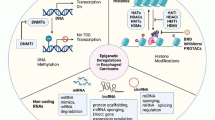
Abstract
Esophageal cancers are a challenging upper gastrointestinal tract tumor entity for interdisciplinary oncology. For the two main histotypes, namely esophageal squamous cell carcinomas and Barrett’s adenocarcinomas, several genetic aberrations have been shown to contribute to carcinogenesis and progression as well as to represent potential novel targets for therapeutic intervention. This is paralleled by growing insight into epigenetic alterations of esophageal cancers. Studies involving the analyses of human tissue specimens predominantly describe altered patterns of miRNA expression, DNA methylation patterns, and histone marks levels. This review provides a critical update on this increasing knowledge of epigenetic alteration in esophageal cancers by specifically focusing on the translational aspects of epigenetic analyses from human tissue specimens.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Esophageal cancers are upper gastrointestinal tract tumors of epithelial cell origin. The two major histotypes of esophageal cancer arise via distinct steps of carcinogenesis. Barrett’s adenocarcinomas (BACs) evolve mainly in the lower esophagus as a consequence of chronic reflux disease via steps of trans-differentiation of normal squamous epithelial cells into columnar/intestinal epithelial cells (intestinal metaplasia), dysplasia, and malignant invasion. In contrast, esophageal squamous cell carcinomas (ESCCs) evolve in all parts of the esophagus via a mere progression from normal squamous epithelial cells to dysplasia and malignant invasion (Fig. 1).
Key epigenetic alterations involved at various stages of esophageal carcinogenesis. The major characteristic morphological features in the carcinogenesis of esophageal squamous cell carcinomas (ESCC) and Barrett’s adenocarcinoma (BAC) are shown (hematoxylin and eosin staining). Selected key epigenetic events and respective genes/proteins regulated by epigenetic mechanisms in the distinct stages are indicated (see text, Tables 1, 2, 3, Supplementary Table S1, and references therein)
Compared with lower gastrointestinal tract tumors, i.e. colorectal cancer, upper gastrointestinal tract tumors are generally less frequent but have a much higher mortality. With a rising incidence of esophageal cancers, particularly adenocarcinomas (Cook et al. 2009), a better understanding of esophageal cancer carcinogenesis and progression is urgently needed to improve patient risk stratification and therapeutic treatment options (Pennathur et al. 2013).
Several key genetic alterations have been identified in the carcinogenesis of esophageal carcinomas, such as mutations of key oncogenes and tumor-suppressor genes (e.g., TP53, MYC) or distinct patterns of chromosome/gene amplifications and deletions (e.g., HER2, FHIT) and tumor cell aneuploidy in general. Whereas some of these genetic alterations are predominantly seen in ESCCs (e.g., EGFR; Lin et al. 2009), others are linked to the majority of BACs (e.g., HER2; Zhang et al. 2009). Importantly, such specific genetic alterations also provide the basis for targeted therapeutic intervention in esophageal cancers, with HER2-targeting inhibitors recently having been approved for first-line therapy of metastatic gastric and esophageal adenocarcinomas (Pennathur et al. 2013). In contrast to other epithelial tumors such as colorectal or lung adenocarcinomas, the treatment options and predictive markers are rather sparse in esophageal cancers. In addition, whether these key genetic alterations alone drive the distinct phenotypes of BAC or ESCC carcinogenesis and therefore also represent valid therapeutic targets for inhibiting esophageal cancer progression remains unclear.
In recent years, because of their role in normal tissue development and differentiation, epigenetic alterations have increasingly been appreciated to contribute to malignant transformation and tumor progression. For example, “environmental damage” imposed by tobacco or alcohol might contribute to the malignant transformation of normal squamous epithelium by the induction of genetic defects or possibly also by the induction of aberrant epigenetic modifications. Moreover, crosstalk appears to occur between (altered) oncogenic signaling pathways and epigenetic modulation (Mohammad and Baylin 2010). Therefore, the past few years have seen an increase in basic and applied research into the field of the epigenetics of esophageal cancer, with publications being registered, for example, via “esophageal cancer, epigenetic, YEAR” from 6 in 2003 to 30 in 2013 and, for example, via “esophageal cancer, methylation, YEAR” from 21 in 2003 to 53 in 2013 [http://www.ncbi.nlm.nih.gov/pubmed/]. True, these seemingly small numbers of publications underline the lack of and necessity for further research into the epigenetics of esophageal cancers. However, (translational) research in esophageal cancer, especially with regard to translational clinical research combining in vitro and in situ analyses, is not as straightforward as in other epithelial tumors: validated cell line models are sparse, and tissue specimens are available from mainly the smallest of biopsies, particularly with respect to interest in precursor lesions and “primary tumors” without prior chemo- or radiotherapy. This rareness of “untouched” esophageal (cancer) tissues limits, in part, the investigational depth. In addition, whereas some methodologies such as microRNA (miRNA) expression and DNA methylation analyses are now readily performed on all kinds of samples, the analysis of histone modifications remains to be further standardized, especially when working with chromatin samples derived from routinely processed tissue specimens. Thus, epigenetic alterations in esophageal cancers are only beginning to be addressed at the level of basic and translational research, and they are still far from being implemented into routine clinico-pathological guidelines.
