Abstract
Cosegregation of markers on chromosome 5q12.3-q14.1 with profound congenital deafness in two Pakistani families (PKDF041 and PKDF141) defines a new recessive deafness locus, DFNB49. A maximum two-point lod score of 4.44 and 5.94 at recombination fraction θ=0 was obtained for markers D5S2055 and D5S424 in families PKDF041 and PKDF141, respectively. Haplotype analysis revealed an 11 cM linkage region flanked by markers D5S647 (74.07 cM) and D5S1501 (85.25 cM). Candidate deafness genes in this region include SLC30A5, OCLN, GTF2H2, and BTF3, encoding solute carrier family 30 (zinc transporter) member 5, occludin, RNA polymerase II transcription initiation factor, and basic transcription factor 3, respectively. Sequence analysis of the coding exons of SLC30A5 in DNA samples from two affected individuals of families PKDF041 and PKDF141 revealed no mutation. The mapping of DFNB49 further confirms the heterogeneity underlying autosomal recessive forms of nonsyndromic deafness.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nonsyndromic forms of deafness transmitted as a recessive trait are the most frequent cause of hereditary hearing loss and often exhibit the most severe hearing phenotype (Cohen et al. 1995). Nonsyndromic autosomal recessive deafness is usually clinically homogeneous and is nonprogressive in nature (Van Camp et al. 1997). As expected from the structural and functional complexity of the inner ear, sensorineural deafness exhibits a high degree of genetic heterogeneity. It has been estimated that at least 1% of human protein-coding genes are involved in the hearing process (Friedman and Griffith 2003). Presently, there are 36 published recessive deafness loci (DFNB), and 20 of the corresponding genes have been identified (Hereditary Hearing Loss Homepage, http://www.uia.ac.be/dnalab/hhh/).
Based on our ongoing genetic studies, a large repertoire of human genes associated with deafness remains to be mapped and identified, and these genes can be used to elucidate the molecular bases of hearing impairment. Because of the high degree of consanguinity, the Pakistani population represents a genetic resource for the identification of these novel deafness loci/genes (Hussain and Bittles 1998). Consanguineous families have contributed significantly to the identification of mutated genes associated with hearing loss (Friedman and Griffith 2003). This communication, describes two such large families segregating recessively inherited, profound congenital deafness, which is linked to markers that lie on chromosome 5q12.3-q14.1 and that define a novel locus, DFNB49 (Fig. 1).
Chromosome 5 markers co-segregating with deafness in families PKDF041 and PKDF141 define DFNB49 (black symbols individuals are deaf). The ‘at risk’ haplotypes are boxed in all generations. The genetic linkage markers and their relative positions in centimorgans (cM) according to the Marshfield map are shown left of each pedigree. a PKDF041; haplotypes showing a region of homozygosity flanked by markers D5S427 (69.23 cM) and D5S2110 (135.25 cM). b PKDF141; haplotypes showing a refined region of 11 cM delimited by markers D5S647 (74.07 cM) and D5S1501 (85.25 cM). Normal hearing individual IV:5 provided the proximal breakpoint at marker D5S647 (74.07 cM). The distal recombination is provided by individuals IV:6, IV:7, V:3, V:4, and V:5 at marker D5S1501 (85.25 cM).
Materials and methods
Family enrollment
Approval for this study was obtained from the IRB at the National Centre of Excellence in Molecular Biology, Lahore, Pakistan (FWA00001758) and the NINDS/NIDCD IRB at the National Institutes of Health, USA (OH-93-N-016). Signed informed consent was obtained from all participants. Genomic DNA was extracted from 10 ml blood samples by using a standard protocol (Grimberg et al. 1989).
Clinical evaluation
Clinical histories were obtained for all of the individuals of families PKDF041 and PKDF141. Physical evaluations were performed to rule out any obvious extra auditory phenotype. Pure tone audiometry tests for air and bone conduction were performed at frequencies from 250 to 8,000 Hz (Fig. 2). Ocular funduscopy and electroretinography (ERG) was performed to detect the presence of retinopathy. Vestibular function was evaluated by testing tandem gait ability and by using the Romberg test.
