Abstract
Background
Recent evidence suggests that the presence of microbiome within human pancreatic ductal adenocarcinoma (PDAC) tissue potentially influences cancer progression and prognosis. However, the significance of tumor-resident microbiome remains unclear. We aimed to elucidate the impact of intratumoral bacteria on the pathophysiology and prognosis of human PDAC.
Methods
The presence of intratumoral bacteria was assessed in 162 surgically resected PDACs using quantitative polymerase chain reaction (qPCR) and in situ hybridization (ISH) targeting 16S rRNA. The intratumoral microbiome was explored by 16S metagenome sequencing using DNA extracted from formalin-fixed paraffin-embedded tissues. The profile of intratumoral bacteria was compared with clinical information, pathological findings including tumor-infiltrating T cells, tumor-associated macrophage, fibrosis, and alterations in four main driver genes (KRAS, TP53, CDKN2A/p16, SMAD4) in tumor genomes.
Results
The presence of intratumoral bacteria was confirmed in 52 tumors (32%) using both qPCR and ISH. The 16S metagenome sequencing revealed characteristic bacterial profiles within these tumors, including phyla such as Proteobacteria and Firmicutes. Comparison of bacterial profiles between cases with good and poor prognosis revealed a significant positive correlation between a shorter survival time and the presence of anaerobic bacteria such as Bacteroides, Lactobacillus, and Peptoniphilus. The abundance of these bacteria was correlated with a decrease in the number of tumor-infiltrating T cells positive for CD4, CD8, and CD45RO.
Conclusions
Intratumoral infection of anaerobic bacteria such as Bacteroides, Lactobacillus, and Peptoniphilus is correlated with the suppressed anti-PDAC immunity and poor prognosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is one of the most aggressive and lethal human cancers, with a 5-year survival rate of less than 10% [1]. PDAC progression and metastasis are not solely determined by genetic alterations in cancer genomes but also by the complex interplay between cancer cells and tumor microenvironment (TME), including tumor-infiltrating lymphocytes (TILs) and dense stromal fibrosis [2, 3]. TILs, particularly tumor-infiltrating CD8+ T cells, critically impact on the prognosis of PDAC as immune effectors capable of eliminating cancer cells [4]. Tumor stroma, densely populated with cancer-associated fibroblasts (CAFs) such as ACTA2 (α-SMA)+ myofibroblasts and collagen fibers, has also been reported to promote resistance to chemotherapy and pancreatic tumor progression by inducing hypoxic TME [5,6,7]. However, the factors that influence the formation of such pancreatic TME have been poorly understood.
Although the pancreas was previously supposed to be sterile, several recent studies have reported the presence of bacteria in the pancreatic tissues. Orally administered bacteria were reportedly detected in the mouse pancreas within 30 min [8], and correlation of the microbiome in the human duodenum and pancreas suggests that bacteria may directly translocate from the duodenum into the pancreas [9]. Furthermore, recent studies have confirmed the presence of bacteria within PDAC and its potential influence on tumor growth [10]. A mouse study demonstrated that microbiota in PDAC promotes oncogenesis by inducing innate and adaptive immune suppression. Corresponding relationship between microbiome and progressive PDAC suggested the oncogenic impact of intratumoral bacteria on PDAC development both in mice and humans [8]. However, another study reported that PDAC with more bacterial diversity showed enhanced immune infiltration and favorable prognosis compared with PDAC with less diverse bacteria [11]. Although tumor-resident bacteria in PDAC have been suggested to affect anti-tumor immunity and tumor prognosis, details remain controversial and unclear. In this study, we investigated bacteria within PDAC tissues that were surgically resected and aimed to elucidate the impact of intratumoral bacteria on TME and post-operative prognosis.
Materials and methods
Sample and data collection
We analyzed formalin-fixed paraffin-embedded (FFPE) sections from 162 PDAC patients who underwent surgery between April 2008 and March 2017 at the Kobe University Hospital. We retrospectively reviewed the patients’ medical information regarding age, gender, body mass index (BMI), carcinoembryonic antigen (CEA, reference range: < 5 ng/mL), and carbohydrate antigen 19-9 (CA19-9, reference range: < 37 ng/mL), white blood cell count, neutrophil count, total leukocytes count, alcohol consumption, current smoking, diabetes mellitus and administration of antibiotics. The tumor characteristics included tumor size and location, pathological stage (Union for International Cancer Control UICC] 8th classification), histological grade, residual tumor status after surgery, and the history of neoadjuvant and adjuvant chemotherapy. Laboratory data were collected within 1 month before surgery.
