Abstract
DNA amplification of the 11q13 region is observed frequently in many carcinomas. Within the amplified region several candidate oncogenes have been mapped, including cyclin D1, TAOS1 and cortactin. Yet, it is unknown which gene(s) is/are responsible for the selective pressure enabling amplicon formation. This is probably due to the use of low-resolution detection methods. Furthermore, the size and structure of the amplified 11q13 region is complex and consists of multiple amplicon cores that differ between different tumor types. We set out to test whether the borders of the 11q13 amplicon are restricted to regions that enable DNA breakage and subsequent amplification. A high-resolution array of the 11q13 region was generated to study the structure of the 11q13 amplicon and analyzed 29 laryngeal and pharyngeal carcinomas and nine cell lines with 11q13 amplification. We found that boundaries of the commonly amplified region were restricted to four segments. Three boundaries coincided with a syntenic breakpoint. Such regions have been suggested to be putatively fragile. Sequence comparisons revealed that the amplicon was flanked by two large low copy repeats known as segmental duplications. These segmental duplications might be responsible for the typical structure and size of the 11q13 amplicon. We hypothesize that the selection for genes through amplification of the 11q13.3 region is determined by the ability to form DNA breaks within specific regions and, consequently, results in large amplicons containing multiple genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genomic instability is an important step in generating the multiple changes required for cancer. It plays a role in tumor initiation, development and progression, and confers various biological features to cancer cells such as response to therapy. DNA amplification is one of the manifestations of genomic instability and is frequently found in human cancer. It is one of the mechanisms by which cells can accomplish overexpression of oncogenes and occurs throughout the human genome at different frequencies. The sequence characteristics of the loci where DNA amplification can occur remain largely unknown. For several decades, the mechanisms of DNA amplification have been extensively studied mainly for genes that are involved in the resistance to cytotoxic drugs. Several models explaining the mechanism of gene amplification have been proposed, including translocation-excision-deletion-amplification, deletion-plus-episome, sister-chromatid-exchange, onionskin, extrachromosomal-doubling-rolling circle and the chromatid-breakage-fusion-bridge (BFB) cycle model (Barr et al. 1996; Ciullo et al. 2002; Coquelle et al. 1997; Schwab 1999; Stark et al. 1989; Toledo et al. 2000; Van Roy et al. 2006; Vogt et al. 2004). One of the main initiators of the proposed mechanisms is DNA double strand breakage.
DNA amplification of the 11q13 region is observed in several types of solid tumors, but most frequently (36%) in carcinomas of the head and neck region (HNSCC) (Schuuring 1995). The involvement in several types of carcinoma and the frequent occurrence in head and neck cancer implicate a selective advantage for cells containing increased mRNA levels for genes within the amplified region. Although the amplification is abundantly present, the clinical significance for increased mRNA levels of a specific target gene is not yet found. Efforts to narrow down the overlapping region of amplification resulted both in the identification of different independent amplicon cores (depending on tumor type) as well as several target genes within each core (Freier et al. 2006; Huang et al. 2002; Hughes-Davies et al. 2003; Hui et al. 2005; Schuuring 1995). We propose that DNA breakage occurs at specific regions explaining the position and size of the amplicon.
Using information from the completed human genome sequence, high-resolution DNA copy number analysis to determine the structure and exact location of an amplicon has only recently become available. Whole genome array CGH (WGA) brought the resolution of conventional chromosomal CGH to a higher level, but the use of high-resolution CGH arrays (HR-aCGH) containing oligonucleotides or genomic (BAC) clones that completely cover and overlap a certain locus improved the analysis enormously (Albertson 2006).
We generated a CGH array designed specifically for the 11q13 region with a higher resolution than any previously described method (Freier et al. 2006; Huang et al. 2002; Hui et al. 2005; Lin et al. 2006), which enables the accurate detection of amplicon structures. We sought to find out whether the formation of 11q13 amplicons is solely due to gene selection or is the result of variation in chromosome structure as reflected in repeat sequences, DNA flexibility and evolutionary breakpoints. For this purpose, we mapped the boundaries of the 11q13 amplicons in a series of 29 HNSCCs and nine head and neck cell lines with previously identified DNA amplification of the 11q13 region using this high density HR-CGH array. By comparing the boundaries of all 38 cases, we found 4 regions flanking the 11q13 amplicons that were common in numerous cases. In silico analysis of the nucleotide sequences at these four common boundaries revealed that three of the boundaries coincided with syntenic breakpoints and the identification of two inverted large low copy repeats (LCRs), known as segmental duplications (SDs) flanking the commonly amplified 11q13.3 region. We hypothesize that the selection of genes in the 11q13.3 region that become amplified is determined by the ability to form DNA breaks within specific regions. This will result in the formation of large amplicons containing many genes that are not at all necessarily relevant for tumorigenesis and tumor progression.
Materials and methods
Head and neck carcinomas and cell lines
For this study, we used primary carcinomas of the larynx (n = 22) and pharynx (n = 8) from 30 patients diagnosed at the University Medical Center Groningen, Leiden University Medical Center (The Netherlands) and the Hospital Valle del Nalon (Oviedo, Spain) (Table 1). These carcinomas were included because all contained 11q13 DNA amplification as determined previously using conventional CGH (Hermsen et al. 2001), Southern blotting (Takes et al. 1997) or WGA analysis (paper in preparation) (Table 1). The percentage of tumor cells present in the DNA sample used for FISH and CGH was established by staining parallel sections with hematoxylin and eosin. DNA from these tumors was isolated using a standard high salt extraction method. All patient samples were primary tumors that had received no therapy prior to surgery.
We also included nine HNSCC cell lines (UMSCC2, UMSCC11A, UMSCC14A, UMSCC14B and UMSCC22B as well as VUSCC59, VUSCC96a, VUSCC147 and VU947T (resp., C01–C09) that contained 11q13 amplification as reported previously (Hermsen et al. 1996; Schuuring et al. 1998). The cell lines were cultured in Dulbecco’s Modified Eagle Medium (DMEM; GIBCO BRL, Gaithersburg, MD) supplemented with 10% FCS (Cambrex Bio Science, Walkersville, Maryland), l-glutamine (GIBCO BRL) and penicillin/streptomycin (50 units/ml and 50 μg/ml) at 37°C in 5% CO2 . DNA from these cell lines was isolated using a standard high salt extraction method.
Generation of a high-resolution CGH array of the chromosome 11q13 region
We have selected 350 overlapping BAC-clones mapped to chromosome 11q13, resulting in an average two-fold coverage of the entire 11q13 region. To increase the resolution within the different 11q13 amplicon cores, various cosmid and PAC clones that were previously mapped within these loci were added to the set (Hughes-Davies et al. 2003; Janssen et al. 2000). In addition, some BAC clones (n = 109) along the chromosome 11q-arm were included on the array. The positioning of the clones was determined using NCBI build 35.1 (June 2004). The BAC clones were purchased from either BACPAC (CHORI, Oakland, USA) or Invitrogen (Breda, Netherlands). All genomic 11q13 clones on this array are listed in Supplementary Table 1. Methods to construct this CGH array were previously described (Tchinda et al. 2004). Briefly, all clones were grown and isolated in a 96-well format, adapted from Dr. M. Rocchi, University of Bari, Italy; (http://www.biologia.uniba.it/rmc/). A 3-DOP-primer based amplification (Fiegler et al. 2003) was applied to the DNA extracted from the genomic clones. These PCR products were spotted five times on epoxy-coated slides (Schott Nexterion, Mainz, Germany) using a Biorobotics Microgrid II Arrayer (Isogen Life Science, IJsselstein, The Netherlands). The array also contains a series of control spots (each five times) including DOP-PCR products from human Cot-1 DNA (Roche), total human DNA DH10B-bacterial DNA, DNA derived from BACs with Drosophila DNA inserts and spot buffer. Furthermore, the array contained 18 human chromosome X-specific clones and three chromosome Y-specific BAC clones as described previously (Kok et al. 2005). The slides were stored at 4°C until further use. Spot quality was verified by hybridization using a random panomer probe (Molecular Probes, Invitrogen, Breda).
