Abstract
Purpose
Mounting evidence suggests that the gut microbiome influences radiotherapy efficacy and toxicity by modulating immune signalling. However, its contribution to radiotherapy outcomes in head and neck cancer (HNC) is yet to be investigated. This study, therefore, aimed to uncover associations between an individual’s pre-therapy gut microbiota and (i) severity of radiotherapy-induced oral mucositis (OM), and (ii) recurrence risk in patients with HNC.
Methods
In this prospective pilot study, 20 patients with HNC scheduled to receive radiotherapy or chemoradiotherapy were recruited. Stool samples were collected before treatment and microbial composition was analysed using 16S rRNA gene sequencing. OM severity was assessed using the NCI-CTCAE scoring system. Patients were also followed for 12 months of treatment completion to assess tumour recurrence.
Results
Overall, 80% of the patients were male with a median age of 65.5 years. Fifty-three percent experienced mild/moderate OM while 47% developed severe OM. Furthermore, 18% experienced tumour relapse within 1 year of treatment completion. A pre-treatment microbiota enriched of Eubacterium, Victivallis, and Ruminococcus was associated with severe OM. Conversely, a higher relative abundance of immunomodulatory microbes Faecalibacterium, Prevotella, and Phascolarctobacterium was associated with a lower risk of tumour recurrence.
Conclusion
Our results indicate that a patient’s gut microbiota composition at the start of treatment is linked to OM severity and recurrence risk. We now seek to validate these findings to determine their ability to predict treatment outcomes in HNC, with the goal of using this data to inform second-generation microbial therapeutics to optimise treatment outcomes for patients with HNC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Head and neck cancer (HNC) is the sixth most common type of cancer with ~930,000 new cases and over 460,000 deaths reported worldwide annually [1]. Given the relative ease by which these tumours can be accessed, radiotherapy is commonly used to treat both early- and advanced-stage HNC [2] with both curative and palliative intent [3]. While largely effective, one of the major challenges in HNC radiotherapy is the heterogeneity in tumour response, recurrence rate, and severity of impactful toxicities.
Oral mucositis (OM), inflammation of the oral/oropharyngeal mucosa, is a common, dose-limiting toxicity in patients treated with radiotherapy for HNC [4]. Curiously, the incidence and severity of OM vary between patients, even in highly homogeneous cohorts [5]. Unfortunately, it remains unclear what drives this variation in OM risk, with traditional risk factors related to patient demographics, disease/treatment variables, and specific genetic variants unable to sensitively identify high-risk patients [6]. The same challenge is faced for radiotherapy efficacy, with the cause of treatment failure and disease recurrence in some patients still largely unexplained [7, 8]. This lack of understanding severely impacts clinical decision-making, patient monitoring, and the provision of optimal supportive care.
Both radiotherapy-induced toxicities and anti-tumour responses are known to be influenced by host immune responses, which are either exaggerated to drive mucosal toxicity or impaired, thus failing to optimally clear residual tumour load [9]. This knowledge has directed attention to how the gut microbiota may contribute to individual treatment responses, with the gut microbiota a profound regulator of immune tone and immunogenic cell death [10]. Due to its immunomodulatory capacity and its impact on pathways/mechanisms critical to cancer treatment efficacy, such as drug metabolism and cell death and repair, the gut microbiota is emerging as a major driver of treatment outcomes in chemotherapy and immunotherapy, with distinct microbial phenotypes predicting the efficacy and toxicity of these therapies [11].
In the context of radiotherapy, the data is limited. However, accumulating evidence strongly suggests that the gut microbiota may also augment both the efficacy and toxicity of radiotherapy [12, 13]. Of note, olfactory signatures reflecting the structure of the gut microbiota community have been associated with gastrointestinal mucositis severity in patients undergoing pelvic radiotherapy [14]. Additionally, recent evidence in preclinical models demonstrates that the gut microbiota can modulate the radiotherapy-induced anti-tumour immune responses and hence impacting its anti-tumour activity [15, 16]. Together, these data indicate that the gut microbiota may similarly control radiotherapy outcomes as in chemotherapy and immunotherapy.
While not directly investigated in HNC, microbiota-dependent modulation of radiotherapy outcomes is supported by anecdotal data. For example, the use of antibiotics is associated with earlier progression and lower survival among patients with locally advanced HNC treated with chemoradiotherapy [17]. , Further, the use of probiotics has shown promising results in reducing the severity of OM among patients with HNC [18]. Despite this, the association between the pre-therapy gut microbiota and treatment outcomes in HNC has yet to be investigated. This study, therefore, aimed to explore the association between the pre-treatment gut microbiota, OM severity, and tumour recurrence in an HNC cohort.
Materials and methods
Ethical approval
This study was approved by the Royal Adelaide Hospital Human Research Ethics Committee (HREC/17/RAH/533 (R20171131)) and was conducted according to the Declaration of Helsinki. The study protocol was sufficiently discussed with participants and informed consent was obtained from each participant before enrolling in the study.
Patients and biospecimen collection
Patients were recruited from the Radiation Oncology Department at the Royal Adelaide Hospital between October 2018 and December 2019. Adult patients diagnosed with HNC and scheduled to receive radiotherapy alone or combined therapies were eligible and underwent screening. Patients were excluded if they had a medical history of chronic gastrointestinal disorders or intestinal symptoms (unrelated to cancer/treatment) or had previous colonic surgery. Pre-treatment stool samples were collected by patients in DNA/RNA Shield Faecal Collection Tubes (Zymo Research, USA) and stored at −80 °C until processing.
