Abstract
Short-term effects of soil physical disturbance by ploughing and nitrogen and phosphate fertilisation on arbuscular mycorrhizal fungal (AMF) communities and on intraspecific populations of Rhizophagus irregularis in a buffer strip surrounded by arable fields were studied. Pre-grown Plantago lanceolata plantlets were transplanted into fertilised and/or ploughed experimental plots. After 3 months, the glomeromycotan communities in the roots of these trap plants were analysed using 454 pyrosequencing of a fragment of the RNA polymerase II gene (RPB1). Intraspecific populations of R. irregularis were studied by restriction fragment length polymorphism (RFLP) analysis of the mitochondrial large ribosomal subunit (mtLSU) gene. Soil disturbance significantly increased the diversity of species-level molecular taxa (MTs) and altered community structure, whilst fertilisation alone had no significant effect, unless coupled with ploughing. At the population level, the expected shift from genotypes of R. irregularis typically found in grasslands to those usually found in arable sites was only partially observed. In conclusion, in the short-term, physical soil disturbance, as well as nitrogen fertilisation when coupled with physical soil disturbance, affected AMF community and to a smaller extent population composition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arbuscular mycorrhiza is a mutualistic interaction known to be instrumental for plant growth under a wide range of environmental conditions as it is able to improve uptake of mineral nutrients, especially phosphorus and nitrogen. It makes arbuscular mycorrhizal fungi (AMF), obligate biotrophs placed in the phylum Glomeromycota, prime candidates for a major role in future sustainable agriculture (Harrison et al. 2002; Jeffries et al. 2003; Barrios 2007; Gianinazzi et al. 2010; Fitter et al. 2011). In order to manage AMF communities in agroecosystems and their benefits in agriculture, a better understanding of factors influencing AMF diversity and composition in agroecosystems is needed. The intensity of agricultural management seems to have a large influence on the community diversity of Glomeromycota (Oehl et al. 2003), and species-level diversity has been shown to be reduced in agricultural soils (Helgason et al. 2002; Öpik et al. 2006) but it is not necessarily low (Hijri et al. 2006). The effect of two common agricultural practices used in conventional and intensive farming, namely physical soil disturbance and the application of fertilisers, has been studied separately in long-term experiments in agroecosystems (e.g. Jansa et al. 2002, 2003; Oehl et al. 2003; Lin et al. 2012; Liu et al. 2012; Thian et al. 2013; Stockinger et al. 2014).
Disturbance in the form of tillage seems to play a major role in AMF community assembly, as it necessitates continuous healing or regrowth of the mycelial network which favours species able to persist under tillage, fertilisation or fungicide treatments (Verbruggen and Kiers 2010). Hence, shifts in AMF communities have been reported in a long-term field experiment in Switzerland (Jansa et al. 2002; Thonar et al. 2012; Stockinger et al. 2014). On the other hand, Lekberg et al. (2012) applied local disturbance to plots in a Danish coastal grassland and did not detect any modification on AMF community structure, which was attributed to a high resilience of the mycorrhizal network. With regard to the effects of fertilisation, it is well established that fungal colonisation levels are reduced in high-nutrient systems (Mäder et al. 2000) and that long-term fertilisation induces shifts in community composition (Toljander et al. 2008; Kim et al. 2014). The latter may involve increased incidence of species having an r-type life history strategy, i.e. low investment in external mycelium and rapid spore production (Verbruggen et al. 2010). However, studies on nitrogen fertilisation effects on AMF community have revealed either positive, neutral or negative effects on AMF diversity (Eom et al. 1999; Egerton-Warburton and Allen 2000; Lin et al. 2012; Liu et al. 2012; Kim et al. 2014), and changes in diversity level do not necessarily imply quantitative shifts of species abundances in the AMF community. For example, Santos et al. (2006) analysed AMF community composition along a nitrogen fertilisation gradient using DGGE and SSU sequencing and found a less diverse community at high N levels but no straightforward qualitative effects.
Evidence shows that AMF taxa can differ in their life history strategies (Sýkorová et al. 2007a). For some, their tolerance to agricultural practices allows to have a broad geographical distribution (Öpik et al. 2006). Differences among species and isolates in the capacity to transport nutrients have been documented (Munkvold et al. 2004; Jansa et al. 2005, 2008). The model fungus Rhizophagus irregularis is currently the only glomeromycotan species for which molecular markers allow the analysis of population structure in colonised roots in the field (Börstler et al. 2008). Its intraspecific diversity has also been studied using molecular markers, which require isolates in root organ culture (Koch et al. 2004). This species has been reported to be ubiquitous, occurring in an extremely wide range of environments, which may be due to ecotypes adapted to different sets of environmental conditions (see Börstler et al. 2010). For example, undisturbed low-nutrient grassland sites were shown to preferentially inhabit a distinct set of mitochondrial large ribosomal subunit (mtLSU) genotypes of R. irregularis, which are very rarely found in arable fields (Börstler et al. 2010). Thus, it could be interesting to link the distribution of this species with that of its populations.
The objective of the study was to better understand the influence of physical soil disturbance and N or/and P fertilisation in the short-term on (i) AMF communities and (ii) R. irregularis populations. A field experiment manipulating these parameters was set up in a buffer strip in a landscape dominated by arable fields to identify relevant factors to community and population assembly. In agricultural landscapes, buffer strips fulfil important functions, e.g. to limit erosion, or as possible diversity reservoir for plants or refuge for the fauna (Vickery et al. 2004), and have therefore been actively supported by the EU regulations since the Common Agricultural Policy (CAP) reform in 2003 (in France, the “Grenelle 2”, law no. 2010-788). It could be hypothesised that they equally constitute reservoirs for AMF. As AMF communities in arable soils are known to be relatively species-poor (Daniell et al. 2001) compared to grasslands (Helgason et al. 1998; Boddington and Dodd 2000), it is useful to understand the transition between the two in order to be able to manage AMF functional diversity and thus optimise their ecosystem services in agroecosystems.
The present approach is unique in its concurrent analysis of two different levels of diversity (population and community). To our knowledge, this is the first direct study of an experimental manipulation of a glomeromycotan population in the field, in which both community and population levels were studied, and the second study using RPB1 gene as community marker gene for AMF. It was hypothesised that the two levels of diversity are both impacted by the manipulations but that they may not respond to all factors in the same manner. Based on previous studies in arable soils, it was expected that the community diversity would decrease and that known disturbance-adapted species such as Funneliformis mosseae would increase in abundance. On the population level, it was expected that mtLSU genotypes of R. irregularis of the type typically found in grasslands would be replaced by types commonly found in arable soils. Previous studies have shown a reduction of AMF richness after fertilisation, however over longer periods of time (Camenzind et al. 2014), and therefore, it was unclear whether our system would respond in a similar way.
Material and methods
Field site and experimental design
The study was conducted in an 8-year-old buffer strip on the experimental site of Epoisses, at Bretenières in the Burgundy region of France. Twenty-four plots (1.5 m × 1.5 m) were delimited within an area of 26 m × 3.5 m. This buffer strip was surrounded by arable lands where no fertiliser, fungicide nor pesticide was applied. The soil of this grassland had the following characteristics: pH (H2O) 7.78, organic carbon content 34.9 g kg−1, total nitrogen 2.68 g kg−1, organic matter 60.3 g kg−1 and phosphorus content (Olsen) 0.121 g kg−1. The experimental set-up corresponded to a split plot design with disturbance as main factor and four levels of fertilisation within each soil disturbance treatment (schematically depicted in Online Resource 1a). Physical soil disturbance consisted of ploughing to a depth of 15 cm using a motor cultivator, resulting in destruction of the grass cover. Fertilisation treatments consisted of local application of either nitrogen alone as NH4NO3 (+N, 100 kg ha−1), phosphorus alone as triple superphosphate (+P, 30 kg ha−1) or in combinations (+N+P), including a non-treated fertilised control (C). Each treatment was replicated three times. In each plot (Online Resource 1b), eight Plantago lanceolata plantlets, pre-grown under controlled conditions in a climate chamber during 1 month, were planted manually in four points (two plantlets by point), each 50 cm inside the plot margins. P. lanceolata was chosen because it occurs naturally in the studied area and is known as an excellent host for AMF.