Hybridization
Hybridization was performed using previously described protocols (Snijders et al. 2001) with small adaptations. Using the manufacture’s protocol the slides were blocked using Nexterion Block E (Schott Nexterion) prior to the hybridization procedure. Reference DNA consisted of a pool of 20 individuals (male or female) with a normal karyotype. Selected test and reference DNA samples (500 ng) were labeled using the bioprime labeling kit (Invitrogen) in combination with either Cy3-dUTP (Perkin-Elmer/NEN, cat. no. NEL 578) or Cy5-dUTP (NEL 579) added to a buffered mixture of dUTP depleted dNTPs and Klenow enzyme. An overnight incubation at 37°C was followed by removal of unincorporated nucleotides using micro-spin G50 columns (Amersham Biosciences, Uppsala, Sweden). The incorporation of labeled nucleotides was validated using a ND-1000 spectrophotometer (Nanodrop, Wilmington, USA). The test and reference DNA were pooled and further concentrated using Microcon YM-30 columns (Millipore, Billerica, MA, USA). Labeled DNA was resolved in 50 μl Nexterion Hyb (Schott Nexterion) complemented with dextran sulfate (5%), 200 μg Cot1 DNA (Roche) and 1 mg yeast t-RNA (Invitrogen). Slides were subjected to a 30–90 min prehybridization at 65°C using 50 μl of hybridization buffer (Schott Nexterion) enriched with dextran sulfate (5%) and salmon sperm DNA (10 μg/μl, Invitrogen). Hereafter, slides were washed in dH2O at 37°C for 5 min. The hybridization mixture was denatured at 100°C and applied to the array using a lifterslip™ (Erie Scientific, Portsmouth, USA). The hybridization was performed at 65°C for 42 h in a Genemachines Mica hybridization chamber (Isogen Life Science, Ijsselstein, The Netherlands). After hybridization, the slides were washed using the manufacturer’s protocol.
Image analysis and quantification
Slides were scanned on an Affymetrix 428 scanner using Jaguar software (Affymetrix, Santa Clara, CA, USA). The acquired images were quantified using Imagene software package 5.6 (Biodiscovery Inc., El Segundo, CA, USA). The mean ratio of replicate spots was quantified using our Analyze Array software version 1,6 as described by Kok et al. (2005). Briefly, a spot was included for normalization when their intensity was >2× background signal (Drosophila DNA). The median ratio of the replicate spots from the same BAC clone was determined and spots differing more than 20% from this value were excluded. Finally, BACs with only two replicate spots were also excluded. For the remaining replicas, the average ratio was calculated. If the standard deviation from the median of these replicated BACs was more than 20% (CV < 0.2), the clones were also excluded. All remaining BACs were included. However, high or low copy signals were only interpreted as possible gain or loss of DNA if at least two consecutive BACs on the array showed the same deviation from the normal ratio.
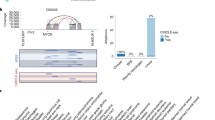
The normalized 2log ratio was used to determine the breakpoint regions with the “aCGH smooth” algorithm (Jong et al. 2004). Cell lines with previously detected copy number changes have been used to determine the parameter settings of aCGH smooth. The following changes were applied: λ = 8.75; pool size = 150 and generations = 80,000. Using these settings, sex mismatch hybridizations using a pool of karyotyped normal males (46, XY) and females (46, XX) were performed to determine technical variation. Furthermore, copy number changes of unknown normalized patients and cell lines were determined. The array-based CGH profile including the aCGH smooth results of all 39 cases are shown in Supplementary Fig. 1 (example is shown in Fig. 1a). The copy number changes (gain, amplification or loss) detected with HR-aCGH of some hybridizations were verified using interphase FISH (examples are shown in Fig. 1).
The detection of 11q13 amplification using array-CGH analysis. Example of two laryngeal carcinomas with both 11q13.3 amplification and 11q deletion (L05 and L01). a A high resolution CGH array of the chromosome 11q13 region was used to determine copy number changes. Normalized 2log ratios from HR-aCGH analysis (Y-axis) were smoothened with the aCGH smooth algorithm (Jong et al. 2004) indicated with gray bars in the CGH-profiles. Each spot represents the median ratio of various replicates from the same genomic clone (see Materials and methods). The position of the genomic clones on human chromosome 11 is indicated on the X-axis in megabases (all clones are listed in Supplementary Table 1). b To confirm the copy number changes observed by aCGH, dual colored fluorescence in situ hybridization (FISH) on interphase nuclei of the same two carcinomas was performed using a FITC labeled probe from a specific region (either I, II or III) combined with a Texas red labeled probe for the chromosome 11 centromere to control ploidy. The most common FISH pattern detected with each region-specific probe (indicated with white dots) is illustrated; the signals of centromere 11-probe are represented by black dots. c Signals of each probe were counted and the various signals were presented as the number of nuclei containing the signal. Both carcinomas contain amplification (probe I > 10 FISH signals and aCGH 2log ratio > 0.5); in both cases a considerable number of admixed normal cells represented by nuclei with two TRITC and two FITC signals were detected (16–20% in L05 and 22–37% in L01) in agreement with the percentage of normal cells estimated in a parallel H&E section (both ∼20%). The centromeric region with 2log ratio detected by aCGH between 0 and 0.5 revealed FISH signals (with probe I) with ratio of 3/3 representing gain of chromosome 11. The telomeric side with a 2log ratio close to but below 0 revealed FISH signal (with probe III) with a ratio of 2/3 representing loss of the q-arm (70% in L05 and 52% in L01) that confirmed the loss predicted by aCGH smooth
Fluorescence in situ hybridization
Fluorescence in situ hybridization (FISH) was performed on either metaphase spreads of the cell lines or interphase nuclei from snap frozen tumor samples following routine procedures (Janssen et al. 2000; Vaandrager et al. 1996). For dual color FISH, probes were labeled with digoxigenin-11-dUTP (Roche) or biotin-16-dUTP (Invitrogen) by standard nick-translation. The hybridization solution contained 50% formamide, 10% dextran sulfate, 50 mM sodium phosphate, pH = 7.0, 2 × SSC, 3 ng/μl of each probe and a 50-fold excess of human Cot-1 DNA (Invitrogen). Immunodetection was performed as described earlier (Vaandrager et al. 1996). Images were captured using a Leica DMRA2 fluorescence microscope (Leica Microsystems, Wetzlar, Germany) equipped with a Leica DC 350Fcharge-coupled device camera. Digital images were processed with Leica CW4000 software. Interfase nuclei were examined by eye and the red (Texas red) and green (FITC) signals were counted separately in approximately 100 nuclei (see Fig. 1b). To confirm copy number changes detected with aCGH, a single BAC from a selected subregion labeled with digoxigenin was co-hybridized with a centromere 11 (plasmid pLC11A) FISH-probe labeled with biotin (or vice versa). The ratio between BAC and centromere signals provides an accurate estimation of copy number differences (see Fig. 1b, c). A ratio of 1.0 indicates normal copy (or polyploidy of chromosome 11), a ratio <1 is loss and >2 is gain. Nuclei were considered to contain amplification of a particular BAC clone when the number of signals exceeded five (and ratio >2.5).