Clinical data collection
Patients were provided with an induction survey to collect demographic information and behavioural/lifestyle factors (see supplementary materials). Clinical data for tumours and treatment characteristics were obtained from medical case notes held at the Royal Adelaide Hospital. OM was scored using the National Cancer Institute Common Terminology Criteria for Adverse Events (NCI-CTCAE) v5.0 [19], which grades OM as grade 1(G1): asymptomatic or mild symptoms; intervention not indicated; grade 2 (G2): moderate pain or ulcer not interfering with oral intake; modified diet indicated; grade 3 (G3): severe pain; interfering with oral intake; grade 4 (G4): life-threatening consequences; urgent intervention indicated; grade 5 (G5): death. Patients were also followed to assess tumour recurrence within 12 months of treatment completion.
DNA extraction and 16S rRNA sequencing
To extract genomic DNA, 2 mL of the sample was first transferred to a sterile microcentrifuge tube and centrifuged at 16,000× g for 20 min at 4 °C. The supernatant was then separated and kept in a tube (not discarded) while the pellet was used for DNA extraction. DNA extraction was performed using Qiagen DNeasy PowerLyzer PowerSoil kit (Qiagen, Germany) as per manufacturer instructions with few modifications. First, Powderbead and C1 solutions were added to the pellet and mixed by brief vortexing. To lyse bacteria cells, the pellet mixture was heated at 65°C for 10 min. Then, the mixture was added into the PowerBead tube and homogenised using QIAGEN Tissuelyser LT (Qiagen, Germany) at 50 oscillation/s for 6 min. The remaining steps were performed as indicated in the kit protocol. The retained supernatant was added back along with the C4 solution during the MB Spin column loading step. To increase the purity of extracted DNA, samples were precipitated using ethanol and sodium chloride, resuspended in nuclease-free water, and stored at −20°C.
DNA concentration was quantified using Qubit 2.0 Fluorometer (Life Technologies, Australia). Samples were sent to the South Australian Genomics Centre for 16S rRNA gene sequencing, performed via Illumina Miseq (San Diego, USA) using primers targeting the hypervariable V3-V4 region:
Forward:
TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG
Reverse: GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC
The bioinformatics analysis was performed using Qiagen CLC Genomics Workbench 21.0.3. Briefly, trimmed and filtered pair-end reads were mapped back to the operational taxonomic units (OTUs) using the Greengenes 97% similarity reference database (v13.8, 2013). The alpha and beta diversity were assessed by the Shannon diversity index and principal coordinates analysis (PCoA) (generalised UniFrac distances) respectively. PERMANOVA analysis was used to measure the significance of beta diversity between groups. Linear discriminant analysis effect size (LEfSe) analysis was conducted using Galaxy online tool using default settings (http://huttenhower.sph.harvard.edu/galaxy/) [20].
Statistical analysis
The statistical analysis was performed using GraphPad prism 9. For quantitative data, unpaired T-test, Mann-Whitney, ANOVA, or Kruskal-Wallis tests were used depending on the Gaussian distribution of the dataset. Fisher’s exact test was used to analyse categorical datasets. Correlation analyses were calculated by Pearson correlation coefficients in Python 3.9.6. A p-value of <0.05 was considered statistically significant.
Results
Patient characteristics
A total of 20 patients were recruited in this study. Patient characteristics are summarised in Table S1. Briefly, 80% of the patients were males with a median age of 65.5 years. Among patients, 75% were either smokers or ex-smokers and 85% reported drinking less than 10 drinks per week. Tumours were located either in the oral cavity (20%), oropharynx (25%), nasal cavity (10%), salivary glands (30%), or HN skin (15%). Half of the patients had early-stage disease (I/II) and the remaining had late-stage disease (III/IV). All patients completed the planned radiotherapy course except for one who discontinued treatment after completing two fractions and hence they were excluded from treatment-related factors analysis. Patients were treated with either radiotherapy alone (31.6%), postoperative radiotherapy (47.4%), chemoradiotherapy (15.8%), or postoperative chemoradiotherapy (5.3%). Overall, patients received an average of 58.62 ± 8.78 Gy cumulative dose in 2.53 ± 1.21 fraction over 5.53 ± 1.46 weeks with 79% treated for curative intent.
Among 19 patients who completed treatment, two received palliative treatment (36 Gy; 6 Gy/F) over 2 weeks. Due to the low exposure, they were excluded from treatment outcomes analyses. Among 17 patients included, 17.7%, 35.3%, 29.4%, and 17.7% experienced G1, G2, G3, and G4 OM respectively. Furthermore, three patients (17.6%) developed recurrence within 12 months post-treatment completion.
Characterisation of HNC patients’ gut microbiota
First, we characterised the gut microbiota of all 20 patients. At the genus level, patients’ gut microbiota was predominantly composed of Bacteroides (39.9%), unclassified Ruminococcaceae (7.4%), Faecalibacterium (6.8%), Parabacteroides (5.6%), and unclassified Lachnospiraceae (4.8%) (Fig. 1A). The average number of positive OTUs was 603.9 [229–864 range] and the average Shannon index value was 3.2 [1.3–4.1 range] (Figs. S1A and S2A).
