Abstract
Over the past few decades, the knowledge on genetic defects causing mental retardation has dramatically increased. In this review, we discuss the importance of balanced chromosomal translocations in the identification of genes responsible for mental retardation. We present a database-search guided overview of balanced translocations identified in patients with mental retardation. We divide those in four categories: (1) balanced translocations that helped to identify a causative gene within a contiguous gene syndrome, (2) balanced translocations that led to the identification of a mental retardation gene confirmed by independent methods, (3) balanced translocations disrupting candidate genes that have not been confirmed by independent methods and (4) balanced translocations not reported to disrupt protein coding sequences. It can safely be concluded that balanced translocations have been instrumental in the identification of multiple genes that are involved in mental retardation. In addition, many more candidate genes were identified with a suspected but (as yet?) unconfirmed role in mental retardation. Some balanced translocations do not disrupt a protein coding gene and it can be speculated that in the light of recent findings concerning ncRNA’s and ultra-conserved regions, such findings are worth further investigation as these potentially may lead us to the discovery of novel disease mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mental retardation is predominantly characterized by an overall intelligence coefficient (IQ) under 70 (Luckasson et al. 2002). It is a common, lifelong disorder, with an estimated frequency of 1–3% of the human population (Leonard and Wen 2002; Roeleveld et al. 1997; WHO 2001). Approximately 0.4% of the population is reported to be moderate/severely handicapped with an IQ < 50. The cause of the mental handicap remains unknown in at least half of the cases. Since genetic causes attributes to about half of the known aetiologies, it is generally believed that about 50% of all mental retardation cases have a genetic origin (Curry et al. 1997; Stevenson et al. 2003; Winnepenninckx et al. 2003). Mutations in several genes have been reported to cause mental retardation, but it has been estimated that the disruption of thousands of different genes may result in mental retardation and the great majority of those are unknown as yet (Ropers 2007).
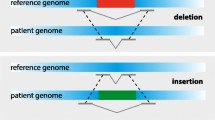
With the exception of families showing X-linked or recessive inheritance, familial clustering is relatively rare in mental retardation pedigrees due to the fact that patients seldom reproduce. Linkage studies that have been so helpful in providing candidate regions to look for disease genes in other types of disorders are therefore rarely feasible in mental retardation research. In order to identify novel genes, we are thus dependent on other hints where to look for disease genes in the genome. One of those leads are balanced translocations, the product of a recombination event between two chromosomes, without loss of any chromosomal material (Fig. 1). These have been postulated to play a causative role in mental retardation disorders a long time ago (Breg et al. 1972), a hypothesis that has been verified by subsequent studies (Jacobs 1974). Balanced translocations are found as de novo events in 1/2,000 live births and are associated with a two- to three-fold increase in risk of inborn abnormalities (Warburton 1991). There are no recent reports on the frequency of balanced translocations in the mentally handicapped, but in one study from 1977, 7 balanced de novo translocations were found in a population of 455 handicapped patients, suggesting that a balanced translocation is responsible for 1.5% of cases of mental retardation (Funderburk et al. 1977). This frequency is in line with a second study reporting that 41 out of 153 balanced translocations were associated with MR and/or congenital malformations (Fryns et al. 1986). A translocation may disrupt a single gene only, and identification of the disrupted sequences may point to a candidate mental retardation gene. A number of genes have been cloned after analysis of such breakpoints in carriers of a balanced translocation (listed in Tables 1a, b, 2). Seemingly contradictory, many balanced translocations have been cloned where no conclusive evidence on the role of the disrupted gene in the aetiology of mental retardation could be obtained or where no gene has been identified at any of the breakpoints (listed in Tables 3, 4).
The aim of this review is to highlight the importance of balanced, gene disrupting translocations in the identification of mental retardation genes. To achieve this, a restrictive search was performed on three major databases of genetic diseases. The Decipher database collects clinical information about submicroscopic imbalances, such as microdeletions, microduplications, microinversions and translocations. The database was accessed on 26 March 2008 and referenced literature for the described syndromes were screened at that time for description of cases with mental retardation and balanced translocations. The term “developmental delay” was intentionally left out of the criteria, because of the much broader definition compared to “mental retardation”. The “Online Mendelian Inheritance In Men” database, provided by NCBI, contains descriptions and research history on most known genetic diseases in humans. It was queried on 24 April 2008 for the terms ‘mental retardation’ AND ‘translocation’. The resulting 230 hits were screened for relevance and redundancy. As a final repository of cytogenetic information, the ‘Mendelian Cytogenetics Network Online Database’ (MCNdb) was accessed on 12 May 2008. With ‘Mental Retardation’ as criterion, 95 entries were retrieved, out of which the de novo, simple balanced translocations were retained. In addition, a few recent individual studies that were not retrieved were included. All cases thus identified are listed in Tables 1, 2, 3, and 4, where multiple translocations through the same gene have been grouped.
Results
A total of 92 individual balanced translocations in patients with a mental handicap were identified. Based on the phenotypic information provided, the cases were split into four categories. A first category was created of patients with a balanced translocation localized within regions of known microdeletion syndromes (Table 1a, b). A second category consists of balanced translocations that led to the unambiguous identification of mental retardation genes, as confirmed by independent methods (Table 2). A third category was created for balanced translocations where candidate genes have been identified at the breakpoints, but where the role of these genes in mental retardation has not been confirmed by independent methods (Table 3). Finally, balanced translocations in patients where no candidate genes were identified at the breakpoints remain as a fourth category (Table 4).
Balanced translocations pinpoint causative genes in contiguous gene syndromes
Microdeletion syndromes are caused by cytogenetically invisible deletions at specific positions in the genome. Classical examples of such mental retardation microdeletion syndromes include the Prader–Willi/Angelman Syndromes (del 15q11-q13), the DiGeorge/velocardiofacial Syndrome (del 22q11.2) and the Williams–Beuren Syndrome (del 7q11.23). On the genomic level, these recurrent deletions are often caused by non-allelic homologous recombination (NAHR) between nearly identical repeat blocks flanking the disease region. More recently, several new microdeletion syndromes have been identified as a result of the rapid development of whole genome copy number variation analysis platforms (Ballif et al. 2007; Koolen et al. 2006; Sharp et al. 2006, 2008; Shaw-Smith et al. 2006). Microdeletion syndromes near the telomeres are commonly referred to as subtelomeric disorders (de Vries et al. 2003). Subtelomeric deletions have the deletion of a series of genes in common with the remainder of the microdeletion syndromes, but the chromosome rearrangements are caused by much more diverse mechanisms and generally result in a more variable size of the deletion (Rooms et al. 2007). Balanced translocations have led to the identification of causative genes within some of the critical deletion regions (Table 1).
Prader–Willi is clinically associated with mental retardation, decreased foetal activity, neonatal hypotonia and feeding difficulties, hyperphagia with obesity, hypogonadism, short stature and small hands and feet (Sun et al. 1996). About 70% of PWS patients have a microdeletion of 3–4 Mb on the paternally derived chromosome 15q11.2 (maternal deletions result in Angelman Syndrome due to the fact that this chromosomal region is subject to genomic imprinting). Most of the remaining cases are due to maternal uniparental disomy, and a few sporadic cases have imprinting defects due to mutations in the imprinting centre.
No intact SNRP mRNA, a gene within the critical region, was detected in a patient with a de novo balanced translocation, and it was thus suggested that full mRNA was necessary for a normal phenotype (Sun et al. 1996) (Fig. 2). Since SNRP expression was unaffected in a second patient with a translocation just downstream of the SNRP-gene (Schulze et al. 1996), the focus of attention shifted downstream to IPW/PAR1, a gene that was silenced in this patient. Expression of SNURF/SNRP, consisting of the SNRP gene together with newly identified exons, extending the original transcript was detected in a translocation patient with PWS, but expression of IPW and PAR1 as well as a novel C/D box snoRNA were absent. Together with atypical microdeletions in this region, these findings put the C/D box snoRNA’s forward as the main candidate (Wirth et al. 2001). Subsequently, the SNRPN locus was shown to consist of 148 exons, spanning from the imprinting centre 5′ of SNURF to the UBE3 gene. The transcript was shown to contain IPW as well as PAR1, besides several snoRNA’s inside its introns. The snoRNA genes are present as two multi-snoRNA gene clusters (PWCR1/HBII-85 and HBII-52), three single-copy snoRNA genes (HBII-436/13/437) and two copies of an additional snoRNA gene (HBII-438A/B) (Runte et al. 2001).
