Abstract
RNA editing is a process that modifies the information in transcripts of almost all angiosperm mitochondrial protein-coding genes. In order to determine the frequency and distribution of mitochondrial RNA editing in Beta vulgaris, cDNAs were sequenced and compared to the published genome sequence. 357 C to U conversions were identified across the 31 known protein genes and pseudogenes in Beta, the fewest so far for a plant mitochondrial genome. Editing patterns in the putative gene orf518 indicate that it is most likely a functional ccmC homolog, indicating that patterns of editing can be a useful determinant of gene functionality. orf518 also contains a triplicated repeat region whose members are nearly identical yet differentially edited, most likely due to differences in the sequence context of the editing sites. In addition, we show that partial editing in Beta is common at silent editing sites but rare at nonsilent editing sites, extending previous observations to a complete plant mitochondrial genome. Finally, the degree of partial editing observed for certain genes was dependent on the choice of primers used, demonstrating that care must be taken when designing primers for use in editing studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA editing, a process that alters the information in a transcript from what is encoded by the genome, occurs in a variety of organisms through insertion, deletion, or modification of nucleotides in RNA (Brennicke et al. 1999; Gott and Emeson 2000). In plants, RNA editing proceeds via an unknown mechanism to convert C to U and U to C in organellar transcripts. Although the mechanism is unknown, the pattern and consequence of editing have been examined in great detail. In some cases, editing can generate new start and/or stop codons (Onodera et al. 1999; Kubo et al. 2000b), increase intron splicing efficiency (Borner et al. 1995), or improve tRNA base pairings in stem regions (Fey et al. 2001). However, most editing sites alter internal amino acids encoded by mRNA (Giege and Brennicke 1999). Editing at these sites usually leads to an increase in protein conservation across species (Covello and Gray 1989; Gualberto et al. 1989; Hiesel et al. 1989). This tendency, along with the fact that editing generally increases the hydrophobicity of proteins (Giege and Brennicke 1999), indicates that these sites are important for the proteins to function properly in their organellar roles. Conversely, silent editing, which has no effect on protein form or function owing to the degeneracy of the genetic code, is less common (Giege and Brennicke 1999) and where present, often occurs in only a subset of transcripts [“partial editing” (Schuster et al. 1990; Kempken et al. 1991)].
The presence of partially edited sites was first detected by direct cDNA sequencing (Gualberto et al. 1989) and soon after by sequencing of cDNA clones (Schuster et al. 1990). In the latter study, 5 out of 16 editing sites were partially edited in the Oenothera nad3 gene. Interestingly, all three silent editing sites were partially edited and each was observed in only one of eight clones. In contrast, only 2 of 13 nonsilent editing sites were partially edited and they were found in 3 or 7 of 8 clones. This was the first indication that silent editing sites are less efficiently edited than nonsilent edits. Numerous subsequent studies have shown similar patterns, but relatively few have discussed this phenomenon (Kempken et al. 1991; Schuster et al. 1991; Kadowaki et al. 1995; Wilson and Hanson 1996) and none have tested this across a complete genome. Other studies have found a high degree of partial editing at both silent and nonsilent sites. In these cases, the incompletely edited transcripts appear to be intermediates that become more fully edited during maturation (Gualberto et al. 1991; Sutton et al. 1991; Yang and Mulligan 1991).
To date, RNA editing content has been assessed for all known protein genes in the mitochondrial genomes of the angiosperms Arabidopsis thaliana (Giege and Brennicke 1999), Brassica napus (Handa 2003), and Oryza sativa (Notsu et al. 2002), and evidence suggests that the liverwort Marchantia polymorpha neither performs nor requires RNA editing (Oda et al. 1992; Ohyama et al. 1993; Steinhauser et al. 1999). To gain a better perspective on the pattern of RNA editing in angiosperms, we have now determined the extent of editing in the mitochondrial genome of the sugar beet Beta vulgaris. In particular, we find that partial editing is much more common at silent sites for almost all protein genes. Furthermore, we show that the examination of partial editing patterns can be useful for determining gene functionality, understanding editing site specificity, and evaluating the effect of primer choice on editing site identification.
Materials and methods
Seeds from cytoplasmic male-fertile B. vulgaris (line TK81-O) were obtained from the National Agricultural Research Center for Hokkaido Region (Hokkaido, Japan). Seedlings were grown for 2 weeks at 20°C in 16 h of light and 8 h of darkness.