The study protocol was reviewed and approved by the ethics committee of the Kobe University Hospital (No.180235). Informed consent was waived due to the retrospective study design and the study information was disclosed on our hospital website, allowing eligible patients to opt out. This study was conducted in accordance with the Declaration of Helsinki. All authors had access to the study data, and reviewed and approved the final manuscript.
DNA extracting and real-time PCR
DNA was extracted from the pancreatic tumoral and adjacent non-tumoral FFPE tissues using the QIAamp DNA FFPE Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s protocol. The extracted DNA was quantified using a fluorometer (Invitrogen Qubit 4.0; Thermo Fisher Scientific, Waltham, MA, USA) and stored at −80 ℃ until further analysis.
Quantitative polymerase chain reaction (qPCR) was performed on Applied Biosystems 7500 real-time PCR system (Applied Biosystems Inc, CA, USA) using SYBR green qPCR assay (Applied Biosystems Inc, CA, USA). The V1–2 regions of 16S rRNA gene were amplified using forward primer 27Fmod (5′-AGRGTTTGATYMTGGCTCAG-3′), and reverse primer 338R (5′-TGCTGCCTCCCGTAGGAGT-3′) [12]. The cycling conditions were 1 cycle at 95 ℃ for 10 min to denature DNA, with amplification proceeding for 40 cycles at 95 ℃ for 15 s, 50 ℃ for 20 s, and 72 ℃ for 1 min, followed by a standard denaturation curve protocol.
In situ hybridization (ISH)
Chromogenic RNA in situ hybridization (ISH) targeting 16S rRNA was performed using RNAscope® 2.5 HD Reagent Kit-RED (Advanced Cell Diagnostics, Hayward, CA, USA) and Fast Red according to the manufacturer’s protocol. The probe used was RNAscope® Probe-EB-16S-rRNA (Cat #464461). The chromogenic reaction within the pancreatic tumor indicated a positive result.
Amplicon sequencing and microbiome analysis
The 16S rDNA V1–2 regions were amplified by PCR and sequenced in the MiSeq platform (Illumina) with MiSeq Reagent kit v2 (500 cycles) using the 250 bp paired-end protocol. The QIIME2 (version 2020.11) pipeline was used to perform microbiome analysis [13]. Demultiplexing and quality filtering were performed on the raw sequence data using the q2-demux plug-in, and amplicon sequence variants (ASVs) were counted after denoising by DADA2 [14].
Taxonomy was assigned to ASVs using reference sequences from Silva (138 SSURef NR99 full-length taxonomy). Bacterial contamination was distinguished using R program package “Decontam” with the parameter “method = frequency” by comparing the data from pancreatic tissues with that derived from 15 samples of FFPE pieces without tissues [15]. Alpha diversity analysis with Shannon index was calculated using QIIME2. Beta diversity analysis using weighted-UniFrac Principal Coordinate Analysis (PCoA) and permutation analysis of variance (PERMANOVA) were also performed using Qiime2. The taxonomic types of intratumoral microbiome to distinguish tumor prognosis were analyzed by linear discriminant analysis (LDA) effect size (LEfSe) calculations using Galaxy Version 1.0 [16].
Immunohistochemistry (IHC) and Elastica van Gieson (EVG) staining
FFPE tissues were sectioned at 5 mm thickness and analyzed by IHC and EVG staining. The following antibodies were used: anti-TP53 (Santa Cruz Biotechnology, Dallas, TX, USA, catalog number: sc-47698), anti-CDKN2A/p16 (Roche Diagnostics, Cat #6695221001), anti-SMAD4 (Santa Cruz Biotechnology, Cat #sc-7966), anti-CD4 (Leica Biosystems, Wetzlar, Germany, Cat #CD4-368-L-CE), anti-CD8 (Roche Diagnostics, Cat #5493846001), anti-FOXP3 (Abcam, Cambridge, UK, Cat #ab20034), anti-CD45RO (BioGenex Laboratories, San Ramon, CA, USA,Cat #AM113-5M), anti-CD68 (ProteinTech Illinois, USA, Cat #66231-2-Ig), anti-CD206 (ProteinTech Illinois, USA, Cat #60143-1-Ig) and anti-α-SMA (Santa Cruz Biotechnology, catalog number: sc-53142). EVG staining was performed using an Elastic Stain Kit (Abcam, Cat #ab150667) according to the manufacturer’s protocol.