Schematical overview of the Prader–Willi/Angelman region. SNURF-SNRPN exons are depicted in blue, snoRNA’s in red and UBE3A exons in green. Described balanced translocations disrupting this region are visualised by black arrows. The locations of the incorporated transcripts PAR-5, IPW and PAR-1 are shown in black
Identification of this long and extremely complex transcribed region subsequently forced a re-evaluation of the previously described cases. The cases described by Conroy, Schulze, Wirth and Kuslich (after reassessment) were in accordance with a critical involvement of the snoRNA clusters, showing no expression of elements downstream of the original SNRP transcript (Conroy et al. 1997; Gallagher et al. 2002; Kuslich et al. 1999; Schulze et al. 1996; Wirth et al. 2001). These observations, together with the assumption of a fusion product in the patient described by Sun et al. (1996) in combination with findings from a deletion case (Greger et al. 1993), allowed a new critical region of 121 kb to be defined, containing the PWCR1/HBII-85 snoRNA cluster and the HBII-438A snoRNA as the only putative functional genes (Gallagher et al. 2002). A translocation detected in a PWS patient, disrupting intron 17 and abolishing PWCR1/HBII-85 and HBII-438A expression (Schüle et al. 2005), and an atypical deletion in another patient, ranging from IPW, which is just 3′ of HBII-85, to UBE3A, without PWS (Runte et al. 2001) confirmed the involvement of these snoRNA’s in the core phenotype of PWS. Conclusive evidence was recently published by Sahoo et al. (2008), describing a PWS patient with a small 174-Kb deletion involving only the PWCR1/HBII-85 cluster. This concluded a search started more than a decade ago, guided by six balanced translocations and leading towards new discoveries in what is considered one of the most complex regions in the human genome.
A more straightforward illustration of the importance of balanced translocations in the identification of mental retardation genes is the identification of the causative gene in the 9q subtelomeric deletion syndrome. After thorough clinical investigation, it was noticed that the phenotype of a patient with a t(X;9)(p11.23;q34.3) showed a striking similarity with that of patients with the subtelomeric del(9qter) syndrome. EHMT1, one of the candidate genes in the commonly deleted region of patients with a subtelomeric deletion was disrupted by the translocation, and considered a good candidate gene (Kleefstra et al. 2005). Twenty-three patients without detectable chromosomal abnormalities and with phenotypic characteristics of the 9q subtelomeric deletion syndrome were further screened for EHMT1 aberrations and mutations were identified in two patients (Kleefstra et al. 2006). This confirmed haplo-insufficiency of EHMT1 as the main cause of the del(9q) syndrome. Similarly, studying patients with a syndromic form of Hirschprung disease known as Mowat–Wilson, two independent labs each found a balanced translocation disrupting the ZEB2 gene. During subsequent mutation screening in patients with the clinical symptoms of Mowat–Wilson, they both confirmed the gene’s involvement in the phenotype of this microdeletion syndrome (Cacheux et al. 2001; Wakamatsu et al. 2001).
Identification of novel mental retardation genes through balanced translocations
One of the first identified mental retardation genes was characterized after studying a female patient with non-syndromic mental retardation and a balanced translocation t(X;12)(q11;q15) (Bienvenu et al. 1997). In contrast to syndromic patients, who are characterized by additional clinical or biochemical features, non-syndromic patients are characterized by mental retardation per se. In females, inactivation of one of the X-chromosomes by Lyonisation is usually a stochastic process. However, as a result of the X:autosome translocation, the intact X-chromosome is preferentially inactivated, silencing one copy of the gene by the X:autosome translocation, and the other by Lyonisation. A gene called oligophrenin-1 (OPHN1) was identified at the translocation breakpoint at Xq11 in the patient described before (Billuart et al. 1998). Disruption of the gene was considered a good indication for involvement in the mental retardation of the patient, but no conclusive evidence was found, and a search for mutations in this gene was initiated in additional patients. The search was facilitated by the availability of large families with X-linked mental retardation mapping to this chromosomal region (Chiurazzi et al. 2008). In one out of four families examined, a frameshift mutation was identified, proving the involvement of oligophrenin-1 in mental retardation. Following basically the same strategy, four additional non-syndromic mental retardation genes have been identified (see Table 2).
Examples of syndromic MRX gene identifications include the discovery of the DCX gene as the causative gene in lissencephaly. The identification of a patient with a balanced (X;2) translocation narrowed down the candidate region to 1 cM (Ross et al. 1997) and greatly facilitated cloning of the gene. The search for mutations in the candidate gene that mapped into this interval was successful in three out of three patients with an X-linked lissencephaly phenotype. However, it is of interest to note that not all cases of balanced autosome/X translocations are due to disruption of the gene that lies on the X-chromosome. The mental retardation in a female patient with a balanced t(X;9)(p11.23;q34.3) translocation that led to the discovery of EHMT1 as the causative gene in the 9q subtelomeric deletion syndrome, was originally proposed to be caused by ZNF81 deficiency, a gene located on the X-chromosome and disrupted by the same translocation (Kleefstra et al. 2004).
Only a handful of autosomal genes have been associated with MR phenotype, mostly involving syndromic forms. Two translocations disrupting the GLI3 gene in patients with Greig Syndrome, initiated mutation analysis on a cohort of patients with a phenotype suggestive of the disorder (Kruger et al. 1989; Tommerup and Nielsen 1983; Wild et al. 1997). Point mutations in GLI3 were shown to result in Greig Syndrome, while frameshift mutations result in a different phenotype, namely the Pallister–Hall Syndrome (Kang et al. 1997). While this is the only example of a new autosomal candidate gene identification, other translocations added to the mutation spectrum in known MR genes. Examples are a t(1;4)(q31;p15.3) translocation disrupting the ASPM gene involved in MCPH5, a t(2;5)(q24.3;q34) translocation disrupting SCN1A associated to MR and SMEI, a t(7;12)(q22;p13) translocation disrupting RELN leading to LIS2, or a t(18;20)(q21.1;q11.2) translocation causing the Pitt–Hopkins Syndrome by disruption of TCF4 (Kalscheuer et al. 2008a; Moller et al. 2008b; Pichon et al. 2004; Zaki et al. 2007). The identification of FOXL2, a transcription factor as the candidate gene responsible for the blepharophimosis/ptosis/epicanthus inversus syndrome (BPES) was identified partially based on translocation data (Boccone et al. 1994; de Almeida et al. 1993; Praphanphoj et al. 2000). Deletion of FOXL2 causes BPES with MR (De Baere et al. 2003), whereas point mutations in the same gene cause BPES without MR.
Identification of candidate genes at breakpoints
In contrast to the causative genes identified at breakpoints that were mentioned above, numerous genes have been identified at translocation breakpoints in patients for which the role in disease could not be confirmed by mutation analysis as yet. Among those, the AUTS2 gene was found disrupted in four unrelated MR patients with balanced translocations (Kalscheuer et al. 2007; Sultana et al. 2002). This apparent correlation of a disruption to MR makes it a strong candidate for a critical role in normal brain function. However, no missense or nonsense mutations have been found in isolated patients. Another example includes the ARHGEF9 gene, encoding collybistin, a gene involved in cell signalling, that was found disrupted at a translocation breakpoint in a female with an X:autosome translocation (Kalscheuer et al. 2008b) The same gene was also found disrupted in a patient with a paracentric inversion of the X-chromosome (Marco et al. 2008). In the latter study, nearly 500 patients with unexplained mental retardation and suggestive X-linked inheritance were screened for mutations, but no unambiguous mutations were identified. A third example is the KIAA1202 gene, found disrupted in two independent female patients with balanced X:autosome translocations (Hagens et al. 2006). Again, a search for mutations in X-linked families revealed no evidence for a clear mutation, though sequence alterations were identified. Are AUTS2, ARHGEF9 and KIAA1202 mental retardation genes or not? For now, this remains a question on interpretation of the data. Some authors do include these genes in the listing for mental retardation genes, while others reserve the term mental retardation gene for genes that have been confirmed by mutation analysis. In addition to genes that have been reported disrupted in multiple patients, numerous reports exist on genes that have been found disrupted in a single case (Table 3). Examples include the DOCK8 gene on 9p24.3 and PAFAH1B3 on 19q13.1 (Griggs et al. 2008).