Total RNA was isolated from whole tissue with the RNeasy Plant Mini Kit (QIAGEN) and then treated with DNase I (Promega) for 30 min at 37°C to remove any contaminating genomic DNA. DNase I was removed by treating with chloroform:isoamyl alcohol (24:1) and RNA was recovered by ethanol precipitation. cDNA was synthesized using M-MulV reverse transcriptase (New England Biolabs) and random hexamers according to manufacturer’s instructions. Gene-specific cDNA products were amplified by polymerase chain reaction (PCR) using primers (Table S1) designed in the 5′ and 3′ untranslated regions (UTR) of each gene. Each reaction was performed using 35 cycles of 45 s at 94°C, 45 s at 50°C, and 2.0–2.5 min at 72°C, with an initial step of 3 min at 94°C and a final step of 10 min at 72°C. No amplification was observed when a cDNA synthesis reaction that excluded reverse transcriptase was used as a template, indicating that there was no contaminating genomic DNA in the cDNA preparation. Amplified cDNA products were purified with the QIAquick PCR Purification Kit (QIAGEN) and then directly sequenced on both strands at the Indiana Molecular Biology Institute. In addition, some products were cloned using the TOPO TA Cloning Kit (Invitrogen). Colonies were screened following the colony PCR protocol provided in the TOPO TA Cloning Kit manual, and positive clones were sequenced.
Sites of RNA editing were determined by comparison of cDNA sequences to the published mitochondrial genome for TK81-O (Kubo et al. 2000a). Sites were scored as partially edited when peaks for both T and C were present and clearly above background in both strands of the directly sequenced cDNA. This approach likely underrepresents the true number of partially edited sites, because sites that are mostly edited (in more than 80–90% of transcripts) will be scored as fully edited and sites that are rarely edited (in fewer than 10–20% of transcripts) will be scored as unedited. Nevertheless, the direct sequence approach used here is preferred over sequencing multiple cDNA clones for three reasons. The latter approach requires a large amount of additional sequencing, cannot accurately determine the degree of partiality unless a large number of clones are analyzed, and will introduce nucleotide changes during the cloning process that can be erroneously scored as rare editing events. Sites of RNA editing in the mitochondrial genomes of Arabidopsis thaliana (GenBank: Y08501, Y08502), Brassica napus (GenBank: AP006444), and Oryza sativa (GenBank: BA000029) were taken from annotations in these GenBank files.
Results and discussion
RNA editing in B. vulgaris mitochondrial genes and pseudogenes
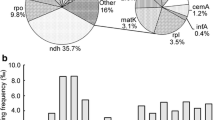
The complete mitochondrial genome of male-fertile B. vulgaris contains 31 known protein genes, including the putative pseudogenes ccmC and sdh4 (Kubo et al. 2000a; Siqueira et al. 2002). Sites of RNA editing have been determined for eight of these genes (Kubo et al. 1993; Senda et al. 1993; Kubo and Mikami 1996; Estiati et al. 1998; Onodera et al. 1999; Kubo et al. 2000b; Itchoda et al. 2002). In order to obtain a more complete picture of the frequency and distribution of RNA editing in this genome, an analysis of RNA editing was undertaken for the remaining 23 genes and pseudogenes. In total, 357 edited sites were found across the 31 genes and pseudogenes in the Beta mitochondrial genome (Table 1), the lowest number of editing sites reported so far for a plant mitochondrial genome. Six additional editing sites were identified in UTR regions. All editing changes are C to U conversions. As described previously, RNA editing generates new start codons for atp6, nad1, and nad4L and new stop codons for atp6 and atp9 (Onodera et al. 1999; Kubo et al. 2000a; Kubo et al. 2000b). No additional start or stop codons are created from editing.
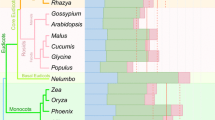
In Beta, the frequency of editing is not consistent across genes, ranging from 0% (0 sites in 1,575 bp) for cox1 to 4.8% (30 sites in only 621 bp) for ccmB (Table 1). Similar inconsistencies in editing frequency as a function of gene length are also observed in Arabidopsis, Brassica, and Oryza (Table S2). However, the frequency of editing does seem to be fairly consistent for each gene across species; genes that are frequently edited in Beta are also frequently edited in Arabidopsis, Brassica, and Oryza (e.g. ccmB and ccmC), while rarely edited Beta genes are also rarely edited in the other species (e.g. atp1 and cox1). There are some obvious differences as well, evidenced by the large standard deviations relative to the mean for some genes, which may be indicative of a history of retroprocessing (reverse transcription of edited mRNA and reinsertion into the genome). Consistent with its low level of RNA editing overall, Beta has the lowest frequency of editing for 16 of 30 genes, while the frequency is highest for only five genes. The similar patterns of editing in homologous genes from Beta, Arabidopsis, Brassica, and Oryza indicate that editing frequencies have remained relatively stable throughout angiosperm history. It further suggests that, barring retroprocessing, these patterns are likely to hold for most, if not all angiosperms and possibly for other plant groups as well.