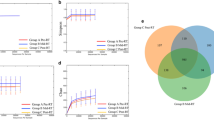
The gut microbiome composition of HNC patients. A The gut microbiota relative abundance at the genus level for all patients. B–C Male patients had a significantly higher number of OTUs (unpaired t-test) and higher alpha diversity (Mann-Whitney test) than female patients. D Female patients have distinctive microbial pattern compared to males. The differential microbial features according to sex (E), age (F), tumour site (G), tumour stage (H), HPV status (I), and treatment type (J). LDA, linear discriminant analysis; CRT, chemoradiotherapy; PORT, postoperative radiotherapy; RT, radiotherapy. *p ≤ 0.05; ***p ≤ 0.001
Sex was the only factor associated with a significant difference in microbial diversity and richness between patients. Female patients had significantly lower OTUs richness (p= 0.0007) and alpha diversity (p= 0.0289). Moreover, the gut microbiota of male and female patients clustered in distinctive patterns as shown by PCoA (p= 0.0052) (Fig. 1B–D). Furthermore, five genera, mainly Prevotella and Phascolarctobacterium, were enriched in males while unclassified Lactobacillales and P-75-a5 were increased in females (Fig. 1E).
Although there was no significant difference in the microbial richness and diversity based on other factors (Figs. S1B-M and S2C-N), specific genera were found to be enriched in specific subgroups. For instance, Faecalibacterium, Paraprevotella, and Ruminococcus-2 were enriched in <50, 55–65, and >65 age groups respectively (Fig. 1F). Furthermore, patients with cutaneous tumours had an increased abundance of unclassified RF32 while SMB35 was increased among patients with salivary gland tumours (Fig. 1G). Phascolarctobacterium was increased in early-stage disease while Enterococcus was enriched in the advanced disease group (Fig. 1H). Moreover, Phascolarctobacterium was enriched in patients with HPV+ tumours (Fig. 1I). The unclassified Enterobacteriaceae was enriched in patients treated with radiotherapy alone while Faecalibacterium and Phascolarctobacterium were increased in those treated with postoperative radiotherapy and chemoradiotherapy respectively (Fig. 1J). Differential compositional changes based on other patient-related factors were also observed (Fig. S3A–H).
Risk factors associated with OM
Three patients were excluded from OM severity analysis (discontinued treatment/received low radiation doses). Patients were divided into either mild/moderate OM (G1–2) or severe OM (G3–4). In this cohort, there was no significant impact of all factors, except treatment type, on OM severity. Expectedly, 75% of patients with tumours in the oral cavity or oropharynx developed severe OM compared to only 22% of patients with tumours in other sites, but the difference was not statistically significant. However, those treated with chemoradiotherapy had significantly more severe OM (100%) compared to those who received radiotherapy without chemotherapy (30.8%) (p= 0.029) (Table 1).
Gut microbiota traits associated with OM
Characterising the gut microbiota based on OM severity, the most abundant genera in the G1–2 OM group were Bacteroides (40%), Parabacteroides (7.8%), Faecalibacterium (6.9%), unclassified Ruminococcaceae (6.8%), and unclassified Clostridiales (4.7%) compared to Bacteroides (41.9%), Faecalibacterium (7.9%), unclassified Ruminococcaceae (7.2%), Prevotella (5.5%), and unclassified Lachnospiraceae (4.2%) in G3–4 OM group (Fig. 2A) (Table S2). Although there was no significant difference in the OTUs richness, alpha, and beta diversity between groups (Fig. S4A–C), Eubacterium, Victivallis, Ruminococcus, Oxalobacter, unclassified Victivallaceae, and unclassified desulfovibrionaceae were significantly increased in patients with G3–4 OM while unclassified RF32, Alistipes, and unclassified ML615J-28 were increased in those with G1–2 OM (Fig. 2B) (Tables S4–S5).
Association between the gut microbiota and OM severity. A The gut microbiota relative abundance at the genus level for G1–2 and G3–4 OM groups. B The differential microbial features for G1–2 and G3–4 OM groups. The relative abundance of Eubacterium (C), Victivallis (D), and Ruminococcus (E) was significantly higher in G3–4 group. F–H Change in the average relative abundance of Eubacterium (F), Victivallis (G), and Ruminococcus (H) according to OM severity grade. I The relative abundance of unclassified RF32 was significantly higher in G1–2 group. J Change in the average relative abundance of unclassified RF32 according to OM severity grade. LDA, linear discriminant analysis. *p ≤ 0.05. Mann-Whitney test; Line represents the median
Among the six genera enriched in the G3–4 OM group, the relative abundance of Eubacterium (p= 0.019), Victivallis (p= 0.016), and Ruminococcus (p= 0.027) was significantly higher in G3–4 compared to G1–2 OM group (Fig. 2C–E). Eubacterium and Ruminococcus genera were most abundant in patients with G3 OM while Victivallis was most abundant among patients with G4 OM (Fig. 2F–H). In contrast, the relative abundance of unclassified RF32 genus (p= 0.032) was significantly higher among patients with G1–2 OM and was most abundant among patients with G2 OM (Fig. 2I–J). Correlation analysis showed a significant positive correlation between the relative abundance of Victivallis and OM severity grade (r= 0.67, p= 0.003) (Fig. 3).
Risk factors associated with tumour recurrence
Among 17 patients included in tumour recurrence analysis, 14 patients did not develop tumour recurrence while 3 patients had recurrence within 12 months of treatment completion. Overall, there was no significant association between any of the patients and treatment-related factors and tumour recurrence (Table 2). Those who developed recurrence had tumours in the oropharynx, nasal cavity, or salivary gland. One of them had early-stage disease and two had advanced-stage disease. All of these patients received similar treatment; however, 2 out of these three patients had treatment breaks or delays.