Plant harvest and DNA extraction
Three months after transplantation, plants in all treatments were harvested in July 2012. Plants were identified to species level where possible. The absence of flowers in several cases necessitated a preliminary classification according to vegetative characters. Plants and rhizosphere soils from each plot were dug out, placed in plastic bags and stored at 4 °C. Shoots were dried and weighed. Roots were washed and weighed, then cut into 1-cm pieces, which were thoroughly mixed within a sample. A small subsample was stained in Trypan blue for quantification of AMF colonisation using the method of Trouvelot et al. (1986). The remaining roots were frozen and stored at −80 °C for subsequent DNA extraction. Seventy milligrammes of fresh roots was ground in a 1.5-mL tube using a pestle in liquid nitrogen and extracted according to the manufacturer’s protocol of the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany). DNA was eluted in 70 and 50 μL AE buffer and stored at −80 °C.
Amplification of mtLSU and restriction fragment length polymorphism analyses
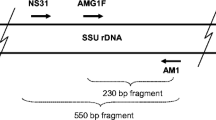
For studying R. irregularis populations, the mtLSU gene was used, which is currently the sole marker gene allowing to genotype this species in colonised roots (Börstler et al. 2008). A first PCR was performed using 0.02 U/μL Phusion polymerase, 1 μL genomic DNA, ×1 Phusion HF Buffer, 0.5 μM of each primer (RNL-28a and RNL-5, Börstler et al. 2008), 0.2 mM of each dNTPs, 0.4 μg/μL BSA and 3 % DMSO in a total volume of 25 μL. Cycling parameters were 30 s at 98 °C, 33 cycles of 10 s at 98 °C, 30 s at 55 °C and 1 min 40 s at 72 °C, followed by 10 min at 72 °C on an Eppendorf Mastercycler epGradient S (Eppendorf, Hamburg, Germany). Success of amplification was checked on 1 % agarose gels. Products of the first PCR were diluted 1:100 and used as templates for the second reaction using the primer pair RNL-29 and RNL-30 (Börstler et al. 2008). One microlitre of diluted PCR product from the first reaction was used together with 0.02 U/μL Phusion polymerase, ×1 Phusion HF Buffer, 0.4 μM of each primer, 0.15 mM of dNTPs, ×4 BSA and 3 % DMSO in a volume of 50 μL. Cycling parameters were 30 s at 98 °C, 36 cycles of 10 s at 98 °C, 20 s at 60 °C and 1 min 40 s at 72 °C, followed by 10 min at 72 °C.
Positive amplifications were cloned according to the instructions of the Strataclone PCR Cloning Kit (Agilent Technology, Courtaboeuf, France). At least 19 clones of each sample were re-amplified using the nested primers. These amplicons were digested using three different enzymes (BsaJI, DraIII, HindIII), according to the method described by Börstler et al. (2008). Digestion products were loaded on a 1.5 % gel (1 % agarose + 0.5 % NuSieve). The combined patterns of the three digestions defined a restriction fragment length polymorphism (RFLP) profile. Each RFLP profile was compared to a reference database of RFLP types (Börstler et al. 2008, 2010) and assigned to known types. For RFLP profiles that were not yet present in the database, the respective amplicons were sequenced by Sanger sequencing (GATC, Mulhouse, France). Maximum likelihood phylogenetic trees were obtained using the RAxML software.
Amplification of the RPB1 gene and 454 pyrosequencing
We chose the gene RPB1 to analyse the AMF community level. This gene was previously identified as a useful marker for phylogenetic analysis of fungi in addition to ribosomal gene regions (James et al. 2006) and especially for the Glomeromycota (Redecker and Raab 2006). In addition, it has been proposed as a secondary DNA barcode for fungi (Schoch et al. 2012). It has the advantage to be generally present as a single copy in the genome, and primers with high specificity for Glomeromycota (>99 %) are available (Stockinger et al. 2014).
Samples were prepared for 454 pyrosequencing using a two-step PCR approach. A first PCR was performed using 0.02 U/μL Phusion polymerase (Thermo-Scientific, Illkirch, France), 1 μL genomic DNA, ×1 Phusion HF Buffer, 0.5 μM of each primer (RPB1-HSm375-mix7 and RPB1-1210r, Stockinger et al. 2014) 0.2 mM of each dNTPs, 0.4 μg/μL BSA and 3 % DMSO in a total volume of 20 μL. Cycling parameters were 30 s at 98 °C, 35 cycles of 10 s at 98 °C, 30 s at 61 °C and 30 s at 72 °C, followed by 10 min at 72 °C on an Eppendorf Mastercycler epGradient S. Amplification success was checked on 1 % agarose gels. Each DNA extract was separately amplified three times. The three PCR products, with a size around 1100 bp, were pooled for each sample and purified using a High Pure PCR Kit (Roche, Basel, Switzerland). Purified PCR products were diluted 1:10 in sterile filtered water (Eurobio, Courtaboeuf, France) and then used as template in the second step of the nested PCR.
In order to add molecular identifiers (MIDs, 10-bp barcodes for post-sequencing sample identification), a short nested PCR was performed using 0.02 U/μL Phusion polymerase, 1 μL of diluted purified PCR product, ×1 Phusion HF Buffer, 0.5 μM of each primer (RPB1-DR160f + mix10 and RPB1-1210 + MID), 0.2 mM of each dNTPs, 0.4 μg/μL BSA and 3 % DMSO in a total volume of 20 μL. Cycling parameters were 30 s at 98 °C, 10 cycles of 10 s at 98 °C, 30 s at 64 °C, 30 s at 72 °C, followed by 10 min at 72 °C. Again, PCR products were purified as described above. Final amplicons (≈700 bp) were quantified using the Quant-iTTM Picogreen® dsDNA Assay Kit (Invitrogen, Life Technologies, Saint-Aubin, France) and mixed equimolarly in one library of 24 MIDs. 454-pyrosequencing was performed on a one-fourth plate by Beckman Coulter Genomics (Grenoble, France) on a 454 GS FLX Roche instrument. Raw data were submitted to the European Nucleotide Archive under the reference ID PRJEB9130.
RPB1 reference database
The glomeromycotan classification follows that of Redecker et al. (2013). Three new sequences of Scutellospora calospora (LK985317–LK985319) were added to the 56 sequences already available in our RPB1 reference database (Stockinger et al. 2014). Sequences were obtained from isolates of the International Bank of Glomeromycota (IBG/BEG) located in Dijon (France) (for details see Online Resource 2 and Online Resource 3). DNA of single or multiple spores was extracted in 5 μL of 5X GoTaq Flexi Buffer (Promega, Charbonnières, France). Gene sequences were amplified as described in the previous paragraph, and amplicons were sequenced by the Sanger method (GATC, Mulhouse, France). Sequences were edited using the Staden package 4 version 1.5 (Bonfield et al. 1995) and aligned in ARB (Ludwig et al. 2004). Phylogenetic relationships were inferred in RAxML using the maximum likelihood method with bootstraps. The two species chosen as outgroup for the reference tree were Mucor hiemalis and Mortierella verticillata.
Sequence analyses and operational taxonomic unit delimitation
Sequence reads were separated based on the MIDs in the library using the Quantitative Insights Into Microbial Ecology (QIIME) pipeline (Caporaso et al. 2010b). Then, primer and MID sequences were removed. Sequences were filtered by quality (25) and length thresholds (200 bp). Sequences having ambiguous bases were removed. Unaligned sequences at 60 % similarity with the reference alignment using PyNAST (Caporaso et al. 2010a) were discarded. Sequences were clustered at 99.7 % similarity using USEARCH (Edgar 2010) in order to perform a UCHIME de novo chimera search on clusters by barcode (Edgar et al. 2011). MACSE (Ranwez et al. 2011) was used to denoise sequences at the codon level. Sequences were classified both based on sequence similarity (operational taxonomic unit—OTUs) and on phylogenetic clustering (molecular taxa). We used both approaches to better support our conclusions for the following reasons: on one hand, using cutoff-based measures as OTUs can be problematic as sequence variation of RPB1 and other marker genes differ between glomeromycotan lineages (Stockinger et al. 2014), resulting in an overestimation of diversity. On the other hand, molecular taxa can be difficult to define in groups with few described species. Using CROP (Hao et al. 2011), an OTU search with a cutoff at 99.2 % sequence identity was performed (Stockinger et al. 2014). OTUs were assigned to a reference tree (Online Resource 3), firstly in QIIME using the RDP-naïve Bayesian classifier method (Wang et al. 2007); secondly the EPA algorithm of RAxML (Berger and Stamatakis 2011) was used for confirmation. According to these phylogenetic assignments, OTUs were grouped in molecular taxa (MTs) at the species level. If the two phylogenetic assignments were not congruent, OTUs were assigned to a higher MT level (genus or family). Before further analysis, singletons and non-glomeromycotan clusters were removed.
