Abstract
In the present study, we developed an efficient method of 1,3-propanediol (1,3-PD) production from glycerol by genetic engineering of Klebsiella pneumoniae AK mutant strains. The proposed approach eliminated by-product formation and IPTG induction resulted in maximal production of 1,3-PD. A series of recombinant strains was designed to constitutively express the dhaB and/or dhaT genes, using the bacteriophage T5 PDE20 promoter and the rho-independent transcription termination signal of the Rahnella aquatilis levansucrase gene. Among these strains, AK/pConT expressing dhaT alone gave the highest yield of 1,3-PD. Fed-batch fermentation resulted in efficient production of 1,3-PD from either pure or crude glycerol, without by-product formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Currently, biodiesel synthesized from animal fat or plant oil is in great demand. However, a large amount of raw glycerol is formed as the main by-product of biodiesel production, which can be as much as 10 % (w/w) of the biodiesel generated [1]. This surplus raw glycerol has not only greatly disturbed the market for traditional glycerol in terms of both preparation and price, but is also a significant environmental problem because glycerol cannot be discharged directly into the environment [2]. Thus, a great deal of research effort has been devoted to the development of methods by which glycerol (a low-cost feedstock) can be refined into industrially valuable materials such as fuels, chemical building blocks, and bioactive substances.
1,3-Propanediol (1,3-PD) is a valuable chemical used mainly in the synthesis of polymethylene terephthalates by polymerization with terephthalates [3]. Applications for these polymers in the manufacture of materials such as textile fibers, films, and plastics are rapidly increasing. The 1,3-PD building block is currently produced by chemical processes such as hydroformylation of ethylene oxide or hydration of acrolein [4, 5]. Recently, a microbial fermentation process using a recombinant Escherichia coli strain containing genes from Klebsiella pneumoniae and Saccharomyces cerevisiae was developed, by which glucose is converted to 1,3-PD [6]. However, the original route to 1,3-PD production involved microbial fermentation using glycerol as substrate. Because glycerol is a major by-product of the biodiesel industry, biological conversion of this chemical is receiving considerable attention.
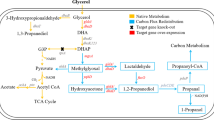
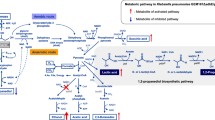
Klebsiella pneumoniae is typical of microorganisms that produce 1,3-PD, and the metabolic pathway responsible has been well studied [7]. Glycerol is first converted to 3-hydroxypropionaldehyde (3-HPA) by a coenzyme B12-dependent glycerol dehyratase (DhaB), and this material is next reduced to 1,3-PD in a reaction catalyzed by a reduced nicotinamide adenine dinucleotide (NADH)-dependent 1,3-PD oxidoreductase (DhaT). In addition to the reductive pathway, glycerol is metabolized by an oxidative pathway, through which glycerol is dehydrogenated to dihydroxyacetone (DHA) by an NAD+-dependent glycerol dehydrogenase (DhaD) and DHA is phosphorylated to dihydroxyacetone phosphate (DHAP) by an ATP-dependent DHA kinase (DhaK). Glycerol assimilated via the oxidative branch results in an increase in biomass, resulting in the generation of many different metabolites such as acetate, ethanol, lactate, succinate, and 2,3-butanediol (2,3-BD).
The genes encoding the functionally linked proteins DhaB and DhaT of the reductive branch and DhaD and DhaK of the oxidative branch are clustered in a region of K. pneumoniae chromosomal DNA termed the dha regulon [8]. Many researchers have sought to enhance 1,3-PD production by metabolic engineering [9–12], either greatly increasing enzyme activities [13–15] or NADH availability [16]. However, minimization of by-product synthesis (which can be as much of 70 % [w/w] of 1,3-PD yield) must be considered when enhanced production of 1,3-PD is desired [17]. 2,3-BD is a major by-product formed during synthesis of 1,3-PD, making downstream purification of 1,3-PD difficult because the two materials have similar boiling points [18].