Pure tone air (right and left ear) and bone conduction (right ear) audiograms (O, X right and left ear, respectively). a Normal hearing individual VI:2 (9 years) and affected individual V:5 (42 years) of PKDF041. b Normal hearing individual IV:5 (35 years) and affected individual V:5 (22 years) of PKDF141. Unaffected individuals have normal hearing, whereas affected individuals are profoundly deaf with hearing thresholds at or above 90 dB at all tested frequencies.
Linkage analysis
Linkage to known DFNB loci was excluded for families PKDF041 and PKDF141 by using at least three microsatellite markers for each of the reported deafness loci (Hereditary Hearing Loss Homepage, http://www.uia.ac.be/dnalab/hhh/). Consequently, a genome-wide scan was performed with 388 fluorescent dye-labeled microsatellite markers from ABI Prism Linkage Mapping Set, Version 2.5 (Applied Biosystems) spaced at an average of 10 cM across the human genome. The microsatellite markers were amplified by the polymerase chain reaction (PCR) on a Gene Amp PCR system 9700 (Perkin Elmer) and were analyzed on an ABI Prism 3100 Genetic Analyzer. The alleles were assigned by using Genescan and Genotyper Software (Applied Biosystems).
Lod score calculations
Lod scores were calculated by using MLINK of the FASTLINK computer package (Schaffer 1996). The disease was coded as fully penetrant, and the disease-allele frequency was set at 0.001. Meiotic recombination frequencies were considered to be equal for males and females. Allele frequencies for microsatellite markers were calculated by genotyping 90 randomly selected unaffected individuals from the same population.
Mutational analysis
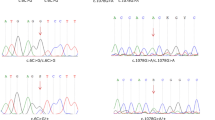
Primers for PCR amplification and subsequent sequencing of SLC30A5 and SLC12A2 were designed by using software at the Primer3 Web site (http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi) to flank all the exon–intron boundaries. PCR amplification, sequencing reactions, and mutation analysis were performed as described elsewhere (Ahmed et al. 2001). The sequence of primers for SLC30A5 and SLC12A2 will be provided upon request.
Nomenclature
DFNB49 was assigned to the new locus reported herein by the Human Genome Organization (HUGO) committee, and all gene symbols follow the recommendations of the HUGO Gene Nomenclature Committee (Povey et al. 2001).
Results
Regardless of age, all affected individuals in families PKDF041 and PKDF141 displayed congenital bilateral profound sensorineural hearing loss, whereas characteristic audiograms of unaffected individuals of both families showed normal hearing (Fig. 2). The unaffected individual of family PKDF141 (IV:5) had a mild conductive hearing loss at lower frequencies. Audiometric evaluations of two other normal hearing individuals IV:3 and IV:4 (data not shown) also showed the same hearing profile, which was different from the profound sensorineural hearing loss observed in the affected individuals. Vestibular function was normal in affected individuals from both families, and clinical evaluations suggested no features of ophthalmologic, skin, or renal anomalies. Funduscopy examinations of affected individuals of PKDF041, viz., V:5 (42 years), VI:3 (16 years), VI:4 (10 years), VI:5 (20 years), and of PKDF141, viz., V:5 (22 years), showed no signs of retinitis pigmentosa (RP). Moreover, ERGs were normal for affected members of PKDF041, viz., V:5 (42 years) and V:7 (30 years), thus excluding a novel Usher syndrome gene segregating in these two families.
The deafness phenotype segregating in families PKDF041 and PKDF141 was not found to be linked to the published deafness loci, thus indicating the involvement of at least one new locus. For deafness segregating in family PKDF041, evidence of linkage was detected on chromosome 5q12.1-q23.3 following a genome-wide scan. Additional markers were genotyped in the DFNB49 interval for all participating members. Haplotype analysis revealed a region of homozygosity of approximately 66 cM (Fig. 1a) flanked by markers D5S427 (69.23 cM) and D5S2110 (135.25 cM). A maximum two-point lod score (Zmax) of 4.44 at recombination fraction θ=0 was obtained for marker D5S2055 (Table 1).
