonucleoside and deoxyribonucleoside 5′-monophosphates by using an engineered protein nanopore equipped with a molecular adapter. J Am Chem Soc 128(5):1705–1710
Clarke J, Wu H-C, Jayasinghe L, Patel A, Reid S, Bayley H (2009) Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4(4):265–270
Ayub M, Hardwick SW, Luisi BF, Bayley H (2013) Nanopore-based identification of individual nucleotides for direct RNA sequencing. Nano Lett 13(12):6144–6150
Butler TZ, Pavlenok M, Derrington IM, Niederweis M, Gundlach JH (2008) Single-molecule DNA detection with an engineered MspA protein nanopore. Proc Natl Acad Sci U S A 105(52):20647–20652
Chen Y-S, Lee C-H, Hung M-Y, Pan H-A, Chiou J-C, Huang GS (2013) DNA sequencing using electrical conductance measurements of a DNA polymerase. Nat Nanotechnol 8(6):452–458
Wang H-Y, Li Y, Qin L-X, Heyman A, Shoseyov O, Willner I, Long Y-T, Tian H (2013) Single-molecule DNA detection using a novel SP1 protein nanopore. Chem Commun 49(17):1741–1743
Ashkenasy N, Sánchez‐Quesada J, Bayley H, Ghadiri MR (2005) Recognizing a single base in an individual DNA strand: a step toward DNA sequencing in nanopores. Angew Chem Int Ed Engl 117(9):1425–1428
Purnell RF, Schmidt JJ (2009) Discrimination of single base substitutions in a DNA strand immobilized in a biological nanopore. ACS Nano 3(9):2533–2538
Liu N, Jiang Y, Zhou Y, Xia F, Guo W, Jiang L (2013) Two‐way nanopore sensing of sequence‐specific oligonucleotides and small‐molecule targets in complex matrices using integrated DNA supersandwich structures. Angew Chem Int Ed Engl 52(7):2007–2011
de Souza N (2014) Sequencing: protein nanopores to detect DNA methylation. Nat Methods 11(1):8–8
Laszlo AH, Derrington IM, Brinkerhoff H, Langford KW, Nova IC, Samson JM, Bartlett JJ, Pavlenok M, Gundlach JH (2013) Detection and mapping of 5-methylcytosine and 5-hydroxymethylcytosine with nanopore MspA. Proc Natl Acad Sci U S A 110(47):18904–18909
Schreiber J, Wescoe ZL, Abu-Shumays R, Vivian JT, Baatar B, Karplus K, Akeson M (2013) Error rates for nanopore discrimination among cytosine, methylcytosine, and hydroxymethylcytosine along individual DNA strands. Proc Natl Acad Sci U S A 110(47):18910–18915
Dudko OK, Mathé J, Szabo A, Meller A, Hummer G (2007) Extracting kinetics from single-molecule force spectroscopy: nanopore unzipping of DNA hairpins. Biophys J 92(12):4188–4195
Vercoutere WA, Winters‐Hilt S, DeGuzman VS, Deamer D, Ridino SE, Rodgers JT, Olsen HE, Marziali A, Akeson M (2003) Discrimination among individual Watson–Crick base pairs at the termini of single DNA hairpin molecules. Nucleic Acids Res 31(4):1311–1318
Lin J, Kolomeisky A, Meller A (2010) Helix-coil kinetics of individual polyadenylic acid molecules in a protein channel. Phys Rev Lett 104(15):158101
Hornblower B, Coombs A, Whitaker RD, Kolomeisky A, Picone SJ, Meller A, Akeson M (2007) Single-molecule analysis of DNA-protein complexes using nanopores. Nat Methods 4(4):315–317
Benner S, Chen RJ, Wilson NA, Abu-Shumays R, Hurt N, Lieberman KR, Deamer DW, Dunbar WB, Akeson M (2007) Sequence-specific detection of individual DNA polymerase complexes in real time using a nanopore. Nat Nanotechnol 2(11):718–724
Hurt N, Wang H, Akeson M, Lieberman KR (2009) Specific nucleotide binding and rebinding to individual DNA polymerase complexes captured on a nanopore. J Am Chem Soc 131(10):3772–3778
Olasagasti F, Lieberman KR, Benner S, Cherf GM, Dahl JM, Deamer DW, Akeson M (2010) Replication of individual DNA molecules under electronic control using a protein nanopore. Nat Nanotechnol 5(11):798–806
Lieberman KR, Cherf GM, Doody MJ, Olasagasti F, Kolodji Y, Akeson M (2010) Processive replication of single DNA molecules in a nanopore catalyzed by phi29 DNA polymerase. J Am Chem Soc 132(50):17961–17972