In silico nucleotide analysis
For this purpose, we used the UCSC database (Kent et al. 2002) (http://genome.ucsc.edu/; hg17, May 2004) and the NCBI human genome browser (http://www.ncbi.nlm.nih.gov/genome/guide/human/; build 35). The May 2004 freeze was used to enable optimal comparison between multiple sequence databases, some of which were not yet available in the March 2006 freeze. The two remaining sequence gaps within the 2004 freeze (chr11:68846378-68848982 and chr11:69437112-69454899) were also still present within the 2006. Thus, within the 11q13 region no significant changes in sequence order have been found between the two assemblies.
Results
Detection of specific boundaries flanking the commonly amplified 11q13.3 region in head and neck cancer
In order to determine the boundaries of the 11q13 amplicon structure we analyzed 22 laryngeal and 8 pharyngeal cancers as well as 9 HNSCC cell lines with previously identified 11q13 amplification on a high-density 11q13 specific CGH array. The array was designed to contain the highest clone density surrounding the 11q13 region (64.1–74.4 Mbp; NCBI build 35), with an average coverage of one clone per 52 kb resulting in average twofold coverage. In addition, various BAC clones along the q-arm outside the 11q13 region were included (Supplementary Table 1).
Accuracy of copy number and size detection was tested by hybridizing head and neck squamous cell carcinoma cell lines and patients containing copy number alterations that were previously identified by interphase FISH. Detecting copy number differences is often performed by comparing patients to a fixed ratio that has been determined by multiple self–self hybridizations (Veltman et al. 2002). Using such a method with fixed ratios hampers the accurate detection of copy number transitions, especially within an admixture of normal cells. Therefore, we preferred the detection of copy number transitions using a smoothing algorithm which enables detection of copy number changes without using a fixed ratio (Jong et al. 2004). Cell lines with previously detected chromosome 11q13 copy number changes were used to determine the parameter settings of aCGH smooth (Jong et al. 2004). Using these settings, copy number changes of unknown normalized patients were determined and verified applying FISH to interphase nuclei (see examples in Fig. 1). Copy numbers exceeding a 2log ratio of 0.5 represented DNA amplification. A 2log ratio between 0 and 0.5 was not considered as amplification but defined as gain because an increase of only one or two extra DNA copies was detected. The loss of DNA copies within a profile resulted in a significant decrease of the smoothened 2log ratio, generally below zero. For example, FISH on the carcinoma cells of patient L05 (Fig. 1) showed that ∼20% of all nuclei had two signals for the centromere 11 probe and two signals for each of the probes from the 11q13 subregion, whereas the majority of nuclei (∼70%) showed three signals of centromere 11 and a varying signal number for the other probes. This indicates that in this particular case ∼70% of the nuclei represent tumor. In these tumor cells the presence of amplification is confirmed by >10 signals for the amplicon probe (probe II). Furthermore, three signals were observed for the proximal probe and two for the distal probe. Because three signals for the centromere 11 probe were detected, the 2log ratio around zero represents loss of the whole 11q-arm starting immediately telomeric of the amplicon (Fig. 1, probe III). Patient L01 showed a similar pattern. The most important finding in this study was that the smoothing algorithm enabled the accurate identification of copy number transitions as well as their location, even in the presence of an admixture of normal cells within the tumor sample. As summarized in Fig. 2, in 29 patients and nine cell lines with previously identified 11q13 amplifications, DNA amplification was also detected with HR-aCGH. One case (P05) with previously detected (low copy) 11q13 amplification did not show amplification using 11q13 aCGH analysis (in Supplementary Fig. 1, the array-based CGH profiles including the aCGH smoothing analysis of all 39 cases are shown). Most amplicons were found within the same region (67–72 Mbp from the p-arm telomere). HR-aCGH and smoothing analysis allowed us to determine the boundaries of an amplicon at a resolution of approximately 52 kb. This analysis revealed that the boundaries are not randomly distributed but clustered within four specific positions (Fig. 3, lower panel). On the centromeric side two boundaries are detected: one in 8/38 cases (boundary 1; ∼400 kb in size) and one more common in 21/38 cases (boundary 2; ∼200 kb in size). On the telomeric side 10/38 cases were clustered in boundary 3 (∼100 kb) and another 12/38 in boundary 4 (∼300 kb in size) (Fig. 3).
The location and size of the 11q13 amplicon in 30 head/neck cancers and nine cell lines detected with 11q13 high-density array CGH. The position and size of the 11q13 amplicon were determined by 11q13 high-density array CGH for each case (in Supplementary Fig. 1 all HR-aCGH-profiles are illustrated). DNA amplification (2log ratio > 0.5) is depicted as thick bars and gain (2log ratio between 0 and 0.5 and not considered as amplification) as thin bars for each of the 39 patients (see Table 1 and Materials and methods) listed on the Y-axis. The position on chromosome 11 is indicated in megabases on the X-axis relative to a representation of the chromosome 11 q-arm (according to NCBI Build 35)
Four common boundaries flanking the 11q13 amplicons coincide with specific repeat sequences and chromosomal transitions in other species. The amplicons within the 67–72 Mbp 11q13.3 region of the 39 cases (from Fig. 2) are represented as thick black bars (thin bars represent gains). The four regions with frequent copy number transitions are indicated as vertical gray blocks and numbered boundary 1 to 4. The position and size of the segmental duplications is indicated by blue boxes. Superimposed, a synteny sequence comparison of the human chromosome 11q13 (position 67–72 Mbp) is shown with sequences from different organisms shown in the phylogenic tree (downloaded from the NET tab at the UCSC genome browser, http://hgwdev.cse.ucsc.edu) (Kent et al. 2002). The synteny comparison shows a sequence comparison (by blastz) between a specific species and the human 11q13 region. Chains of sequence are shown as colored boxes referring to the homologous chromosome from the compared species. Gaps in chains of sequence are displayed as threads and angle brackets (< or >) mark the orientation. All of the tracks and color codes can be found at http://hgwdevbaertsch. cse.ucsc.edu/cgi-bin/hgTracks. The arrows mark the position of the syntenic transitions between human 11q13.3 sequences and mouse (also see Fig. 4) and cow. The gaps and/or discontinuity at positions 67.2–67.5 and 71.0–71.2 mark the regions that are not present in most other organism; those regions indicate the presence of segmental duplications (see text). The position and size of the segmental duplications is indicated by blue boxes
In addition, HR-aCGH analysis revealed that 30/38 cases had 11q13 amplification with accompanying deletion of 11q-ter. This 11q-deletion started immediately adjacent to the amplicon at 11q13.3 in 25/38 cases and more distal from the amplicon in 10/38 cases (Supplementary Fig. 1 and examples in Fig. 1).