Genome-wide linkage analysis of family PKDF141 segregating nonsyndromic deafness also showed linkage to markers at chromosome 5q12.3-q14.1, and haplotype analysis refined the linked region to 11 cM (Fig. 1b) delimited by markers D5S647 (74.07 cM) and D5S1501 (85.25 cM). Individual IV:5 of PKDF141 had normal hearing and was homozygous for markers in the proximal portion of the linked haplotype, with a recombination at D5S647 defining the proximal breakpoint of the linked region at 74.07 cM. Individuals IV:6, IV:7, V:3, V:4, and V:5 provided the distal breakpoint of the critical linked region at D5S1501. The parents of affected individuals V:6 and V:7 were unrelated. The deafness phenotype in these individuals may have been the result of compound heterozygosity for allelic mutations. A Zmax of 5.94 at θ=0 was found for D5S424 (Table 1).
Discussion
The region of homozygosity on 5q12.3-q14.1 for affected individuals of family PKDF041 overlaps with the reported interval for Usher syndrome (USH2C; Pieke-Dahl et al. 2000). Usher syndrome is an autosomal recessive disorder characterized by sensorineural hearing loss, RP and variable vestibular dysfunction (Ahmed et al. 2003b). Mutations of the gene encoding VLGR1 (MIM 602851), the very large G-protein-coupled receptor-1, have recently been reported in affected individuals of several USH2C-linked families (Weston et al. 2004). Although affected members of families PKDF041 do not have RP, VLGR1 cannot be excluded as a candidate gene. Mutant alleles of several Usher syndrome genes are reported to be associated with nonsyndromic hearing loss (Weil et al. 1997). We have previously shown that mutations of CDH23, USH1C, and PCDH15 can cause nonsyndromic hearing loss (Bork et al. 2001; Ahmed et al. 2002, 2003a).
Assuming the deafness segregating in these two families is caused by allelic mutations, the linkage region for DFNB49 is 11 cM in length and is delimited by D5S647 (74.07 cM) and D5S1501 (85.25 cM), which excludes VLGR1 as a candidate gene. Family PKDF141 delineates this smallest linkage region, which contains a number of candidate genes (Fig. 3). As the haplotypes of both families differ from each other, most likely the two mutant alleles are not the same.
Deafness segregating in families PKDF041 and PKDF141 show linkage to a region of human chromosome 5q12.3-q14.1. Candidate genes for DFNB49 deafness are derived from data available on the UCSC Genome Browser (http://genome.ucsc.edu/cgi-bin/hgGateway, July 2003 assembly).
Three deafness loci (DFNA42, DFNA1, and DFNA15) have been previously mapped to chromosome 5q, but their genetic intervals do not overlap with the larger interval for DFNB49 defined by family PKDF041 (Xia et al. 2002; Leon et al. 1992; Vahava et al. 1998).
One of the plausible candidate genes in the region of DFNB49 locus is solute carrier family member SLC30A5. Mutations in a solute carrier family member, SLC26A4, are responsible for DFNB4 (Li et al. 1998), Pendred syndrome (Everett et al. 1997), and Enlarged Vestibular Aqueduct syndrome (Usami et al. 1999). SLC26A4 mutations account for approximately 10% of hereditary deafness in diverse populations that include eastern and southern Asians (Park et al. 2003). However, all 16 protein-coding exons of SLC30A5 have been sequenced by using DNA samples from two affected individuals of families PKDF041and PKDF141, and no mutation has been found.