Statistical analysis of pyrosequencing reads and RFLPs
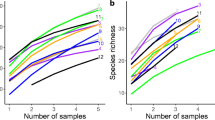
To determine whether sequence depth was sufficient, OTU accumulation curves were generated using the QIIME pipeline. Prior to statistical analyses, the number of sequence reads per sample was standardised by OTU subsampling to a 1400-sequences threshold, which was determined by OTU accumulation analysis. Three samples were removed from analyses as their sequence numbers were too low: In these cases (disturbed and undisturbed controls and +N/+P/undisturbed condition), the remaining two replicates were used (see Online Resource 5). Representative OTU sequences were aligned in order to obtain a tree used in unweighted (based on phylogenetic structure) and weighted (based on phylogenetic structure and weighted by OTU abundances) UniFrac distance matrices (Lozupone and Knight 2005) were visualised by principal coordinates analyses (PCoAs) using QIIME. In addition, non-phylogenetic Bray-Curtis dissimilarity distance matrices were calculated based on OTU abundance, MT abundances at species, genus and family levels for ordination analyses based by non-metric multidimensional scaling (NMDS) using the vegan package (Oksanen et al. 2013) of the software (R development core team 2013), and only the most informative is shown.
Treatment effects (physical soil disturbance, nitrogen and phosphorus fertilisation) as well as block effects (experimental design) on AMF community were statistically tested by nested permutation multivariate analysis of variance (PERMANOVA) with 2000 permutations in QIIME based on Bray-Curtis distances using MTs. Alpha diversity was evaluated using the Shannon index per sample on abundance matrices. As the design was unbalanced, linear mixed models were performed, using the nlme package (Pinheiro et al. 2014) of R software, to compare across treatments the mean Shannon diversity both on species-level and genus-level MT abundances. Those models were randomised by block, and comparisons among fertilisation treatments (both nitrogen and phosphorus) were tested within each disturbance treatment, as they were nested factors. Indicator species for treatments were identified using the indicspecies package of R software (De Caceres and Legendre 2009), and only species having both sensitivity and specificity values higher than 0.75 were retained. Significant results are shown only for species with more than 100 sequences over the whole experiment to avoid overestimating the influence of rare species.
Prior to analyses of the RFLP data, clone numbers per sample were standardised by random subsampling to 19 clones, the lowest number of analysed clones per sample. The vegan package of R was used to compare mtLSU RFLP-type abundances by performing a nested mixed model of variance, as described above in the community analysis part.
Results
Sequences, operational taxonomic units and molecular taxa
A total of 51,106 sequence reads were recovered after filtering and removal of chimeric sequences, which overall generated 62 OTUs at 99.2 % nucleotide sequence identity, ranging from 12 to 33 OTUs per sample. Based on OTU accumulation curve analysis, a subsampling to 1400 sequences per sample (Online Resource 4) was performed, as the curves indicated that the great majority of AMF OTUs was obtained using this threshold. Twenty-one samples of 1400 sequences each were used for downstream analysis, with OTU numbers per sample ranging from 10 to 30. Overall, representatives of five out of the 11 families of the phylum Glomeromycota were detected as the OTU assignation to the reference tree generated 25 species-level molecular taxa (see Online Resource 5) which belonged to 13 genus-level MTs. In all samples, Glomerales accounted for more than 85 % of the sequence reads. Often, one species in a genus was highly overrepresented (see Online Resource 5).
Plant diversity, biomass and root colonisation
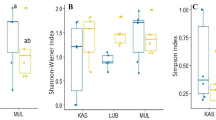
Alpha diversity, expressed by the Shannon index, was increased significantly in disturbed plots (F = 32.986; P < 0.001) and under nitrogen fertilisation within the disturbance treatment (F = 6.559; P = 0.013), whereas no effect of phosphorus fertilisation within disturbance treatments was found according to nested linear mixed model analysis (Fig. 1, Table 1). An increase of P. lanceolata root and shoot biomass was found in disturbed plots (F = 35.805 and F = 52.715, P < 0.001, respectively). Moreover, the increase of shoot biomass within disturbed plots was larger under nitrogen fertilisation (F = 4.650; P = 0.034, data in Online Resource 6). P. lanceolata plantlets showed a reduction of the intensity of mycorrhization by disturbance (F = 11.729, P = 0.005) and among non-disturbed plots by phosphorus fertilisation (F = 8.881, P = 0.005, data in Online Resource 6). A positive effect of physical soil disturbance was observed on plant richness (F = 91.769, P < 0.001, data in Online Resource 6).
Mean Shannon diversity index based on species-level molecular taxa (MTs) across treatments of physical soil disturbance (ND non disturbed, D disturbed), and fertilisation treatments within the physical soil disturbance treatment (+N nitrogen, +P phosphorus, +N+P nitrogen and phosphorus, C no fertilisation). Different letters indicate significant differences of variance (P < 0.05) between treatments according to the nested linear mixed model randomised by block. Error bars represent standard error
Shifts in glomeromycotan community composition
In this study, statistical analysis by nested PERMANOVA (Table 2) showed that physical soil disturbance (R 2 = 0.364, P: 0.001) had a significant effect on AMF community composition and that nitrogen fertilisation had a significant effect within the soil disturbance treatment (R 2 = 0.194, P = 0.022). The NMDS ordination of Bray-Curtis dissimilarity distance matrix on species-level MTs revealed differences of the symbiotic AMF communities from physically disturbed and undisturbed plots (grey polygons, Fig. 2). Both fresh root and dry shoot biomasses (blue arrows) were negatively correlated (P < 0.05) with the axis NMDS1 that segregated physical soil disturbance treatments. Abundances of R. irregularis and Claroideoglomus sp1 were positively correlated with this axis, whereas F. mosseae and Septoglomus viscosum were negatively correlated (red crosses in Fig. 2).
Non-metric multidimensional scaling (NMDS) ordination of arbuscular mycorrhizal fungal community composition based on a Bray-Curtis dissimilarity distance matrix of species-level molecular taxa (MTs). The two polygons link samples in the same soil disturbance treatment (disturbed and undisturbed plots, respectively). Colours indicate fertilisation treatments: nitrogen fertilisation (brown), phosphorus fertilisation (green), nitrogen and phosphorus fertilisation (blue) and no fertilisation (black). Red crosses correspond to species vectors. Significant (P < 0.05) projected vectors are displayed as blue arrows
Using the De Caceres-Jansen index after 2000 iterations, five species were identified as indicator species for physical soil disturbance (Online Resource 7, Fig. 3), i.e. F. mosseae, Funneliformis sp3, Glomeraceae sp6, Septoglomus sp3 and S. viscosum-like, whereas none of the species was identified as indicator of non-disturbance. Concerning F. mosseae, a positive effect of phosphorus fertilisation in interaction with soil disturbance was observed. In addition, a negative effect of physical soil disturbance and nitrogen fertilisation was observed for R. irregularis (Online Resource 8). The abundance of the species Claroideoglomus sp2 was reduced in disturbed plots, where its abundance increased when nitrogen fertiliser was applied. The abundance of Paraglomus sp2 was generally higher in disturbed plots, especially under nitrogen fertilisation (Online Resource 8). The same tendencies were found at the genus level (data not shown).
Mean relative abundance as percentage of the different species-, genus- and family level molecular taxa (MTs) in the community after subsampling to a 1400 sequence threshold across treatments of disturbance (ND non-disturbed plots, D disturbed plots) and fertilisations (+N nitrogen, +P phosphorus, +N+P nitrogen and phosphorus, C no fertilisation). Dollar symbols show indicator species identified by the De Caceres-Jansen index using 2000 permutations
Thus, soil disturbance was a strong factor influencing MT relative abundance (at genus and species levels), but it seems to be largely modulated by fertilisation, especially nitrogen. No spatial (block) effect was identified in any of the performed diversity analyses.