Cockroft SL, Chu J, Amorin M, Ghadiri MR (2008) A single-molecule nanopore device detects DNA polymerase activity with single-nucleotide resolution. J Am Chem Soc 130(3):818–820
Cherf GM, Lieberman KR, Rashid H, Lam CE, Karplus K, Akeson M (2012) Automated forward and reverse ratcheting of DNA in a nanopore at 5-A precision. Nat Biotechnol 30(4):344–348
Li J, Gershow M, Stein D, Brandin E, Golovchenko J (2003) DNA molecules and configurations in a solid-state nanopore microscope. Nat Mater 2(9):611–615
Fologea D, Gershow M, Ledden B, McNabb DS, Golovchenko JA, Li J (2005) Detecting single stranded DNA with a solid state nanopore. Nano Lett 5(10):1905–1909
Kim MJ, Wanunu M, Bell DC, Meller A (2006) Rapid fabrication of uniformly sized nanopores and nanopore arrays for parallel DNA analysis. Adv Mater 18(23):3149–3153
Kuan AT, Golovchenko JA (2012) Nanometer-thin solid-state nanopores by cold ion beam sculpting. Appl Phys Lett 100(21):213104
Smeets RM, Keyser UF, Krapf D, Wu M-Y, Dekker NH, Dekker C (2006) Salt dependence of ion transport and DNA translocation through solid-state nanopores. Nano Lett 6(1):89–95
Chang H, Kosari F, Andreadakis G, Alam M, Vasmatzis G, Bashir R (2004) DNA-mediated fluctuations in ionic current through silicon oxide nanopore channels. Nano Lett 4(8):1551–1556
Luan B, Stolovitzky G (2013) An electro-hydrodynamics-based model for the ionic conductivity of solid-state nanopores during DNA translocation. Nanotechnology 24(19):195702
Skinner GM, van den Hout M, Broekmans O, Dekker C, Dekker NH (2009) Distinguishing single-and double-stranded nucleic acid molecules using solid-state nanopores. Nano Lett 9(8):2953–2960
Wanunu M, Dadosh T, Ray V, Jin J, McReynolds L, Drndić M (2010) Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nat Nanotechnol 5(11):807–814
Venta K, Shemer G, Puster M, Rodriguez-Manzo JA, Balan A, Rosenstein JK, Shepard K, Drndic M (2013) Differentiation of short, single-stranded DNA homopolymers in solid-state nanopores. ACS Nano 7(5):4629–4636
Kowalczyk SW, Hall AR, Dekker C (2009) Detection of local protein structures along DNA using solid-state nanopores. Nano Lett 10(1):324–328
Venkatesan BM, Shah AB, Zuo JM, Bashir R (2010) DNA sensing using nanocrystalline surface‐enhanced Al2O3 nanopore sensors. Adv Funct Mater 20(8):1266–1275
Venkatesan BM, Dorvel B, Yemenicioglu S, Watkins N, Petrov I, Bashir R (2009) Highly sensitive, mechanically stable nanopore sensors for DNA analysis. Adv Mater 21(27):2771–2776
Steinbock LJ, Otto O, Chimerel C, Gornall J, Keyser UF (2010) Detecting DNA folding with nanocapillaries. Nano Lett 10(7):2493–2497
Steinbock L, Otto O, Skarstam D, Jahn S, Chimerel C, Gornall J, Keyser U (2010) Probing DNA with micro-and nanocapillaries and optical tweezers. J Phys Condens Matter 22(45):454113
Larkin J, Henley R, Bell DC, Cohen-Karni T, Rosenstein JK, Wanunu M (2013) Slow DNA transport through nanopores in hafnium oxide membranes. ACS Nano 7(11):10121–10128
Larkin JW, Henley R, Bell DC, Cohen-Karni T, Rosenstein JK, Wanunu M (2014) Detection of single biopolymers at high current bandwidth with hafnium oxide nanopores. Biophys J 106(2):413a–414a
Larkin J, Henley RY, Muthukumar M, Rosenstein JK, Wanunu M (2014) High-bandwidth protein analysis using solid-state nanopores. Biophys J 106(3):696–704
Ivankin A, Henley RY, Larkin J, Carson S, Toscano ML, Wanunu M (2014) Label-free optical detection of biomolecular translocation through nanopore arrays. ACS Nano 8(10):10774–10781
Garaj S, Hubbard W, Reina A, Kong J, Branton D, Golovchenko JA (2010) Graphene as a subnanometre trans-electrode membrane. Nature 467(7312):190–193