Evaluation of tumor-infiltrating lymphocytes (TILs), macrophage, and tumor fibrosis
TILs positive for CD4, CD8, FOXP3, CD45RO and macrophage for CD68, CD206 were assessed by immunohistochemical staining on slides with the maximum divided surface of tumors. Each subset of TILs and macrophage were counted at 200 × magnification (counts/mm2) using Image J (Java image processing program inspired by National Institute of Health (NIH), USA). Three fields separated by at least 5 mm each were counted and the mean value was calculated for each case. The cases were classified as high density or low density based on the median value.
To assess fibrosis within pancreatic cancer, tumor stromal collagen and myofibroblasts were evaluated by EVG staining and immunostaining for α-SMA, respectively. The stained sections were digitally scanned and analyzed using Adobe Photoshop CC2019 software (Adobe Inc., San Jose, CA). The red area in EVG-stained sections and the brown area in α-SMA-stained sections were quantified as tumor stromal collagen and αSMA+ myofibroblasts, respectively. Each area was divided by the whole tumor area analyzed and defined as the area proportion of “tumor stromal collagen” and “α-SMA+ fibroblast,” respectively. Additionally, all cases were classified into two groups (high and low) based on the median value of the area proportion. Representative IHC images for TILs, CD68, CD206, α-SMA, and EVG staining image are shown in Supplementary Fig. 1.
Evaluation of driver gene alterations
Alterations of KRAS, TP53, CDKN2A/p16, and SMAD4 genes in the tumor were determined by next-generation sequencing (NGS) analysis, droplet digital PCR (ddPCR) and IHC using DNA extracted from FFPE as reported previously [17]. In brief, KRAS mutations were determined by NGS. TP53 mutations were determined based on a combination of NGS, ddPCR, and IHC. CDKN2A/p16 and SMAD4 mutations were determined using IHC. IHC sections were evaluated by two experienced pathologists (M.K. and T.I.) who were unaware of the clinical data. TP53, CDKN2A/p16, and SMAD4 were evaluated with Kappa values of 0.982 (P < 0.0001), 0.964 (P < 0.0001), 0.942 (P < 0.0001), respectively, and the agreement was high between the pathologists.
Statistical analysis
SPSS (SPSS Inc., Chicago, IL, USA) and GraphPad Prism 8 (GraphPad Software, La Jolla, CA, USA) were used for statistical analyses. The chi-square test or Fisher’s exact test, when applicable, was used to compare frequencies, and the Wilcoxon rank-sum test was used to compare skewed continuous variables. Overall survival (OS) was estimated using the Kaplan–Meier method and compared using a log-rank test. Hazard ratios (HRs) and the corresponding 95% confidence intervals (CIs) were estimated using Cox proportional-hazards models. The multivariate analyses included factors with statistical significance in univariate analysis. All statistical tests were two-tailed, and statistical significance was set at P < 0.05.
Result
Identification of bacteria within PDAC
To identify tumors that contained intratumoral bacteria (Fig. 1A), we initially screened 162 human PDAC samples by qPCR targeting bacterial 16S rRNA gene, which detected positive amplification in 107 samples. These samples were further evaluated by ISH for 16S rRNA (Fig. 1B), and we confirmed 52 PDACs (32.1% among 162 tumors) that contained tumor-resident bacteria both on qPCR and ISH. Clinical information of the 52 cases with PDACs definitely positive for intratumoral bacteria were shown in Table 1.