Many translocation breakpoints do not disrupt a candidate gene
The last and potentially largest category of balanced translocations, however, consists of breakpoints that have been identified, but where no coding gene has been identified at any of the breakpoints. This does not only apply to the cases that have been reported several years ago, when genomic maps and molecular technologies did not allow gene identification in many cases, but also to cases reported most recently. Published examples are listed in Table 4, but the number of reported cases is presumably only a fraction of the cases that are detected. The great majority of these class has likely not been reported as no gene was found interrupted and it is generally assumed that only disturbance of a protein coding gene can contribute to disease. Whether this assumption is correct is unknown at present. For instance, we ourselves recently cloned a balanced t(2;6)(p15;p22.3) de novo translocation in a patient with West Syndrome who showed no other chromosomal abnormalities. West Syndrome, also known as the “Infantile Spasms Syndrome” is a genetically heterogeneous mental retardation syndrome associated with epilepsy and hypsarithmia. Mutations in ARX and CDKL5 are both known to be causative, but no autosomal candidate genes have been identified so far (Kalscheuer et al. 2003; Kato 2006). Using marker analysis on flow-sorted derivatives, the breakpoint region provided by FISH analysis was refined and ultimately sequenced. Spanning the breakpoint on chromosome 6, a non coding RNA transcript, expressed in the foetal brain, was detected. Although such findings are hard to interpret at the moment, these should not be discarded.
Discussion
Over the past years, different labs have used balanced translocations to identify causative genes for mental retardation. The results ranged from the identification of mental retardation genes, like OPHN1, to characterisation of complex regulatory domains as seen for the Prader–Willi Syndrome. It can thus be safely concluded that balanced translocations are useful tools for the identification responsible for mental retardation genes including the identification of causative genes within regions deleted in contiguous gene syndromes. Some microdeletion syndromes have been “reduced” to a single gene disorder, with mutations in a single gene responsible for most of the clinical symptoms, while in other microdeletion syndromes, such genes have not been identified, neither through a balanced translocation nor through any other method. It can be questioned whether single candidate genes exist for all contiguous gene syndromes, or whether some microdeletion syndromes are truly contiguous, and as such the consequence of the combined absence of a multitude of genes.
Balanced translocations have been most successful in characterising candidate genes for X-linked non-syndromic mental retardation. Of the 16 genes identified for this specific type of a mental handicap, 5 identifications have been guided by a balanced translocation (see Table 2) (Chiurazzi et al. 2008). Genes responsible for autosomal, syndromic mental retardation have also been identified at the position of translocation breakpoints observed in a single patient. In both cases, the identification of mutations in the causative genes was facilitated by an enrichment of the initial study population based on linkage data or phenotype, enabling selection of a population of patients significantly enriched for the specific phenotype, enormously increasing the odds ratio of finding a disease related mutation. In X-linked mental retardation, large collections of nuclear families have been assembled and the number of genes potentially involved in any specific X-linked disorder has been further reduced by linkage analysis (Chiurazzi et al. 2008). In syndromic disorders, a selection on the basis of the clinical phenotype seems a prerequisite for gene cloning, as only genes responsible for specific disorders characterized by a combination of mental retardation and a discriminating set of congenital abnormalities have been successfully identified up till now. Sometimes, mutations in genes involved in syndromic mental retardation were also found in patients with non-syndromic mental retardation. Thus, studying syndromal cases may also shed light on the diverse genetic defects that may lead to brain dysfunction in general.
In absence of clear selection criteria, mutations in very few, if any autosomal genes responsible for mental retardation have been reported despite the fact that numerous gene disrupting translocations have been identified. There are multiple plausible explanations for this discrepancy. First, rather than the gene disrupted by the balanced translocation, a positional effect affecting the dosage of a multitude of genes on the rearranged chromosomes is causative for the mental handicap. A second possibility is that disruption of the function of the gene, either by truncation or by fusion with a second transcript, is causative for the observed clinical symptoms, but mutations are extremely rare and can only be detected after screening populations of thousands and thousands of patients. Using current technologies, screening of such large populations is not feasible. Thirdly, there is the possibility that the balanced translocation might actually be coincidental, and the patient carries a causative mutation or microaberration somewhere else in the genome. Multiple studies on the nature of balanced chromosomal aberrations using array based methods have revealed that novel microaberrations occur in about one-third of the investigated subjects. In more than half of these cases, the aberrations are located at the seemingly balanced breakpoints and may take away a series of genes, making them less feasible for candidate gene identification (Baptista et al. 2008; Fantes et al. 2008; Gribble et al. 2005; Higgins et al. 2008). The remaining cases usually show no aberrations at the array level of analysis, but may contain imbalances of a few bases when sequenced (Higgins et al. 2008). Despite the fact that most of the reported balanced translocations are not analyzed at such a high resolution, it has to be stressed that candidate genes identified so far were directly disrupted by the breakpoint.
The last category, in which no coding gene has been identified at the breakpoints, may potentially become the most interesting one. Only 1–2% of the human genome encodes for proteins, and the function of the remaining DNA, if any, is largely unknown (Lander et al. 2001). For some time this remaining DNA was seen as junk, but with recent findings on ultra-conserved regions and ncRNA’s, these noncoding regions are being intensely studied. So although this last category may have seemed uninteresting in the past, the findings on possible physiological roles of ncRNA’s can place them back into the centre of attention (Mercer et al. 2008; Satterlee et al. 2007). Several forms of ncRNA’s have been identified, and have been shown involved in neuronal development, gene expression and RNA editing. The mechanism on which their function is based is not yet fully understood, and the possibility of chromatin remodelling is currently under active investigation (Attanasio et al. 2008). Furthermore, the involvement of snoRNA’s in the phenotype of PWS has stated a clear example of their putative role in MR syndromes. Characterisation of expressed RNA molecules spanning breakpoints in patients with specific phenotypes may help increase our understanding of this new family of potentially important genetic players.
Collecting the available knowledge on balanced translocations, we noticed that this information is scattered across different databases such as OMIM, DECIPHER, ECARUCA, MCNdb, or DGV and that some cases described in individual research papers accessible through PubMed are not listed in any of these. A comprehensive record curated by a central dedicated database of all known balanced disruptions, including high-resolution mapping of copy number variants, mapping data of the breakpoint and a detailed description of the clinical phenotype would be an enormous advantage. This should allow future researchers to relate the translocation breakpoints with aberrations found using high-resolution copy number analysis. Novel technology, including the array-CGH and dense SNP analysis is creating a continuous stream of new chromosomal aberrations involved in MR and to be able to correlate the aberrations found with these technologies with genes disrupted by translocation breakpoints would be of enormous help in the interpretation of these new findings. When correlating genomic imbalances to a phenotype, it is extremely important to have a close cooperation between the molecular laboratories and the clinic in order to be able to cluster patients according to their phenotype. This is clearly illustrated by the translocation that led to identification of EHMT1 in del(9q) as described before. As shown in Table 1b, three genes are thought to be involved in the phenotype of the contiguous 3pter deletion syndrome based on translocations, and ITPN1 was proposed based on an atypical deletion patient (Cargile et al. 2002). Although all three gene disruptions seem to cause some aspects of the phenotype, none of them can explain the whole set of mental and growth retardation and dysmorphic features, and none of the three is confirmed by independent mutations in similar patients (Endris et al. 2002; Cargile et al. 2002; Frints et al. 2003; Fernandez et al. 2004). The observation of partial phenotypes for single gene disruptions points to a contiguous syndrome for del(3pter), but to the best of our knowledge, no explicit decision has been made for the 2q37 deletion syndrome, on the whether or not it is a contiguous syndrome. Nearly a 100 patients are characterized, and yet no general genotype to phenotype link has been established (Falk and Casas 2007).
With the development of recent techniques, the cloning of many more breakpoints can be awaited in the years to come. For instance, next generation sequencing technology has already been proven to be a reliable method to identify the breakpoints from flow-sorted or laser-dissected chromosomes in a single experiment (Kundakovic et al. 2008). This technique can thus substantially decrease the efforts necessary to clone translocation breakpoints. A further breakthough that can be anticipated is the massive parallel sequencing of multiple candidate genes in thousands of patients with mental retardation. This may enable the detection of mutations in genes that are mutated in a very low percentage of patients. Thus, novel technology in combination with an improved database annotation of disease related genomic abnormalities hold great promise for the elucidation of the role of the sequences interrupted in patients with mental retardation in the near future.