R. irregularis population study
Overall, 24 different mtLSU RFLP profiles were detected. The corresponding sequences fall into the two intraspecific clades (Online Resource 9) of genotypes typically found in arable fields and in grasslands, respectively (Börstler et al. 2008), and a third more basal clade related to the isolate BEG 141. The RFLP profiles were classified into these three groups (Fig. 4). In the three replicates of the non-disturbed and non-fertilised treatments, exclusively, RFLP profiles corresponding to the “grassland clade” were found. The same was the case under N fertilisation combined with soil disturbance. In the other treatments, at least half of all RFLP profiles were also of this type; in addition, arable field genotypes were detected to varying extents in the remaining disturbed and/or fertilised treatments, but the effects of the treatments were not statistically significant (P > 0.05). No clear distribution pattern was found for RFLP types corresponding to the BEG 141 clade. Overall, R. irregularis mtLSU genotypes were relatively patchily distributed with different replicates of the same treatment strongly varying.
Combined diagram of the mean sequence number of the species-level MT Rhizophagus irregularis (bar chart with standard deviation of mean abundance per treatment) and the relative abundance of three mtLSU genotype groups: the BEG 141 group, “arable genotypes” and “grassland genotypes”. mtLSU genotypes are shown in the proportions of the clones detected for each group, with relation to the species-level MT abundance in the respective treatment, set as 100 %. Physical soil disturbance treatments are ND non-disturbed plots and D disturbed plots. Fertilisation treatments are +N nitrogen, +P phosphorus, +N+P nitrogen and phosphorus, C no fertilisation
Discussion
The usefulness of the protein-encoding gene RPB1 as molecular marker, proposed as secondary fungal DNA barcode (Schoch et al. 2012), has been previously demonstrated in an AMF community analysis of an agroecosystem (Stockinger et al. 2014). In the present study, molecular taxon richness is in the same range as expected from previous spore- or rDNA sequence-based studies (Oehl et al. 2003; Öpik et al. 2006), and all expected lineages of glomeromycotan fungi were detected. Considering the major known disadvantage of the multitude of rDNA-based systems for AMF diversity analysis, namely the strong sequence polymorphism within the organism; RPB1 therefore offers an interesting new alternative.
The present study revealed strong effects by physical soil disturbance on AMF community composition and diversity and by nitrogen fertilisation within the soil disturbance treatment. Soil disturbance has previously been identified as a factor influencing AMF communities’ composition in natural plant communities (Husband et al. 2002; Sýkorová et al. 2007b; Schnoor and Olsson 2010), as well as in agroecosystems in long-term experiments (Jansa et al. 2002; Rosendahl 2008; Borriello et al. 2012; Stockinger et al. 2014). The negative effect of physical soil disturbance on plant root colonisation found here has been previously reported in heathland and forest, but not in an annual pasture (Jasper et al. 1991). In contrast to the present findings, no significant change in AMF communities was observed in a recent study of a soil disturbance experiment over 4.5 months in a Danish coastal grassland (Lekberg et al. 2012). The authors explained this finding by a high resilience of local AMF communities. These authors also used P. lanceolata as a bait plant and 454 sequencing to analyse the AMF community. Other studies have been conducted in agriculturally used fields subjected to different tillage treatments and are therefore more difficult to compare. For example, Stockinger et al. (2014) detected significant shifts in AMF community composition according to tillage treatments. Nevertheless, the present results are in agreement with studies which have revealed an influence of nitrogen fertilisation on AMF community composition in long-term experiments, both in agroecosystems (Avio et al. 2013; Lin et al. 2012) and in a semiarid grassland (Porras-Alfaro et al. 2007). Borriello et al. (2012) concluded that AMF communities are more strongly structured by nitrogen fertilisation than by soil disturbance. However, it was not possible to strictly test this hypothesis in the present study, due to the nested experimental design.
A significant increase in AMF diversity, as expressed by the Shannon index, was detected in the disturbed plots, which could come as a surprise because some other studies have shown that agricultural practices, and especially soil disturbance in the form of tillage, negatively affect AMF OTU richness, as for example in a 2-year experiment in a grassland (Schnoor et al. 2011). It seems quite possible that diversity increases initially after disturbance before decreasing in the long-term. It would therefore be interesting to further monitor the kinetics of this kind of system under repeated disturbance and/or long-term fertilisation.
Increased phosphorus fertilisation levels have long been known to reduce AMF root colonisation levels (see e.g. Gianinazzi et al. 2002), which is what was found also in this study in undisturbed plots. Changes in AMF community structure or diversity could also be expected in this situation, as previously reported in some studies (e.g. Alguacil et al. 2010). However, this was not the case, in line with other studies (Smilauer 2001; Jansa et al. 2014), which could indicate that downregulation of colonisation affects AMF taxa rather globally, not in a specific manner. This hypothesis would be interesting to verify by analyses of artificial communities under phosphorus fertilisation under controlled conditions.
The present study detected shifts in AMF community composition and diversity in the short-term, which are driven by soil physical disturbance and intensified by nitrogen fertilisation. A simultaneous increase root and shoot biomass of the studied plantlets was found, which could in turn have had an influence on the AMF community composition. Soil disturbance also affects abiotic properties of the soil like nutrient availability (Fraterrigo et al. 2005) or soil moisture (Robertson et al. 1993). Soil disturbance is also known to decrease plant species competition in grasslands (Wilson and Tilman 1993), creating new patches for plant colonisation (Grubb 1977) and for competition for light (De Cauwer et al. 2006), resulting in an increase in plant biodiversity (Schnoor and Olsson 2010). Specific interactions have been reported between pairs of plant and fungal symbionts (Vandenkoornhuyse et al. 2003; Scheublin et al. 2007), and a decisive influence of AMF on plant communities was demonstrated (van der Heijden et al. 1998, 2003, 2008). It might be therefore an interesting hypothesis to address the possible effects that an AMF community modified by disturbance has on plant diversity.
The types of treatments applied may involve direct and plant-mediated effects. By disturbing and fertilising the plots at the small scale, the opportunity was left open for intact hyphal networks to recolonize the bait plants from surrounding host plants, which led to an increase in plant species richness in the disturbed plots. Disturbance has a direct impact on the fungal network by disrupting it, thereby interacting directly with fungal traits of mycelium architecture and the ability to restore hyphal connections and homogenise within plot the inoculum potential. Correlations between plant species and AMF richness have been reported as positive (Landis et al. 2004; Hiiesalu et al. 2014) and negative (Öpik et al. 2008; Lekberg et al. 2013). On the other hand, the space liberated by disturbance alters plant growth by competitive release, which could again directly affect fungal symbionts. Fertilisation is likely to have complex plant-mediated effects, as the community of symbiotic fungi may be altered when the plant downregulates its root colonisation level. This could lead to altered competition between the fungi or to preferential nutrient allocation to selected fungal species by the plant (Kiers and van der Heijden 2006). As the grass cover was destroyed in our disturbed plots, this obviously allowed dormant components of the soil seed bank to establish and may have had an effect on AMF assemblages by competitive release. Equally, we assume that the observed effects on shoot and root biomass are resulting from soil physical disturbance and fertilisation treatments.
The root samples of P. lanceolata were strongly dominated firstly by Glomeraceae, with at least 45 % of sequences per sample and a total number of 15 species-level MTs, followed by Claroideoglomeraceae (representing 7 to 33 % of sequences per sample, with a total number of five MTs at species level). The dominance of Glomeraceae has been reported in numerous studies, whereas a dominance of these two families together has only been reported in grasslands (Dai et al. 2013). What is notable is the almost complete absence of Gigasporaceae and Acaulosporaceae. However, those two families have often been described to be relatively rare in grasslands (Sýkorová et al. 2007a; Dumbrell et al. 2010), with few exceptions (Lin et al. 2012), and have also been shown to have slow root colonisation rates. In addition, the Gigasporaceae are known to form extensive mycelium in the soil rather than in the roots (Hart and Reader 2002). Previous studies have paid little attention to Claroideoglomeraceae, which was relatively recently split off from the Glomeraceae, possibly because the family was less prominent in many molecular studies due to mismatches with the frequently used primer AM1 (Alguacil et al. 2008; Dai et al. 2013). Generally, PCR primer biases have been shown to influence detected community patterns (Kohout et al. 2014).