Recently, we engineered a mutant strain of K. pneumoniae (termed AK-VOT) to eliminate by-product formation during production of 1,3-PD from glycerol, by inactivation of the oxidative branch of the glycerol metabolic pathway [19]. In such recombinant strains, by-product formation, with the exception of acetate, was eliminated, resulting in a higher 1,3-PD yield relative to that of the wild-type strain. In the present study, we further improved 1,3-PD yield by constitutive expression of a dha gene to increase the economy of 1,3-PD production.
Materials and methods
Strains, plasmids, and media
The K. pneumoniae strains described in our previous study [18] were grown either in LB [yeast extract (Difco), 0.5 % (w/v); Bacto-trypton (Difco), 1.0 % (w/v); and NaCl, 1.0 % (w/v)] or in a defined glycerol-containing medium with 20 g/l of glycerol, 2 g/l (NH4)2SO4, 3.4 g/l K2HPO4, 1.3 g/l KH2PO4, 0.2 g/l MgSO4, 0.002 g/l CaCl2 2H2O, 1 g/l yeast extract, 1 ml Fe solution [5 g/l FeSO4 7H2O and 4 ml/l HCl (37 %, w/v)], and 1 ml trace element solution [70 mg/l ZnCl2, 100 mg/l MnCl2 4H2O, 60 mg/l H3BO3, 200 mg/l CoCl2 4H2O, 20 mg/l CuCl2 2H2O, 25 mg/l NiCl2 6H2O, 35 mg/l Na2MoO4 2H2O, with 4 ml HCl (37 %, w/v)]. Tetracycline was added to a final concentration of 50 μg/ml.
Construction of recombinant plasmids
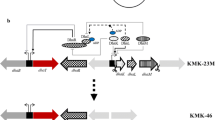
A schematic representation of the strategy used to construct pConT, pConB, pConBT, pConK, and pConTK is shown in Fig. 1. The primers used for amplification of DNA sequences by polymerase chain reaction (PCR) are shown in Table 1. The 75-bp promoter PD/E20 of bacteriophage T5 (GenBank accession no. M11599) and the 80-bp sequence of the rho-independent transcription terminator (TlsrA) from the levansucrase gene of Rahnella aquatilis (GenBank accession no. AAC36458) were synthesized by GenoTech (Daejeon, Korea). These DNA sequences were cloned into the pUC118 vector to generate pUC-PD/E20-TlsrA. The open reading frames (orfs) of the following genes were amplified from chromosomal DNA of K. pneumoniae MGH78578 using the primers shown in Table 1: dhaB α (1.67 kb, GenBank accession no. ABR78884), dhaB β (0.59 kb, ABR78883), and dhaBγ (0.43 kb, ABR78882) encoding glycerol dehydratase subunits (α, β and γ); dhaB RA1 (1.82 kb, ABR78881) and dhaB RA2 (0.35 kb, ABR78887) encoding glycerol dehydratase reactivators (1 and 2); and dhaT (1.16 kb, ABR78886) and yqhD K (1.16 kb, ABR78827), encoding 1,3-PD oxidoreductase and an isozyme thereof [20], respectively. Amplified DNA fragments were cloned into the pGEM T-Easy (Promega, Madison, WI) vector followed by nucleotide sequencing to confirm the absence of any errors. The PshBI and NdeI–SalI fragments including dhaB βγ (pGEM-dhaB βγ ) and dhaB RA1 were serially inserted into the NdeI and SacI sites of pGEM-dhaB α to create pGEM-dhaB. The pGEM-dhaB, -dhaT, -yqhDk, and -dhaB RA2 plasmids were treated with EcoRI, NdeI–XhoI, BglII–XhoI, and NdeI–XhoI, respectively. After gap filling using the Klenow fragment, each DNA molecule was inserted into the EcoRV site of pUC-PD/E20-TlsrA. To generate pConBT, the NotI and SpeI-XbaI fragments, including dhaB (pUC-dhaB), dhaB RA2 (pUC-dhaB RA2 ), and dhaT (pUC-dhaT), were serially inserted into the NotI and XbaI sites of pBR322, respectively. Plasmids pConB, pConT, pConK, and pConTK were similarly constructed, as shown in Fig. 1. The resultant plasmids were transformed into K. pneumoniae AK by electroporation [19].