References
Attanasio C, Reymond A, Humbert R, Lyle R, Kuehn MS, Neph S, Sabo PJ, Goldy J, Weaver M, Haydock A, Lee K, Dorschner M, Dermitzakis ET, Antonarakis SE, Stamatoyannopoulos JA (2008) Assaying the regulatory potential of mammalian conserved non-coding sequences in human cells. Genome Biol 9:R168
Attree O, Olivos IM, Okabe I, Bailey LC, Nelson DL, Lewis RA, McInnes RR, Nussbaum RL (1992) The Lowe’s oculocerebrorenal syndrome gene encodes a protein highly homologous to inositol polyphosphate-5-phosphatase. Nature 358:239–242
Ayme S, Mattei MG, Mattei JF, Giraud F (1979) Abnormal childhood phenotypes associated with the same balanced chromosome rearrangements as in the parents. Hum Genet 48:7–12
Ballif BC, Hornor SA, Jenkins E, Madan-Khetarpal S, Surti U, Jackson KE, Asamoah A, Brock PL, Gowans GC, Conway RL, Graham JM, Medne L, Zackai EH, Shaikh TH, Geoghegan J, Selzer RR, Eis PS, Bejjani BA, Shaffer LG (2007) Discovery of a previously unrecognized microdeletion syndrome of 16p11.2–p12.2. Nat Genet 39:1071–1073
Baptista J, Mercer C, Prigmore E, Gribble SM, Carter NP, Maloney V, Thomas NS, Jacobs PA, Crolla JA (2008) Breakpoint mapping and array CGH in translocations: comparison of a phenotypically normal and an abnormal cohort. Am J Hum Genet 82:927–936
Bhalla K, Phillips HA, Crawford J, McKenzie OL, Mulley JC, Eyre H, Gardner AE, Kremmidiotis G, Callen DF (2004) The de novo chromosome 16 translocations of two patients with abnormal phenotypes (mental retardation and epilepsy) disrupt the A2BP1 gene. J Hum Genet 49:308–311
Bienvenu T, Der-Sarkissian H, Billuart P, Tissot M, Des Portes V, Bruls T, Chabrolle JP, Chauveau P, Cherry M, Kahn A, Cohen D, Beldjord C, Chelly J, Cherif D (1997) Mapping of the X-breakpoint involved in a balanced X;12 translocation in a female with mild mental retardation. Eur J Hum Genet 5:105–109
Billuart P, Bienvenu T, Ronce N, des Portes V, Vinet MC, Zemni R, Roest Crollius H, Carrie A, Fauchereau F, Cherry M, Briault S, Hamel B, Fryns J-P, Beldjord C, Kahn A, Moraine C, Chelly J (1998) Oligophrenin-1 encodes a rhoGAP protein involved in X-linked mental retardation. Nature 392:923–926
Boccone L, Meloni A, Falchi AM, Usai V, Cao A (1994) Blepharophimosis, ptosis, epicanthus inversus syndrome, a new case associated with de novo balanced autosomal translocation [46, XY, t(3;7)(q23;q32)]. Am J Med Genet 51:258–259
Bonaglia MC, Giorda R, Borgatti R, Felisari G, Gagliardi C, Selicorni A, Zuffardi O (2001) Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am J Hum Genet 69:261–268
Borg I, Freude K, Kubart S, Hoffmann K, Menzel C, Laccone F, Firth H, Ferguson-Smith MA, Tommerup N, Ropers HH, Sargan D, Kalscheuer VM (2005) Disruption of Netrin G1 by a balanced chromosome translocation in a girl with Rett syndrome. Eur J Hum Genet 13:921–927
Breg WR, Miller DA, Allderdice PW, Miller OJ (1972) Identification of translocation chromosomes by quinacrine fluorescence. Am J Dis Child 123:561–564
Cacheux V, Dastot-Le Moal F, Kaariainen H, Bondurand N, Rintala R, Boissier B, Wilson M, Mowat D, Goossens M (2001) Loss-of-function mutations in SIP1 Smad interacting protein 1 result in a syndromic Hirschsprung disease. Hum Mol Genet 10:1503–1510
Cargile CB, Goh DL, Goodman BK, Chen XN, Korenberg JR, Semenza GL, Thomas GH (2002) Molecular cytogenetic characterization of a subtle interstitial del(3)(p25.3p26.2) in a patient with deletion 3p syndrome. Am J Med Genet 109:133–138
Chitayat D, Fagerstrom CL, Kalousek DK, Rootman J, Taylor GP, Hall JG (1989) De novo reciprocal 1p;2q translocation in a child with multiple congenital anomalies/mental retardation syndrome. Am J Med Genet 32:36–41
Chiurazzi P, Schwartz CE, Gecz J, Neri G (2008) XLMR genes: update 2007. Eur J Hum Genet 16:422–434
Coco R, Penchaszadeh VB (1982) Cytogenetic findings in 200 children with mental retardation and multiple congenital anomalies of unknown cause. Am J Med Genet 12:155–173
Conroy JM, Grebe TA, Becker LA, Tsuchiya K, Nicholls RD, Buiting K, Horsthemke B, Cassidy SB, Schwartz S (1997) Balanced translocation 46, XY, t(2;15)(q37.2;q11.2) associated with atypical Prader–Willi syndrome. Am J Hum Genet 61:388–394
Curry CJ, Stevenson RE, Aughton D, Byrne J, Carey JC, Cassidy S, Cunniff C, Graham JM Jr, Jones MC, Kaback MM, Moeschler J, Schaefer GB, Schwartz S, Tarleton J, Opitz J (1997) Evaluation of mental retardation: recommendations of a consensus conference. Am J Med Genet 72:468–477
Daoud H, Gruchy N, Constans JM, Moussaoui E, Saumureau S, Bayou N, Amy M, Vedrine S, Vu PY, Rotig A, Laumonnier F, Vourc’h P, Andres CR, Leporrier N, Briault S (2009) Haploinsufficiency of the GPD2 gene in a patient with nonsyndromic mental retardation. Hum Genet 124:649–658
de Almeida JC, Llerena Junior JC, Goncalves Neto JB, Jung M, Martins RR (1993) Another example favouring the location of BPES at 3q2. J Med Genet 30:86
De Baere E, Beysen D, Oley C, Lorenz B, Cocquet J, De Sutter P, Devriendt K, Dixon M, Fellous M, Fryns JP, Garza A, Jonsrud C, Koivisto PA, Krause A, Leroy BP, Meire F, Plomp A, Van Maldergem L, De Paepe A, Veitia R, Messiaen L (2003) FOXL2 and BPES: mutational hotspots, phenotypic variability, and revision of the genotype–phenotype correlation. Am J Hum Genet 72:478–487
de Vries BBA, Winter R, Schinzel A, van Ravenswaaij-Arts C (2003) Telomeres: a diagnosis at the end of the chromosomes. J Med Genet 40:385–398
des Portes V, Pinard JM, Billuart P, Vinet MC, Koulakoff A, Carrie A, Gelot A, Dupuis E, Motte J, Berwald-Netter Y, Catala M, Kahn A, Beldjord C, Chelly J (1998) A novel CNS gene required for neuronal migration and involved in X-linked subcortical laminar heterotopia and lissencephaly syndrome. Cell 92:51–61
Dockery HE, Neale HC, Fitzgerald PH (1982) Gross congenital abnormality associated with an apparently balanced chromosomal translocation g(9;17)(q34;q11). J Med Genet 19:380–383
Duba HC, Doll A, Neyer M, Erdel M, Mann C, Hammerer I, Utermann G, Grzeschik KH (2002) The elastin gene is disrupted in a family with a balanced translocation t(7;16)(q11.23;q13) associated with a variable expression of the Williams–Beuren syndrome. Eur J Hum Genet 10:351–361
Dubos A, Pannetier S, Hanauer A (2008) Inactivation of the CDKL3 gene at 5q31.1 by a balanced t(X;5) translocation associated with nonspecific mild mental retardation. Am J Med Genet A 146A:1267–1279
Egemen A, Ulger Z, Ozkinay F, Gulen F, Cogulu O (2005) A de novo t (X;8)(p11.2;q24.3) demonstrating Cornelia de Lange syndrome phenotype. Genet Couns 16:27–30
Endris V, Wogatzky B, Leimer U, Bartsch D, Zatyka M, Latif F, Maher ER, Tariverdian G, Kirsch S, Karch D, Rappold GA (2002) The novel Rho-GTPase activating gene MEGAP/srGAP3 has a putative role in severe mental retardation. Proc Natl Acad Sci USA 99:11754–11759