Gut microbiota traits associated with tumour recurrence
Characterising the gut microbiota based on tumour recurrence, the most abundant genera among patients with no recurrence (no REC) were Bacteroides (39%), Faecalibacterium (8.9%), unclassified Ruminococcaceae (7.2%), Parabacteroides (5.9%), and Prevotella (4.9%) compared to Bacteroides (50%), unclassified Clostridiales (6.4%), unclassified Ruminococcaceae (6.0%), Parabacteroides (5.7%), and Blautia (4.5%) in recurrence (REC) group (Fig. 4A) (Table S3). Generally, there was no significant difference in the number of OTUs, alpha, and beta diversity between groups (Fig. S5A-B and Fig. 4B). However, Faecalibacterium, Prevotella, and Phascolarctobacterium were enriched in patients with no recurrence, and Adlercreutzia, Pseudoramibacter_Eubacterium, Desulfitobacter, Eggerthella, Megasphaera, and p-75-a5 were increased in patients with recurrence (Fig. 4C) (Tables S4–S5). The relative abundance of Faecalibacterium (p= 0.029), Prevotella (p= 0.031), and Phascolarctobacterium (p= 0.019) was significantly higher in patients with no recurrence (Fig. 4D–F). Furthermore, patients who did not develop recurrence also had a significantly higher Prevotella to Bacteroides (P/B) ratio (p= 0.047) (Fig. 4G). Conversely, the relative abundance of Adlercreutzia (p= 0.006) and Eggerthella (p= 0.006) genera was significantly higher in patients with recurrence (Fig. 4H–I). There was no significant difference between recurrence and no recurrence groups in the relative abundance of other genera (Fig. S5C–F).
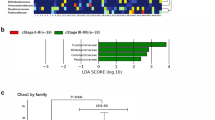
Association between the gut microbiota and tumour recurrence at 12 months. A The relative abundance of the gut microbiota at the genus level for No REC and REC groups. B PCoA of No REC and REC groups. C LEfSe analysis showing the differential genera enriched in No REC and REC groups. D–G The relative abundance of Faecalibacterium (D), Phascolarctobacterium (E), Prevotella (F), and P/B ratio (G) was significantly higher in No REC group. H–I The relative abundance of Eggerthella (H) and Adlercreutzia (I) was significantly higher in REC group. LDA, linear discriminant analysis. *p≤ 0.05; **p≤ 0.001; Mann-Whitney test; Line represents the median
Discussion
Despite the recent technological advances in cancer radiotherapy, variability in radiotherapy outcomes in terms of efficacy and toxicity remains a key challenge. Here, we build on the growing consensus that an individual’s unique, pre-treatment gut microbiota is associated with radiotherapy responses, identifying enrichment and reduction in key taxa linked with distinct treatment outcomes.
Although there was no difference between patients with mild/moderate or severe OM in both the microbial richness and diversity, six bacterial genera were enriched in patients with severe OM. Among these microbes, Eubacterium (E. biforme species), Victivallis, and Ruminococcus genera were the most significantly increased. Eubacterium, a genus of gram-positive anaerobic bacteria belongs to the Erysipelotrichaceae family with Eubacterium biforme (E. biforme) classified as main species within this genus [21]. Eubacterium has been recently reclassified as Holdemanella and E. biforme as Holdemanella biformis (H. biformis) [22]. We refer to them here as Eubacterium and E. biforme based on the reference database used for the analysis. Both beneficial and detrimental effects of this bacterium have been reported. It has been reported that E. biforme can produce C18-3OH, a free long-chain fatty acid with potential anti-inflammatory properties, which in turn reduces colitis severity in mice [23]. Conversely, other studies have reported that an increase in Eubacterium is associated with severe cystic fibrosis [24], nonalcoholic steatohepatitis [25], irritable bowel syndrome [26], and HIV infection [27]. In vitro incubation of peripheral blood mononuclear cells from HIV positive and negative subjects with E. biforme bacterial lysates was associated with a higher tumour necrosis factor α (TNF-α) to interleukin 10 ratio as compared to incubating cells with lipopolysaccharides or three other bacterial species, suggesting a pro-inflammatory property of this species [27]. Another genus that showed a strong correlation with OM severity is Victivallis. Victivallis, a genus of gram-negative anaerobic bacteria, belongs to the Victivallaceae family. It is the only genus in the Victivallaceae family and includes one well-characterised species, Victivallis vadensis [28]. Currently, little is known about the function and impact of this on the human gastrointestinal tract; however, an increase in the abundance of the Victivallaceae family or its genus and species has been linked to inflammatory conditions including colorectal cancer [29], Hashimoto’s thyroiditis [30], and cerebral ischemic stroke [31]. Although this genus is present in a low abundance, the detection rate (OTUs>0) was 62.5% of patients with severe OM compared to only 11.1% of those with mild/moderate OM. This suggests that Victivallis may contribute to OM severity despite its low abundance and warrants further investigation. Ruminococcus, a genus of strictly anaerobic gram-positive cocci of the Lachnospiraceae family [32], was also increased in patients with severe OM. It comprises five species including R. gnavus and R. torques [33]. Both species are mucolytic and have been linked to the pathogenesis of chronic inflammatory conditions including inflammatory bowel disease [34]. R. gnavus can also secrete a pro-inflammatory polysaccharide inducing the production of TNF-α through the toll-like receptor 4-dependent pathway; hence, contributing to Crohn’s disease pathogenesis [35]. This suggests that these mucolytic and pro-inflammatory species potentially contribute to OM pathogenesis through degradation of mucus layer and activating systemic inflammation. Together, present results suggest that these three genera could contribute to OM severity, potentially due to their pro-inflammatory properties. Further studies are needed to validate this association and to determine the mechanism by which these microbes may influence OM pathogenesis.