Amplicon boundaries are flanked by synteny transitions
In order to study whether the four common boundaries were associated with specific sequences related to DNA fragility, we performed an in silico analysis of the region containing most 11q13.3 amplifications (from position 67–72 Mb). Fusion of chromosomes during evolution might preferably occur at fragile regions and can be considered as possible breakpoint regions in the process of DNA amplification (Yuan et al. 1997). Therefore, we compared human chromosome 11 (hChr11) to homologous sequences of a number of sequenced mammals that are phylogenetically close to human (Fig. 3). No synteny breakpoints between human and chimpanzee (Pan trochlodytes) were observed because hChr11 is highly conserved in higher apes. However, when the q13 region of hChr11 was compared to the mouse genome (Mus musculus), regions of high similarity were found for mouse chromosome 7 (mChr7) and 19 (mChr19) (Figs. 3, 4). Two separate regions on mChr7 were found to be homologous to a continuous region on hChr11 (regions II and III in Fig. 4). The transition between these mouse chromosomes is located at approximately 68.6 Mbp on hChr11 (NCBI B35, UCSC hg17) that coincides with boundary 2 detected in 21/38 cases (Fig. 3). The transition between mChr7 region II and mChr19 region III is located at approximately 70.9 Mb and coincides with boundary 4 observed in 12/38 cases (Fig. 3). This comparison reveals that the telomeric locus in the mouse genome conserved during evolution coincides with the commonly amplified locus at human 11q13.3 between boundaries 2 and 4 (region II in Fig. 4). Boundary 3, which is determined by 10/38 cases, coincides with a transition in the cow genome (Fig. 3). Immediately flanking boundary 4 on the telomeric side and boundary 1 on the centromeric side, the region on hChr11 does not contain any homology to other species (Fig. 3). After this “synteny gap”, there is a chromosomal transition between hChr11 and its homologous counterparts in macaque, mouse, rat, dog and cow. In summary, within the region between 67 and 72 Mb on hChr11 we identified four regions containing syntenic transitions. These transitions coincided with three of the four common amplicon boundaries, implying that these regions are prone to break.
Schematic representation of the syntenic homology between the human chromosome 11q13.3 region and the mouse genome. Nucleotide sequence comparison revealed homology between human chromosome 11 (hChr11) and mouse chromosome 7 (mChr7) and 19 (mChr19) using the ENSEMBL synteny viewer (http://www.ensembl.org/Homo_sapiens/syntenyview). The homology between the large arm of hChr11 and mChr 7 and 19 can be subdivided into three regions (I, II and III). The human chromosome 11q13.3 region that is commonly amplified (between boundary 2 and 4; marked as II) shows homology with a telomeric part region of the mChr7 telomere. The sequence on the centromeric side of the human 11q13.3 amplicon (marked I) is homologous to sequence from mChr19. The sequence on the telomeric side of the human 11q13.3 amplicon (marked III) is homologous to a region on mChr7 and does not flank the region homologous to region II
Segmental duplications flank the 11q13.3 amplicon
We have found two large chromosomal structures flanking the 11q13.3 amplicon that can be identified as a “synteny gap”, i.e. there is no high level homology between this part of hChr11 and other species (Fig. 3). Further analysis revealed that these regions (67.20–67.55 Mbp and 70.95–71.30 Mbp) consisted of large repeats with an inverted orientation (Fig. 5). Such large repeats are known as low copy repeats (LCRs) or segmental duplications (SDs) (Bailey and Eichler 2006; Locke et al. 2004; Stankiewicz et al. 2004). Both segmental duplications at 11q13.3 are part of a family of segmental duplications that is known to contain clusters of olfactory receptors (Newman and Trask 2003; Olender et al. 2004). These clusters often reside near syntenic breakpoint regions (Bailey et al. 2004; Yue and Haaf 2006). We tested for homologous regions of these SDs in silico by nucleotide sequence comparisons using the BLAT search program (http://genome.ucsc.edu) and two different fragments of 25 kb within the SDs. This comparison revealed a homology >95% between both regions (Table 2). Furthermore, the use of the segmental duplications tab at http://hgwdev.cse.ucsc.edu containing experimental tracks of the UCSC Genome Bioinformatics Group revealed high homology between the telomeric segmental duplication at 11q13 and segmental duplications on chromosome 4p16, 8p23 and 3q21. These regions were also found in the in silico sequence comparison (Table 2, the telomeric SD). We performed dual color FISH on metaphase preparations from lymphocytes of a healthy individual using a BAC clone from the telomeric SD (RP11-167J8) in combination with either the chromosome 11 centromere probe (Fig. 6a) or the chromosome centromere 8 probe (Fig. 6b). Multiple copies of the telomeric SD were detected on different chromosomes in agreement with the sequence comparison analysis (Table 2, the telomeric SD).
The structure of the segmental duplications flanking the 11q13 amplicon. Alignment of 500 kb sequence containing the centromeric segmental duplication (top 67.1–67.6 Mbp) compared to the telomeric segmental duplication (bottom 70.9–71.4 Mbp) was performed using the MAUVE multiple genome alignment tool (http://gel.ahabs.wisc.edu/mauve/). The interconnecting lines between both sequences (boxes) indicate blocks with high homology located in an inversed orientation
Segmental duplication detected by metaphase FISH. We have performed a dual color metaphase FISH on normal lymphocytes using a BAC clone located within the centromeric segmental duplication (RP11-167j8) labeled with FITC in combination with the centromere 11 probe (pLC11A) labeled with Texas red (asterisk in a) or the chromosome centromere 8 probe (D8Z2) labeled with Texas red (asterisk in b). The segmental duplication signals are not restricted to chromosome 11 and 8. The telomeric segmental duplication BAC clone (RP11-684b2) gave similar results indicating its overlap with the centromeric segmental duplication (data not shown)
Segmental duplications have been shown to be involved in the formation of genetic disorders (Stankiewicz and Lupski 2002), but their involvement in cancer is yet to be proven. The repeat structure of these SDs surrounding the 11q13.3 amplicon might underlie formation of secondary DNA structures creating loops and fragility. Because fragility has been associated with repeat content also, we investigated the 11q13.3 region for differences in GC-content, and the presence of LINE, SINE and SATR1/SATR2 repeats (http://hgwdev.cse.ucsc.edu). In this region we observed no clear differences in GC or repeat content, except that the segmental duplications at 11q13 are flanked by the clusters of SATR1 and SATR2 satellite repeats (data not shown) as has previously been reported for olfactory receptor containing segmental duplications (Newman and Trask 2003; Stankiewicz et al. 2004).