Falk RE, Casas KA (2007) Chromosome 2q37 deletion: clinical and molecular aspects. Am J Med Genet 145C:357–371
Fantes JA, Boland E, Ramsay J, Donnai D, Splitt M, Goodship JA, Stewart H, Whiteford M, Gautier P, Harewood L, Holloway S, Sharkey F, Maher E, van Heyningen V, Clayton-Smith J, Fitzpatrick DR, Black GC (2008) FISH mapping of de novo apparently balanced chromosome rearrangements identifies characteristics associated with phenotypic abnormality. Am J Hum Genet 82:916–926
Fernandez T, Morgan T, Davis N, Klin A, Morris A, Farhi A, Lifton RP, State MW (2004) Disruption of contactin 4 (CNTN4) results in developmental delay and other features of 3p deletion syndrome. Am J Hum Genet 74:1286–1293
FitzPatrick DR, Carr IM, McLaren L, Leek JP, Wightman P, Williamson K, Gautier P, McGill N, Hayward C, Firth H, Markham AF, Fantes JA, Bonthron DT (2003) Identification of SATB2 as the cleft palate gene on 2q32–q33. Hum Mol Genet 12:2491–2501
Friedrich U, Dalby M, Staehelin-Jensen T, Bruun-Petersen G (1982) Chromosomal studies of children with developmental language retardation. Dev Med Child Neurol 24:645–652
Frints SGM, Marynen P, Hartmann D, Fryns J-P, Steyaert J, Schachner M, Rolf B, Craessaerts K, Snellinx A, Hollanders K, D’Hooge R, De Deyn PP, Froyen G (2003) CALL interrupted in a patient with non-specific mental retardation: gene dosage-dependent alteration of murine brain development and behavior. Hum Mol Genet 12:1463–1474
Fryns JP, Hendrickx G (1996) Malformative syndrome with trigonocephaly, shallow orbits, ptosis, growth and mental retardation. De novo autosomal reciprocal t(9;13)(Q32;Q22) in a male patient. Ann Genet 39:51–53
Fryns JP, Kleczkowska A, Kubień E, Berghe H (1986) Excess of mental retardation and/or congenital malformation in reciprocal translocations in man. Hum Genet 72:1–8
Fryns JP, Kleczkowska A, Smeets E, Thiry P, Geutjens J, Van den Berghe H (1990) Cohen syndrome and de novo reciprocal translocation t(5;7)(q33.1;p15.1). Am J Med Genet 37:546–547
Funderburk SJ, Spence MA, Sparkes RS (1977) Mental retardation associated with “balanced” chromosome rearrangements. Am J Hum Genet 29:136–141
Gallagher RC, Pils B, Albalwi M, Francke U (2002) Evidence for the role of PWCR1/HBII-85 C/D box small nucleolar RNAs in Prader–Willi syndrome. Am J Hum Genet 71:669–678
Garcia-Sagredo JM, San Roman C, Gallego Gomez ME, Lledo G (1983) Fragile chromosome 16(q22) cause a balanced translocation at the same point. Hum Genet 65:211–213
Gecz J, Barnett S, Liu J, Hollway G, Donnelly A, Eyre H, Eshkevari HS, Baltazar R, Grunn A, Nagaraja R, Gilliam C, Peltonen L, Sutherland GR, Baron M, Mulley JC (1999) Characterization of the human glutamate receptor subunit 3 gene (GRIA3), a candidate for bipolar disorder and nonspecific X-linked mental retardation. Genomics 62:356–368
Greger V, Woolf E, Lalande M (1993) Cloning of the breakpoints of a submicroscopic deletion in an Angelman syndrome patient. Hum Mol Genet 2:921–924
Gribble SM, Prigmore E, Burford DC, Porter KM, Ng BL, Douglas EJ, Fiegler H, Carr P, Kalaitzopoulos D, Clegg S, Sandstrom R, Temple IK, Youings SA, Thomas NS, Dennis NR, Jacobs PA, Crolla JA, Carter NP (2005) The complex nature of constitutional de novo apparently balanced translocations in patients presenting with abnormal phenotypes. J Med Genet 42:8–16
Griggs BL, Ladd S, Saul RA, DuPont BR, Srivastava AK (2008) Dedicator of cytokinesis 8 is disrupted in two patients with mental retardation and developmental disabilities. Genomics 91:195–202
Hagens O, Dubos A, Abidi F, Barbi G, Van Zutven L, Hoeltzenbein M, Tommerup N, Moraine C, Fryns JP, Chelly J, van Bokhoven H, Gecz J, Dollfus H, Ropers HH, Schwartz CE, s Santos R, Kalscheuer V, Hanauer A (2006) Disruptions of the novel KIAA1202 gene are associated with X-linked mental retardation. Hum Genet 118:578–590
Higgins AW, Alkuraya FS, Bosco AF, Brown KK, Bruns GA, Donovan DJ, Eisenman R, Fan Y, Farra CG, Ferguson HL, Gusella JF, Harris DJ, Herrick SR, Kelly C, Kim HG, Kishikawa S, Korf BR, Kulkarni S, Lally E, Leach NT, Lemyre E, Lewis J, Ligon AH, Lu W, Maas RL, MacDonald ME, Moore SD, Peters RE, Quade BJ, Quintero-Rivera F, Saadi I, Shen Y, Shendure J, Williamson RE, Morton CC (2008) Characterization of apparently balanced chromosomal rearrangements from the developmental genome anatomy project. Am J Hum Genet 82:712–722
Histinx TW, Gabreels FJ, Rutten FJ, Korten II, Scheres JM, Joosten EM (1975) A mentally retarded child with a translocation involving chromosomes 12 and 19. J Med Genet 12:207–210
Holmes SE, Riazi MA, Gong W, McDermid HE, Sellinger BT, Hua A, Chen F, Wang Z, Zhang G, Roe B, Gonzalez I, McDonald-McGinn DM, Zackai E, Emanuel BS, Budarf ML (1997) Disruption of the clathrin heavy chain-like gene (CLTCL) associated with features of DGS/VCFS: a balanced (21;22)(p12;q11) translocation. Hum Mol Genet 6:357–367
Ishikawa-Brush Y, Powell JF, Bolton P, Miller AP, Francis F, Willard HF, Lehrach H, Monaco AP (1997) Autism and multiple exostoses associated with an X;8 translocation occurring within the GRPR gene and 3′ to the SDC2 gene. Hum Mol Genet 6:1241–1250
Israel J, Lessick M, Szego K, Wong P (1991) Translocation 19;Y in a child with Bannayan–Zonana phenotype. J Med Genet 28:427–428
Jacobs PA (1974) Correlation between euploid structural chromosome rearrangements and mental subnormality in humans. Nature 249:164–165
Jenkins EC, Curcuru-Giordano FM, Krishna SG, Cantarella J (1975) De novo occurrence of 46, XX, t(4;13) (q31;q14) in a mentally retarded girl. Ann Genet 18:117–120
Johnston JJ, Olivos-Glander I, Killoran C, Elson E, Turner JT, Peters KF, Abbott MH, Aughton DJ, Aylsworth AS, Bamshad MJ, Booth C, Curry CJ, David A, Dinulos MB, Flannery DB, Fox MA, Graham JM, Grange DK, Guttmacher AE, Hannibal MC, Henn W, Hennekam RC, Holmes LB, Hoyme HE, Leppig KA, Lin AE, Macleod P, Manchester DK, Marcelis C, Mazzanti L, McCann E, McDonald MT, Mendelsohn NJ, Moeschler JB, Moghaddam B, Neri G, Newbury-Ecob R, Pagon RA, Phillips JA, Sadler LS, Stoler JM, Tilstra D, Walsh Vockley CM, Zackai EH, Zadeh TM, Brueton L, Black GC, Biesecker LG (2005) Molecular and clinical analyses of Greig cephalopolysyndactyly and Pallister–Hall syndromes: robust phenotype prediction from the type and position of GLI3 mutations. Am J Hum Genet 76:609–622
Kähkönen M, Leisti J, Thoden CJ, Autio S (1986) Frequency of rare fragile sites among mentally subnormal schoolchildren. Clin Genet 30:234–238
Kalscheuer VM, Feenstra I, Van Ravenswaaij-Arts CM, Smeets DF, Menzel C, Ullmann R, Musante L, Ropers HH (2008a) Disruption of the TCF4 gene in a girl with mental retardation but without the classical Pitt–Hopkins syndrome. Am J Med Genet 146A:2053–2059