Overall, few qualitative differences among treatments were observed here. As analysis depth of the high-throughput sequencing method increased, compared to cloning/sequencing approaches, a pronounced presence/absence pattern is less likely to be expected. The use of pyrosequencing data as a semi-quantitative measure of taxon/OTU abundance has been demonstrated in some studies but has also been disputed (Amend et al. 2010; Harris et al. 2010). We nevertheless choose to carefully discuss some quantitative shifts in the proportions of MTs/OTUs across treatments, for the following reasons: even if the inherent PCR biases leading to high or low sequence numbers are difficult to control, they are likely to remain the same across treatments, leaving differences in sequence read abundance as a valid working proxy of fungal abundances. Nevertheless, the necessity for true quantitative analysis such as qPCR to analyse glomeromycotan communities should be underlined, but this would require extensive knowledge of the taxa present to be able to design qPCR primers.
Among the shifts in community composition observed in the present study, some were in accordance with previous studies, which were mostly long-term experiments, whilst others concerned taxa that have never been studied before in this context. Soil disturbance causes hyphal network disruption, which favours species with r-type life history strategies, having large capacities to quickly sporulate and to colonise soil and roots, e.g. F. mosseae (Sýkorová et al. 2007a; Rosendahl 2008). In the present study, the abundance of the MT F. mosseae was indeed highly correlated with the soil disturbance treatment for which it was identified as indicator species. Also, the relative abundance of Septoglomus and especially the species S. viscosum increased significantly in disturbed plots, which identified it as an indicator species for physical soil disturbance. To our knowledge, this is the first time that such a response has been reported for this species. In some previous studies, Paraglomus has been reported to be relatively rare in agricultural soils compared to pasture soil (Lumini et al. 2010) and was shown to increase with plant growth (Yu et al. 2012). In spore-based studies, this genus was likely to be overlooked because of its small, colourless spores (Oehl et al. 2003, 2005). In other studies, it might have been missed because of primer mismatches as mentioned by Stockinger et al. (2009, 2010). Interestingly, the genus Paraglomus seems to be more easily detected in soil than in roots (Hempel et al. 2007). It seems to be widespread in agricultural soils in England (Gosling et al. 2014) where it responded to organic management and phosphorus fertilisation. Dai et al. (2013) reported the genus Paraglomus to be a dominant OTU in the maritime ecozone of Canada and to be rare in grasslands. In the field situation studied here, the genus Paraglomus (especially the molecular species Paraglomus sp2) was significantly more frequent in disturbed plots. These results suggest that the distribution of this genus in agricultural systems and its response to environmental factors merit further study.
The present study indicates a global decrease of the genus Claroideoglomus and of Claroideoglomus sp1 upon soil disturbance, an effect, however, that appears to be reversed by nitrogen fertilisation. This species was one of the most prominent species in terms of explaining the variability of the community composition of samples. Interestingly, according to treatments, contrasting trends were observed in relative abundance of the genera Claroideoglomus and Rhizophagus (i.e. R. irregularis). One possible explanation would be that the members of these two genera act in a complementary fashion, as demonstrated between Claroideoglomus claroideum and R. irregularis (Jansa et al. 2008). As R. irregularis has been frequently discussed as a generalist (Sýkorová et al. 2007a), the shift of R. irregularis relative abundance is somewhat surprising but has to be discussed with regard to the population structure of this species, which was analysed for the first time in parallel in the present study.
About half of the mtLSU genotypes of R. irregularis detected here are new to science. Genotypes typically found in grasslands have been distinguished from those characteristic of arable fields (Börstler et al. 2008). It has not been clear which cultural practices are mainly responsible for the division between the two population subsets, arable fields vs. grassland types, and what may trigger a transition between them. As intraspecific diversity in the Glomeromycota may account for a fair proportion of functional diversity (Leake et al. 2004; Koch et al. 2006; Angelard et al. 2010), and this phenomenon could be relevant for plant growth in agricultural systems. We therefore hypothesised that arable genotypes of R. irregularis would replace the grassland types under agricultural practices. Contrary to this expectation, a clear shift was not observed from grassland to arable RFLP types under the present conditions. Also, the experimental replicates showed large variation in their population composition. Such a pattern might reflect tolerance of grassland RFLP types to soil disturbance/fertilisation, perhaps favoured by the fact that the experiment was relatively small-scaled and allowed ingrowth of mycelia from the surrounding grasslands. It could however also be a possible consequence of the low quantity of inoculum of arable-type R. irregularis in the initial grassland, which may also partly explain the surprising decrease of the species-level MT R. irregularis in the physical soil disturbance treatment. Confirmation of the link between population and species distributions of a generalist species such as R. irregularis to understand their widespread distribution will be an interesting question to assess in further studies from both an ecological and an agricultural point of view.
In conclusion, physical soil disturbance and fertilisation of a buffer strip, even within less than a season, resulted in clear shifts in the structure of AMF communities. Physical soil disturbance was the dominant factor although it was clearly modulated by nitrogen fertilisation. Changes in the population structure of R. irregularis were less pronounced. These results will contribute to a better understanding of the dynamics of AMF communities and populations in agroecosystems. Future studies will have to show whether these effects are reversible and/or whether over longer periods of time these patterns will even become more pronounced, consisting in a reduced AMF species diversity and a clear shift of arable genotypes of R. irregularis.
References
Alguacil MM, Lumini E, Roldan A, Salinas-Garcia JR, Bonfante P, Bianciotto V (2008) The impact of tillage practices on arbuscular mycorrhizal fungal diversity in subtropical crops. Ecol Appl 18:527–536. doi:10.1890/07-0521.1
Alguacil MM, Lozano Z, Campoy JM, Roldán A (2010) Phosphorus fertilisation management modifies the biodiversity of AM fungi in a tropical savanna forage system. Soil Biol Biochem 1114–1122. doi:10.1016/j.soilbio.2010.03.012
Amend AS, Seifert KA, Bruns TD (2010) Quantifying microbial communities with 454 pyrosequencing: does read abundance count? Mol Ecol 19:5555–5565. doi:10.1111/j.1365-294X.2010.04898.x
Angelard C, Colard A, Niculita-Hirzel H, Croll D, Sanders IR (2010) Segregation in a mycorrhizal fungus alters rice growth and symbiosis-specific gene transcription. Curr Biol 20:1216–1221. doi:10.1016/j.cub.2010.05.031
Avio L, Castaldini M, Fabiani A, Bedini S, Sbrana C, Turrini A, Giovannetti M (2013) Impact of nitrogen fertilisation and soil tillage on arbuscular mycorrhizal fungal communities in a Mediterranean agroecosystem. Soil Biol Biochem 67:285–294. doi:10.1016/j.soilbio.2013.09.005
Barrios E (2007) Soil biota, ecosystem services and land productivity. Ecol Econ 64:269–285. doi:10.1016/j.ecolecon.2007.03.004