Schematic representation of the strategies used for construction of pConT, pConB, pConBT, pConK, and pConTK. P and T indicate the 75 bp of promoter PD/E20 of bacteriophage T5 and the 80 bp of the rho-independent transcription terminator (TlsrA) of the levansucrase gene of Rahnella aquatilis, respectively. Abbreviations for restriction enzymes: Bg, BglII; E, EcoRV; EI, EcoRI; N, NotI; Nd, NdeI; P, PshBI; Sc, SacI; Sl, SalI; Sp, SpeI; Xb, XbaI; Xh, XhoI. Kle, Klenow fragment
Fermentation by recombinant K. pneumoniae strains
Seed cells for fermentation were prepared in a 250-ml flask containing 50 ml of LB medium. Flasks were incubated at 37 °C for 12 h and the cultures subsequently inoculated into the fermentor at 2.5 % (v/v). Both batch and fed-batch fermentations were conducted at 37 °C, with stirring at 200 rpm and aeration at 0.5 vvm in a 5-l vessel (Kobiotech. Co., Ltd, Korea) containing 2 l of fermentation medium with tetracycline (10 μg/ml). pH was controlled by automatic addition of 28 % (v/v) NH4OH. The carbon sources used were pure glycerol (purity 99 %, w/w) and crude glycerol (purity 70 %, w/w) derived from the biodiesel industry (ECO Solutions, Korea).
Enzyme activity assay
Cells were harvested by centrifugation at 6,000g for 15 min, and washed twice with 50 mM Tris buffer, pH 8.0. The cell pellet was resuspended in a small amount of buffer and cells were disrupted by ultrasonication for 5 min on ice. Cell debris was removed by brief centrifugation. Glycerol dehydratase (DhaB) activity was estimated using the 3-methyl-2-benzothiazolinon hydrazine (MBTH) method [16]. The activity of 1,3-PD oxidoreductase (DhaT or YqhD) was determined using the reverse reaction (conversion of 1,3-propanediol to 3-hydroxypropionaldehyde) according to the technique of Sulzenbacher et al. [17]. One unit of enzyme activity was defined as the amount of enzyme consuming 1 μmol substrate per minute. Protein levels were determined using a protein assay kit (Bio-Rad), with bovine serum albumin (Sigma, St Louis, MO) as standard, according to the method of Bradford. Data are expressed as mean values ± SDs. Each experiment was performed three times, and the results showed no statistically significant difference, as determined using the Student’s t test.
Metabolite analysis
Culture broth concentrations of metabolites including glycerol, acetate, and 1,3-PD were determined using a high-performance liquid chromatography system (Agilent 1200) equipped with a refractive index detector and an organic acid analysis column (300 × 78 mm; Aminex HPX-87H; Bio-Rad) [18]. The mobile phase was 0.005 mol/l H2SO4 and a flow rate of 0.8 ml per minute was maintained during elution. The column temperature was maintained at 65 °C.