RNA editing as a determinant of gene functionality
The ccmC gene was initially reported to be a pseudogene in the mitochondrial genome of Beta (Kubo et al. 2000a), and this assessment has been perpetuated in comparisons of mitochondrial gene content across species (Notsu et al. 2002; Handa 2003; Turmel et al. 2003; Clifton et al. 2004; Sugiyama et al. 2005). It seems that the pseudogene status was originally assigned based on the absence of a start codon at the same position as in other plants (Satoh et al. 2004). However, because the ccmC pseudogene does not contain any stop codons, Satoh et al. (2004) renamed it as orf518 and orf496 in male-fertile and male-sterile Beta mitochondria, respectively. The 3′ halves of orf518 and orf496 are homologous to ccmC from other species, and the 5′ ends contain a fragment of atp9 (Fig. 1a; Satoh et al. 2004).
The orf518 gene in Beta vulgaris. a Structure of the orf518 gene. Shaded areas indicate regions homologous to known mitochondrial genes. Possible translational start codons are indicated by arrows; the second start codon is homologous to the start site of atp9. Stop codon is indicated by a star. The three tandem repeats that comprise the TR3 region are bracketed. b RNA editing in orf518. RNA editing in the entire orf518 gene and in the TR3 region was analyzed by direct sequencing of amplified cDNA. Thin horizontal lines indicate the sizes of the analyzed cDNA products. Primers positions are indicated by open boxes. Open and closed ovals represent partially and fully edited sites, respectively. Unedited sites in the TR3 region are marked by the letter X. RNA editing in the TR3 region was also analyzed by sequencing 18 clones. The number of times each clone type was found is indicated to the right of each type. Nucleotide changes observed in only a single clone are not shown due to the possibility that they were introduced during PCR and/or cloning. c Nucleotide sequence of TR3 and the atp9-homologous region. Nucleotide states for all sequences are shown in editing columns (marked by arrows); for all other columns, nucleotides identical with orf518 or the orf518 repeat 1 sequence are plotted as periods. Columns of interest within the repeat region are numbered according to their position in the repeat. Edited cytidines are represented by the letter E; frequency of editing at each site is shown in (b). Putative start codons are boxed. orf518 begins at the first start codon, while atp9 begins at the second start codon
We designed primers flanking the entire orf518 gene and verified that it is edited at 28 sites in the ccmC-homologous region (Table 1, Fig. 1a). Editing patterns indicate that the pseudogene designation is not warranted. Only 6 of the 28 editing sites are silent, consistent with the low levels of silent editing observed across angiosperm mitochondrial genes. Furthermore, all 22 nonsilent edits increase the similarity of the predicted protein to CCMC proteins from other angiosperms (data not shown); such conservation is a hallmark of RNA editing in functional genes but not necessarily pseudogenes (see below). Finally, all 22 nonsilent editing sites are fully edited, whereas 4 of 6 silent editing sites are partially edited. The dichotomous frequency of partial editing at nonsilent vs. silent editing sites found in the ccmC-homologous region mirrors the pattern observed across the other functional genes in the Beta mitochondrial genome. Taken together, these results indicate that orf518 is an intact and likely functional ccmC homolog in Beta, and they demonstrate that patterns of editing can provide important clues regarding the functionality of plant mitochondrial genes.
If ccmC in Beta is indeed functional, it is not clear where translation starts since it does not share the position of the start codon found in other plants. However, five other ATG codons could serve as the translational start site (Fig. 1a), all of which are found in both male-fertile and male-sterile Beta (an in-frame stop codon upstream of the first putative start codon precludes any additional possibilities). Of these possibilities, the most likely candidates seem to be the second and the fifth start site. The second putative start codon is in the duplicated atp9 region, and is in fact homologous to the start codon for atp9. Thus, ccmC in Beta may have co-opted the promoter and ribosomal binding site of atp9. The fifth putative start codon is found only 11-13 codons downstream from the reported start codons for Arabidopsis, Brassica, Pisum, Oenothera, Oryza, Triticum, Zea, and Marchantia (data not shown). Interestingly, the above species, as well as the green alga Chara and the jakobid Reclinomonas, also code for a methionine at this downstream position, raising the possibility that it may actually serve as the translational start site for all of these species.