Study outline. A Flow chart to determine the presence of bacteria in human PDAC tissues. Of the 162 cases, 55 cases in which 16SrRNA could not be extracted by qPCR were excluded. In addition, of the remaining 107 cases, 55 cases in which 16SrRNA could not be identified in the tumor by ISH were excluded. Finally, 52 cases were included in this study. B Representative images of in situ hybridization targeting bacterial 16S rRNA gene. Red arrow heads indicate positive signals. Scale bars, 50 μm. PDAC pancreatic ductal adenocarcinoma, PCR polymerase chain reaction, ISH in situ hybridization
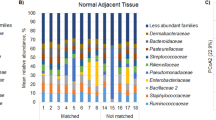
To characterize the microbiome within PDAC, the 52 bacteria-positive PDACs were analyzed by 16S metagenome sequencing. As their non-tumoral counterparts, 26 non-tumoral tissues that were sufficiently distant from the tumor and positive for 16S rRNA amplification in qPCR were collected among the 52 cases with bacteria-positive PDAC and examined in parallel. To carefully exclude reads resulting from inevitable contamination of environmental bacteria, we assessed the sequencing data derived from FFPE tissues in comparison with those from non-tissue control pieces (see “Materials and methods”) using the R-package “Decontam.” The composition at the genus level before and after the use of Decontam are shown in Supplementary Fig. 2A and Fig. 2A, respectively. After the process excluding contamination with Decontam, many taxonomic levels of bacteria (1315 species, 723 genera, 306 families, 170 orders, 74 steels, 38 phylum) were identified in the tumor and non-tumoral tissues. At the phylum level (Supplementary Fig. 2B), Proteobacteria and Firmicutes were highly abundant in PDAC tissues, which was consistent with previous reports [10]. At the genus level, the genera Pseudomonas, Curvibacter, Streptococcus, Sphingomonas, and Corynebacterium were abundantly detected in PDACs (Fig. 2A). Subsequently, to investigate changes in the microbiome composition within PDAC due to administration of antibiotics, a diversity analysis was performed between 11 cases where antibiotics were used within 1 month before surgery and 41 cases where they were not used. There were no significant differences in alpha diversity measures by Shannon Index (P = 0.42) and beta diversity analyzed by weighted-UniFrac PcoA (P = 0.25, using PERMANOVA) in the two groups (Supplementary Fig. 3). These results indicate that the administration of antibiotics did not significantly alter the composition of the microbiome within PDAC in this study.
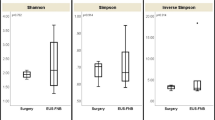
Microbiome composition in PDAC and non-tumoral tissues. A Taxonomic profiles of predominant bacterial genera by mean relative abundance (%) in PDAC tissue and non-tumoral tissues. B Comparative analysis of the alpha diversity of the microbiome communities between PDAC and non-tumoral tissues using Shannon index. C Principal coordinate analysis (PCoA) using weighted-UniFrac distance of beta diversity among PDAC and non-tumoral tissues. PDAC pancreatic ductal adenocarcinoma
Alpha diversity measures by Shannon Index were significantly lower (P < 0.01) in PDAC than non-tumoral tissues (Fig. 2B). Beta diversity analyzed by weighted-UniFrac PcoA was also significantly different between PDAC and non-tumoral tissues (P < 0.01, using PERMANOVA) (Fig. 2C). These findings suggested that microbiome within PDAC tissues were unique and different from those in non-tumoral tissues. At least approximately one third of human PDACs were suggested to have tumor-resident bacteria, and our metagenome sequencing successfully detected the characteristics of microbiome within PDAC while eliminating the effects of contamination.
Identification of PDAC microbiome associated with prognosis
To identify the intratumoral microbiome involved in prognosis, we first assessed the survival of the 52 patients with bacteria-infected PDAC (Fig. 3A). The median follow-up period of the cohort was 23.6 months, and a majority of the patients (71.2%, 37/52) did not survive during the follow-up period. The 52 patients were effectively divided into two groups based on the median follow-up period: “short-term survival group” and “long-term survival group” (Fig. 3B). All patients in the short-term survival group died within the observation period, and the overall survival between the two groups showcased significant difference (P < 0.001). A comparison of the patients’ backgrounds of the two groups showed significant differences in pathological stage and residual tumor status (Supplementary Table 1).
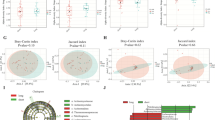
Identification of intratumoral bacteria associated with prognosis in pancreatic cancer patients. A Kaplan–Meier curve of overall survival of patients included in this study. B Overall survival of patients with PDAC in short-term survival group and long-term survival group. The patients were divided into two groups based on the median follow-up period (23.6 months). The survival curves were compared between them by the log-rank test. C LEfSe calculations between the short-term survival group and the long-term survival group were performed using a threshold of 2.0. D Forest plots showing the hazard ratio of prognosis by the presence of each genus detected by LEfSe. PDAC pancreatic ductal adenocarcinoma, LEfSe linear discriminant analysis effect size
Subsequently, we compared the tumor-resident microbiome between the two groups. Alpha diversity tended to be lower in the long-term survival group, but was not significantly different (P = 0.06, using Shannon index). There was no significant difference in beta diversity between the two groups (P = 0.58, using permutational multivariate analysis of variance, Supplementary Fig. 4). LEfSe analysis was performed to explore the differences in the predominance of bacterial communities between the two groups, which demonstrated that one genus and ten genera were dominant in the long-term survival group and the short-term survival group, respectively (Fig. 3C). The presence of any of these genera identified other than Staphylococcus was associated with an increased HR for the prognosis of PDAC in the univariate analysis (Supplementary Table 2). Further multivariate analysis of the ten genera that were significantly different in the univariate analysis revealed that the presence of Bacteroides, Lactobacillus, and Peptoniphilus were significantly associated with poor prognosis (Fig. 3D).