Discussion
Chromosomal amplification is initiated by double strand breaks
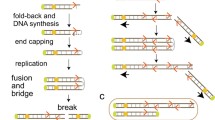
On the basis of their finding that the 11q13 amplicon is flanked by inverted duplicated segments, Shuster et al. (2000) suggested that 11q13 amplification is most likely to arise through the mechanism of breakage-fusion-bridge (BFB) cycles. This model is used to explain the occurrence of DNA amplification of many other chomosomal loci (Toledo et al. 1992; Windle et al. 1991). Even though BFB cycles seem to underlie the formation of amplicons (Toledo et al. 1992; Trask and Hamlin 1989), the exact mechanism is yet to be proven. Many clues for the involvement of the BFB mechanism have been elucidated in in vitro systems, because it is difficult to identify and modify parts of the mechanism in vivo (Coquelle et al. 2002; Kuo et al. 1998; Okuno et al. 2004; Pipiras et al. 1998; Schimke 1988; Toledo et al. 1992; Trask and Hamlin 1989). One of the most important prerequisites for the induction of amplification is the involvement of two double strand breaks surrounding the key gene(s) within an amplicon (Coquelle et al. 2002; Difilippantonio et al. 2002; Kuo et al. 1998; Mills et al. 2003; Pipiras et al. 1998; Windle et al. 1991). Since amplification plays a potent role in carcinogenesis, we have tried to gain insight into the mechanism of amplification of the 11q13 region by investigating the genomic structure at the position of copy number transitions in head and neck cancer that frequently shows amplification of the 11q13 region (Schuuring 1995). We used CGH array for the detection of 11q13 amplifications, which enabled us to map the boundaries of all amplicons at high resolution. For this purpose, we generated a CGH array for the 11q13 region (position 64.1–77.4 Mbp) using a set of overlapping genomic clones that cover this region at least twice with an average coverage of one clone per 52 kb. The resolution of our array is higher than any other mapping strategy previously reported for this region (Freier et al. 2006; Huang et al. 2002; Hui et al. 2005; Lin et al. 2006). The boundaries in 29 human carcinomas and nine cell lines clustered in four specific regions (Fig. 2), implicating a local susceptibility for DNA breakage. In silico analysis revealed that three of these regions co-localized with syntenic chromosomal transitions in the mouse and cow genome. Remarkably, the region containing amplicons was flanked by two SDs.
The mechanism of BFB requires the occurrence of at least two double strand breaks. The first break would lead to the loss of the telomere. The second break occurs at the bridge phase, when cell division forces the dicentric chromosome to break apart (Toledo et al. 1992). It is the position of the first break and the subsequent attempt to repair it, which determines whether BFB occurs. Therefore, telomeric loss should be detected frequently if BFB is the mechanism of amplification. Our aCGH data revealed that in 25/38 cases with 11q13 amplification telomeric 11q-sequences are indeed deleted with a breakpoint adjacent to the region of amplification (Fig. 1 and Supplementary Fig. 1). Using FISH and conventional CGH analysis in cell lines with amplification at 11q13.3, loss of telomeric DNA sequences were reported previously (Hermsen et al. 1997; Jin et al. 1998). This implies that the break causing telomeric loss is probably also responsible for initiating multiple rounds of amplification. Recovering from this cycle of BFB requires either the re-activation of telomerase or DNA recombination to capture telomeres from intact chromosomes (Maser and DePinho 2002). Both mechanisms serve to alleviate the activation of checkpoints similar as DNA repair allows cells to re-enter growth after DNA damage (Maser and DePinho 2004). Interestingly, several DNA repair genes are located on the chromosome 11q-arm including MRE11A, ATM, H2AFX and CHK1. Therefore, deletion of the 11q telomere might further enhance genomic instability by the functional loss of these DNA repair genes (Lobachev et al. 2002; Reshmi and Gollin 2005).
Studies on clinical specimens have shown that overexpression of the cyclin D1 gene is associated with chromosomal abnormalities (Jin et al. 2002; Nimeus et al. 2004). Increased expression of cyclin D1 in normal hepatocytes in vitro triggered marked chromosomal abnormalities (Nelsen et al. 2005). In a considerable number of head and neck carcinomas with 11q13 amplification and concomitant cyclin D1 overexpression, increased expression was also detected in the premalignant lesions adjacent to the carcinoma, however, without 11q13 amplification (Izzo et al. 1998). This indicated that increased cyclin D1 expression is also involved in the initiation of genetic instability including gene amplification in early head and neck cancer. Because the cyclin D1 gene itself is located within the 11q13 amplicon, its overexpression due to 11q13 amplification might maintain and/or enhance further genomic instability in head and neck cancer. (Izzo et al. 1998; Nimeus et al. 2004). Thus, genetic instability could be an explanation for the frequent 11q13.3 amplification in multiple tumor types (Albertson 2006).
Previously, syntenic transitions surrounding the 11q13.3 amplicon were suggested to be transitions in replication timing (Watanabe et al. 2002, 2004; Woodfine et al. 2004). The change in replication timing is indicative of differences in DNA structure (e.g. GC and repeat content), but has not been assigned to specific sequences. Yet, the change in replication timing as described by Watanabe et al. (2004) overlaps with boundary 2 in our analysis. A replication timing difference between fragile sites and surrounding regions has also been suggested (Le Beau et al. 1998; Palakodeti et al. 2004; Sen et al. 1989). Fragile sites are potential sites for DNA double strand breakage and possible initiators for BFB-mediated amplification (Kuo et al. 1998). Although it is known that clastogenic drugs cause DNA breakage, it is unknown why these breaks are not sequence specific and occur within regions of up to several megabases (Richards 2001). Within the chromosome 11 cytogenetic band 13 three fragile sites have been mapped previously. Even though the exact location is unknown, FRA11A and FRA11H are mapped further centromeric (Hagemeijer et al. 1991; Perucca-Lostanlen et al. 1997) and FRA11F further telomeric to 11q13.3 (Craig-Holmes et al. 1987). We propose that these 11q13 fragile sites are not involved in the fragility of chromosomal breakage necessary for 11q13 amplification, whereas the specific chromosomal structure of the 11q13.3 region marked by the presence of both syntenic transitions and segmental duplications is.
The role of segmental duplications in amplification
The specific homology between human chromosome 11q13.3 and the telomeric part of mouse chromosome 7 provides a possible evolutionary explanation for the structural difference between 11q13.3 and flanking regions. With palindromic segmental duplications flanking the 11q13 region, large secondary DNA structures might be formed due to sequence homology. Segmental duplications could cause fragility similar to palindromic sequences and alu repeats, which have previously been implicated in double strand break mediated amplification (Narayanan et al. 2006; Tanaka et al. 2002). Segmental duplications have been reported to be involved in many genetic aberrations (Lupski 1998) and in some hematological malignancies (Barbouti et al. 2004). Amplification at 17p11.2∼p12 has been linked to the presence of SDs (van Dartel and Hulsebos 2004), but a general role for SD mediated amplification is yet to be proven. However, the involvement of SDs in chromosomal deletions involved in genomic disorders (Shaw and Lupski 2004) as well as chromosomal duplications mediated by segmental duplications has been established (De Gregori et al. 2005). For instance, congenital heart defects are associated with deletions on distal 8p (del8p) (Devriendt et al. 1999) and recurrent translocations have been identified between 4p16 and 8p23 (Giglio et al. 2002). Strikingly, these aberrations are found within regions that are flanked by two SDs sharing high homology with the 11q13 SDs (Table 2). Furthermore, Saunders et al. (2000) have described dicentric chromosomes in oral squamous cell carcinoma containing 11q13 amplifications as well as translocations between chromosome 11, 4 and 8 and 3. These translocations could be mediated by homology between the segmental duplications. The mechanism underlying amplicon formation involving EGFR (Vogt et al. 2004), MYCN (Savelyeva and Schwab 2001) and ERBB2 (Coquelle et al. 1997) seems structurally different from the complex amplicon that is formed at 11q13 (Albertson 2006). The involvement of the segmental duplications might thus be a unique feature of amplification at 11q13. This genomic structure at the chromosomal 11q13 region will have important implications for the identification of a key gene in the 11q13 amplicon. Since the amplicon size of the 11q13 region in head and neck cancer is determined by unique genomic structures, not a single gene but a set of genes will be selected in most carcinomas. This is reflected by the fact that our current analysis does not lead to a region containing a specific gene of interest (J.H. Gibcus et al., unpublished data) in agreement with previous findings by others (Freier et al. 2006; Huang et al. 2002; Hui et al. 2005) including a recently published paper by Huang et al. (2006). Therefore, several genes remain candidates for driving the amplification, including cyclin D1, TAOS1, FGF19, FADD, PPFIA1 and cortactin. Functional analysis of the candidate genes is needed to identify the gene(s) relevant to tumor progression.