Irrespective of this, an interest is arising in the therapeutic targeting of epigenetic modifiers, including the inhibition of DNA methyltransferases (DNMTs; e.g., 5-Azacytidine, Decitabine) or histone deacetylases (HDACs; e.g., Vorinostat) in ongoing clinical trials involving patients with esophageal cancers (e.g., Azacytidine/NCT01386346, Depsipeptide/NCT00098644; http://www.cancer.gov/clinicaltrials). These largely build not only on promising studies of these inhibitors in other cancer entities, but also on the above-mentioned growing number of basic epigenetic research studies of esophageal cancers. Therefore, this review will reveal the current knowledge of epigenetic alterations in esophageal cancers from a translational view, by focusing on alterations of miRNAs, DNA methylation, and histone modifications investigated in patient tissue specimens of esophageal carcinomas and associated precursor lesions.
miRNAs in esophageal carcinogenesis
Interestingly, the topic of “microRNAs” (miRNAs or miRs) has been widely addressed in human tissue specimens of esophageal cancers, including candidate miR approaches or broad microarray-based screening approaches (see Table 1 for selected recent studies):
Individual studies, mainly in ESCCs, have pointed out (in part, contradictory) differences of individual miR expression levels in normal esophageal epithelial cells versus esophageal cancer cells (e.g., down-regulated in esophageal cancer: miR-133, miR-145, miR-203, miR-205, miR-375; up-regulated in esophageal cancer: miR-21, miR-145, miR-223). Low expression of selected miRs in esophageal cancer cells (mainly ESCC) has been found to be associated with more aggressive tumors and lymph node or distant metastasis (e.g., let-7/let-7c, miR-375) and/or with poor survival (e.g., let-7/let-7c, miR-150, miR-375). Similarly, other selected miRs showing high expression in esophageal cancer cells (mainly ESCC, some BAC) are associated with advanced tumor stages (e.g., miR-21, miR-145) and poor survival (e.g., miR-103/107, miR-200 family, miR-223). This is complemented by several studies that have measured miRs in the serum of ESCC patients (Supplementary Table S1). Indeed, the results of these independent analyses mirror the findings of human tissue specimen-based analyses only in a few aspects, e.g., the detection of high levels of miR-21 or low levels of miR-375 in the serum of ESCC patients. Since these studies have used different (mainly small) groups of patients with different stages of cancer and treatment histories and have involved different techniques and approaches for the measurement of miRs, some debate clearly remains concerning the functional role of the miRs in esophageal cancers. To illustrate this, the following will describe two selected miRs, namely miR-21 and miR-375, in more detail, since several studies have been published for each of these two miRs.
miR-21 was one of the earlier discovered miRs, with a gene location on chromosome 17q23, i.e., a chromosomal region frequently altered by gene copy number gains, especially in BACs. The impact of altered miR-21 expression is mainly seen on tumor suppressor genes, such as PTEN, as shown in other cancer entities.