Kalscheuer VM, FitzPatrick D, Tommerup N, Bugge M, Niebuhr E, Neumann LM, Tzschach A, Shoichet SA, Menzel C, Erdogan F, Arkesteijn G, Ropers HH, Ullmann R (2007) Mutations in autism susceptibility candidate 2 (AUTS2) in patients with mental retardation. Hum Genet 121:501–509
Kalscheuer VM, Musante L, Fang C, Hoffmann K, Fuchs C, Carta E, Deas E, Venkateswarlu K, Menzel C, Ullmann R, Tommerup N, Dalpra L, Tzschach A, Selicorni A, Luscher B, Ropers HH, Harvey K, Harvey RJ (2008b) A balanced chromosomal translocation disrupting ARHGEF9 is associated with epilepsy, anxiety, aggression, and mental retardation. Hum Mutat
Kalscheuer VM, Tao J, Donnelly A, Hollway G, Schwinger E, Kubart S, Menzel C, Hoeltzenbein M, Tommerup N, Eyre H, Harbord M, Haan E, Sutherland GR, Ropers HH, Gecz J (2003) Disruption of the serine/threonine kinase 9 gene causes severe X-linked infantile spasms and mental retardation. Am J Hum Genet 72:1401–1411
Kang S, Graham JM Jr, Olney AH, Biesecker LG (1997) GLI3 frameshift mutations cause autosomal dominant Pallister–Hall syndrome. Nat Genet 15:266–268
Kato M (2006) A new paradigm for West syndrome based on molecular and cell biology. Epilepsy Res 70(Suppl 1):S87–S95
Kleczkowska A, Fryns JP, Vinken L, van den Berghe H (1985) Effect of balanced X/autosome translocations on sexual and physical development. A personal experience in 4 patients. Clin Genet 27:147–152
Kleefstra T, Yntema HG, Oudakker AR, Banning MJ, Kalscheuer VM, Chelly J, Moraine C, Ropers HH, Fryns JP, Janssen IM, Sistermans EA, Nillesen WN, de Vries LB, Hamel BC, van Bokhoven H (2004) Zinc finger 81 (ZNF81) mutations associated with X-linked mental retardation. J Med Genet 41:394–399
Kleefstra T, Smidt M, Banning MJG, Oudakker AR, Van Esch H, de Brouwer APM, Nillesen W, Sistermans EA, Hamel BCJ, de Bruijn D, Fryns J-P, Yntema HG, Brunner HG, de Vries BBA, van Bokhoven H (2005) Disruption of the gene euchromatin histone methyl transferase1 (Eu-HMTase1) is associated with the 9q34 subtelomeric deletion syndrome. J Med Genet 42:299–306
Kleefstra T, Brunner HG, Amiel J, Oudakker AR, Nillesen WM, Magee A, Genevieve D, Cormier-Daire V, van Esch H, Fryns JP, Hamel BCJ, Sistermans EA, de Vries BBA, van Bokhoven H (2006) Loss-of-function mutations in euchromatin histone methyl transferase 1 (EHMT1) cause the 9q34 subtelomeric deletion syndrome. Am J Hum Genet 79:370–377
Koolen DA, Vissers LELM, Pfundt R, de Leeuw N, Knight SJL, Regan R, Kooy RF, Reyniers E, Romano C, Fichera M, Schinzel A, Baumer A, Anderlid B-M, Schoumans J, Knoers NV, Geurts van Kessel A, Sistermans EA, Veltman J, Brunner HG, de Vries BBA (2006) A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet 38:999–1001
Kruger G, Gotz J, Kvist U, Dunker H, Erfurth F, Pelz L, Zech L (1989) Greig syndrome in a large kindred due to reciprocal chromosome translocation t(6;7)(q27;p13). Am J Med Genet 32:411–416
Kundakovic M, Chen Y, Guidotti A, Grayson DR (2008) The reelin and GAD67 promoters are activated by epigenetic drugs that facilitate the disruption of local repressor complexes. Mol Pharmacol
Kurotaki N, Imaizumi K, Harada N, Masuno M, Kondoh T, Nagai T, Ohashi H, Naritomi K, Tsukahara M, Makita Y, Sugimoto T, Sonoda T, Hasegawa T, Chinen Y, Tomita Ha HA, Kinoshita A, Mizuguchi T, Yoshiura Ki K, Ohta T, Kishino T, Fukushima Y, Niikawa N, Matsumoto N (2002) Haploinsufficiency of NSD1 causes Sotos syndrome. Nat Genet 30:365–366
Kuslich CD, Kobori JA, Mohapatra G, Gregorio-King C, Donlon TA (1999) Prader–Willi syndrome is caused by disruption of the SNRPN gene. Am J Hum Genet 64:70–76
Kutsche K, Yntema H, Brandt A, Jantke I, Nothwang HG, Orth U, Boavida MG, David D, Chelly J, Fryns JP, Moraine C, Ropers HH, Hamel BC, van B, Gal A (2000) Mutations in ARHGEF6, encoding a guanine nucleotide exchange factor for Rho GTPases, in patients with X-linked mental retardation. Nat Genet 26:247–250
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Leonard H, Wen X (2002) The epidemiology of mental retardation: challenges and opportunities in the new millennium. Ment Retard Dev Disabil Res Rev 8:117–134
Li DY, Toland AE, Boak BB, Atkinson DL, Ensing GJ, Morris CA, Keating MT (1997) Elastin point mutations cause an obstructive vascular disease, supravalvular aortic stenosis. Hum Mol Genet 6:1021–1028
Lin T, Orrison BM, Leahey AM, Suchy SF, Bernard DJ, Lewis RA, Nussbaum RL (1997) Spectrum of mutations in the OCRL1 gene in the Lowe oculocerebrorenal syndrome. Am J Hum Genet 60:1384–1388
Luckasson R, Borthwick-Duffy S, Buntinx WHE, Coulter DL, Craig PM, Reeve A, Schalock RL, Snell MA, Spitalnik DM, Spreat S, Tassé MJ (2002) Mental retardation: definition, classification, and systems of supports, 10th edn. American Association on Mental Retardation, Washington, DC
Mansouri MR, Marklund L, Gustavsson P, Davey E, Carlsson B, Larsson C, White I, Gustavson KH, Dahl N (2005) Loss of ZDHHC15 expression in a woman with a balanced translocation t(X;15)(q13.3;cen) and severe mental retardation. Eur J Hum Genet 13:970–977
Marco EJ, Abidi FE, Bristow J, Dean WB, Cotter P, Jeremy RJ, Schwartz CE, Sherr EH (2008) ARHGEF9 disruption in a female patient is associated with X linked mental retardation and sensory hyperarousal. J Med Genet 45:100–105
Masuno M, Imaizumi K, Fukushima Y, Tanaka Y, Ishii T, Nakamura M, Kuroki Y (1997) Median cleft of upper lip and pedunculated skin masses associated with de novo reciprocal translocation 46, X, t(X;16)(q28;q11.2). J Med Genet 34:952–954
Mattei MG, Mattei JF, Ayme S, Malpuech G, Giraud F (1978) A dynamic study in two new cases of X chromosome translocations. Hum Genet 41:251–257
McGhee EM, Klump CJ, Bitts SM, Cotter PD, Lammer EJ (2000) Candidate region for Coffin–Siris syndrome at 7q32→34. Am J Med Genet 93:241–243
McPherson EW, Laneri G, Clemens MM, Kochmar SJ, Surti U (1997) Apparently balanced t(1;7)(q21.3;q34) in an infant with Coffin–Siris syndrome. Am J Med Genet 71:430–433
Mercer TR, Dinger ME, Mariani J, Kosik KS, Mehler MF, Mattick JS (2008) Noncoding RNAs in long-term memory formation. Neuroscientist 14:434–445
Moller RS, Kubart S, Hoeltzenbein M, Heye B, Vogel I, Hansen CP, Menzel C, Ullmann R, Tommerup N, Ropers HH, Tumer Z, Kalscheuer VM (2008a) Truncation of the Down syndrome candidate gene DYRK1A in two unrelated patients with microcephaly. Am J Hum Genet 82:1165–1170
Moller RS, Schneider LM, Hansen CP, Bugge M, Ullmann R, Tommerup N, Tumer Z (2008b) Balanced translocation in a patient with severe myoclonic epilepsy of infancy disrupts the sodium channel gene SCN1A. Epilepsia 49:1091–1094