Other interesting candidate genes within the region of the DFNB49 locus include OCLN, GTF2H2, and BTF3, encoding the tight-junction protein occludin, RNA polymerase II transcription initiation factor, and basic transcription factor 3, respectively. Tight-junction proteins contribute to ion-selective paracellular barriers, and mutant alleles of the claudin 14 gene CLDN14 are responsible for nonsyndromic deafness in DFNB29 linked families (Wilcox et al. 2001; Ben-Yosef et al. 2003). OCLN is, therefore, a good candidate for the deafness gene segregating in our two families. The occludin knock-out mouse has not been reported to have hearing loss, but its hearing status does not seem to have been evaluated (Saitou et al. 2000). Genes encoding transcription factors EYA4, POU3F4, POU4F3, and TFCP2L3 are reported to be mutated in DFNA10 (Wayne et al. 2001), DFN3 (De Kok et al. 1995), DFNA15 (Vahava et al. 1998), and DFNA28 (Peters et al. 2002), respectively. TFCP2L3 is expressed in the epithelial cells of many tissues, and yet the only phenotype of a dominant mutant allele of TFCP2L3 is progressive loss of hearing (Peters et al. 2002).
An alternative to a one-locus hypothesis explaining the segregation of deafness in our two families is that there are two different, closely linked, deafness loci. In the linkage interval defined by family PKDF041, there are more than 100 genes including KCNN2, a member of the potassium channel genes. KCNN2 is an attractive candidate, because KCNQ4, another potassium channel gene, is mutated in dominant deafness DFNA2 (Kubisch et al. 1999). However, SLC12A2 is perhaps the most interesting candidate gene in this interval, because there is a deaf mouse model for SLC12A2 (Delpire et al. 1999). The 27 protein-coding exons of SLC12A2 have been sequenced by using DNA from affected individuals of family PKDF041, but no mutation has been found.
Rather than continuing to screen the two families for mutations that define DFNB49, we propose that it would be more efficient to identify additional families segregating recessive hearing loss mapping to this interval and to use these families to refine the critical region(s). Localization of locus DFNB49 is a first step toward the identification of a novel gene that is necessary for the development and/or maintenance of normal hearing.
Electronic database information
URLs for data in this article are as follows:
Center for Medical Genetics, Marshfield Medical Research Foundation: http://research.marshfieldclinic.org/genetics/
Hereditary Hearing Loss Homepage: http://www.uia.ac.be/dnalab/hhh/
Online Mendelian Inheritance in Man (OMIM): http://www.ncbi.nlm.nih.gov/entrez/OMIM
Primer3 Web-Based Server (primer3_www.cgi v 0.2): http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi
The Human Genome Organization (HUGO): http://www.gene.ucl.ac.uk/hugo/
UCSC Genome Bioinformatics: http://genome.ucsc.edu/
References
Ahmed ZM, Riazuddin S, Bernstein SL, Ahmed Z, Khan S, Griffith AJ, Morell RJ, Friedman TB, Wilcox ER (2001) Mutations of the protocadherin gene PCDH15 cause Usher syndrome type 1F. Am J Hum Genet 69:25–34
Ahmed ZM, Smith TN, Riazuddin S, Makishima T, Ghosh M, Bokhari S, Menon PSN, Deshmukh D, Griffith AJ, Riazuddin S, Friedman TB, Wilcox ER (2002) Nonsyndromic recessive deafness DFNB18 and Usher syndrome type IC are allelic mutations of USHIC. Hum Genet 110:527–531