Several studies have shown that miR-21 is up-regulated in tumor cells of both ESCCs and BACs when compared with normal esophageal epithelium (Table 1; Feber et al. 2008; Akagi et al. 2011; Hamano et al. 2011; Garman et al. 2013; Wang et al. 2013). Feber et al. (2008) analyzed a small group of samples derived from fresh-frozen normal epithelium (n = 9), ESCCs (n = 10), and BACs (n = 10) with associated precursor lesions (n = 6) by microarray-based profiling and bioinformatics-clustering analyses. They detected an increase of up to five-fold of miR-21 expression in tumor versus normal esophageal epithelial cells (Feber et al. 2008). In a second microarray-based study focusing on BAC cases (Garman et al. 2013), samples of normal epithelium (n = 11), Barrett’s esophagus (n = 14), and esophageal adenocarcinoma (n = 11) were first screened by microarray-based analysis. Selected candidate miRs (including miR-21) were then validated by quantitative reverse transcription plus the polymerase chain reaction (q-RT-PCR) in a second group of cases (n = 18), but only for a comparison of normal squamous epithelium with BAC. Again, miR-21 was one of the candidate miRs up-regulated in BACs. Three subsequent studies focused on ESCCs. Akagi et al. (2011) used q-RT-PCR to measure five selected miRs and found that miR-21 was elevated in ESCCs, especially in those ESCCs with lymph node metastasis. In addition, Hamano et al. (2011) examined selected miRs by q-RT-PCR, but this time in ESCCs with previous neoadjuvant chemotherapy (n = 98). In this group of patients, miR-21 expression was also higher in tumor than in normal esophageal epithelial cells. Moreover, miR-21 expression was marginally associated with poor survival. Similarly, Wang et al. (2013) detected overexpression of miR-21, again by q-RT-PCR, in 16 cases.
Thus, the above studies indicate that the up-regulation of miR-21 occurs in both histotypes of esophageal cancers. This fits well with the known effect of miR-21 on tumor suppressors. However, interestingly, one large study has also detected miR-21 in ESCC “associated stroma” (Mathe et al. 2009). In their study, Mathe and colleagues investigated 100 BACs and 70 ESCCs, derived from various countries, by microarray-based expression profiling (smaller discovery case set of a total of n = 76) and validation in a separate case set (n = 94) by q-RT-PCR. The data support the finding that miR-21 is generally up-regulated in ESCCs and BACs compared with normal epithelium. However, in ESCCs, the data raised the issue that miR-21 expression in the normal esophageal epithelium was associated with poor patient prognosis; the authors here comment on a potential contribution of the “stroma” to miR-21 function. Indeed, miR-21 is also associated with the immune system (Tili et al. 2013). Since both ESCCs and BACs are composed not only of tumor cells, but also of, for example, various proportions of infiltrating lymphocytes, the specificity of the miR-21 measurements in all of the above studies needs to be reconsidered in terms of the degree of precision with which the tumor cells were selected for miR isolation and down-stream analyses. Without prior microdissection to enrich for pure cell populations of normal esophageal epithelial and invasive ESCC and BAC cells, some vagueness remains as to whether the measured miR-21 levels are tumor-associated or additionally driven by the level of the inflammatory component of the lesions.
As seen for miR-21, several studies have addressed miR-375 in ESCCs and BACs (Table 1). The gene of miR-375 is located on chromosome 2 and has been associated with the regulation of oncogenes such as MYC and TP53. In agreement with this, miR-375 expression has been found to be down-regulated in ESCCs and BACs, possibly thereby reflecting/supporting the activity of the tumor cells (Mathe et al. 2009; Li et al. 2011; Kong et al. 2012; Leidner et al. 2012; Wu et al. 2013). Kong et al. (2012) not only detected miR-375 down-regulation in ESCCs and its correlation to advanced stage ESCCs and to poor survival, but also showed that miR-375 could interact with the 3’-untranslated region of the insulin-like growth factor receptor 1 (IGFR) mRNA. This interaction led to the down-regulation of IGFR in in vitro model systems. Moreover, miR-375 and IGFR expression in ESCCs was inversely correlated. In the same study, the expression of miR-375 was shown to be regulated by promoter methylation (Kong et al. 2012), thereby also supporting a higher level of epigenetic complexity with the concept that DNA methylation and miRNA regulation are able to cross-talk (Lujambio et al. 2008; Suzuki et al. 2013).
DNA methylation in esophageal carcinogenesis and esophageal cancer therapy
As seen for other epithelial tumors, DNA methylation has been quite widely characterized by analysis of specific CpG-island promoter regions of selected candidate genes in DNA obtained from human tissue specimens of esophageal carcinomas and/or associated precursor lesions. The reader is referred to two recent excellent reviews of DNA methylation in ESSCs (Baba et al. 2013; Chen et al. 2013). An overview of studies for ESCCs, BACs, and associated pre-cursor lesions is provided in Table 2.
In ESCCs, hyper-methylation of genes associated with DNA replication (e.g., MGMT, FHIT), DNA repair genes (e.g., MLH1, MSH2), and genes involved in cell cycle progression, oncogenic signaling, cell differentiation, or motility (e.g., p16/CDKN2A, IGF2, E-Cadherin/CDH1, claudins) have all been associated with a poor prognosis (see references listed in Table 2). Of interest, the intestinal/columnar differentiation gene caudal-related homeobox gene (CDX2) was found to be silenced by DNA methylation in ESCCs (Guo et al. 2007), thus underlining the importance of CDX2 in the various mechanisms of carcinogenesis and diverse histology of ESCCs and BACs (see below).