Musio A, Selicorni A, Focarelli ML, Gervasini C, Milani D, Russo S, Vezzoni P, Larizza L (2006) X-linked Cornelia de Lange syndrome owing to SMC1L1 mutations. Nat Genet 38:528–530
Nielsen J, Krag-Olsen B (1981) Follow-up of 32 children with autosomal translocations found among 11, 148 consecutively newborn children from 1969 to 1974. Clin Genet 20:48–54
Nothwang HG, Kim HG, Aoki J, Geisterfer M, Kubart S, Wegner RD, van Moers A, Ashworth LK, Haaf T, Bell J, Arai H, Tommerup N, Ropers HH, Wirth J (2001) Functional hemizygosity of PAFAH1B3 due to a PAFAH1B3-CLK2 fusion gene in a female with mental retardation, ataxia and atrophy of the brain. Hum Mol Genet 10:797–806
Patel ZM, Mulye VR, Raghavan K, Shah SB (1987) 12;14 translocation in Coffin–Siris syndrome. Indian Pediatr 24:435–438
Petrij F, Dorsman JC, Dauwerse HG, Giles RH, Peeters T, Hennekam RC, Breuning MH, Peters DJ (2000) Rubinstein–Taybi syndrome caused by a de novo reciprocal translocation t(2;16)(q36.3;p13.3). Am J Med Genet 92:47–52
Petrij F, Gilles RH, Dauwerse HG, Saris JJ, Hennekam RCM, Masuno M, Tommerup N, van Ommen G-JB, Goodman RH, Peters DJM, Breuning MH (1995) Rubinstein–Taybi syndrome caused by mutations in the transcriptional co-activator CBP. Nature 376:348–351
Pichon B, Vankerckhove S, Bourrouillou G, Duprez L, Abramowicz MJ (2004) A translocation breakpoint disrupts the ASPM gene in a patient with primary microcephaly. Eur J Hum Genet 12:419–421
Praphanphoj V, Goodman BK, Thomas GH, Niel KM, Toomes C, Dixon MJ, Geraghty MT (2000) Molecular cytogenetic evaluation in a patient with a translocation (3;21) associated with blepharophimosis, ptosis, epicanthus inversus syndrome (BPES). Genomics 65:67–69
Prieto-Carrasquero M, Rojas-Atencio A, Gonzalez S, Pineda-Del Villar L, Soto M, Quintero M, Miranda LE, Canizales J (1995) Clinical experience with balanced reciprocal translocations. Genet Couns 6:349–354
Ramocki MB, Dowling J, Grinberg I, Kimonis VE, Cardoso C, Gross A, Chung J, Martin CL, Ledbetter DH, Dobyns WB, Millen KJ (2003) Reciprocal fusion transcripts of two novel Zn-finger genes in a female with absence of the corpus callosum, ocular colobomas and a balanced translocation between chromosomes 2p24 and 9q32. Eur J Hum Genet 11:527–534
Rasmussen K, Nielsen J, Dahl G (1982) The prevalence of chromosome abnormalities among mentally retarded persons in a geographically delimited area of Denmark. Clin Genet 22:244–255
Reilly DS, Lewis RA, Ledbetter DH, Nussbaum RL (1988) Tightly linked flanking markers for the Lowe oculocerebrorenal syndrome, with application to carrier assessment. Am J Hum Genet 42:748–755
Roeleveld N, Zielhuis GA, Gabreels F (1997) The prevalence of mental retardation: a critical review of recent literature. Dev Med Child Neurol 39:125–132
Rooms L, Reyniers E, Kooy RF (2007) Diverse chromosome breakage mechanisms underlie subtelomeric rearrangements, a common cause of mental retardation. Hum Mutat 28:177–182
Ropers HH (2007) New perspectives for the elucidation of genetic disorders. Am J Hum Genet 81:199–207
Ross ME, Allen KM, Srivastava AK, Featherstone T, Gleeson JG, Hirsch B, Harding BN, Andermann E, Abdullah R, Berg M, Czapansky-Bielman D, Flanders DJ, Guerrini R, Motte J, Mira AP, Scheffer I, Berkovic S, Scaravilli F, King RA, Ledbetter DH, Schlessinger D, Dobyns WB, Walsh CA (1997) Linkage and physical mapping of X-linked lissencephaly/SBH (XLIS): a gene causing neuronal migration defects in human brain. Hum Mol Genet 6:555–562
Runte M, Huttenhofer A, Gross S, Kiefmann M, Horsthemke B, Buiting K (2001) The IC-SNURF-SNRPN transcript serves as a host for multiple small nucleolar RNA species and as an antisense RNA for UBE3A. Hum Mol Genet 10:2687–2700
Sahoo T, del Gaudio D, German JR, Shinawi M, Peters SU, Person RE, Garnica A, Cheung SW, Beaudet AL (2008) Prader–Willi phenotype caused by paternal deficiency for the HBII-85 C/D box small nucleolar RNA cluster. Nat Genet 40:719–721
Santos CB, Discepoli G, Pigliapoco F, Boy R, Pimentel MM (2003) De novo balanced translocation (2;10)(q24;q22) associated with mental retardation. Ann Genet 46:471–473
Satterlee JS, Barbee S, Jin P, Krichevsky A, Salama S, Schratt G, Wu DY (2007) Noncoding RNAs in the brain. J Neurosci 27:11856–11859
Schüle B, Albalwi M, Northrop E, Francis DI, Rowell M, Slater HR, McKinlay Gardner RJ, Francke U (2005) Molecular breakpoint cloning and gene expression studies of a novel translocation t(4;15)(q27;q11.2) associated with Prader–Willi syndrome. BMC Med Genet 6:18
Schulze A, Hansen C, Skakkebaek NE, Brondum-Nielsen K, Ledbeter DH, Tommerup N (1996) Exclusion of SNRPN as a major determinant of Prader–Willi syndrome by a translocation breakpoint. Nat Genet 12:452–454
Sharp AJ, Hansen S, Selzer RR, Cheng Z, Regan R, Hurst JA, Stewart H, Price SM, Blair E, Hennekam RC, Fitzpatrick CA, Segraves R, Richmond TA, Guiver C, Albertson DG, Pinkel D, Eis PS, Schwartz S, Knight SJL, Eichler EE (2006) Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 38:1038–1042
Sharp AJ, Mefford HC, Li K, Baker C, Skinner C, Stevenson RE, Schroer RJ, Novara F, De Gregori M, Ciccone R, Broomer A, Casuga I, Wang Y, Xiao C, Barbacioru C, Gimelli G, Dalla Bernardina B, Torniero C, Giorda R, Regan R, Murday V, Mansour S, Fichera M, Castiglia L, Failla P, Ventura M, Jiang Z, Cooper GM, Knight SJL, Romano C, Zuffardi O, Chen C, Schwartz CE, Eichler EE (2008) A recurrent 15q13.3 microdeletion syndrome associated with mental retardation and seizures. Nat Genet 40:322–328
Shaw-Smith C, Pittman AM, Willatt L, Martin H, Rickman L, Gribble S, Curley R, Cumming S, Dunn C, Kalaitzopoulos D, Porter K, Prigmore E, Krepischi-Santos ACV, Varela MC, Koiffmann CP, Lees AJ, Rosenberg C, Firth HV, de Silva R, Carter NP (2006) Microdeletion encompassing MAPT at chromosome 17q21.3 is associated with developmental delay and learning disability. Nat Genet 38:1032–1037
Shoichet SA, Duprez L, Hagens O, Waetzig V, Menzel C, Herdegen T, Schweiger S, Dan B, Vamos E, Ropers HH, Kalscheuer VM (2006) Truncation of the CNS-expressed JNK3 in a patient with a severe developmental epileptic encephalopathy. Hum Genet 118:559–567
Shoichet SA, Hoffmann K, Menzel C, Trautmann U, Moser B, Hoeltzenbein M, Echenne B, Partington M, Van Bokhoven H, Moraine C, Fryns JP, Chelly J, Rott HD, Ropers HH, Kalscheuer VM (2003) Mutations in the ZNF41 gene are associated with cognitive deficits: identification of a new candidate for X-linked mental retardation. Am J Hum Genet 73:1341–1354
Skovby F, Niebuhr E (1974) Presumably balanced translocations involving the same band of chromosome No. 4 found in two mentally retarded, dysmorphic individuals. Ann Genet 17:243–249