The sdh4 gene was originally described as absent from the mitochondrial genome of Beta (Kubo et al. 2000a). However, it was later shown to be present as a pseudogene, containing a 5 bp insertion that creates multiple premature stop codons (Siqueira et al. 2002). We found sdh4 to be transcribed and edited at four positions (Table 1). All four editing sites are downstream of the first introduced stop codon and are partially edited. After correcting for the frameshift caused by the 5 bp insertion, two editing sites are silent while the other two are nonsilent and increase protein similarity to homologous proteins from other species (data not shown). The two nonsilent editing sites are also found in Lycopersicon esculentum and Solanum tuberosum (Adams et al. 2001; Siqueira et al. 2002).
The relaxation of constraints on editing sites appears to be a common phenomenon for mitochondrial pseudogenes. Similar to Beta sdh4, the rps14 pseudogene from Triticum contains one conserved nonsilent editing site that is only partially edited (Sandoval et al. 2004). In some pseudogenes, such as rps12 from Oenothera and rps14 from Solanum, an evolutionarily conserved editing site has been completely lost (Grohmann et al. 1992; Quinones et al. 1996). The most spectacular example of relaxed editing constraints in pseudogenes comes from Oenothera rps19, which shows not only two losses of editing sites that increase protein conservation, but also five gains of editing sites that decrease protein conservation (Schuster and Brennicke 1991). For a number of genes, unusual editing patterns are the only indication that they are not functional (Mundel and Schuster 1996; Handa 2003; Mower 2005). Although these genes are intact, in-frame, and transcribed, they have lost editing at evolutionarily conserved positions, suggesting pseudogenization of their mitochondrial copies and the presence of functional copies in the nucleus.
Differential editing in a triplicated repeat region
Within the Beta mitochondrial genome are a number of VNTR (variable number of tandem repeats) regions (Nishizawa et al. 2000). One such VNTR, TR3, is 66 bp in length and is repeated, nearly perfectly, three times near the 5′ end of orf518 in male-fertile Beta (Fig. 1a; Satoh et al. 2004). The first 20 bp of each repeat is the 3′ end of the partial atp9 duplication found at the beginning of orf518. During our investigation of the extent of RNA editing in the orf518 gene, we were surprised to find differential editing among the three copies of this tandem repeat region. To explore this result in more detail, we generated cDNA covering the entire TR3 region and analyzed editing in 18 clones (Fig. 1b). The first nucleotide in each repeat is a cytidine that is homologous to a fully edited cytidine in the native atp9 gene. This site is also found to be fully edited (18/18 clones) in the duplicated atp9 region of the first repeat, but is completely unedited (0/18) in the second and third repeats. A second editing site is located at position 32 in each repeat and is usually edited (14/18) in the first repeat but rarely edited in the second and third repeats (3/18 and 2/18, respectively).
The differential editing of the first editing site is almost certainly the result of the different flanking sequences in repeat 1 versus repeats 2 and 3 (Fig. 1c). The upstream sequence of the first editing site in repeat 1 is identical for 35 bp to the upstream sequence of the homologous atp9 editing site, and both sites are fully edited. In contrast, the upstream region in repeats 2 and 3 is the end of the previous repeat. Since these editing sites share none of the homology with the upstream atp9 sequence and are completely unedited, it seems likely that site specificity is contained in the upstream atp9 region. It is also possible that the downstream sequence has an effect on conferring specificity. Repeats 2 and 3 have a C only 2 bp downstream of the editing site, while repeat 1 and atp9 have an A at this same position. These findings are consistent with in vitro and in organello RNA editing assays that have shown that editing site specificity lies in the sequences immediately flanking editing sites (Choury et al. 2004; Neuwirt et al. 2005).