Only a few studies have addressed or detected DNA hypo-methylation in ESCCs (Table 2), including a recent study regarding the global hypo-methylation of GADD45a, which is associated with poor tumor differentiation and prognosis (Wang et al. 2012). Moreover, the loss of imprinting of IGF2 has been recently described in ESCC, and this loss of methylation is associated with a shorter length of survival (Murata et al. 2013). In addition, LINE-1 element hypo-methylation is associated with poor prognosis (Iwagami et al. 2013) and is apparently detectable even in “normal” esophageal epithelium of patients with a strong smoking history (Shigaki et al. 2012).
In BACs, CpG promoter DNA methylation patterns of selected genes have been investigated, including the detection of DNA hyper-methylation in genes involved in cell cycle progression (p16/CDKN2A), in apoptosis (e.g., SFRP1), in the WNT/β-catenin pathway (e.g., APC), and in protease inhibitors (e.g., TIMPs; see references listed in Table 2). The use of candidate gene approaches has demonstrated DNA hypo-methylation in genes involved in intestinal/columnar differentiation (e.g., CDX1; Wong et al. 2005).
However, the most recent and broad data on DNA hypo-methylation in ESCCs and BACs (including precursor lesions) have evolved from two recent studies on microarray-based or genome-wide DNA methylation patterns (Alvarez et al. 2011; Lima et al. 2011). Lima and colleagues (2011) selected 10 cases of ESCCs and analyzed DNA methylation patterns of samples derived from fresh-frozen tissue specimens of invasive tumor cells, “tissue surrounding the tumor” (not further specified), and normal squamous epithelium. Samples were analyzed first by the Illumina GoldenGate methylation assay, focusing on 807 cancer-related genes, and candidates were then further confirmed by pyrosequencing in a larger cohort of about 100 ESCCs and 27 non-case-matched normal esophageal epithelial controls. The data revealed an increase of general DNA methylation from samples of normal epithelium via “surrounding tissue” to tumor, including methylation of genes observed previously in ESSCs (e.g., p16/CDKN2A, MGMT). Hyper-methylated genes that were specifically emphasized in the study in terms of tumor evolvement were BCL3, belonging to the inhibitory κB (IκB)-family and regulating NFκB activity, and TFF1, a trefoil factor family member (Lima et al. 2011). The functional consequences of altered BCL3 or TTF1 DNA methylation and protein expression remain to be elucidated. Moreover, details of the cell populations (mixtures) present in the samples denoted as “surrounding tumor” were lacking (Lima et al. 2011). In the second genome-wide DNA methylation study in BACs, Alvarez and colleagues (2011) used a small group of histologically proven fresh-frozen tissue samples of BACs and associated precursor lesions for parallel analysis of (1) genome-wide cytosine methylation, (2) RNA profiling, and (3) array-based comparative genomic hybridization (aCGH). DNA methylation analysis was based on the “HpaII tiny fragment Enrichment by Ligation-mediated PCR (HELP) assay”, and candidate loci were validated by mass spectrometry-based high-throughput quantitative methylation PCR analyses (Sequenome). Of note, this study revealed a predominance of DNA hypo-methylation rather than DNA hyper-methylation at early stages of BAC carcinogenesis (Alvarez et al. 2011). Moreover, the authors of this study detected DNA hypo-methylation in a series of genes associated with the immune system, such as chemokines (e.g., CXCL1, CXCL3). Another novel finding of Alvarez et al. (2011) was that, in BAC carcinogenesis, DNA methylation also occurred in genomic regions outside of CpG-islands. Furthermore, by combining DNA methylation, RNA profiling, aCGH analyses, and functional in vitro experiments, the study by Alvarez et al. (2011) nicely supports the fact that single candidate loci and/or DNA methylation analysis alone is insufficient to unravel the complex interaction between epigenetic, genetic, and functional consequences.
In this context, the cross-talk of DNA methylation and miR expression mentioned above (Lujambio et al. 2008; Suzuki et al. 2013) is further underlined by a recent study in ESSCs, showing by bisulfite-sequencing PCR (BSP) and methylation-specific PCR (MSP) that the low expression of several miRs (miR-34a, miR-34b/c, miR-129-2) is attributable to DNA methylation (Lujambio et al. 2008). Moreover, this can be reversed by treatment with the DNA methyltransferase (DNMT) inhibitor 5-aza-2′-deoxycytidine/decitabine (DAC).