In this manuscript, we have compared the breakpoint patterns of 38 head and neck carcinomas and cell lines with 11q13 amplification in detail. Within the 11q13.3 region amplifications occur at specific regions that coincide with regions of synteny transition in the mouse and cow genome. Furthermore, the transitions are flanked by segmental duplications. Other studies have reported enrichment of segmental duplications near synteny breaks (Armengol et al. 2003), indicating that the same DNA structure that enabled chromosomal breakage during evolution and the insertion of a SD might also be involved in the initiation of amplification.
References
Albertson DG (2006) Gene amplification in cancer. Trends Genet 22(8):447–455
Armengol L, Pujana MA, Cheung J, Scherer SW, Estivill X (2003) Enrichment of segmental duplications in regions of breaks of synteny between the human and mouse genomes suggest their involvement in evolutionary rearrangements. Hum Mol Genet 12:2201–2208
Atayar C, Kok K, Kluiver J, Bosga A, van den Berg E, van d V, Blokzijl T, Harms G, Davelaar I, Sikkema-Raddatz B, Martin-Subero JI, Siebert R, Poppema S, van den Berg A (2006) BCL6 alternative breakpoint region break and homozygous deletion of 17q24 in the nodular lymphocyte predominance type of Hodgkin’s lymphoma-derived cell line DEV. Hum Pathol 37:675–683
Bailey JA, Eichler EE (2006) Primate segmental duplications: crucibles of evolution, diversity and disease. Nat Rev Genet 7:552–564
Bailey JA, Baertsch R, Kent WJ, Haussler D, Eichler EE (2004) Hotspots of mammalian chromosomal evolution. Genome Biol 5:R23
Barbouti A, Stankiewicz P, Nusbaum C, Cuomo C, Cook A, Hoglund M, Johansson B, Hagemeijer A, Park SS, Mitelman F, Lupski JR, Fioretos T (2004) The breakpoint region of the most common isochromosome, i(17q), in human neoplasia is characterized by a complex genomic architecture with large, palindromic, low-copy repeats. Am J Hum Genet 74:1–10
Barr FG, Nauta LE, Davis RJ, Schafer BW, Nycum LM, Biegel JA (1996) In vivo amplification of the PAX3-FKHR and PAX7-FKHR fusion genes in alveolar rhabdomyosarcoma. Hum Mol Genet 5:15–21
Ciullo M, Debily MA, Rozier L, Autiero M, Billault A, Mayau V, El Marhomy S, Guardiola J, Bernheim A, Coullin P, Piatier-Tonneau D, Debatisse M (2002) Initiation of the breakage-fusion-bridge mechanism through common fragile site activation in human breast cancer cells: the model of PIP gene duplication from a break at FRA7I. Hum Mol Genet 11:2887–2894
Coquelle A, Pipiras E, Toledo F, Buttin G, Debatisse M (1997) Expression of fragile sites triggers intrachromosomal mammalian gene amplification and sets boundaries to early amplicons. Cell 89:215–225
Coquelle A, Rozier L, Dutrillaux B, Debatisse M (2002) Induction of multiple double-strand breaks within an hsr by meganucleaseI-SceI expression or fragile site activation leads to formation of double minutes and other chromosomal rearrangements. Oncogene 21:7671–7679
Craig-Holmes AP, Strong LC, Goodacre A, Pathak S (1987) Variation in the expression of aphidicolin-induced fragile sites in human lymphocyte cultures. Hum Genet 76:134–137
De Gregori M, Pramparo T, Memo L, Gimelli G, Messa J, Rocchi M, Patricelli MG, Ciccone R, Giorda R, Zuffardi O (2005) Direct duplication 12p11.21–p13.31 mediated by segmental duplications: a new recurrent rearrangement? Hum Genet 118:207–213
Devriendt K, Matthijs G, Van Dael R, Gewillig M, Eyskens B, Hjalgrim H, Dolmer B, McGaughran J, Brondum-Nielsen K, Marynen P, Fryns JP, Vermeesch JR (1999) Delineation of the critical deletion region for congenital heart defects, on chromosome 8p23.1. Am J Hum Genet 64:1119–1126
Difilippantonio MJ, Petersen S, Chen HT, Johnson R, Jasin M, Kanaar R, Ried T, Nussenzweig A (2002) Evidence for replicative repair of DNA double-strand breaks leading to oncogenic translocation and gene amplification. J Exp Med 196:469–480
Fiegler H, Carr P, Douglas EJ, Burford DC, Hunt S, Smith J, Vetrie D, Gorman P, Tomlinson IP, Carter NP (2003) DNA microarrays for comparative genomic hybridization based on DOP-PCR amplification of BAC and PAC clones. Genes Chromosomes Cancer 36:361–374
Freier K, Sticht C, Hofele C, Flechtenmacher C, Stange D, Puccio L, Toedt G, Radlwimmer B, Lichter P, Joos S (2006) Recurrent coamplification of cytoskeleton-associated genes EMS1 and SHANK2 with CCND1 in oral squamous cell carcinoma. Genes Chromosomes Cancer 45:118–125
Giglio S, Calvari V, Gregato G, Gimelli G, Camanini S, Giorda R, Ragusa A, Guerneri S, Selicorni A, Stumm M, Tonnies H, Ventura M, Zollino M, Neri G, Barber J, Wieczorek D, Rocchi M, Zuffardi O (2002) Heterozygous submicroscopic inversions involving olfactory receptor-gene clusters mediate the recurrent t(4;8)(p16;p23) translocation. Am J Hum Genet 71:276–285
Hagemeijer A, Lafage M, Mattei MG, Simonetti J, Smit E, de LO, Birnbaum D (1991) Localization of the HST/FGFK gene with regard to 11q13 chromosomal breakpoint and fragile site. Genes Chromosomes Cancer 3:210–214
Hermsen MA, Joenje H, Arwert F, Welters MJ, Braakhuis BJ, Bagnay M, Westerveld A, Slater R (1996) Centromeric breakage as a major cause of cytogenetic abnormalities in oral squamous cell carcinoma. Genes Chromosomes Cancer 15:1–9
Hermsen MA, Joenje H, Arwert F, Braakhuis BJ, Baak JP, Westerveld A, Slater R (1997) Assessment of chromosomal gains and losses in oral squamous cell carcinoma by comparative genomic hybridisation. Oral Oncol 33:414–418
Hermsen M, Guervos MA, Meijer G, Baak J, van Diest P, Marcos CA, Sampedro A (2001) New chromosomal regions with high-level amplifications in squamous cell carcinomas of the larynx and pharynx, identified by comparative genomic hybridization. J Pathol 194:177–182
Huang X, Gollin SM, Raja S, Godfrey TE (2002) High-resolution mapping of the 11q13 amplicon and identification of a gene, TAOS1, that is amplified and overexpressed in oral cancer cells. Proc Natl Acad Sci USA 99:11369–11374
Huang X, Godfrey TE, Gooding WE, McCarty KS Jr, Gollin SM (2006) Comprehensive genome and transcriptome analysis of the 11q13 amplicon in human oral cancer and synteny to the 7F5 amplicon in murine oral carcinoma. Genes Chromosomes Cancer 45:1058–1069