Results and discussion
IPTG induction is essential for 1,3-PD production by K. pneumoniae AK/pVOT
In the recombinant K. pneumoniae AK/pVOT strain, the expression of both dhaB on chromosomal DNA and dhaT on plasmid DNA was under the control of the E. coli lacZ gene promoter [19], which allowed gene expression to be regulated by addition of an inducer such as isopropyl-d-thiogalactoside (IPTG). During cultivation of K. pneumoniae AK/pVOT, addition of IPTG (0.5 mM) induced 1,3-PD production from glycerol without by-product formation. Although IPTG is an efficient inducer of the lacZ promoter, development of an industrial 1,3-PD production process might be compromised by the high price of this chemical. We examined the effect of IPTG on glycerol metabolism by K. pneumoniae AK/pVOT upon cultivation in a 5-l bioreactor. The glycerol consumption rates by the recombinant strain were similar in both the absence and presence of IPTG (Fig. 2a), but cell growth was reduced in the presence of IPTG (Fig. 2b), probably because the chemical has some intrinsic toxicity. However, 1,3-PD production was enhanced in the presence of IPTG (Fig. 2c), indicating that IPTG induction is necessary for efficient production of 1,3-PD from glycerol by K. pneumoniae AK/pVOT. It is well known that lactose can usually substitute for IPTG as an inducer of the lacZ promoter. We thus examined whether lactose could replace IPTG during fermentative cultivation of K. pneumoniae AK/pVOT. However, as shown in Fig. 2, lactose negatively affected all of glycerol consumption rate, cell growth, and 1,3-PD production, to extents greater than those seen in the absence of IPTG.
Construction of recombinant K. pneumoniae AK strains constitutively expressing dha genes
We constructed a series of recombinant strains expressing dha genes in a constitutive manner, using the constitutive PDE20 gene promoter of bacteriophage T5 and an efficient rho-independent transcription termination signal from the Rahnella aquatilis levansucrase gene [21]. K. pneumoniae AK strains harboring pConB, pConT, or pConBT were prepared as described in Methods; these strains constitutively expressed dhaB, dhaT, or both the dhaB and dhaT genes, respectively. We also constructed K. pneumoniae AK strains harboring pConK or pConTK, thus with the yqhD k gene encoding an 1,3-PD oxidoreductase isozyme or with both dhaT and yqhD k (encoding two 1,3-PD oxidoreductases), respectively. The activities of DhaB, DhaT, and YqhDk in the recombinant strains were as expected (Fig. 3).
Fermentation of glycerol by recombinant K. pneumoniae AK strains producing 1,3-PD
Upon batch fermentation of K. pneumoniae AK recombinant strains constitutively expressing dhaB (pConB), dhaT (pConT), or both dhaB and dhaT (pConBT), in the absence of IPTG induction, the AK/pConT strain showed a faster glycerol consumption rate, a higher cell growth rate, and a greater 1,3-PD production level compared with the AK/pVOT strain with IPTG induction (Fig. 4). The AK/pConB and AK/pConBT strains showed the opposite effects, which may be attributable to an imbalance in metabolic flow, with accumulation of a toxic intermediate, 3-HPA, resulting from the increased activity of DhaB. Unexpectedly, high-level expression of the yqhD k gene encoding an 1,3-PD oxidoreductase isozyme negatively affected glycerol metabolism. This may be due to low DhaT activity, although it is unclear why DhaT activity should be reduced upon high-level expression of YqhDk. Glycerol metabolism was recovered by co-expression of dhaT and yqhDk. Thus, K. pneumoniae AK harboring pConT constitutively expressing dhaT alone produced the highest level of 1,3-PD from glycerol.
Fed-batch fermentation for 1,3-PD production by K. pneumoniae AK/pConT using pure or crude glycerol as carbon source
We examined 1,3-PD production from glycerol by K. pneumoniae AK/pConT, using fed-batch fermentation. The maximum levels of 1,3-PD production and productivity from pure glycerol were 26.6 g/l and 0.51 g/l/h in the absence of IPTG induction (Fig. 5a). Similar values (25.9 g/l and 0.54 g/l/h) were obtained using crude glycerol derived from the biodiesel industry as a carbon source (Fig. 5b). Although the maximum levels of 1,3-PD synthesized by AK/pConT were lower than those reported elsewhere [22, 23], the recombinant strain has one crucial advantage, namely, that no by-product, especially 2,3-PD, was formed by the AK/pConT strain, with the exception of acetate, thus rendering downstream 1,3-PD purification economical.