Together, the above studies rise the question as to how DNA methylation itself is regulated by DNMTs, especially since (clinical) inhibitors to DNMTs are available (Christman 2002). Few studies have addressed DNMT expression or alterations in esophageal cancers (Kassis et al. 2006; Fan et al. 2010). In our own studies (unpublished data), we find frequent loss or down-regulation of DNMT1 in invasive tumor cells of ESSC (Fig. 2), which is accompanied by the reduction of 5me-Cytosine labeling. This effect is also seen in BACs, but here it is not as prominent (Fig. 2). Moreover, as seen by DNMT1 knockdown experiments in the above-mentioned study by Kassis et al. (2006), our recent data show that the inhibition of DNMTs by azacytidine in esophageal cancer cells induces the loss of cell viability, particularly in ESCCs (unpublished data).
DNA methylation in human tissue specimens of esophageal squamous cell carcinoma (ESCC) and Barrett’s adenocarcinoma (BAC). Serial analysis of DNA methyltransferase (DNMT1) expression and levels of 5-methyl-Cytosine (5me-Cyt) by immunohistochemistry of case-matched tissue specimens of ESCCs (a–i) and BACs (j–r), including normal esophageal epithelium (normal) and precursor lesions (dysplasia). Hematoxylin and eosin (HE) staining is shown in a–c for ESCC and in j–l for BAC. Note the loss of DNMT1 (f) and detectable 5me-Cytosine levels (i) in ESCC. Note also DNMT1 expression (o) and low 5me-Cytosine levels (r) in BAC. Bar 200 μm
In conclusion, although numerous individual genes regulated by DNA methylation have been identified, comprehensive in situ and functional analysis of DNA methylation and associated biological effects in ESSCs and BACs are urgently needed for a better understanding of their carcinogenesis from a molecular pathological view and for potential future work into the action of drugs targeting DNA methylation.
Histone modifications in esophageal carcinogenesis and esophageal cancer therapy
The investigation of histone modifications in human tissue specimens of esophageal cancers has been restricted to a series of studies exclusively on ESSCs and has involved the analysis of specific histone marks by immunohistochemical staining (Table 3). Two more comprehensive studies combining several histone marks in ESCCs have been performed. Tzao et al. (2009) have examined histone 3 lysine 18 (H3K18ac), acetylated histone 4 lysine 12 (H4K12ac), dimethylated histone 4 arginine 3 (H4R3me2), dimethylated histone 3 lysine 4 (H3K4me2), and trimethylated histone 3 lysine 27 (H3K27me3). In their study, correlations of histone marks have been detected with tumor differentiation (H3K18ac, H4K3me2, H3K27me3), with tumor stage and lymph node status (H3K27me3), and with improved prognosis (low H3K18ac, low H3K27me3). In the study by I and colleagues (2010), the levels of H3K18ac and H4R3me2 have been found to be of prognostic value. The genes and associated cellular functions regulated by the above-detected “aberrant” histone marks remain unknown, as chromatin immunoprecipitation (ChIP) from formalin-fixed and paraffin-embedded tissue specimens derived from diagnostic procedures is not yet routinely established (Fanelli et al. 2010).
In view of their potential for improved esophageal cancer treatment, HDACs are starting to play a more prominent role, since several inhibitors (with different specificities) are available (Bolden et al. 2013). Interestingly, a pubmed-based literature research retrieved only two studies that examined HDAC expression in esophageal cancers. In ESCCs, Toh and colleagues (2003) described the down-regulation of HDAC1 in ESCC as compared with normal squamous epithelium. In a far more comprehensive study on BACs, Langer and colleagues (2010) investigated about 180 BACs, with 2/3 being primary resected tumors and 1/3 having received prior chemotherapy treatment. In general, reduced HDAC1 and HDAC2 expression was observed in up to 50 % and 30 % of BACs, respectively. Only HDAC2 was associated with pathological parameters (tumor differentiation, lymph node stage), and neither HDAC1 nor HDAC2 showed a prognostic impact (Langer et al. 2010). In our recent study of ESCCs and BACs, including precursor lesions (unpublished data), we have detected generally prominent and strong HDAC1 and HDAC2 expression in basal cells/glands of normal esophageal epithelium, dysplastic lesions, and invasive tumor cells (Fig. 3). Tumor-cell-specific reduction of HDAC1 and HDAC2 is only seen in 10–20 % of ESCC or BAC cases. Interestingly, serial analysis of pan H3 acetylation reveals a similarly strong level of this mark in dysplasia and BAC (Fig. 3). In contrast, ESCC cells show a decrease of pan H3 acetylation, suggesting that HDAC activity, but not mere expression, is differentially altered (especially in ESCCs). Inhibition of HDACs might hence be a potential future tool for the therapeutic targeting of esophageal cancer. Indeed, preliminary evidence has been presented showing that the inhibition of HDAC1 and HDAC2 by the specific HDAC inhibitor depsipeptide/FK228 has an antitumor effect on ESCC cells in vitro and in xenograft models in vivo (Hoshino et al. 2005).