For the second editing site, the difference in editing frequency is most likely due to the single nucleotide difference found 10 bp upstream of the editing site (Fig. 1c). Repeat 1 has a T at this position, whereas repeats 2 and 3 have a C. A second difference between repeat 1 and repeats 2 and 3 lies 29 bp upstream and may also contribute to the observed frequency differential, although this difference may be too far from the editing site to have an effect. A downstream difference, at position 60, is unique to repeat 3 and thus cannot account for the differential editing in repeat 1 versus repeats 2 and 3. The fact that only one or two changes in the sequence surrounding this editing site can have such a large effect on the frequency of editing indicates that the editing machinery is highly sensitive to specific sequence context.
Abundance of partial editing at silent editing sites
In Beta, the proportion of partial editing at silent and nonsilent editing sites is highly biased (Table 1). In total, 47 out of 330 editing sites are partially edited (editing sites for atp1, nad4, and nad9 were not counted since partial editing information was not reported for these genes). Of the 41 silent editing sites for which partial editing data are available, 24 (58.5%) are partially edited. In contrast, only 23 of the 289 nonsilent editing sites (8.0%) are partially edited. Most of these partially edited nonsilent sites occur in three genes: mttB, matR, and sdh4. For all other genes, partial editing at nonsilent editing sites is extremely rare. If partially edited sites were distributed without regard to their effect on the amino acids produced, then we would expect 5.8 silent (= 41 × 47/330) and 41.2 nonsilent ( = 289 × 47/330) editing sites to be partially edited. The observed numbers of partially edited silent and nonsilent sites differ significantly from random expectations (P = 9.7e-16; Χ 2 = 64.5; d.f. = 1).
Thus, the nonrandom distribution of partial editing extends to most, if not all (see next section) genes in the mitochondrial genome of Beta (Table 1). The predominance of partial editing at many silent sites may simply represent an inefficiency of editing at these sites, with translation proceeding irrespective of their editing status. Alternatively, partially edited silent sites, despite having no effect on protein sequence, may nevertheless serve a functional role. For example, editing of these sites may be one of the final steps in mRNA maturation, signaling that the transcript is ready for translation.
Effect of primer choice on RNA editing site detection
Initial assessment of partial editing in Beta mitochondrial genes revealed four genes (matR, nad6, mttB, rps12) for which many editing sites appeared to be partially edited (Fig. 2). Because the cDNA primers were designed based on the genomic sequence, it is possible that we preferentially selected for editing intermediates during PCR. This preferential amplification could occur if editing sites are located in the primer binding sites, which would create mismatches between the primer and edited transcripts but not unedited or incompletely edited transcripts. It is also possible that processing of these transcripts during maturation may proceed into or beyond the primer binding sites (which are in the UTR regions); thus, only immature transcripts would be amplified.
Effect of primer choice on editing site detection. Symbol usage follows that of Fig.1. RNA editing in all four transcripts was analyzed by direct sequencing of amplified cDNA. The upper cDNA sequence for each gene was the result of the initial analysis. The lower cDNA sequence was the result of the reanalysis using internal primers that were designed in regions that lack editing. The mttB gene is shown starting at an ATA codon based on genomic annotations (GenBank: BA000009)
To eliminate these possibilities, we reevaluated the editing patterns of these four genes (Fig. 2) using an internal set of primers designed in regions that are free of editing. In our initial analysis of editing in nad6, seven editing sites were identified and all were partially edited. Upon reanalysis, however, all previously identified sites are fully edited. In addition, four new editing sites were found, two of which are silent and only partially edited. For rps12, we initially evaluated the entire nad3/rps12 cotranscription product and observed six partially edited sites in rps12 (nad3, on the other hand, was partially edited at only one silent editing site). Using new primers, all six editing sites in rps12 are fully edited. This suggests that the primers initially used for nad6 and rps12 were selecting for incompletely edited transcripts, and implies that primer choice is critical in studies on RNA editing.
In contrast, the frequency and completeness of editing did not change upon reevaluation of matR and mttB (Fig. 2). It is not clear why matR and mttB continue to have such a high degree of partial editing at nonsilent sites. Partial editing seems to be the rule for these two genes, as this is observed at many sites in matR from Glycine, Solanum, Triticum, and Zea (Thomson et al. 1994; Begu et al. 1998) and in Arabidopsis mttB (Sunkel et al. 1994). Although little is known about the actual mitochondrial function of matR and mttB, they are most likely essential because they have not been lost from the mitochondrial genome in any of the 280 angiosperms examined (Adams et al. 2002) and because they are intact and in-frame in all angiosperms from which they’ve been isolated. Interestingly, both of these genes have uncertain translational start sites for many angiosperms (Sunkel et al. 1994; Thomson et al. 1994; Begu et al. 1998). Although non-canonical start sites may be used, it is possible that further maturation steps (e.g. processing or trans-splicing) are required. Thus, the high degree of partial editing for matR and mttB may also be due to amplification of immature transcripts. However, it remains a possibility that matR and mttB are in fact pseudogenes that have simply not acquired the usual identifying characteristics, such as in-frame stop codons or frameshifting indels.