Merchant CA, Healy K, Wanunu M, Ray V, Peterman N, Bartel J, Fischbein MD, Venta K, Luo Z, Johnson ATC, Drndić M (2010) DNA translocation through graphene nanopores. Nano Lett 10(8):2915–2921
Schneider GF, Kowalczyk SW, Calado VE, Gg P, Zandbergen HW, Vandersypen LMK, Dekker C (2010) DNA translocation through graphene nanopores. Nano Lett 10(8):3163–3167
Liu S, Lu B, Zhao Q, Li J, Gao T, Chen Y, Zhang Y, Liu Z, Fan Z, Yang F (2013) Boron nitride nanopores: highly sensitive DNA single‐molecule detectors. Adv Mater 25(33):4549–4554
Liu K, Feng J, Kis A, Radenovic A (2014) Atomically thin molybdenum disulfide nanopores with high sensitivity for DNA translocation. ACS Nano 8(3):2504–2511
Schneider GF, Xu Q, Hage S, Luik S, Spoor JN, Malladi S, Zandbergen H, Dekker C (2013) Tailoring the hydrophobicity of graphene for its use as nanopores for DNA translocation. Nat Commun 4:2619
Luan B, Peng H, Polonsky S, Rossnagel S, Stolovitzky G, Martyna G (2010) Base-by-base ratcheting of single stranded DNA through a solid-state nanopore. Phys Rev Lett 104(23):238103
Shim J, Rivera JA, Bashir R (2013) Electron beam induced local crystallization of HfO 2 nanopores for biosensing applications. Nanoscale 5(22):10887–10893
Lagerqvist J, Zwolak M, Di Ventra M (2006) Fast DNA sequencing via transverse electronic transport. Nano Lett 6(4):779–782
Ivanov AP, Instuli E, McGilvery CM, Baldwin G, McComb DW, Albrecht T, Edel JB (2010) DNA tunneling detector embedded in a nanopore. Nano Lett 11(1):279–285
Tsutsui M, Rahong S, Iizumi Y, Okazaki T, Taniguchi M, Kawai T (2011) Single-molecule sensing electrode embedded in-plane nanopore. Sci Rep 1:46
Fanget A, Traversi F, Khlybov S, Granjon P, Magrez A, Forró L, Radenovic A (2013) Nanopore integrated nanogaps for DNA detection. Nano Lett 14(1):244–249
Tsutsui M, Taniguchi M, Yokota K, Kawai T (2010) Identifying single nucleotides by tunnelling current. Nat Nanotechnol 5(4):286–290
Pathak B, Löfås H, Prasongkit J, Grigoriev A, Ahuja R, Scheicher RH (2012) Double-functionalized nanopore-embedded gold electrodes for rapid DNA sequencing. Appl Phys Lett 100(2):023701
Liang F, Li S, Lindsay S, Zhang P (2012) Synthesis, physicochemical properties, and hydrogen bonding of 4 (5)‐substituted 1‐H‐imidazole‐2‐carboxamide, a potential universal reader for DNA sequencing by recognition tunneling. Chem Eur J 18(19):5998–6007
Huang S, He J, Chang S, Zhang P, Liang F, Li S, Tuchband M, Fuhrmann A, Ros R, Lindsay S (2010) Identifying single bases in a DNA oligomer with electron tunnelling. Nat Nanotechnol 5(12):868–873
Xie P, Xiong Q, Fang Y, Qing Q, Lieber CM (2012) Local electrical potential detection of DNA by nanowire-nanopore sensors. Nat Nanotechnol 7(2):119–125
Traversi F, Raillon C, Benameur S, Liu K, Khlybov S, Tosun M, Krasnozhon D, Kis A, Radenovic A (2013) Detecting the translocation of DNA through a nanopore using graphene nanoribbons. Nat Nanotechnol 8(12):939–945
Saha KK, Drndić M, Nikolic BK (2011) DNA base-specific modulation of microampere transverse edge currents through a metallic graphene nanoribbon with a nanopore. Nano Lett 12(1):50–55
Min SK, Kim WY, Cho Y, Kim KS (2011) Fast DNA sequencing with a graphene-based nanochannel device. Nat Nanotechnol 6(3):162–165
Liang L, Cui P, Wang Q, Wu T, Ågren H, Tu Y (2013) Theoretical study on key factors in DNA sequencing with graphene nanopores. RSC Adv 3(7):2445–2453
Meller A, Nivon L, Branton D (2001) Voltage-driven DNA translocations through a nanopore. Phys Rev Lett 86(15):3435
Kowalczyk SW, Wells DB, Aksimentiev A, Dekker C (2012) Slowing down DNA translocation through a nanopore in lithium chloride. Nano Lett 12(2):1038–1044
Zhang Y, Liu L, Sha J, Ni Z, Yi H, Chen Y (2013) Nanopore detection of DNA molecules in magnesium chloride solutions. Nanoscale Res Lett 8(1):1–8