Among the compositional changes observed is the increased abundance of Unclassified RF32 in patients with mild/moderate OM. Since this genus was also increased in patients with HN skin tumours, and all developed mild/moderate OM, we believe that this genus is associated with tumour site rather than OM severity. Although the LEFSe analysis revealed the Alistipes, and unclassified ML615J-28 were also enriched in patients with mild/moderate OM, the comparison of relative abundance did not yield a significant difference between groups. Overall, this study did not identify any bacterial taxa to be specifically associated with mild/moderate OM.
In terms of tumour recurrence, there was no difference in microbial richness and diversity between patients. Interestingly, patients who did not develop recurrence had a significantly higher abundance of Faecalibacterium, Prevotella, and Phascolarctobacterium. Additional analysis at the species level identified that Faecalibacterium Prausnitzii (F. prausnitzii) and Prevotella Copri (P. copri) were enriched in patients with no recurrence. Generally, these three genera comprise gram-negative bacteria and have been linked to better immunotherapy outcomes in patients with melanoma and non-small cell lung cancer [36,37,38]. For instance, in patients with melanoma treated with anti-PD-1 immunotherapy, responders had an increased abundance of Faecalibacterium and Phascolarctobacterium, with Faecalibacterium associated with prolonged progression-free survival [36, 38]. Furthermore, an increase in P. copri was associated with a preferred response in a cohort of patients with non-small cell lung cancer treated with anti-PD-1 immunotherapy [37]. We also noticed that those who did not develop recurrence had a significantly higher P/B ratio, which is an enterotype associated with a favourable response to anti-PD-1/PD-L1 immunotherapy in patients with gastrointestinal cancers [39]. Current evidence suggests that these microbes modulate immunotherapy anti-tumour response through enhancing CD8+ T cell expansion and function [37, 36]. This could be similar in the context of radiotherapy as anti-tumour immune response also plays a central role in radiotherapy-induced tumour control [13]. In a preclinical study, targeting gram-positive bacteria with vancomycin improved radiotherapy anti-tumour activity by enhancing tumour-associated antigen presentation to CD8+ T cells [15]. Conversely, Adlercreutzia and Eggerthella (E. Lenta), both belonging to the Eggerthellaceae family, were increased in those who developed recurrence. Previous studies have reported that these genera are enriched in non-responders treated with immunotherapy for metastatic melanoma [38]. Together, the current results suggest that certain gut microbes are positively or negatively associated with the risk of recurrence in HNC patients.
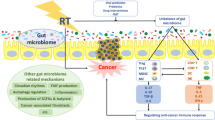
This study aimed to assess the feasibility of utilising the baseline gut microbiota to predict radiotherapy outcomes in order to identify patients at higher risk of developing severe OM or experiencing tumour recurrence and hence allows for taking prophylactic actions to improve treatment outcomes for these groups of patients. However, gut microbiota may undergo compositional changes throughout the course of treatment, and this could also affect HNC radiotherapy toxicity and efficacy. The composition of the gut microbiota at baseline or during treatment could influence the pathogenesis of OM by regulating systemic inflammatory responses. For instance, pro-inflammatory gut microbes present in the baseline gut microbiota or enriched after exposure to treatment could exacerbate the inflammatory processes in the oral cavity and therefore result in more severe OM. This could be mediated by changes in the local intestinal environment such as intestinal inflammation and disruption of the intestinal barrier integrity or by the systemic effects of the gut microbiota on the host’s immune system [40]. Conversely, gut microbes with anti-inflammatory properties positively influence OM by promoting intestinal homeostasis and reducing systemic inflammatory signals resulting in a mild OM (Fig. 5). The gut microbiota could also impact radiotherapy efficacy by modulating the systemic immunity and subsequently the anti-tumour immune responses (Fig. 5) [13, 41]. Although the gut microbiota effect on radiotherapy efficacy has not been investigated in the context of HNC, current preclinical data indicate that the gut microbiota can modulate radiotherapy-induced anti-tumour immune responses including dendritic cells function and activation of tumour-specific CD8+ T cell and interferon-γ pathway [15, 16] and as such enhancing tumour control and reducing the risk of recurrence.
Although this study mainly focused on the gut microbiota, the oral microbiota could also have an impact on radiotherapy outcomes in HNC, particularly the development and severity of OM [42]. Furthermore, emerging evidence indicates that there is an association between oral and gut microbiota. Oral microbiota can translocate into the gut, particularly in the presence of oral pathological conditions (e.g. periodontitis or oral mucositis) or systemic inflammation [40, 43]. This leads to gut microbial dysbiosis and disruption of intestinal homeostasis and this in turn could negatively impact OM outcomes. As such, studying both oral and gut microbiota together will help identify the oral and gut microbial signatures of those with unfavourable radiotherapy outcomes. Moreover, both oral and gut microbiota could be targeted to improve radiotherapy response and reduce its toxicities in patients with HNC.
Overall, this is the first study to characterise the association between gut microbiota and radiotherapy outcomes in patients with HNC. It demonstrates that specific gut microbes are associated with OM severity and risk of tumour recurrence and, as such, supports that the gut microbiota could be exploited to predict radiotherapy outcomes. Another strength of the study is that it assessed microbial signatures associated with both efficacy and toxicity of radiotherapy, which is a critical approach to achieving optimal outcomes for cancer treatments [44]. However, the study is not without limitations. We recognise the small sample size of our cohort and the presence of different confounding factors at baseline and, therefore, emphasise our results must be interpreted with caution. The small sample size may result in biases in the association between OM severity, tumour recurrence, and patients and treatment-related risk factors as well as the microbial signature. Moreover, baseline confounding factors including heterogeneity in tumour primary sites and type of treatment received could impact OM severity and recurrence risk analysis. Therefore, future studies should validate these findings in a larger cohort with minimal variation in the baseline factors. Ideally, the effect of the gut microbiota could be investigated in a large cohort of patients with tumours in a specific site, e.g. oral cavity or oropharynx, and treated with a specific treatment, e.g. radiotherapy alone or chemoradiotherapy. This will allow studying the gut microbiome impact on both OM and treatment outcomes in a more homogeneous cohort. Other factors known to influence the gut microbiota such as age range, comorbidities, and lifestyle-related factors should be taken into account when designing these studies. Additionally, longitudinal studies to analyse the gut microbiome throughout the course of treatment will give an insight into the treatment-induced alterations in the gut microbiota and its association with therapeutic outcomes.