Genetic stability of the K. pneumoniae AK/pConT strain
Finally, we measured the stability of the plasmid expressing dhaT in K. pneumoniae AK/pConT in the absence of selective antibiotics. After culture for 56 h in the 5-l fermentor (thus, for about 150 generations), samples were collected and bacteria cultivated on LB plates. Next, 100 single colonies were replica plated to screen for tetracycline-resistant transformants. The plasmid was maintained in 93 % of cells of the recombinant strain, indicating that pConT is stable in K. pneumoniae AK.
Conclusion
In the present study, we developed a series of recombinant K. pneumoniae AK strains engineered to eliminate by-product formation, in which the dhaB and/or dhaT genes encoding enzymes catalyzing the synthesis of 1,3-PD were constitutively expressed. Among these strains, AK/pConT expressing dhaT alone yielded the highest level of 1,3-PD. Fed-batch fermentation resulted in efficient production of 1,3-PD from either pure or crude glycerol, without by-product formation. Although the current level of 1,3-PD production by the recombinant strain is lower than those reported elsewhere, further engineering to enhance 1,3-PD yield will likely result in an economical biological process for production of this valuable material.
References
Nakamura CE, Whited GM (2003) Metabolic engineering for the microbial production of 1,3-propanediol. Curr Opin Biotechnol 14:454–459
Biebl H, Menzel K, Zeng AP, Deckwer WD (1999) Microbial production of 1,3-propanediol. Appl Microbiol Biotechnol 52:289–297
Saxena RK, Anand P, Saran S, Isar J (2009) Microbial production of 1,3-propanediol: recent developments and emerging opportunities. Biotechnol Adv 27:895–913
Homann T, Tag C, Biebl H, Deckwer WD, Schink B (1990) Fermentation of glycerol to 1,3-propanediol by Klebsiella and Citrobacter strains. Appl Microbiol Biotechnol 33:121–126
Daniel R, Stuertz K, Gottschalk G (1995) Biochemical and molecular characterization of the oxidative branch of glycerol utilization by Citrobacter freundii. J Bacteriol 177:392–401
El-Ziney MG, Debevere JM (1998) The effect of reuterin on Listeria monocytogenes and Escherichia coli O157:h7 in milk and cottage cheese. J Food Prot 61:1275–1280
Marçal D, Rêgo AT, Carrondo MA, Enguita FJ (2009) 1,3-Propanediol dehydrogenase from Klebsiella pneumoniae: decameric quaternary structure and possible subunit cooperativity. J Bacteriol 191:1143–1151
Bhatia SK, Kurian JV (2008) Biological characterization of sorona polymer from corn-derived 1,3-propanediol. Biotechnol Lett 30:619–623
Ma J, Rao Z, Xu L, Liao X, Fang H, Zhuge B, Zhuge J (2009) Expression of dha operon required for 1,3-PD formation in Escherichia coli and Saccharomyces cerevisiae. Curr Microbiol 60:191–198
Gellissen G (2000) Heterologous protein production in methylotrophic yeasts. Appl Microbiol Biotechnol 54:741–750
Sohn JH (1997) Characterization of telomere-associated ARSs (autonomously replicating sequences) and their use for multiple gene integration in Hansenula polymorpha. PhD thesis. Korea Advanced Institute of Science and Technology, Taejon, Korea
Heo JH, Hong WK, Cho EY, Kim MW, Kim JY, Kim CH, Rhee SK, Kang HA (2003) Properties of the Hansenula polymorpha-derived constitutive GAP promoter, assessed using an HSA reporter gene. FEMS Yeast Res 4:175–184