Belkin M, Maffeo C, Wells DB, Aksimentiev A (2013) Stretching and controlled motion of single-stranded DNA in locally heated solid-state nanopores. ACS Nano 7(8):6816–6824
McNally B, Singer A, Yu Z, Sun Y, Weng Z, Meller A (2010) Optical recognition of converted DNA nucleotides for single-molecule DNA sequencing using nanopore arrays. Nano Lett 10(6):2237–2244
Larkin J, Foquet M, Korlach J, Wanunu M (2013) 207 Nanopore immobilization of DNA polymerase enhances single-molecule sequencing. J Biomol Struct Dyn 31(sup1):134–135
Hyun C, Rollings R, Li J (2012) Probing access resistance of solid‐state nanopores with a scanning‐probe microscope tip. Small 8(3):385–392
Hyun C, Kaur H, Rollings R, Xiao M, Li J (2013) Threading immobilized DNA molecules through a solid-state nanopore at > 100 μs per base rate. ACS Nano 7(7):5892–5900
Hall AR, Scott A, Rotem D, Mehta KK, Bayley H, Dekker C (2010) Hybrid pore formation by directed insertion of [alpha]-haemolysin into solid-state nanopores. Nat Nanotechnol 5(12):874–877
Langecker M, Arnaut V, Martin TG, List J, Renner S, Mayer M, Dietz H, Simmel FC (2012) Synthetic lipid membrane channels formed by designed DNA nanostructures. Science 338(6109):932–936
Burns JR, Stulz E, Howorka S (2013) Self-assembled DNA nanopores that span lipid bilayers. Nano Lett 13(6):2351–2356
Hernández-Ainsa S, Misiunas K, Thacker VV, Hemmig EA, Keyser UF (2014) Voltage-dependent properties of DNA origami nanopores. Nano Lett 14(3):1270–1274
Plesa C, Ananth AN, Linko V, Gülcher C, Katan AJ, Dietz H, Dekker C (2013) Ionic permeability and mechanical properties of DNA origami nanoplates on solid-state nanopores. ACS Nano 8(1):35–43
Hernández-Ainsa S, Bell NA, Thacker VV, Göpfrich K, Misiunas K, Fuentes-Perez ME, Moreno-Herrero F, Keyser UF (2013) DNA origami nanopores for controlling DNA translocation. ACS Nano 7(7):6024–6030
Smeets RM, Keyser UF, Dekker NH, Dekker C (2008) Noise in solid-state nanopores. Proc Natl Acad Sci U S A 105(2):417–421
Smeets R, Dekker N, Dekker C (2009) Low-frequency noise in solid-state nanopores. Nanotechnology 20(9):095501
Tabard-Cossa V, Trivedi D, Wiggin M, Jetha NN, Marziali A (2007) Noise analysis and reduction in solid-state nanopores. Nanotechnology 18(30):305505
Wildeson IH, Ewoldt DA, Colby R, Stach EA, Sands TD (2010) Controlled growth of ordered nanopore arrays in GaN. Nano Lett 11(2):535–540
Kleefen A, Pedone D, Grunwald C, Wei R, Firnkes M, Abstreiter G, Rant U, Tampé R (2010) Multiplexed parallel single transport recordings on nanopore arrays. Nano Lett 10(12):5080–5087
Baaken G, Ankri N, Schuler A-K, Jr R, Behrends JC (2011) Nanopore-based single-molecule mass spectrometry on a lipid membrane microarray. ACS Nano 5(10):8080–8088
Eisenstein M (2012) Oxford nanopore announcement sets sequencing sector abuzz. Nat Biotechnol 30(4):295–296
Data from pocket-sized genome sequencer unveiled (2014) Nature Publishing Group. http://www.nature.com/news/data-from-pocket-sized-genome-sequencer-unveiled-1.14724
Zhang N, Hu Y-X, Gu Z, Ying Y-L, He P-G, Long Y-T (2014) An integrated software system for analyzing nanopore data. Chin Sci Bull 59(35):4942–4945
Gao R, Ying Y-L, Yan B-Y, Long Y-T (2014) An integrated current measurement system for nanopore analysis. Chin Sci Bull 59(35):4968–4973
Acknowledgments
This work was supported by the National Natural Science Foundation of China (21322302, 21303024, 21365003 and 31400731).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Shi, J., Hou, J. & Fang, Y. Recent advances in nanopore-based nucleic acid analysis and sequencing. Microchim Acta 183, 925–939 (2016). https://doi.org/10.1007/s00604-015-1503-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-015-1503-y