Berger SA, Stamatakis A (2011) Aligning short reads to reference alignments and trees. Bioinformatics 27:2068–2075. doi:10.1093/bioinformatics/btr320
Boddington CL, Dodd JC (2000) The effect of agricultural practices on the development of indigenous arbuscular mycorrhizal fungi. II. Studies in experimental microcosms. Plant Soil 218:145–157. doi:10.1023/A:1014911318284
Bonfield JK, Smith KF, Staden R (1995) A new DNA sequence assembly program. Nucl Acids Res 23:4992–4999. doi:10.1093/nar/23.24.4992
Borriello R, Lumini E, Girlanda M, Bonfante P, Bianciotto V (2012) Effects of different management practices on arbuscular mycorrhizal fungal diversity in maize fields by a molecular approach. Biol Fertil Soils 48:911–922. doi:10.1007/s00374-012-0683-4
Börstler B, Raab PA, Thiéry O, Morton JB, Redecker D (2008) Genetic diversity of the arbuscular mycorrhizal fungus Glomus intraradices as determined by mitochondrial large subunit rRNA gene sequences is considerably higher than previously expected. New Phytol 180:452–465. doi:10.1111/j.1469-8137.2008.02574.x
Börstler B, Thiéry O, Sýkorová Z, Berner A, Redecker D (2010) Diversity of mitochondrial large subunit rDNA haplotypes of Glomus intraradices in two agricultural field experiments and two semi-natural grasslands. Mol Ecol 19:1497–1511. doi:10.1111/j.1365-294X.2010.04590.x
Camenzind T, Hempel S, Homeier J, Horn S, Velescu A, Wilcke W, Rillig MC (2014) Nitrogen and phosphorus additions impact arbuscular mycorrhizal abundance and molecular diversity in a tropical montane forest. Glob Chang Biol 20:3546–3659. doi:10.1111/gcb.12618
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010a) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Caporaso JG, Kuczynski J, Stombaugh J et al (2010b) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. doi:10.1038/nmeth.f.303
Dai M, Bainard LD, Hamel C, Gan Y, Lynch D (2013) Impact of land use on arbuscular mycorrhizal fungal communities in rural Canada. Appl Environ Microbiol 79:6719–6729. doi:10.1128/AEM.01333-13
Daniell TJ, Husband R, Fitter AH, Young JPW (2001) Molecular diversity of arbuscular mycorrhizal fungi colonising arable crops. FEMS Microbiol Ecol 36:203–209. doi:10.1111/j.1574-6941.2001.tb00841.x
De Caceres M, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology, URL http://sites.google.com/site/miqueldecaceres/
De Cauwer B, Reheul D, D’Hooghe K, Nijs I, Milbau A (2006) Disturbance effects on early succession of field margins along the shaded and unshaded side of a tree lane. Agr Ecosyst Environ 112:78–86. doi:10.1016/j.agee.2005.07.006
Dumbrell AJ, Nelson M, Helgason T, Dytham C, Fitter AH (2010) Relative roles of niche and neutral processes in structuring a soil microbial community. ISME J 4:325–337. doi:10.1038/ismej.2010.48
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. doi:10.1093/bioinformatics/btq461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Egerton-Warburton LM, Allen EB (2000) Shifts in arbuscular mycorrhizal communities along an anthropogenic nitrogen deposition gradient. Ecol Appl 10:484–496. doi:10.1890/1051-0761(2000)010[0484:SIAMCA]2.0.CO;2
Eom AH, Hartnett DC, Wilson GWT, Figge DAH (1999) The effect of fire, mowing and fertilizer amendment on arbuscular mycorrhizas in tallgrass prairie. Am Midl Nat 142:55–70. doi:10.1674/0003-0031(1999)142[0055:TEOFMA]2.0.CO;2
Fitter AH, Helgason T, Hodge A (2011) Nutritional exchanges in the arbuscular mycorrhizal symbiosis: implications for sustainable agriculture. Fungal Biol Rev 25:68–72. doi:10.1016/j.fbr.2011.01.002
Fraterrigo JM, Turner MG, Pearson SM, Dixon P (2005) Effects of past land use on spatial heterogeneity of soil nutrients in southern appalachian forests. Ecol Monogr 75:215–230. doi:10.1890/03-0475
Gianinazzi S, Schüepp H, Barea JM, Haselwandter K (eds) (2002) Mycorrhizal technology in agriculture: from genes to bioproducts. Birkhäuser Verlag, Switzerland
Gianinazzi S, Gollotte A, Binet MN, van Tuinen D, Redecker R, Wipf D (2010) Agroecology: the key role of arbuscular mycorrhizas in ecosystem services. Mycorrhiza 20:519–530. doi:10.1007/s00572-010-0333-3
Gosling P, Proctor M, Jones J, Bending GD (2014) Distribution and diversity of Paraglomus spp. in tilled agricultural soils. Mycorrhiza 24:1–11. doi:10.1007/s00572-013-0505-z
Grubb PJ (1977) Maintenance of species-richness in plant communities—importance of regeneration niche. Biol Rev Camb Philos 52:107–145. doi:10.1111/j.1469-185X.1977.tb01347.x
Hao XL, Jiang R, Chen T (2011) Clustering 16S rRNA for OTU prediction: a method of unsupervised Bayesian clustering. Bioinformatics 27:611–618. doi:10.1093/bioinformatics/btq725
Harris JK, Sahl JW, Castoe TA, Wagner BD, Pollock DD, Spear JR (2010) Comparison of normalization methods for construction of large, multiplex amplicon pools for next-generation sequencing. Appl Environ Microbiol 76:3863–3868. doi:10.1128/AEM.02585-09
Harrison MJ, Dewbre GR, Liu JY (2002) A phosphate transporter from Medicago truncatula involved in the acquisition of phosphate released by arbuscular mycorrhizal fungi. Plant Cell 14:2413–2429. doi:10.1105/tpc.004861
Hart MM, Reader RJ (2002) Taxonomic basis for variation in the colonization strategy of arbuscular mycorrhizal fungi. New Phytol 153:335–344. doi:10.1046/j.0028-646X.2001.00312.x
Helgason T, Daniell TJ, Husband R, Fitter AH, Young JPW (1998) Ploughing up the wood-wide web? Nature 394:431. doi:10.1038/28764
Helgason T, Merryweather JW, Denison J, Wilson P, Young JPW, Fitter AH (2002) Selectivity and functional diversity in arbuscular mycorrhizas of co-occurring fungi and plants from a temperate deciduous woodland. J Ecol 90:371–384. doi:10.1046/j.1365-2745.2001.00674.x
Hempel S, Renker C, Buscot F (2007) Differences in the species composition of arbuscular mycorrhizal fungi in spore, root and soil communities in a grassland ecosystem. Environ Microbiol 9:1930–1938. doi:10.1111/j.1462-2920.2007.01309.x
Hiiesalu I, Partel M, Davison J, Gerhold P, Metsis M, Moora M, Öpik M, Vasar M, Zobel M, Wilson SD (2014) Species richness of arbuscular mycorrhizal fungi: associations with grassland plant richness and biomass. New Phytol 203:233–244. doi:10.1111/nph.12765
Hijri I, Sýkorová Z, Oehl F, Ineichen K, Mäder P, Wiemken A, Redecker D (2006) Communities of arbuscular mycorrhizal fungi in arable soils are not necessarily low in diversity. Mol Ecol 15:2277–2289. doi:10.1111/j.1365-294X.2006.02921.x
Husband R, Herre EA, Turner SL, Gallery R, Young JPW (2002) Molecular diversity of arbuscular mycorrhizal fungi and patterns of host association over time and space in a tropical forest. Mol Ecol 11:2669–2678. doi:10.1046/j.1365-294X.2002.01647.x
James TY, Kauff F, Schoch CL et al (2006) Reconstructing the early evolution of Fungi using six-gene phylogeny. Nature 443:818–824. doi:10.1038/nature05110
Jansa J, Mozafar A, Anken T, Ruh R, Sanders IR, Frossard E (2002) Diversity and structure of AMF communities as affected by tillage in a temperate soil. Mycorrhiza 12:225–234. doi:10.1007/s005572-002-0163-z
Jansa J, Mozafar A, Kuhn G, Anken T, Ruh R, Sanders IR, Frossard E (2003) Soil tillage affects the community structure of mycorrhizal fungi in maize roots. Ecol Appl 13:1164–1176. doi:10.1890/1051-0761(2003)13[1164:STATCS]2.0.CO;2
Jansa J, Mozafar A, Frossard E (2005) Phosphorus acquisition strategies within arbuscular mycorrhizal fungal community of a single field site. Plant Soil 276:163–176. doi:10.1007/s11104-005-4274-0
Jansa J, Smith FA, Smith SE (2008) Are there benefits of simultaneous root colonization by different arbuscular mycorrhizal fungi? New Phytol 177:779–789. doi:10.1111/j.1469-8137.2007.02294.x
Jansa J, Erb A, Oberholzer HS, Smilauer P, Egli S (2014) Soil and geography are more important determinants of indigenous arbuscular mycorrhizal communities than management practices in Swiss agricultural soils. Mol Ecol 23:2118–2135. doi:10.1111/mec.12706