Conclusion
Our study demonstrates that a gut microbiota enriched of Eubacterium, Victivallis, and Ruminococcus is associated with severe OM. Additionally, enrichment for Faecalibacterium, Prevotella, and Phascolarctobacterium confers lower recurrence risk. These pilot data, therefore, reinforce the emerging hypothesis that an individual’s unique microbiota can be used to predict treatment outcomes and be used to direct the provision of proactive supportive care. Moving forward, these data should be used to identify candidate microbes suitable for second-generation probiotics aimed at pre-conditioning the microbiota to optimise treatment outcomes.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Yeh SA (2010) Radiotherapy for head and neck cancer. Semin Plast Surg 24(2):127–136. https://doi.org/10.1055/s-0030-1255330
Gooi Z, Fakhry C, Goldenberg D, Richmon J, Kiess AP (2016) AHNS Series: do you know your guidelines? Principles of radiation therapy for head and neck cancer: a review of the National Comprehensive Cancer Network guidelines. Head Neck 38(7):987–992. https://doi.org/10.1002/hed.24448
Maria OM, Eliopoulos N, Muanza T (2017) Radiation-induced oral mucositis. Front Oncol 7:89. https://doi.org/10.3389/fonc.2017.00089
Bachour PC, Sonis ST (2018) Predicting mucositis risk associated with cytotoxic cancer treatment regimens: rationale, complexity, and challenges. Curr Opin Support Palliat Care 12(2):198–210. https://doi.org/10.1097/spc.0000000000000339
Wardill HR, Sonis ST, Blijlevens NMA, Van Sebille YZA, Ciorba MA, Loeffen EAH, Cheng KKF, Bossi P, Porcello L, Castillo DA, Elad S, Bowen JM, On behalf of The Mucositis Study Group of the Multinational Association of Supportive Care in Cancer/International Society of Oral O (2020) Prediction of mucositis risk secondary to cancer therapy: a systematic review of current evidence and call to action. Supp Care Cancer 28(11):5059–5073. https://doi.org/10.1007/s00520-020-05579-7
Begg AC, Stewart FA, Vens C (2011) Strategies to improve radiotherapy with targeted drugs. Nature Reviews Cancer 11(4):239–253
Orth M, Lauber K, Niyazi M, Friedl AA, Li M, Maihöfer C, Schüttrumpf L, Ernst A, Niemöller OM, Belka C (2014) Current concepts in clinical radiation oncology. Radiation and environmental biophysics 53(1):1–29. https://doi.org/10.1007/s00411-013-0497-2
Cytlak UM, Dyer DP, Honeychurch J, Williams KJ, Travis MA, Illidge TM (2021) Immunomodulation by radiotherapy in tumour control and normal tissue toxicity. Nat Rev Immunol. https://doi.org/10.1038/s41577-021-00568-1
Roy S, Trinchieri G (2017) Microbiota: a key orchestrator of cancer therapy. Nature Reviews Cancer 17:271–285
Alexander JL, Wilson ID, Teare J, Marchesi JR, Nicholson JK, Kinross JM (2017) Gut microbiota modulation of chemotherapy efficacy and toxicity. Nature Reviews Gastroenterology and Hepatology 14(6):356–365
Al-Qadami G, Van Sebille Y, Le H, Bowen J (2019) Gut microbiota: implications for radiotherapy response and radiotherapy-induced mucositis. Expert Review of Gastroenterology & Hepatology 13:485–496. https://doi.org/10.1080/17474124.2019.1595586
Tonneau M, Elkrief A, Pasquier D, Socorro TPD, Chamaillard M, Bahig H, Routy B (2021) The role of the gut microbiome on radiation therapy efficacy and gastrointestinal complications: a systematic review. Radiotherapy and Oncology 156:1–9. https://doi.org/10.1016/j.radonc.2020.10.033
Covington JA, Wedlake L, Andreyev J, Ouaret N, Thomas MG, Nwokolo CU, Bardhan KD, Arasaradnam RP (2012) The detection of patients at risk of gastrointestinal toxicity during pelvic radiotherapy by electronic nose and FAIMS: a pilot study. Sensors (Basel) 12(10):13002–13018. https://doi.org/10.3390/s121013002
Uribe-Herranz M, Rafail S, Beghi S, Gil-de-Gómez L, Verginadis I, Bittinger K, Pustylnikov S, Pierini S, Perales-Linares R, Blair IA, Mesaros CA, Snyder NW, Bushman F, Koumenis C, Facciabene A (2020) Gut microbiota modulate dendritic cell antigen presentation and radiotherapy-induced antitumor immune response. J Clin Invest 130(1):466–479. https://doi.org/10.1172/jci124332
Yang K, Hou Y, Zhang Y, Liang H, Sharma A, Zheng W, Wang L, Torres R, Tatebe K, Chmura SJ, Pitroda SP, Gilbert JA, Fu Y-X, Weichselbaum RR (2021) Suppression of local type I interferon by gut microbiota–derived butyrate impairs antitumor effects of ionizing radiation. J Exp Med 218(3):e20201915. https://doi.org/10.1084/jem.20201915
Nenclares P, Bhide SA, Sandoval-Insausti H, Pialat P, Gunn L, Melcher A, Newbold K, Nutting CM, Harrington KJ (2020) Impact of antibiotic use during curative treatment of locally advanced head and neck cancers with chemotherapy and radiotherapy. Eur J Cancer 131:9–15. https://doi.org/10.1016/j.ejca.2020.02.047