Association of intratumoral bacteria with prognostic factors in PDAC
Notably, the three genera (i.e., Bacteroides, Lactobacillus, and Peptoniphilus) that were significantly associated with poor prognosis in multivariate analysis (hereafter referred to as “prognostic bacteria”) were all anaerobes. The abundance among these bacteria were shown in Fig. 4A. A low (r = 0.28) but significant positive correlation was found between the abundance of Bacteroides and Lactobacillus (P = 0.048). While there was no significant correlation between Peptoniphilus and Bacteroides, as well as Peptoniphilus and Lactobacillus, there was a positive trend in the abundance of both. Thus, we hypothesized that a hypoxic environment conducive to the growth of these prognostic bacteria may influence the progression of PDAC. To test this hypothesis, we compared clinical and tumor characteristics between 24 cases positive for at least one of the three prognostic bacteria and 28 cases negative for all of them (Table 2). There were no significant differences in clinical characteristics including the tumor size and pathological stage between the two groups. Features in tumor genomes such as alterations in KRAS, TP53, CDKN2A/p16, and SMAD4 genes were also comparable between the two groups. Quantification of the area of tumor stromal collagen and αSMA+ fibroblasts showed no significant difference in intra-tumor fibrosis as well. In marked contrast, the degree of tumor-infiltrating T cells positive for CD4, CD8, and CD45RO were significantly lower in the tumors with prognostic bacteria than their counterparts (P = 0.005, P = 0.03, and P = 0.005, respectively). We also evaluated tumor-associated macrophage using CD68 and CD206 as markers of M1-like and M2-like macrophages, respectively, and found that the abundance of intra-tumor immunosuppressive M2 macrophages tended to be higher in the group with prognostic bacteria than their counterpart although the difference was not significant.
The comparison of overall survival between the groups with and without prognostic bacteria confirmed that the survival of the group with prognostic bacteria was significantly poorer than that without prognostic bacteria (P < 0.001, Fig. 4B). Multivariate COX hazard analysis showed that the presence of prognostic bacteria was an independent prognostic factor (P < 0.001, HR = 3.48, 95%CI 1.65–7.33, adjusted for clinical characteristics as follow: prognostic bacteria, pathological stage and residual tumor status). These findings suggested that the presence of the prognostic bacteria was associated with suppressed anti-tumor immunity leading to poor prognosis.
An in-depth investigation of the correlation between the presence of the prognostic bacteria and TILs was performed. The significant negative correlations were detected between the abundance of the prognostic bacteria and the number of CD4+, CD8+, CD45RO+ T cells in PDAC (P = 0.006, P < 0.001, P < 0.001, respectively, Fig. 5). Such significant negative correlations between the prognostic bacteria and tumor-suppressive TILs were also observed when each genus was examined (Supplementary Fig. 5); CD45 RO+ T cells in Bacteroides (P < 0.001), CD8+ and CD45RO+ T cells in Lactobacillus (P = 0.009, P = 0.007, respectively), and CD4+ and CD8+ cells in Peptoniphilus (P = 0.004, P = 0.017, respectively). On the contrary, there was no significant correlation between the amount of immune-suppressive FOXP3+ T cells infiltrating in tumors and the abundance of the prognostic bacteria (Fig. 5). These findings support the fact that TME with the three prognostic bacteria is associated with the suppression of anti-tumor immunity and poor prognosis.
Discussion
In this study, we observed a correlation between the presence of three anaerobic bacteria, Bacteroides, Lactobacillus, and Peptoniphilus, within PDAC with a suppressed anti-tumor immunity, and poor prognosis. In contrast, there was no significant correlation between these bacteria and intratumoral fibrosis or alterations in major driver genes in tumor genomes.