Speleman F, Van Roy N, Wiegant J, Verschraegen-Spae MR, Benoit Y, Govaert P, Goossens L, Leroy JG (1992) Detection of subtle reciprocal translocations by fluorescence in situ hybridization. Clin Genet 41:169–174
Stevenson RE, Procopio-Allen AM, Schroer RJ, Collins JS (2003) Genetic syndromes among individuals with mental retardation. Am J Med Genet 123A:29–32
Striano P, Elia M, Castiglia L, Galesi O, Pelligra S, Striano S (2005) A t(4;9)(q34;p22) translocation associated with partial epilepsy, mental retardation, and dysmorphism. Epilepsia 46:1322–1324
Sultana R, Yu CE, Yu J, Munson J, Chen D, Hua W, Estes A, Cortes F, de la Barra F, Yu D, Haider ST, Trask BJ, Green ED, Raskind WH, Disteche CM, Wijsman E, Dawson G, Storm DR, Schellenberg GD, Villacres EC (2002) Identification of a novel gene on chromosome 7q11.2 interrupted by a translocation breakpoint in a pair of autistic twins. Genomics 80:129–134
Sun Y, Nicholls RD, Butler MG, Saitoh S, Hainline BE, Palmer CG (1996) Breakage in the SNRPN locus in a balanced 46, XY, t(15;19) Prader–Willi syndrome patient. Hum Mol Genet 5:517–524
Talisetti A, Forrester SR, Gregory D, Johnson L, Schneider MC, Kimonis VE (2003) Temtamy-like syndrome associated with translocation of 2p24 and 9q32. Clin Dysmorphol 12:175–177
Tao J, Van Esch H, Hagedorn-Greiwe M, Hoffmann K, Moser B, Raynaud M, Sperner J, Fryns JP, Schwinger E, Gecz J, Ropers HH, Kalscheuer VM (2004) Mutations in the X-linked cyclin-dependent kinase-like 5 (CDKL5/STK9) gene are associated with severe neurodevelopmental retardation. Am J Hum Genet 75:1149–1154
Tommerup N, Nielsen F (1983) A familial reciprocal translocation t(3;7) (p21.1;p13) associated with the Greig polysyndactyly-craniofacial anomalies syndrome. Am J Med Genet 16:313–321
Tranebjaerg L, Sjo O, Warburg M (1986) Retinal cone dysfunction and mental retardation associated with a de novo balanced translocation 1;6(q44;q27). Ophthalmic Paediatr Genet 7:167–173
van der Maarel SM, Scholten IH, Huber I, Philippe C, Suijkerbuijk RF, Gilgenkrantz S, Kere J, Cremers FP, Ropers HH (1996) Cloning and characterization of DXS6673E, a candidate gene for X-linked mental retardation in Xq13.1. Hum Mol Genet 5:887–897
van Hemel JO, van Biervliet JP, der Grift PW (1975) A girl with 46, XX, t(1;15) karyotype. Cytogenetic and clinical observations. Clin Genet 8:213–217
Vervoort VS, Viljoen D, Smart R, Suthers G, DuPont BR, Abbott A, Schwartz CE (2002) Sorting nexin 3 (SNX3) is disrupted in a patient with a translocation t(6;13)(q21;q12) and microcephaly, microphthalmia, ectrodactyly, prognathism (MMEP) phenotype. J Med Genet 39:893–899
Wakamatsu N, Yamada Y, Yamada K, Ono T, Nomura N, Taniguchi H, Kitoh H, Mutoh N, Yamanaka T, Mushiake K, Kato K, Sonta S, Nagaya M (2001) Mutations in SIP1, encoding Smad interacting protein-1, cause a form of Hirschsprung disease. Nat Genet 27:369–370
Walter S, Sandig K, Hinkel GK, Mitulla B, Ounap K, Sims G, Sitska M, Utermann B, Viertel P, Kalscheuer V, Bartsch O (2004) Subtelomere FISH in 50 children with mental retardation and minor anomalies, identified by a checklist, detects 10 rearrangements including a de novo balanced translocation of chromosomes 17p13.3 and 20q13.33. Am J Med Genet 128A:364–373
Warburton D (1991) De novo balanced chromosome rearrangements and extra marker chromosomes identified at prenatal diagnosis: clinical significance and distribution of breakpoints. Am J Hum Genet 49:995–1013
Weaving LS, Christodoulou J, Williamson SL, Friend KL, McKenzie OL, Archer H, Evans J, Clarke A, Pelka GJ, Tam PP, Watson C, Lahooti H, Ellaway CJ, Bennetts B, Leonard H, Gecz J (2004) Mutations of CDKL5 cause a severe neurodevelopmental disorder with infantile spasms and mental retardation. Am J Hum Genet 75:1079–1093
WHO (2001) World health report 2001. Mental health: new understanding, new hope, Geneva, Switzerland
Wilbur L, Curcuru-Giordano FM, Krishna SG, Kardon NB, Jenkins EC (1977) A case of 46, XY, t(1;13) (q24;q32) with mental retardation. Hum Genet 37:239–242
Wild A, Kalff-Suske M, Vortkamp A, Bornholdt D, Konig R, Grzeschik KH (1997) Point mutations in human GLI3 cause Greig syndrome. Hum Mol Genet 6:1979–1984
Winnepenninckx B, Rooms L, Kooy RF (2003) Mental retardation: a review of the genetic causes. Brit J Dev Disabil 49:29–44
Wirth J, Back E, Huttenhofer A, Nothwang HG, Lich C, Gross S, Menzel C, Schinzel A, Kioschis P, Tommerup N, Ropers HH, Horsthemke B, Buiting K (2001) A translocation breakpoint cluster disrupts the newly defined 3′ end of the SNURF-SNRPN transcription unit on chromosome 15. Hum Mol Genet 10:201–210
Wirth J, Nothwang HG, van der Maarel S, Menzel C, Borck G, Lopez-Pajares I, Brondum-Nielsen K, Tommerup N, Bugge M, Ropers HH, Haaf T (1999) Systematic characterisation of disease associated balanced chromosome rearrangements by FISH: cytogenetically and genetically anchored YACs identify microdeletions and candidate regions for mental retardation genes. J Med Genet 36:271–278
Wu Y, Arai AC, Rumbaugh G, Srivastava AK, Turner G, Hayashi T, Suzuki E, Jiang Y, Zhang L, Rodriguez J, Boyle J, Tarpey P, Raymond FL, Nevelsteen J, Froyen G, Stratton M, Futreal A, Gecz J, Stevenson R, Schwartz CE, Valle D, Huganir RL, Wang T (2007) Mutations in ionotropic AMPA receptor 3 alter channel properties and are associated with moderate cognitive impairment in humans. Proc Natl Acad Sci USA 104:18163–18168
Yue Y, Stout K, Grossmann B, Zechner U, Brinckmann A, White C, Pilz DT, Haaf T (2006) Disruption of TCBA1 associated with a de novo t(1;6)(q32.2;q22.3) presenting in a child with developmental delay and recurrent infections. J Med Genet 43:143–147
Zaki M, Shehab M, El-Aleem AA, Abdel-Salam G, Koeller HB, Ilkin Y, Ross ME, Dobyns WB, Gleeson JG (2007) Identification of a novel recessive RELN mutation using a homozygous balanced reciprocal translocation. Am J Med Genet A 143A:939–944
Zemni R, Bienvenu T, Vinet MC, Sefiani A, Carrie A, Billuart P, McDonell N, Couvert P, Francis F, Chafey P, Fauchereau F, Friocourt G, des Portes V, Cardona A, Frints S, Meindl A, Brandau O, Ronce N, Moraine C, van Bokhoven H, Ropers HH, Sudbrak R, Kahn A, Fryns JP, Beldjord C, Chelly J (2000) A new gene involved in X-linked mental retardation identified by analysis of an X;2 balanced translocation. Nat Genet 24:167–170
Acknowledgments
Our experimental work is supported by the Belgian National Fund for Scientific Research, Flanders (FWO).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vandeweyer, G., Kooy, R.F. Balanced translocations in mental retardation. Hum Genet 126, 133–147 (2009). https://doi.org/10.1007/s00439-009-0661-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-009-0661-6