Ahmed ZM, Riazuddin S, Ahmad J, Bernstein SL, Guo Y, Sabar MF, Sieving P, Riazuddin S, Griffith AJ, Friedman TB, Belyantseva IA, Wilcox ER (2003a) PCDH15 is expressed in the neurosensory epithelium of the eye and ear and mutant alleles are responsible for both USH1F and DFNB23. Hum Mol Genet 12:3215–3223
Ahmed ZM, Riazuddin S, Riazuddin S, Wilcox ER (2003b) The molecular genetics of Usher syndrome. Clin Genet 63:431–444
Ben-Yosef T, Belyantseva IA, Saunders TL, Hughes ED, Kawamoto K, Van Itallie CM, Beyer LA, Halsey K, Gardner DJ, Wilcox ER, Rasmussen J, Anderson JM, Dolan DF, Forge A, Raphael Y, Camper SA, Friedman TB (2003) Claudin 14 knockout mice, a model for autosomal recessive deafness DFNB29, are deaf due to cochlear hair cell degeneration. Hum Mol Genet 12:2049–2061
Bork JM, Peters LM, Riazuddin S, Bernstein SL, Ahmed ZM, Ness SL, Polomeno R, Ramesh A, Schloss M, Srisailpathy CR, Wayne S, Bellman S, Desmukh D, Ahmed Z, Khan SN, Kaloustian VM, Li XC, Lalwani A, Riazuddin S, Bitner-Glindzicz M, Nance WE, Liu XZ, Wistow G, Smith RJ, Griffith AJ, Wilcox ER, Friedman TB, Morell RJ (2001) Usher syndrome 1D and nonsyndromic autosomal recessive deafness DFNB12 are caused by allelic mutations of the novel cadherin-like gene CDH23. Am J Hum Genet 68:26–37
Cohen MM, Gorlin RJ (1995) Epidemiology, etiology and genetic patterns. In: Gorlin RJ, Toriello HV, Cohen MM (eds) Hereditary hearing loss and its syndromes. Oxford University Press, Oxford, pp 43–61
De Kok YJ, Van der Maarel SM, Bitner-Glindzicz M, Huber I, Monaco AP, Malcolm S, Pembrey ME, Ropers HH, Cremers FP (1995) Association between X-linked mixed deafness and mutations in the POU domain gene POU3F4. Science 267:685–688
Delpire E, Lu J, England R, Dull C, Thorne T (1999) Deafness and imbalance associated with inactivation of the secretory Na-K-2Cl co-transporter. Nat Genet 22:192–195
Everett LA, Glaser B, Beck JC, Idol JR, Buchs A, Heyman M, Adawi F, Hazani E, Nassir E, Baxevanis AD, Sheffield VC, Green ED (1997) Pendred syndrome is caused by mutations in a putative sulphate transporter gene (PDS). Nat Genet 17:411–422
Friedman TB, Griffith AJ (2003) Human nonsyndromic sensorineural deafness. Annu Rev Genomics Hum Genet 4:341–402
Grimberg J, Nawoschik S, Belluscio L, McKee R, Turck A, Eisenberg A (1989) A simple and efficient non-organic procedure for the isolation of genomic DNA from blood. Nucleic Acids Res 17:8390
Hussain R, Bittles AH (1998) The prevalence and demographic characteristics of consanguineous marriages in Pakistan. J Biosoc Sci 30:261–275
Kubisch C, Schroeder BC, Friedrich T, Lutjohann B, El-Amraoui A, Marlin S, Petit C, Jentsch TJ (1999) KCNQ4, a novel potassium channel expressed in sensory outer hair cells, is mutated in dominant deafness. Cell 96:437–446
Leon PE, Raventos H, Lynch E, Morrow J, King MC (1992) The gene for an inherited form of deafness maps to chromosome 5q31. Proc Natl Acad Sci USA 89:5181–5184
Li XC, Everett LA, Lalwani AK, Desmukh D, Friedman TB, Green ED, Wilcox ER (1998) A mutation in PDS causes non-syndromic recessive deafness. Nat Genet 18:215–217
Park HJ, Shaukat S, Liu XZ, Hahn SH, Naz S, Ghosh M, Kim HN, Moon SK, Abe S, Tukamoto K, Riazuddin S, Kabra M, Erdenetungalag R, Radnaabazar J, Khan S, Pandya A, Usami SI, Nance WE, Wilcox ER, Riazuddin S, Griffith AJ (2003) Origins and frequencies of SLC26A4 (PDS) mutations in east and south Asians: global implications for the epidemiology of deafness. J Med Genet 40:242–248
Peters LM, Anderson DW, Griffith AJ, Grundfast KM, San Agustin TB, Madeo AC, Friedman TB, Morell RJ (2002) Mutation of a transcription factor, TFCP2L3, causes progressive autosomal dominant hearing loss, DFNA28. Hum Mol Genet 11:2877–2885