Jasper DA, Abbott LK, Robson AD (1991) The effect of soil disturbance on vesicular-arbuscular mycorrhizal fungi in soils from different vegetation types. New Phytol 118:471–476. doi:10.1111/j.1469-8137.1991.tb00029.x
Jeffries P, Gianinazzi S, Perotto S, Turanu K, Barea M (2003) The contribution of arbuscular mycorrhizal fungi in sustainable maintenance of plant health and soil fertility. Biol Fertil Soils 37:1–16. doi:10.1007/s00374-002-0546-5
Kiers ET, van der Heijden MGA (2006) Mutualistic stability in the arbuscular mycorrhizal symbiosis: exploring hypotheses of evolutionary cooperation. Ecology 87:1627–1636. doi:10.1890/0012-9658(2006)87[1627:MSITAM]2.0.CO;2
Kim YC, Gao C, Zheng Y, He XH, Chen L, Wan SQ, Guo LD (2014) Arbuscular mycorrhizal fungal community response to warming and nitrogen addition in a semi-arid steppe ecosystem. Mycorrhiza. doi:10.1007/s00572-014-0608-1
Koch AM, Kuhn G, Fontanillas P, Fumagalli L, Goudet J, Sanders IR (2004) High genetic variability and low local diversity in a population of arbuscular mycorrhizal fungal. Proc Natl Acad Sci U S A 101:2369–2371. doi:10.1073/pnas.0306441101
Koch AM, Croll D, Sanders IR (2006) Genetic variability in a population of arbuscular mycorrhizal fungi causes variation in plant growth. Ecol Lett 9:103–110. doi:10.1111/j.1461-0248.2005.00853.x
Kohout P, Sudova R, Janouskova M, Ctvrtlikova M, Hejda M, Pankova H, Slavikova R, Stajerova K, Vosatka M, Sýkorová Z (2014) Comparison of commonly used primer sets for evaluating arbuscular mycorrhizal fungal communities: is there a universal solution? Soil Biol Biochem 68:482–493. doi:10.1016/j.soilbio.2013.08.027
Landis FC, Gargas A, Givnish TJ (2004) Relationships among arbuscular mycorrhizal fungi, vascular plants and environmental conditions in oak savannas. New Phytol 164:493–504. doi:10.1111/j.1469-8137.2004.01202.x
Leake JR, Johnson D, Donnelly DP, Muckle GE, Boddy L, Read DJ (2004) Networks of power and influence: the role of mycorrhizal mycelium in controlling plant communities and agroecosystem functioning. Can J Bot 82:1016–1045. doi:10.1139/B04-060
Lekberg Y, Schnoor T, Kjoller R, Gibbons SM, Hansen LH, Al-Soud WA, Sorensen SJ, Rosendahl S (2012) 454-sequencing reveals stochastic local reassembly and high disturbance tolerance within arbuscular mycorrhizal fungal communities. J Ecol 100:151–160. doi:10.1111/j.1365-2745.2011.01894.x
Lekberg Y, Gibbons SM, Rosendahl S, Ramsey PW (2013) Severe plant invasions can increase mycorrhizal fungal abundance and diversity. ISME J 7:1424–1433. doi:10.1038/ismej.2013.41
Lin X, Feng Y, Zhang H, Chen R, Wang J, Zhang J, Chu H (2012) Long-term balanced fertilization decreases arbuscular mycorrhizal fungal diversity in an arable soil in north China revealed by 454 pyrosequencing. Environ Sci Technol 46:5764–5771. doi:10.1021/es3001695
Liu YJ, Shi GX, Mao L, Cheng G, Jiang SJ, Ma XJ, An LZ, Du GZ, Johnson NC, Feng HY (2012) Direct and indirect influences of 8 yr of nitrogen and phosphorus fertilization on Glomeromycota in an alpine meadow ecosystem. New Phytol 194:523–535. doi:10.1111/1574-6941.12361
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235. doi:10.1128/AEM.71.12.8228-8235.2005
Ludwig W, Strunk O, Westram R et al (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371. doi:10.1093/nar/gkh293
Lumini E, Orgiazzi A, Borriello R, Bonfante P, Bianciotto V (2010) Disclosing arbuscular mycorrhizal fungal biodiversity in soil through a land-use gradient using a pyrosequencing approach. Environ Microbiol 12:2165–2179. doi:10.1111/j.1462-2920.2009.02099.x
Mäder P, Edenhofer S, Boller T, Wiemken A, Niggli U (2000) Arbuscular mycorrhizae in a long-term field trial comparing low-input (organic, biological) and high-input (conventional) farming systems in a crop rotation. Biol Fertil Soils 31:150–156. doi:10.1007/s003740050638
Munkvold L, Kjoller R, Vestberg M, Rosendahl S, Jakobsen I (2004) High functional diversity within species of arbuscular mycorrhizal fungi. New Phytol 164:357–364. doi:10.1111/j.1469-8137.2004.01169.x
Oehl F, Sieverding E, Ineichen K, Mäder P, Boller T, Wiemken A (2003) Impact of land use intensity on the species diversity of arbuscular mycorrhizal fungi in agroecosystems of Central Europe. Appl Environ Microbiol 69:2816–2824. doi:10.1128/AEM.69.5.2816-2824.2003
Oehl F, Sieverding E, Ineichen K, Ris EA, Boller T, Wiemken A (2005) Community structure of arbuscular mycorrhizal fungi at different soil depths in extensively and intensively managed agroecosystems. New Phytol 165:273–283. doi:10.1111/j.1469-8137.2004.01235.x
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2013) Vegan: community ecology package. R package version 2.0-9. http://CRAN.R-project.org/package=vegan
Öpik M, Moora M, Liira J, Zobel M (2006) Composition of root-colonizing arbuscular mycorrhizal fungal communities in different ecosystems around the globe. J Ecol 94:778–790. doi:10.1111/j.1365-2745.2006.01136.x
Öpik M, Moora M, Zobel M, Saks U, Wheatley R, Wright F, Daniell T (2008) High diversity of arbuscular mycorrhizal fungi in a boreal herb-rich coniferous forest. New Phytol 179:867–876. doi:10.1111/j.1469-8137.2008.02515.x
Pinheiro J, Bates D, Debroy S, Sarkar D, R core team (2014) Package nmle. Linear and nonlinear mixed effects models. http://CRAN.R-project.org/package=nlme
Porras-Alfaro A, Herrera J, Natvig DO, Sinsabaugh RL (2007) Effect of long-term nitrogen fertilization on mycorrhizal fungi associated with a dominant grass in a semiarid grassland. Plant Soil 296:65–75. doi:10.1007/s11104-007-9290-9
R development core team (2013) R: a language and environment for statistical computing. R foundation for statistical computing, Vienna, ISBN 3-900051-07-0, URL http://www.R-project.org/
Ranwez V, Harispe S, Delsuc F, Douzery EJP (2011) MACSE: Multiple Alignment of Coding SEquences accounting for frameshifts and stop codons. PLoS One 6, e22594. doi:10.1371/journal.pone.0022594
Redecker D, Raab P (2006) Phylogeny of the Glomeromycota (arbuscular mycorrhizal fungi): recent developments and new gene markers. Mycologia 98:885–895. doi:10.3852/mycologia.98.6.885
Redecker D, Schüßler A, Stockinger H, Stürmer SL, Morton JB, Walker C (2013) An evidence-based consensus for the classification of arbuscular mycorrhizal fungi (Glomeromycota). Mycorrhiza 23:515–531. doi:10.1007/s00572-013-0486-y
Robertson GP, Crum JR, Ellis BG (1993) The spatial variability of soil resources following long-term disturbance. Oecologia 96:451–456. doi:10.1007/BF00320501
Rosendahl S (2008) Communities, populations and individuals of arbuscular mycorrhizal fungi. New Phytol 178:253–266. doi:10.1111/j.1469-8137.2008.02378.x
Santos JC, Finlay RD, Tehler A (2006) Molecular analysis of arbuscular mycorrhizal fungi colonising a semi-natural grassland along a fertilisation gradient. New Phytol 172:159–168. doi:10.1111/j.1469-8137.2006.01799.x
Scheublin TR, van Logtestijn RSP, van der Heijden MGA (2007) Presence and identity of arbuscular mycorrhizal fungi influence competitive interactions between plant species. J Ecol 95:631–638. doi:10.1111/j.1365-2745.2007.01244.x
Schnoor TK, Olsson PA (2010) Effects of soil disturbance on plant diversity of calcareous grasslands. Agr Ecosyst Environ 139:714–719. doi:10.1016/j.agee.2010.10.018
Schnoor TK, Lekberg Y, Rosendahl S, Olsson PA (2011) Mechanical soil disturbance as a determinant of arbuscular mycorrhizal fungal communities in semi-natural grassland. Mycorrhiza 21:211–220. doi:10.1007/s00572-010-0325-3
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding Consortium (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A 109:6241–6246. doi:10.1073/pnas.1117018109
Smilauer P (2001) Communities of arbuscular mycorrhizal fungi in grassland: seasonal variability and effects of environment and host plants. Folia Geobot 36:243–263. doi:10.1007/BF02803179