Histone alterations in human tissue specimens of esophageal squamous cell carcinoma (ESCC) and Barrett’s adenocarcinoma (BAC). Serial analysis of histone deacetylase 1 (HDAC1), histone deacetylase 2 (HDAC2) and pan histone H3 acetylation (pan-H3Ac) by immunohistochemistry of case-matched tissue specimens of ESCCs (a–l) and BACs (m–x), including normal esophageal epithelium and precursor lesions. Hematoxylin and eosin (HE) staining is shown in a–c for ESCC and m–o for BAC. Note the general strong expression of HDAC1, HDAC2, and strong levels of pan-H3Ac in BACs and weaker in ESCCs. Bar 200 μm
Thus, very little is still known about the role of histone modifications in esophageal cancer, clearly supporting further investigations.
Concluding remarks
Esophageal cancers still represent a major challenge of interdisciplinary oncology. Numerous translational approaches have yielded better insight into the genetic aberrations driving esophageal cancer carcinogenesis and progression, and the identification of novel (genetic) targets for therapeutic intervention is advancing. Recently, these aspects have been paralleled by a growing interest in epigenetics of esophageal cancers, with several exciting studies reporting the alterations of miRNAs, DNA methylation, and histone modifications in patient tissue specimens of esophageal carcinomas and associated precursor lesions. Increases in our knowledge of epigenetic regulation of esophageal cancers will clearly contribute potential further biomarkers and treatment options in future. The gained insight into the epigenetics and genetics of esophageal cancer might open a wide window for improved patient care.
References
Akagi I, Miyashita M, Ishibashi O, Mishima T, Kikuchi K, Makino H, Nomura T, Hagiwara N, Uchida E, Takizawa T (2011) Relationship between altered expression levels of MIR21, MIR143, MIR145, and MIR205 and clinicopathologic features of esophageal squamous cell carcinoma. Dis Esophagus 24:523–530
Alvarez H, Opalinska J, Zhou L, Sohal D, Fazzari MJ, Yu Y, Montagna C, Montgomery EA, Canto M, Dunbar KB, Wang J, Roa JC, Mo Y, Bhagat T, Ramesh KH, Cannizzaro L, Mollenhauer J, Thompson RF, Suzuki M, Meltzer SJ, Melnick A, Greally JM, Maitra A, Verma A (2011) Widespread hypomethylation occurs early and synergizes with gene amplification during esophageal carcinogenesis. PLoS Genet 7:e1001356
Baba Y, Watanabe M, Baba H (2013) A review of the alterations in DNA methylation in esophageal squamous cell carcinoma. Surg Today 43:1355–1364
Bolden JE, Shi W, Jankowski K, Kan CY, Cluse L, Martin BP, MacKenzie KL, Smyth GK, Johnstone RW (2013) HDAC inhibitors induce tumor-cell-selective pro-apoptotic transcriptional responses. Cell Death Dis 4:e519
Chen J, Kwong DL, Cao T, Hu Q, Zhang L, Ming X, Chen J, Fu L, Guan X (2013) Esophageal squamous cell carcinoma (ESCC): advance in genomics and molecular genetics. Dis Esophagus (in press)
Christman JK (2002) 5-Azacytidine and 5-aza-2′-deoxycytidine as inhibitors of DNA methylation: mechanistic studies and their implications for cancer therapy. Oncogene 21:5483–5495
Cook MB, Chow WH, Devesa SS (2009) Oesophageal cancer incidence in the United States by race, sex, and histologic type, 1977–2005. Br J Cancer 101:855–859
Fan H, Liu D, Qiu X, Qiao F, Wu Q, Su X, Zhang F, Song Y, Zhao Z, Xie W (2010) A functional polymorphism in the DNA methyltransferase-3A promoter modifies the susceptibility in gastric cancer but not in esophageal carcinoma. BMC Med 8:12