In recent years, the presence of microbiome in PDAC has been demonstrated by several studies [18]. We showed that bacteria within PDAC exhibited a distinct profile from that within non-tumoral pancreatic tissues and were predominantly composed of the phyla Proteobacteria and Firmicutes, which was consistent with previous reports [10, 19]. Although it has been controversial whether the bacterial profile within PDAC is different from that in non-tumoral pancreatic tissues [8, 9, 20], comparative analysis using samples that exhibited the evidence of bacterial presence in the present study demonstrated the colonization of distinct bacterial species within PDAC that were different from those in the surrounding tissue. The unique TME of PDAC such as extensive desmoplasia leading to hypoxia may allow the colonization of a characteristic microbiome that is distinct from the surrounding tissue environment [21]. A microbiota analysis with a larger cohort would be necessary to draw definitive conclusions.
Intratumoral infiltration of immune cells plays a crucial role in regulating tumor progression [22,23,24]. We previously demonstrated a positive correlation between increased infiltration of CD4+, CD8+, and CD45RO+ T cells within the PDAC and favorable prognosis of PDAC patients [17]. Importantly, several studies have reported that the gut and intra-tumor microbiome are involved in the regulation of intratumoral immune ecosystem in various malignancies including PDAC [8, 11, 25,26,27,28], although, the precise role of the microbiome in PDAC remains largely unclear. In the present study, we demonstrated that the increased abundance of three anaerobic bacteria including Bacteroides, Lactobacillus, and Peptoniphilus within PDAC was significantly correlated with the suppressed intratumoral infiltration of effector T cells as well as a poor prognosis of PDAC. PDAC is well-known to create a hypoxic microenvironment due to its limited cellularity and compressed, desmoplastic stroma [29, 30]. Hypoxic conditions in PDAC reportedly enhanced the intracellular survival of an anaerobic bacteria, Porphyromonas gingivalis [31]. Notably, a recent study showed that the hypoxic environment was enhanced by intestinal bacteria vice versa and modulated tissue-resident lymphocytes in mice [32]. Moreover, some anaerobic bacteria such as Fusobacterium nucleatum are associated with immune suppression [33]. Thus, tumor stroma as well as intratumoral colonization of bacteria in itself, may establish the hypoxic microenvironment that allows the growth of anaerobic bacteria in a subset of PDAC, which might lead to poor prognosis via immune suppression. It also remains unclear whether the intra-tumor bacteria themselves are directly involved in suppressing the intra-tumor immune system. Tryptophan metabolites derived from Lactobacillus have recently been reported to induce immunosuppression by affecting intra-tumor macrophages [34]. In line with this, we found that intra-tumor immunosuppressive M2 macrophages tended to be higher in the prognostic bacteria group. Moreover, some studies have also reported direct involvement of certain bacteria in tumor immunosuppression. F. nucleatum within PDAC was demonstrated to promote pancreatic cancer progression through autocrine and paracrine mechanisms of the CXCL1–CXCR2 axis [35]. Intratumoral P. gingivalis was also reported to promote PDAC progression via elevating the secretion of neutrophilic chemokines and neutrophil elastase [36]. Further studies are warranted to investigate whether certain specific anaerobic bacteria are involved in inducing immune suppression through some mechanism or whether anaerobic bacteria just tend to colonize in PDAC with poor prognosis.
One of the major challenges in microbiome analysis using human pancreatic tissues is the inevitable presence of contamination, as it is difficult to aseptically extract bacteria from human pancreatic samples. By evaluating the effect of environmental contamination with data derived from non-tissue FFPE pieces and by analyzing only bacteria for which possibilities of contamination were statistically excluded [15], we addressed this issue and obtained unique bacterial profiles in tumoral and non-tumoral pancreatic tissues. Additionally, some clinical events such as preoperative endoscopic procedures might have affected the intra-PDAC microbiome in the present study. Indeed, endoscopic ultrasound-guided fine needle biopsy or endoscopic retrograde cholangiopancreatography was performed for diagnosis and drainage purpose in all cases analyzed. If endoscopic procedures could enhance the colonization of bacteria in PDAC tissues and intratumoral presence of anaerobic bacteria might lead to the suppression of anti-PDAC immunity, prophylaxis against such anaerobic bacteria during endoscopic procedures could be important to improve the treatment efficiency against PDAC.