Hill J, Donald KA, Griffiths DE (1991) DMSO-enhanced whole cell yeast transformation. Nucleic Acids Res 9:5791
Hong WK, Kim CH, Heo SY, Luo LH, Oh BR, Seo JW (2010) Enhanced production of ethanol from glycerol by engineered Hansenula polymorpha expressing pyruvate decarboxylase and aldehyde dehydrogenase genes from Zymomonas mobilis. Biotechnol Lett 32:1077–1082
Gonzalez-Pajuelo M, Meynial-Salles I, Mendes F, Soucaille P, Vasconcelos I (2006) Microbial conversion of glycerol to 1,3-propanediol: physiological comparison of a natural producer, Clostridium butyricum VPI 3266, and an engineered strain, Clostridium acetobutylicum DG1(pSPD5). Appl Environ Microbiol 72:96–101
Olivier C, Aline LF (2001) Primers and a specific DNA probe for detecting lactic acid bacteria producing 3-hydroxypropionaldehyde from glycerol in spoiled ciders. J Food Prot 64:833–837
Sulzenbacher G, Alvarez K, Van Den Heuvel RH, Versluis C, Spinelli S, Campanacci V, Valencia C, Cambillau C, Eklund H, Tegoni M (2004) Crystal structure of E. coli alcohol dehydrogenase YqhD: evidence of a covalently modified NADP coenzyme. J Mol Biol 342:489–502
Oh BR, Seo JW, Choi MH, Kim CH (2008) Optimization of culture conditions for 1,3-propanediol production from crude glycerol by Klebsiella pneumoniae using response surface methodology. Biotechnol Bioprocess Eng 13:524–532
Seo MY, Seo JW, Heo SY, Baek JO, Rairakhwada D, Oh BR, Seo PS, Choi MH, Kim CH (2009) Elimination of by-product formation during production of 1,3-propanediol in Klebsiella pneumoniae by inactivation of glycerol oxidative pathway. Appl Microbiol Biotechnol 84:527–534
Seo JW, Seo MY, Oh BR, Heo SY, Baek JO, Rairakhwada D, Luo LH, Hong WK, Kim CH (2010) Identification and utilization of a 1,3-propanediol oxidoreductase isoenzyme for production of 1,3-propanediol from glycerol in Klebsiella pneumoniae. Appl Microbiol Biotechnol 85:659–666
Seo JW, Hong WK, Rairakhwada R, Seo PS, Choi MH, Song KB, Rhee SK, Kim CH (2009) An efficient plasmid vector for constitutive high-level expression of foreign genes in Escherichia coli. Biotechnol Lett 31:877–881
Xu YZ, Guo NN, Zheng ZM, Ou XJ, Liu HJ, Liu DH (2009) Metabolism in 1,3-propanediol fed-batch fermentation by a d-lactate deficient mutant of Klebsiella pneumoniae. Biotechnol Bioeng 104:965–972
Zhuge B, Zhang C, Fang H, Zhuge J, Permaul K (2010) Expression of 1,3-propanediol oxidoreductase and its isoenzyme in Klebsiella pneumoniae for bioconversion of glycerol into 1,3-propanediol. Appl Microbiol Biotechnol 87:2177–2184
Acknowledgments
This subject was supported by Korea Ministry of Environment as “Converging technology project” and by the New & Renewable Energy Technology Development Program of the Korea Institute of Energy Technology Evaluation and Panning (KETEP) Grant (2010T100100690) funded by the Korea government Ministry of Knowledge Economy.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oh, BR., Seo, JW., Heo, SY. et al. Efficient production of 1,3-propanediol from glycerol upon constitutive expression of the 1,3-propanediol oxidoreductase gene in engineered Klebsiella pneumoniae with elimination of by-product formation. Bioprocess Biosyst Eng 36, 757–763 (2013). https://doi.org/10.1007/s00449-013-0901-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-013-0901-y