Shu Z, Li P, Yu B, Huang S, Chen Y (2020) The effectiveness of probiotics in prevention and treatment of cancer therapy-induced oral mucositis: a systematic review and meta-analysis. Oral Oncol 102:104559. https://doi.org/10.1016/j.oraloncology.2019.104559
Trotti A, Colevas AD, Setser A, Basch E (2007) Patient-reported outcomes and the evolution of adverse event reporting in oncology. J Clin Oncol 25(32):5121–5127. https://doi.org/10.1200/jco.2007.12.4784
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12(6):R60. https://doi.org/10.1186/gb-2011-12-6-r60
Wade WG (2006) The genus Eubacterium and related genera. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes. Springer US, New York, NY, pp 823–835. https://doi.org/10.1007/0-387-30744-3_28
De Maesschalck C, Van Immerseel F, Eeckhaut V, De Baere S, Cnockaert M, Croubels S, Haesebrouck F, Ducatelle R, Vandamme P (2014) Faecalicoccus acidiformans gen. nov., sp. nov., isolated from the chicken caecum, and reclassification of Streptococcus pleomorphus (Barnes et al. 1977), Eubacterium biforme (Eggerth 1935) and Eubacterium cylindroides (Cato et al. 1974) as Faecalicoccus pleomorphus comb. nov., Holdemanella biformis gen. nov., comb. nov. and Faecalitalea cylindroides gen. nov., comb. nov., respectively, within the family Erysipelotrichaceae. Int J Syst Evol Microbiol 64(Pt_11):3877–3884. https://doi.org/10.1099/ijs.0.064626-0
Pujo J, Petitfils C, Le Faouder P, Eeckhaut V, Payros G, Maurel S, Perez-Berezo T, Van Hul M, Barreau F, Blanpied C, Chavanas S, Van Immerseel F, Bertrand-Michel J, Oswald E, Knauf C, Dietrich G, Cani PD, Cenac N (2020) Bacteria-derived long chain fatty acid exhibits anti-inflammatory properties in colitis. Gut 70:1088–1097. https://doi.org/10.1136/gutjnl-2020-321173
Schippa S, Iebba V, Santangelo F, Gagliardi A, De Biase RV, Stamato A, Bertasi S, Lucarelli M, Conte MP, Quattrucci S (2013) Cystic fibrosis transmembrane conductance regulator (CFTR) allelic variants relate to shifts in faecal microbiota of cystic fibrosis patients. PLoS One 8(4):e61176. https://doi.org/10.1371/journal.pone.0061176
Rau M, Rehman A, Dittrich M, Groen AK, Hermanns HM, Seyfried F, Beyersdorf N, Dandekar T, Rosenstiel P, Geier A (2018) Fecal SCFAs and SCFA-producing bacteria in gut microbiome of human NAFLD as a putative link to systemic T-cell activation and advanced disease. United European Gastroenterol J 6(10):1496–1507. https://doi.org/10.1177/2050640618804444
Maity C, Gupta AK, Saroj DB, Biyani A, Bagkar P, Kulkarni J, Dixit Y (2020) Impact of a gastrointestinal stable probiotic supplement Bacillus coagulans LBSC on human gut microbiome modulation. Journal of Dietary Supplements 18:577–596. https://doi.org/10.1080/19390211.2020.1814931
Lozupone CA, Li M, Campbell TB, Flores SC, Linderman D, Gebert MJ, Knight R, Fontenot AP, Palmer BE (2013) Alterations in the gut microbiota associated with HIV-1 infection. Cell host & microbe 14(3):329–339. https://doi.org/10.1016/j.chom.2013.08.006
Plugge CM, Zoetendal EG (2014) The family Victivallaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: other major lineages of bacteria and the archaea. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 1019–1021. https://doi.org/10.1007/978-3-642-38954-2_150
Sánchez-Alcoholado L, Ordóñez R, Otero A, Plaza-Andrade I, Laborda-Illanes A, Medina JA, Ramos-Molina B, Gómez-Millán J, Queipo-Ortuño MI (2020) Gut microbiota-mediated inflammation and gut permeability in patients with obesity and colorectal cancer. Int J Mol Sci 21(18):6782. https://doi.org/10.3390/ijms21186782
Cornejo-Pareja I, Ruiz-Limón P, Gómez-Pérez AM, Molina-Vega M, Moreno-Indias I, Tinahones FJ (2020) Differential microbial pattern description in subjects with autoimmune-based thyroid diseases: a pilot study. Journal of personalized medicine 10(4):192. https://doi.org/10.3390/jpm10040192
Li N, Wang X, Sun C, Wu X, Lu M, Si Y, Ye X, Wang T, Yu X, Zhao X, Wei N, Wang X (2019) Change of intestinal microbiota in cerebral ischemic stroke patients. BMC Microbiol 19(1):191–191. https://doi.org/10.1186/s12866-019-1552-1
La Reau AJ, Meier-Kolthoff JP, Suen G (2016) Sequence-based analysis of the genus Ruminococcus resolves its phylogeny and reveals strong host association. Microbial genomics 2(12):e000099. https://doi.org/10.1099/mgen.0.000099