Pieke-Dahl S, Moller CG, Kelley PM, Astuto LM, Cremers CWRJ, Gorin MB, Kimberling WJ (2000) Genetic heterogeneity of Usher syndrome type II: localisation to chromosome 5q. J Med Genet 37:256–262
Povey S, Lovering R, Bruford E, Wright M, Lush M, Wain H (2001) The HUGO Gene Nomenclature Committee (HGNC). Hum Genet 109:678–680
Saitou M, Furuse M, Sasaki H, Schulzke JD, Fromm M, Takano H, Noda T, Tsukita S (2000) Complex phenotype of mice lacking occludin, a component of tight junction strands. Mol Biol Cell 11:4131–4142
Schaffer AA (1996) Faster linkage analysis computations for pedigrees with loops or unused alleles. Hum Hered 46:226–235
Usami S, Abe S, Weston M D, Shinkawa H, Van Camp G, Kimberling WJ (1999) Nonsyndromic hearing loss associated with enlarged vestibular aqueduct is caused by PDS mutations. Hum Genet 104:188–192
Vahava O, Morell R, Lynch ED, Weiss S, Kagan ME, Ahituv N, Morrow JE, Lee MK, Skvorak AB, Morton CC, Blumenfeld A, Frydman M, Friedman TB, King MC, Avraham KB (1998) Mutation in transcription factor POU4F3 associated with inherited progressive hearing loss in humans. Science 279:1950–1954
Van Camp G, Willems PJ, Smith RJH (1997) Nonsyndromic hearing impairment unparalleled heterogeneity. Am J Hum Genet 60:758–764
Wayne S, Robertson NG, DeClau F, Chen N, Verhoeven K, Prasad S, Tranebjarg L, Morton CC, Ryan A F, Van Camp G, Smith RJH (2001) Mutations in the transcriptional activator EYA4 cause late-onset deafness at the DFNA10 locus. Hum Mol Genet 10:195–200
Weil D, Kussel P, Blanchard S, Levy G, Levi-Acobas F, Drira M, Ayadi H, Petit C (1997) The autosomal recessive isolated deafness, DFNB2, and the Usher 1B syndrome are allelic defects of the myosin-VIIA gene. Nat Genet 16:191–193
Weston MD, Luijendijk MWJ, Humphrey KD, Moller C, Kimberling WJ (2004) Mutations in the VLGR1 gene implicate G-protein signaling in the pathogenesis of Usher syndrome type II. Am J Hum Genet 74:357–366
Wilcox ER, Burton QL, Naz S, Riazuddin S, Smith TN, Ploplis B, Belyantseva I, Ben-Yosef T, Liburd NA, Morell RJ, Kachar B, Wu DK, Griffith AJ, Riazuddin S, Friedman TB (2001) Mutations in the gene encoding tight junction claudin-14 cause autosomal recessive deafness DFNB29. Cell 104:165–172
Xia J, Deng H, Feng Y, Zhang H, Pan Q, Dai H, Long Z, Tang B, Deng H, Chen Y, Zhang R, Zheng D, He Y, Xia K (2002) A novel locus for autosomal dominant nonsyndromic hearing loss identified at 5q31.1–32 in a Chinese pedigree. J Hum Genet 47:635–640
Acknowledgements
We are grateful to the families for their participation in this study. We thank Dr. Dennis Drayna and Dr. Robert Morell for their comments on this manuscript, and Sabiha Nazli and Muhammad Awais for their technical assistance. This study was supported by the Higher Education Commission, Islamabad, Pakistan; the Ministry of Science and Technology, Islamabad, Pakistan; and the International Center for Genetic Engineering and Biotechnology, Trieste, Italy under project CRP/PAK02-01 (contract no. 02/013). Part of this study in USA was supported by intramural funds from the National Institute on Deafness and Other Communication Disorders (1ZO1 DC000035-07 and 1ZO1 DC000039-07).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ramzan, K., Shaikh, R.S., Ahmad, J. et al. A new locus for nonsyndromic deafness DFNB49 maps to chromosome 5q12.3-q14.1. Hum Genet 116, 17–22 (2005). https://doi.org/10.1007/s00439-004-1205-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-004-1205-8