In conclusion, our findings demonstrate that the presence of anaerobic genera such as Bacteroides, Lactobacillus, and Peptoniphilus within PDAC potentially have prognostic relevance. These genera might be implicated in immune-mediated prognostic deterioration of tumors. This study emphasizes the significance of anaerobic bacteria colonization in PDAC in the clinical management of PDAC, although further comprehensive investigations are necessary to fully understand its implications.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19:1423–37.
Yao W, Maitra A, Ying H. Recent insights into the biology of pancreatic cancer. EBioMedicine. 2020;53:102655.
Foucher ED, Ghigo C, Chouaib S, et al. Pancreatic ductal adenocarcinoma: a strong imbalance of good and bad immunological cops in the tumor microenvironment. Front Immunol. 2018;9:1044.
Neesse A, Algül H, Tuveson DA, et al. Stromal biology and therapy in pancreatic cancer: a changing paradigm. Gut. 2015;64:1476–84.
Chen Y, Kim J, Yang S, et al. Type I collagen deletion in αSMA(+) myofibroblasts augments immune suppression and accelerates progression of pancreatic cancer. Cancer Cell. 2021;39:548–65.e6.
Yu M, Tannock IF. Targeting tumor architecture to favor drug penetration: a new weapon to combat chemoresistance in pancreatic cancer? Cancer Cell. 2012;21:327–9.
Pushalkar S, Hundeyin M, Daley D, et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 2018;8:403–16.
Del Castillo E, Meier R, Chung M, et al. The microbiomes of pancreatic and duodenum tissue overlap and are highly subject specific but differ between pancreatic cancer and noncancer subjects. Cancer Epidemiol Biomarkers Prev. 2019;28:370–83.
Geller LT, Barzily-Rokni M, Danino T, et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357:1156–60.
Riquelme E, Zhang Y, Zhang L, et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell. 2019;178:795–806.e12.
Kim SW, Suda W, Kim S, et al. Robustness of gut microbiota of healthy adults in response to probiotic intervention revealed by high-throughput pyrosequencing. DNA Res. 2013;20:241–53.
Bolyen E, Rideout JR, Dillon MR, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
Callahan BJ, McMurdie PJ, Rosen MJ, et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Davis NM, Proctor DM, Holmes SP, et al. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome. 2018;6:226.
Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12:R60.
Tanaka T, Masuda A, Inoue J, et al. Integrated analysis of tertiary lymphoid structures in relation to tumor-infiltrating lymphocytes and patient survival in pancreatic ductal adenocarcinoma. J Gastroenterol. 2023;58:277–91.
Chen Z, Zhang S, Dong S, et al. Association of the microbiota and pancreatic cancer: opportunities and limitations. Front Immunol. 2022;13: 844401.
Nejman D, Livyatan I, Fuks G, et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science. 2020;368:973–80.
Thomas RM, Gharaibeh RZ, Gauthier J, et al. Intestinal microbiota enhances pancreatic carcinogenesis in preclinical models. Carcinogenesis. 2018;39:1068–78.
Lafaro KJ, Melstrom LG. The paradoxical web of pancreatic cancer tumor microenvironment. Am J Pathol. 2019;189:44–57.
Zhang Y, Zoltan M, Riquelme E, et al. Immune cell production of interleukin 17 induces stem cell features of pancreatic intraepithelial neoplasia cells. Gastroenterology. 2018;155:210–23.e3.
Das S, Shapiro B, Vucic EA, et al. Tumor cell-derived Il1β promotes desmoplasia and immune suppression in pancreatic cancer. Cancer Res. 2020;80:1088–101.
Balachandran VP, Łuksza M, Zhao JN, et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature. 2017;551:512–6.
Gopalakrishnan V, Spencer CN, Nezi L, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359:97–103.
Sivan A, Corrales L, Hubert N, et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350:1084–9.
Ma C, Han M, Heinrich B, et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science. 2018;360:eaan5931.
Sethi V, Kurtom S, Tarique M, et al. Gut microbiota promotes tumor growth in mice by modulating immune response. Gastroenterology. 2018;155:33–7.e6.
Ye LY, Zhang Q, Bai XL, et al. Hypoxia-inducible factor 1alpha expression and its clinical significance in pancreatic cancer: a meta-analysis. Pancreatology. 2014;14:391–7.
Tan Z, Xu J, Zhang B, et al. Hypoxia: a barricade to conquer the pancreatic cancer. Cell Mol Life Sci. 2020;77:3077–83.