Togo AH, Diop A, Bittar F, Maraninchi M, Valero R, Armstrong N, Dubourg G, Labas N, Richez M, Delerce J, Levasseur A, Fournier P-E, Raoult D, Million M (2018) Description of Mediterraneibacter massiliensis, gen. nov., sp. nov., a new genus isolated from the gut microbiota of an obese patient and reclassification of Ruminococcus faecis, Ruminococcus lactaris, Ruminococcus torques, Ruminococcus gnavus and Clostridium glycyrrhizinilyticum as Mediterraneibacter faecis comb. nov., Mediterraneibacter lactaris comb. nov., Mediterraneibacter torques comb. nov., Mediterraneibacter gnavus comb. nov. and Mediterraneibacter glycyrrhizinilyticus comb. nov. Antonie Van Leeuwenhoek 111(11):2107–2128. https://doi.org/10.1007/s10482-018-1104-y
Png CW, Lindén SK, Gilshenan KS, Zoetendal EG, McSweeney CS, Sly LI, McGuckin MA, Florin TH (2010) Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am J Gastroenterol 105(11):2420–2428. https://doi.org/10.1038/ajg.2010.281
Henke MT, Kenny DJ, Cassilly CD, Vlamakis H, Xavier RJ, Clardy J (2019) Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc Natl Acad Sci U S A 116(26):12672–12677. https://doi.org/10.1073/pnas.1904099116
Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV, Prieto PA, Vicente D, Hoffman K, Wei SC, Cogdill AP, Zhao L, Hudgens CW, Hutchinson DS, Manzo T, Petaccia de Macedo M, Cotechini T, Kumar T, Chen WS et al (2018) Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359(6371):97–103. https://doi.org/10.1126/science.aan4236
Jin Y, Dong H, Xia L, Yang Y, Zhu Y, Shen Y, Zheng H, Yao C, Wang Y, Lu S (2019) The diversity of gut microbiome is associated with favorable responses to anti–programmed death 1 immunotherapy in Chinese patients with NSCLC. Journal of Thoracic Oncology 14(8):1378–1389. https://doi.org/10.1016/j.jtho.2019.04.007
Limeta A, Ji B, Levin M, Gatto F, Nielsen J (2020) Meta-analysis of the gut microbiota in predicting response to cancer immunotherapy in metastatic melanoma. JCI insight 5(23):e140940. https://doi.org/10.1172/jci.insight.140940
Peng Z, Cheng S, Kou Y, Wang Z, Jin R, Hu H, Zhang X, Gong J-f, Li J, Lu M, Wang X, Zhou J, Lu Z, Zhang Q, Tzeng DTW, Bi D, Tan Y, Shen L (2020) The gut microbiome is associated with clinical response to anti–PD-1/PD-L1 immunotherapy in gastrointestinal cancer. Cancer Immunology Research 8(10):1251–1261. https://doi.org/10.1158/2326-6066.cir-19-1014
Al-Qadami G, Van Sebille Y, Bowen J, Wardill H (2022) Oral-gut microbiome axis in the pathogenesis of cancer treatment-induced oral mucositis. Front Oral Health 3:881949. https://doi.org/10.3389/froh.2022.881949
Liu J, Liu C, Yue J (2021) Radiotherapy and the gut microbiome: facts and fiction. Radiat Oncol 16(1):9. https://doi.org/10.1186/s13014-020-01735-9
Reis Ferreira M, Pasto A, Ng T, Patel V, Guerrero Urbano T, Sears C, Wade WG (2022) The microbiota and radiotherapy for head and neck cancer: what should clinical oncologists know? Cancer Treat Rev 109:102442. https://doi.org/10.1016/j.ctrv.2022.102442
Olsen I, Yamazaki K (2019) Can oral bacteria affect the microbiome of the gut? J Oral Microbiol 11(1):1586422. https://doi.org/10.1080/20002297.2019.1586422
Wardill HR, Chan RJ, Chan A, Keefe D, Costello SP, Hart NH (2022) Dual contribution of the gut microbiome to immunotherapy efficacy and toxicity: supportive care implications and recommendations. Supportive Care in Cancer 30(8):6369–6373. https://doi.org/10.1007/s00520-022-06948-0
Acknowledgements
We would like to sincerely thank the nurses and radiation oncologists at the Department of Radiation Oncology (Royal Adelaide Hospital) for their assistance during the recruitment process.
Funding
This study was supported by the Royal Adelaide Hospital Clinical Project Grant (CI Wardill).
Author information
Authors and Affiliations
Contributions
GA, JB, YVS, KS, and HL made substantial contributions to study conceptualisation and design (GA, JB, and HL); patient recruitment and sample collection (GA and HL); DNA extraction and microbial data analysis (GA and KS); and drafting (GA) and revising the manuscript (GA, JB, YVS, KS, and HL). HW is PI for the study, securing relevant ethical approvals and seed funding as well as contributing to data analysis and revising the manuscript. MD and JV contributed to the data analysis and interpretation and reviewing of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Royal Adelaide Hospital Human Research Ethics Committee (HREC/17/RAH/533 (R20171131)).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Hannah Wardill and Hien Le have shared senior authorship.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Al-Qadami, G., Bowen, J., Van Sebille, Y. et al. Baseline gut microbiota composition is associated with oral mucositis and tumour recurrence in patients with head and neck cancer: a pilot study. Support Care Cancer 31, 98 (2023). https://doi.org/10.1007/s00520-022-07559-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00520-022-07559-5