Fanelli M, Amatori S, Barozzi I, Soncini M, Dal Zuffo R, Bucci G, Capra M, Quarto M, Dellino GI, Mercurio C, Alcalay M, Viale G, Pelicci PG, Minucci S (2010) Pathology tissue-chromatin immunoprecipitation, coupled with high-throughput sequencing, allows the epigenetic profiling of patient samples. Proc Natl Acad Sci U S A 107:21535–21540
Feber A, Xi L, Luketich JD, Pennathur A, Landreneau RJ, Wu M, Swanson SJ, Godfrey TE, Litle VR (2008) MicroRNA expression profiles of esophageal cancer. J Thorac Cardiovasc Surg 135:255–260
Garman KS, Owzar K, Hauser ER, Westfall K, Anderson BR, Souza RF, Diehl AM, Provenzale D, Shaheen NJ (2013) MicroRNA expression differentiates squamous epithelium from Barrett’s esophagus and esophageal cancer. Dig Dis Sci 58:3178–3188
Guo M, House MG, Suzuki H, Ye Y, Brock MV, Lu F, Liu Z, Rustgi AK, Herman JG (2007) Epigenetic silencing of CDX2 is a feature of squamous esophageal cancer. Int J Cancer 121:1219–1226
Hamano R, Miyata H, Yamasaki M, Kurokawa Y, Hara J, Moon JH, Nakajima K, Takiguchi S, Fujiwara Y, Mori M, Doki Y (2011) Overexpression of miR-200c induces chemoresistance in esophageal cancers mediated through activation of the Akt signaling pathway. Clin Cancer Res 17:3029–3038
Hoshino I, Matsubara H, Hanari N, Mori M, Nishimori T, Yoneyama Y, Akutsu Y, Sakata H, Matsushita K, Seki N, Ochiai T (2005) Histone deacetylase inhibitor FK228 activates tumor suppressor Prdx1 with apoptosis induction in esophageal cancer cells. Clin Cancer Res 11:7945–7952
I H, Ko E, Kim Y, Cho EY, Han J, Park J, Kim K, Kim DH, Shim YM (2010) Association of global levels of histone modifications with recurrence-free survival in stage IIB and III esophageal squamous cell carcinomas. Cancer Epidemiol Biomark Prev 19:566–573
Iwagami S, Baba Y, Watanabe M, Shigaki H, Miyake K, Ishimoto T, Iwatsuki M, Sakamaki K, Ohashi Y, Baba H (2013) LINE-1 hypomethylation is associated with a poor prognosis among patients with curatively resected esophageal squamous cell carcinoma. Ann Surg 257:449–455
Kassis ES, Zhao M, Hong JA, Chen GA, Nguyen DM, Schrump DS (2006) Depletion of DNA methyltransferase 1 and/or DNA methyltransferase 3b mediates growth arrest and apoptosis in lung and esophageal cancer and malignant pleural mesothelioma cells. J Thorac Cardiovasc Surg 131:298–306
Kong KL, Kwong DL, Chan TH, Law SY, Chen L, Li Y, Qin YR, Guan XY (2012) MicroRNA-375 inhibits tumour growth and metastasis in oesophageal squamous cell carcinoma through repressing insulin-like growth factor 1 receptor. Gut 61:33–42
Langer R, Mutze K, Becker K, Feith M, Ott K, Hofler H, Keller G (2010) Expression of class I histone deacetylases (HDAC1 and HDAC2) in oesophageal adenocarcinomas: an immunohistochemical study. J Clin Pathol 63:994–998
Leidner RS, Ravi L, Leahy P, Chen Y, Bednarchik B, Streppel M, Canto M, Wang JS, Maitra A, Willis J, Markowitz SD, Barnholtz-Sloan J, Adams MD, Chak A, Guda K (2012) The microRNAs, MiR-31 and MiR-375, as candidate markers in Barrett’s esophageal carcinogenesis. Genes Chromosomes Cancer 51:473–479
Li X, Lin R, Li J (2011) Epigenetic silencing of microRNA-375 regulates PDK1 expression in esophageal cancer. Dig Dis Sci 56:2849–2856
Lima SC, Hernandez-Vargas H, Simao T, Durand G, Kruel CD, Le Calvez-Kelm F, Ribeiro Pinto LF, Herceg Z (2011) Identification of a DNA methylome signature of esophageal squamous cell carcinoma and potential epigenetic biomarkers. Epigenetics 6:1217–1227