Hughes-Davies L, Huntsman D, Ruas M, Fuks F, Bye J, Chin SF, Milner J, Brown LA, Hsu F, Gilks B, Nielsen T, Schulzer M, Chia S, Ragaz J, Cahn A, Linger L, Ozdag H, Cattaneo E, Jordanova ES, Schuuring E, Yu DS, Venkitaraman A, Ponder B, Doherty A, Aparicio S, Bentley D, Theillet C, Ponting CP, Caldas C, Kouzarides T (2003) EMSY links the BRCA2 pathway to sporadic breast and ovarian cancer. Cell 115:523–535
Hui AB, Or YY, Takano H, Tsang RK, To KF, Guan XY, Sham JS, Hung KW, Lam CN, van Hasselt CA, Kuo WL, Gray JW, Huang DP, Lo KW (2005) Array-based comparative genomic hybridization analysis identified cyclin D1 as a target oncogene at 11q13.3 in nasopharyngeal carcinoma. Cancer Res 65:8125–8133
Izzo JG, Papadimitrakopoulou VA, Li XQ, Ibarguen H, Lee JS, Ro JY, El Naggar A, Hong WK, Hittelman WN (1998) Dysregulated cyclin D1 expression early in head and neck tumorigenesis: in vivo evidence for an association with subsequent gene amplification. Oncogene 17:2313–2322
Janssen JW, Vaandrager JW, Heuser T, Jauch A, Kluin PM, Geelen E, Bergsagel PL, Kuehl WM, Drexler HG, Otsuki T, Bartram CR, Schuuring E (2000) Concurrent activation of a novel putative transforming gene, myeov, and cyclin D1 in a subset of multiple myeloma cell lines with t(11;14)(q13;q32). Blood 95:2691–2698
Jin Y, Hoglund M, Jin C, Martins C, Wennerberg J, Akervall J, Mandahl N, Mitelman F, Mertens F (1998) FISH characterization of head and neck carcinomas reveals that amplification of band 11q13 is associated with deletion of distal 11q. Genes Chromosomes Cancer 22:312–320
Jin Y, Jin C, Wennerberg J, Hoglund M, Mertens F (2002) Cyclin D1 amplification in chromosomal band 11q13 is associated with overrepresentation of 3q21–q29 in head and neck carcinomas. Int J Cancer 98:475–479
Jong K, Marchiori E, Meijer G, Van der Vaart A, Ylstra B (2004) Breakpoint identification and smoothing of array comparative genomic hybridization data. Bioinformatics 20:3636–3637
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Kok K, Dijkhuizen T, Swart YE, Zorgdrager H, van d V, Fehrmann R, te Meerman GJ, Gerssen-Schoorl KB, van ET, Sikkema-Raddatz B, Buys CH (2005) Application of a comprehensive subtelomere array in clinical diagnosis of mental retardation. Eur J Med Genet 48:250–262
Kuo MT, Sen S, Hittelman WN, Hsu TC (1998) Chromosomal fragile sites and DNA amplification in drug-resistant cells. Biochem Pharmacol 56:7–13
Le Beau MM, Rassool FV, Neilly ME, Espinosa R, III, Glover TW, Smith DI, McKeithan TW (1998) Replication of a common fragile site, FRA3B, occurs late in S phase and is delayed further upon induction: implications for the mechanism of fragile site induction. Hum Mol Genet 7:755–761
Lin M, Smith LT, Smiraglia DJ, Kazhiyur-Mannar R, Lang JC, Schuller DE, Kornacker K, Wenger R, Plass C (2006) DNA copy number gains in head and neck squamous cell carcinoma. Oncogene 25:1424–1433
Lobachev KS, Gordenin DA, Resnick MA (2002) The Mre11 complex is required for repair of hairpin-capped double-strand breaks and prevention of chromosome rearrangements. Cell 108:183–193
Locke DP, Segraves R, Nicholls RD, Schwartz S, Pinkel D, Albertson DG, Eichler EE (2004) BAC microarray analysis of 15q11–q13 rearrangements and the impact of segmental duplications. J Med Genet 41:175–182
Lupski JR (1998) Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet 14:417–422
Maser RS, DePinho RA (2002) Connecting chromosomes, crisis, and cancer. Science 297:565–569
Maser RS, DePinho RA (2004) Telomeres and the DNA damage response: why the fox is guarding the henhouse. DNA Repair (Amst) 3:979–988
Mills KD, Ferguson DO, Alt FW (2003) The role of DNA breaks in genomic instability and tumorigenesis. Immunol Rev 194:77–95
Narayanan V, Mieczkowski PA, Kim HM, Petes TD, Lobachev KS (2006) The pattern of gene amplification is determined by the chromosomal location of hairpin-capped breaks. Cell 125:1283–1296
Nelsen CJ, Kuriyama R, Hirsch B, Negron VC, Lingle WL, Goggin MM, Stanley MW, Albrecht JH (2005) Short term cyclin D1 overexpression induces centrosome amplification, mitotic spindle abnormalities, and aneuploidy. J Biol Chem 280:768–776
Newman T, Trask BJ (2003) Complex evolution of 7E olfactory receptor genes in segmental duplications. Genome Res 13:781–793
Nimeus E, Baldetorp B, Bendahl PO, Rennstam K, Wennerberg J, Akervall J, Ferno M (2004) Amplification of the cyclin D1 gene is associated with tumour subsite, DNA non-diploidy and high S-phase fraction in squamous cell carcinoma of the head and neck. Oral Oncol 40:624–629
Okuno Y, Hahn PJ, Gilbert DM (2004) Structure of a palindromic amplicon junction implicates microhomology-mediated end joining as a mechanism of sister chromatid fusion during gene amplification. Nucleic Acids Res 32:749–756
Olender T, Feldmesser E, Atarot T, Eisenstein M, Lancet D (2004) The olfactory receptor universe–from whole genome analysis to structure and evolution. Genet Mol Res 3:545–553
Palakodeti A, Han Y, Jiang Y, Le Beau MM (2004) The role of late/slow replication of the FRA16D in common fragile site induction. Genes Chromosomes Cancer 39:71–76
Perucca-Lostanlen D, Hecht BK, Courseaux A, Grosgeorge J, Hecht F, Gaudray P (1997) Mapping FRA11A, a folate-sensitive fragile site in human chromosome band 11q13.3. Cytogenet Cell Genet 79:88–91
Pipiras E, Coquelle A, Bieth A, Debatisse M (1998) Interstitial deletions and intrachromosomal amplification initiated from a double-strand break targeted to a mammalian chromosome. EMBO J 17:325–333
Reshmi SC, Gollin SM (2005) Chromosomal instability in oral cancer cells. J Dent Res 84:107–117
Richards RI (2001) Fragile and unstable chromosomes in cancer: causes and consequences. Trends Genet 17:339–345
Saunders WS, Shuster M, Huang X, Gharaibeh B, Enyenihi AH, Petersen I, Gollin SM (2000) Chromosomal instability and cytoskeletal defects in oral cancer cells. Proc Natl Acad Sci USA 97:303–308
Savelyeva L, Schwab M (2001) Amplification of oncogenes revisited: from expression profiling to clinical application. Cancer Lett 167:115–123
Schimke RT (1988) Gene amplification in cultured cells. J Biol Chem 263:5989–5992
Schuuring E (1995) The involvement of the chromosome 11q13 region in human malignancies: cyclin D1 and EMS1 are two new candidate oncogenes–a review. Gene 159:83–96