Gnanasekaran J, Binder Gallimidi A, Saba E, et al. Intracellular Porphyromonas gingivalis promotes the tumorigenic behavior of pancreatic carcinoma cells. Cancers (Basel). 2020;12:2331.
Harada Y, Sujino T, Miyamoto K, et al. Intracellular metabolic adaptation of intraepithelial CD4+CD8αα+ T lymphocytes. iScience. 2022;25:104021.
Mima K, Sukawa Y, Nishihara R, et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 2015;1:653–61.
Hezaveh K, Shinde RS, Klötgen A, et al. Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity. 2022;55:324–40.e8.
Hayashi M, Ikenaga N, Nakata K, et al. Intratumor Fusobacterium nucleatum promotes the progression of pancreatic cancer via the CXCL1-CXCR2 axis. Cancer Sci. 2023;114:3666–78.
Tan Q, Ma X, Yang B, et al. Periodontitis pathogen Porphyromonas gingivalis promotes pancreatic tumorigenesis via neutrophil elastase from tumor-associated neutrophils. Gut Microbes. 2022;14:2073785.
Acknowledgements
The authors would like to thank Nobuyuki Kakiuchi (Department of Pathology and Tumor Biology, Kyoto University) and Yohei Masugi (Department of Pathology, Keio University School of Medicine) for their valuable comments. We would like to thank Editage (www.editage.jp) for the English language editing.
Funding
Open Access funding provided by Kobe University. This work was supported by JSPS KAKENHI (Grants-in-Aid for Scientific Research, 19K08444: A.M., 19H03698,22H03058: Y.K.), Pancreas Research Foundation of Japan (A.M.), Japan Science and Technology Agency (JST)-Moonshot R&D under grant number JPMJMS2022.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
535_2023_2069_MOESM1_ESM.tif
Supplementary file1 Supplementary Fig. 1 Representative images of immunohistochemistry and Elastica van Gieson staining showing high or low levels of CD4+, CD8+, FOXP3+, CD45RO+, CD68, CD206 and tumor stromal collagen. Scale bars, 50 μm. TILs tumor-infiltrating lymphocytes (TIF 89838 KB)
535_2023_2069_MOESM2_ESM.tif
Supplementary file2 Supplementary Fig. 2 Microbiome composition in PDAC, non-tumoral tissues and non-tissue FFPE pieces. A Taxonomic profiles of predominant bacterial genera by mean relative abundance (%) in PDAC tissues, non-tumoral tissues, and non-tissue FFPE pieces before using Decontam. B Taxonomic profiles of predominant bacterial phylum by mean relative abundance (%) in PDAC and non-tumoral tissues after filtering with Decontam (TIF 14528 KB)
535_2023_2069_MOESM3_ESM.tif
Supplementary file3 Supplementary Fig. 3 Diversity analysis of the group of administration of antibiotics and not administration of antibiotics. A Comparative analysis of the alpha diversity of the microbiome communities between the group of administration of antibiotics and no administration of antibiotics using Shannon index. B Principal coordinate analysis (PCoA) using weighted-UniFrac distance of beta diversity between the group of administration of antibiotics and no administration of antibiotics. PDAC pancreatic ductal adenocarcinoma (TIF 4608 KB)
535_2023_2069_MOESM4_ESM.tif
Supplementary file4 Supplementary Fig. 4 Diversity analysis of long-term survival group and short-term survival group. A Comparative analysis of the alpha diversity of the microbiome communities between long-term and short-term survival groups using Shannon index. B Principal coordinate analysis (PCoA) using weighted-UniFrac distance of beta diversity between long-term and short-term survival groups. PDAC pancreatic ductal adenocarcinoma (TIF 4515 KB)
535_2023_2069_MOESM5_ESM.tif
Supplementary file5 Supplementary Fig. 5 Correlation between TME findings and the abundance of Bacteroides, Lactobacillus, and Peptoniphilus in tumors. P values and correlation coefficient (r) values were analyzed by Spearman correlation. TME tumor microenvironment, TILs tumor-infiltrating lymphocytes, α-SMA alpha-smooth muscle actine (TIF 6828 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abe, S., Masuda, A., Matsumoto, T. et al. Impact of intratumoral microbiome on tumor immunity and prognosis in human pancreatic ductal adenocarcinoma. J Gastroenterol 59, 250–262 (2024). https://doi.org/10.1007/s00535-023-02069-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-023-02069-5