Stockinger H, Walker C, Schüßler A (2009) ‘Glomus intraradices DAOM197198’, a model fungus in arbuscular mycorrhiza research, is not Glomus intraradices. New Phytol 183:1176–1187, http://www.jstor.org/stable/40302144
Stockinger H, Krüger M, Schüßler A (2010) DNA barcoding of arbuscular mycorrhizal fungi. New Phytol 187:461–474. doi:10.1111/j.1469-8137.2010.03262.x
Stockinger H, Peyret-Guzzon M, Bouffaud ML, Koegel S, Redecker D (2014) The largest subunit of RNA polymerase II as a new marker gene to study assemblages of arbuscular mycorrhizal fungi in the field. PLoS One 9, e107783. doi:10.1371/journal.pone.0107783
Sýkorová Z, Ineichen K, Wiemken A, Redecker D (2007a) The cultivation bias: different communities of arbuscular mycorrhizal fungi detected in roots from the field, from bait plants transplanted to the field, and from a greenhouse trap experiment. Mycorrhiza 18:1–14. doi:10.1007/s00572-007-0147-0
Sýkorová Z, Wiemken A, Redecker D (2007b) Cooccurring Gentiana verna and Gentiana acaulis and their neighboring plants in two Swiss upper montane meadows harbor distinct arbuscular mycorrhizal fungal communities. Appl Environ Microbiol 73:5426–5434. doi:10.1128/AEM.00987-07
Thian H, Drijber RA, Zhang JL, Li XL (2013) Impact of long-term nitrogen fertilization and rotation with soybean on the diversity and phosphorus metabolism of indigenous arbuscular mycorrhizal fungi within the roots of maize (Zea mays L.). Agric Ecosyst Environ 164:53–61. doi:10.1016/j.agee.2012.09.007
Thonar C, Erb A, Jansa J (2012) Real‐time PCR to quantify composition of arbuscular mycorrhizal fungal communities—marker design, verification, calibration and field validation. Mol Ecol Resour 12:219–232. doi:10.1111/j.1755-0998.2011.03086.x
Toljander JF, Santos-Gonzalez JC, Tehler A, Finlay RD (2008) Community analysis of arbuscular mycorrhizal fungi and bacteria in the maize mycorrhizosphere in a long-term fertilization trial. FEMS Microbiol Ecol 65:323–338. doi:10.1111/j.1574-6941.2008.00512.x
Trouvelot A, Kough JL, Gianinazzi-Pearson V (1986) Mesure du taux de mycorhization VA d’un systeme radiculaire. Recherche de methodes d’estimation ayant une signification fonctionnelle. Physiological and genetical aspects of mycorrhizae. Proceedings of the 1st European Symposium on Mycorrhizae. INRA, Paris, pp 217–221
van der Heijden MGA, Klironomos JN, Ursic M, Moutoglis P, Streitwolf-Engel R, Boller T, Wiemken A, Sanders IR (1998) Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 396:69–72. doi:10.1038/23932
van der Heijden MGA, Wiemken A, Sanders IR (2003) Different arbuscular mycorrhizal fungi alter coexistence and resource distribution between co-occurring plant. New Phytol 157:569–578. doi:10.1046/j.1469-8137.2003.00688.x
van der Heijden MGA, Bardgett RD, van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310. doi:10.1111/j.1461-0248.2007.01139.x
Vandenkoornhuyse P, Ridgway KP, Watson IJ, Fitter AH, Young JPM (2003) Co-existing grass species have distinctive arbuscular mycorrhizal communities. Mol Ecol 12:3085–3095. doi:10.1046/j.1365-294X.2003.01967.x
Verbruggen E, Kiers ET (2010) Evolutionary ecology of mycorrhizal functional diversity in agricultural systems. Evol Appl 3:547–560. doi:10.1111/j.1752-4571.2010.00145.x
Verbruggen E, Roling WFM, Gamper HA, Kowalchuk GA, Verhoef HA, van der Heijden MGA (2010) Positive effects of organic farming on below-ground mutualists: large-scale comparison of mycorrhizal fungal communities in agricultural soils. New Phytol 186:968–979. doi:10.1111/j.1469-8137.2010.03230.x
Vickery JA, Bradbury RB, Henderson IG, Eaton MA, Grice PV (2004) The role of agri-environment schemes and farm management practices in reversing the decline of farmland birds in England. Biol Conserv 119:19–39. doi:10.1016/j.biocon.2003.06.004
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. doi:10.1128/AEM.00062-07
Wilson SD, Tilman D (1993) Plant competition and resource availability in response to disturbance and fertilization. Ecology 74:599–611. doi:10.2307/1939319
Yu L, Nicolaisen M, Larsen J, Ravnskov S (2012) Succession of root-associated fungi in Pisum sativum during a plant growth cycle as examined by 454 pyrosequencing. Plant Soil 358:216–224. doi:10.1007/s11104-012-1188-5
Acknowledgments
The authors are grateful to Marie-Hélène Bernicaut, Philippe Chamois, Annie Colombet, Odile Chatagnier, Bernard Alixant, Eric Bernaud, Diederik van Tuinen, Romain Barnard, Aymé Spor and Nassybat Binti Said Ismaïl for their helpful participation and comments in this study, which was funded by the gratefully acknowledged Burgundy Region (FABER programme no. 2010-9201AAO047S01404).
Conflict of interest
The authors declare that they have no competing interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Schematic representation of the experimental set-up used at the experimental domain of Epoisses at Bretenières, Burgundy region, France. a. Overview of the experimental design: unploughed plots are shown in grey, ploughed plots in white. Fertilisation treatments (+N, 100 kg ha−1 of nitrogen; +P, 30 kg ha−1 of phosphorus; +N+P, combined fertilisation; C, no fertilisation) were randomly arranged within each strip of the ploughing treatment. b. Zoom on a plot showing planting pattern of Plantago lanceolata trap plant, pre-grown in a climate chamber during 1 month. (PDF 322 kb)
Online Resource 2
Sequences used for the RPB1 reference tree. Bold characters refer to new sequences. (PDF 508 kb)
Online Resource 3
Maximum likelihood tree of RPB1 sequences used as reference tree to assign operational taxonomic units to molecular taxa (MTs). Branch support values are based on 1000 bootstrap replicates using RAxML software. Mortierella verticillata and Mucor hiemalis were chosen as outgroups to root the tree. (PDF 333 kb)
Online Resource 4
Accumulation curve analysis of observed OTU numbers versus cumulative sequence number by sample (different colours) before (a) and after (b) subsampling to a 1400 sequence threshold (represented by the dotted line). (PDF 569 kb)
Online Resource 5
Number of RPB1 sequence reads retained per sample after sub-sampling to a threshold of 1400 sequence reads at the species-level molecular taxa (MT). Numbers of OTUs found for each MT are in brackets. (PDF 492 kb)
Online Resource 6
Data table of Plantago lanceolata fresh shoot weight, dry root weight and root colonisation parameters (estimation by Trouvelot et al. 1986) and estimation of plant species richness found in each plot (other than P. lanceolata) according to soil treatment (physical soil disturbance and fertilisation treatments). (PDF 169 kb)
Online Resource 7
Statistical significance (P value) of indicator species analysis based on species-level molecular taxa (MT) after sub-sampling to a threshold of 1400 sequences. Only species with more than 100 sequences over the total read number and having both sensitivity and specificity values higher than 0.75 were kept. (PDF 261 kb)
Online Resource 8
Variance of species abundance reads using a nested linear mixed model showing effects of nitrogen and phosphorus fertilisation within the physical soil disturbance treatments randomised by block. Significant results (P < 0.05) are in bold type. Red dollar symbols show indicator species identified by the De Caceres-Jansen index using 2000 permutations. (PDF 297 kb)
Online Resource 9
Restriction fragment length polymorphism patterns (=RFLP types) of the mitochondrial rRNA subunit gene region from root-colonising Rhizophagus irregularis, obtained using primers RNL-29/ RNL-30. Numbers of clones of each RFLP type are listed per treatment. (PDF 455 kb)
Rights and permissions
About this article
Cite this article
Peyret-Guzzon, M., Stockinger, H., Bouffaud, ML. et al. Arbuscular mycorrhizal fungal communities and Rhizophagus irregularis populations shift in response to short-term ploughing and fertilisation in a buffer strip. Mycorrhiza 26, 33–46 (2016). https://doi.org/10.1007/s00572-015-0644-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-015-0644-5