Schuuring E, van DH, Schuuring-Scholtes E, Verhoeven E, Michalides R, Geelen E, de BC, Brok H, van B, V, Kluin P (1998) Characterization of the EMS1 gene and its product, human Cortactin. Cell Adhes Commun 6:185–209
Schwab M (1999) Oncogene amplification in solid tumors. Semin Cancer Biol 9:319–325
Sen S, Hittelman WN, Teeter LD, Kuo MT (1989) Model for the formation of double minutes from prematurely condensed chromosomes of replicating micronuclei in drug-treated Chinese hamster ovary cells undergoing DNA amplification. Cancer Res 49:6731–6737
Shaw CJ, Lupski JR (2004) Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet 13 Spec No 1:R57–R64
Shuster MI, Han L, Le Beau MM, Davis E, Sawicki M, Lese CM, Park NH, Colicelli J, Gollin SM (2000) A consistent pattern of RIN1 rearrangements in oral squamous cell carcinoma cell lines supports a breakage-fusion-bridge cycle model for 11q13 amplification. Genes Chromosomes Cancer 28:153–163
Snijders AM, Nowak N, Segraves R, Blackwood S, Brown N, Conroy J, Hamilton G, Hindle AK, Huey B, Kimura K, Law S, Myambo K, Palmer J, Ylstra B, Yue JP, Gray JW, Jain AN, Pinkel D, Albertson DG (2001) Assembly of microarrays for genome-wide measurement of DNA copy number. Nat Genet 29:263–264
Stankiewicz P, Lupski JR (2002) Molecular-evolutionary mechanisms for genomic disorders. Curr Opin Genet Dev 12:312–319
Stankiewicz P, Shaw CJ, Withers M, Inoue K, Lupski JR (2004) Serial segmental duplications during primate evolution result in complex human genome architecture. Genome Res 14:2209–2220
Stark GR, Debatisse M, Giulotto E, Wahl GM (1989) Recent progress in understanding mechanisms of mammalian DNA amplification. Cell 57:901–908
Takes RP, Baatenburg de Jong RJ, Schuuring E, Hermans J, Vis AA, Litvinov SV, Van Krieken JH (1997) Markers for assessment of nodal metastasis in laryngeal carcinoma. Arch Otolaryngol Head Neck Surg 123:412–419
Tanaka H, Tapscott SJ, Trask BJ, Yao MC (2002) Short inverted repeats initiate gene amplification through the formation of a large DNA palindrome in mammalian cells. Proc Natl Acad Sci USA 99:8772–8777
Tchinda J, Dijkhuizen T, van der Vlies P, Kok K, Horst J (2004) Translocations involving 6p22 in acute myeloid leukaemia at relapse: breakpoint characterization using microarray-based comparative genomic hybridization. Br J Haematol 126:495–500
Toledo F, Le Roscouet D, Buttin G, Debatisse M (1992) Co-amplified markers alternate in megabase long chromosomal inverted repeats and cluster independently in interphase nuclei at early steps of mammalian gene amplification. EMBO J 11:2665–2673
Toledo F, Coquelle A, Svetlova E, Debatisse M (2000) Enhanced flexibility and aphidicolin-induced DNA breaks near mammalian replication origins: implications for replicon mapping and chromosome fragility. Nucleic Acids Res 28:4805–4813
Trask BJ, Hamlin JL (1989) Early dihydrofolate reductase gene amplification events in CHO cells usually occur on the same chromosome arm as the original locus. Genes Dev 3:1913–1925
Vaandrager JW, Schuuring E, Zwikstra E, de Boer CJ, Kleiverda KK, Van Krieken JH, Kluin-Nelemans HC, van Ommen GJ, Raap AK, Kluin PM (1996) Direct visualization of dispersed 11q13 chromosomal translocations in mantle cell lymphoma by multicolor DNA fiber fluorescence in situ hybridization. Blood 88:1177–1182
van Dartel M., Hulsebos TJ (2004) Amplification and overexpression of genes in 17p11.2 ∼ p12 in osteosarcoma. Cancer Genet Cytogenet 153:77–80
Van Roy N., Vandesompele J, Menten B, Nilsson H, De SE, Rocchi M, De PA, Pahlman S, Speleman F (2006) Translocation-excision-deletion-amplification mechanism leading to nonsyntenic coamplification of MYC and ATBF1. Genes Chromosomes Cancer 45:107–117
Veltman JA, Schoenmakers EF, Eussen BH, Janssen I, Merkx G, van Cleef B, van Ravenswaaij CM, Brunner HG, Smeets D, van Kessel AG (2002) High-throughput analysis of subtelomeric chromosome rearrangements by use of array-based comparative genomic hybridization. Am J Hum Genet 70:1269–1276
Vogt N, Lefevre SH, Apiou F, Dutrillaux AM, Cor A, Leuraud P, Poupon MF, Dutrillaux B, Debatisse M, Malfoy B (2004) Molecular structure of double-minute chromosomes bearing amplified copies of the epidermal growth factor receptor gene in gliomas. Proc Natl Acad Sci USA 101:11368–11373
Watanabe Y, Fujiyama A, Ichiba Y, Hattori M, Yada T, Sakaki Y, Ikemura T (2002) Chromosome-wide assessment of replication timing for human chromosomes 11q and 21q: disease-related genes in timing-switch regions. Hum Mol Genet 11:13–21
Watanabe Y, Ikemura T, Sugimura H (2004) Amplicons on human chromosome 11q are located in the early/late-switch regions of replication timing. Genomics 84:796–805
Windle B, Draper BW, Yin YX, O’Gorman S, Wahl GM (1991) A central role for chromosome breakage in gene amplification, deletion formation, and amplicon integration. Genes Dev 5:160–174
Woodfine K, Fiegler H, Beare DM, Collins JE, McCann OT, Young BD, Debernardi S, Mott R, Dunham I, Carter NP (2004) Replication timing of the human genome. Hum Mol Genet 13:575
Yuan B, Oechsli MN, Hendler FJ (1997) A region within murine chromosome 7F4, syntenic to the human 11q13 amplicon, is frequently amplified in 4NQO-induced oral cavity tumors. Oncogene 15:1161–1170
Yue Y, Haaf T (2006) 7E olfactory receptor gene clusters and evolutionary chromosome rearrangements. Cytogenet Genome Res 112:6–10
Acknowledgment
We would like to thank Trijnie Dijkhuizen, Pieter van der Vlies and Itty Oostendorp for technical assistance. We are grateful to Jane Briggs for reading the manuscript. This work is financially supported by the foundations of “Nijbakker-Morra”, “De Drie Lichten”, “Maurits en Anna de Kock” and the Groningen University Institute for Drug Exploration (GUIDE) in the Netherlands.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below are the links to the electronic supplementary material
Rights and permissions
About this article
Cite this article
Gibcus, J.H., Kok, K., Menkema, L. et al. High-resolution mapping identifies a commonly amplified 11q13.3 region containing multiple genes flanked by segmental duplications. Hum Genet 121, 187–201 (2007). https://doi.org/10.1007/s00439-006-0299-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-006-0299-6