Lin DC, Du XL, Wang MR (2009) Protein alterations in ESCC and clinical implications: a review. Dis Esophagus 22:9–20
Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D, Montuenga LM, Rossi S, Nicoloso MS, Faller WJ, Gallagher WM, Eccles SA, Croce CM, Esteller M (2008) A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci U S A 105:13556–13561
Mathe EA, Nguyen GH, Bowman ED, Zhao Y, Budhu A, Schetter AJ, Braun R, Reimers M, Kumamoto K, Hughes D, Altorki NK, Casson AG, Liu CG, Wang XW, Yanaihara N, Hagiwara N, Dannenberg AJ, Miyashita M, Croce CM, Harris CC (2009) MicroRNA expression in squamous cell carcinoma and adenocarcinoma of the esophagus: associations with survival. Clin Cancer Res 15:6192–6200
Mohammad HP, Baylin SB (2010) Linking cell signaling and the epigenetic machinery. Nat Biotechnol 28:1033–1038
Murata A, Baba Y, Watanabe M, Shigaki H, Miyake K, Ishimoto T, Iwatsuki M, Iwagami S, Yoshida N, Oki E, Morita M, Nakao M, Baba H (2013) IGF2 DMR0 methylation, loss of imprinting, and patient prognosis in esophageal squamous cell carcinoma. Ann Surg Oncol 21:1166–1174
Pennathur A, Gibson MK, Jobe BA, Luketich JD (2013) Oesophageal carcinoma. Lancet 381:400–412
Shigaki H, Baba Y, Watanabe M, Iwagami S, Miyake K, Ishimoto T, Iwatsuki M, Baba H (2012) LINE-1 hypomethylation in noncancerous esophageal mucosae is associated with smoking history. Ann Surg Oncol 19:4238–4243
Suzuki H, Maruyama R, Yamamoto E, Kai M (2013) Epigenetic alteration and microRNA dysregulation in cancer. Front Genet 4:258
Tili E, Michaille JJ, Croce CM (2013) MicroRNAs play a central role in molecular dysfunctions linking inflammation with cancer. Immunol Rev 253:167–184
Toh Y, Yamamoto M, Endo K, Ikeda Y, Baba H, Kohnoe S, Yonemasu H, Hachitanda Y, Okamura T, Sugimachi K (2003) Histone H4 acetylation and histone deacetylase 1 expression in esophageal squamous cell carcinoma. Oncol Rep 10:333–338
Tzao C, Tung HJ, Jin JS, Sun GH, Hsu HS, Chen BH, Yu CP, Lee SC (2009) Prognostic significance of global histone modifications in resected squamous cell carcinoma of the esophagus. Mod Pathol 22:252–260
Wang B, Yin BL, He B, Chen C, Zhao M, Zhang W, Xia ZK, Pan Y, Tang J, Zhou X, Yin N (2012) Overexpression of DNA damage-induced 45 alpha gene contributes to esophageal squamous cell cancer by promoter hypomethylation. J Exp Clin Cancer Res 31:11
Wang N, Zhang CQ, He JH, Duan XF, Wang YY, Ji X, Zang WQ, Li M, Ma YY, Wang T, Zhao GQ (2013) MiR-21 down-regulation suppresses cell growth, invasion and induces cell apoptosis by targeting FASL, TIMP3, and RECK genes in esophageal carcinoma. Dig Dis Sci 58:1863–1870
Wong NA, Wilding J, Bartlett S, Liu Y, Warren BF, Piris J, Maynard N, Marshall R, Bodmer WF (2005) CDX1 is an important molecular mediator of Barrett’s metaplasia. Proc Natl Acad Sci U S A 102:7565–7570
Wu X, Ajani JA, Gu J, Chang DW, Tan W, Hildebrandt MA, Huang M, Wang KK, Hawk E (2013) MicroRNA expression signatures during malignant progression from Barrett’s esophagus to esophageal adenocarcinoma. Cancer Prev Res (Phila) 6:196–205
Zhang HY, Spechler SJ, Souza RF (2009) Esophageal adenocarcinoma arising in Barrett esophagus. Cancer Lett 275:170–177
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 62 kb)
Rights and permissions
About this article
Cite this article
Ahrens, T.D., Werner, M. & Lassmann, S. Epigenetics in esophageal cancers. Cell Tissue Res 356, 643–655 (2014). https://doi.org/10.1007/s00441-014-1876-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-014-1876-y