References
Adams KL, Qiu Y-L, Stoutemyer M, Palmer JD (2002) Punctuated evolution of mitochondrial gene content: high and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc Natl Acad Sci USA 99:9905–9912
Adams KL, Rosenblueth M, Qiu Y-L, Palmer JD (2001) Multiple losses and transfers to the nucleus of two mitochondrial succinate dehydrogenase genes during angiosperm evolution. Genetics 158:1289–1300
Begu D, Mercado A, Farre JC, Moenne A, Holuigue L, Araya A, Jordana X (1998) Editing status of mat-r transcripts in mitochondria from two plant species: C-to-U changes occur in putative functional RT and maturase domains. Curr Genet 33:420–428
Borner GV, Morl M, Wissinger B, Brennicke A, Schmelzer C (1995) RNA editing of a group II intron in Oenothera as a prerequisite for splicing. Mol Gen Genet 246:739–744
Brennicke A, Marchfelder A, Binder S (1999) RNA editing. FEMS Microbiol Rev 23:297–316
Clifton SW, Minx P, Fauron CM, Gibson M, Allen JO, Sun H, Thompson M, Barbazuk WB, Kanuganti S, Tayloe C, Meyer L, Wilson RK, Newton KJ (2004) Sequence and comparative analysis of the maize NB mitochondrial genome. Plant Physiol 136:3486–3503
Choury D, Farre JC, Jordana X, Araya A (2004) Different patterns in the recognition of editing sites in plant mitochondria. Nucleic Acids Res 32:6397–6406
Covello PS, Gray MW (1989) RNA editing in plant mitochondria. Nature 341:662–666
Estiati A, Kubo T, Mikami T (1998) The ribosomal protein S7 gene is transcribed and edited in sugar beet mitochondria. Physiol Plant 102:325–327
Fey J, Weil JH, Tomita K, Cosset A, Dietrich A, Small I, Marechal-Drouard L (2001) Editing of plant mitochondrial transfer RNAs. Acta Biochim Pol 48:383–389
Giege P, Brennicke A (1999) RNA editing in Arabidopsis mitochondria effects 441 C to U changes in ORFs. Proc Natl Acad Sci USA 96:15324–15329
Gott JM, Emeson RB (2000) Functions and mechanisms of RNA editing. Annu Rev Genet 34:499–531
Grohmann L, Brennicke A, Schuster W (1992) The mitochondrial gene encoding ribosomal protein S12 has been translocated to the nuclear genome in Oenothera. Nucleic Acids Res 20:5641–5646
Gualberto JM, Bonnard G, Lamattina L, Grienenberger JM (1991) Expression of the wheat mitochondrial nad3-rps12 transcription unit: correlation between editing and mRNA maturation. Plant Cell 3:1109–1120
Gualberto JM, Lamattina L, Bonnard G, Weil JH, Grienenberger JM (1989) RNA editing in wheat mitochondria results in the conservation of protein sequences. Nature 341:660–662
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res 31:5907–5916
Hiesel R, Wissinger B, Schuster W, Brennicke A (1989) RNA editing in plant mitochondria. Science 246:1632–1634
Itchoda N, Nishizawa S, Nagano H, Kubo T, Mikami T (2002) The sugar beet mitochondrial nad4 gene: an intron loss and its phylogenetic implication in the Caryophyllales. Theor Appl Genet 104:209–213
Kadowaki K, Ozawa K, Kazama S, Kubo N, Akihama T (1995) Creation of an initiation codon by RNA editing in the coxI transcript from tomato mitochondria. Curr Genet 28:415–422
Kempken F, Mullen JA, Pring DR, Tang HV (1991) RNA editing of sorghum mitochondrial atp6 transcripts changes 15 amino acids and generates a carboxy-terminus identical to yeast. Curr Genet 20:417–422
Kubo T, Mikami T (1996) A duplicated sequence in sugarbeet mitochondrial transcripts is differentially edited: analysis of orfB and its derivative orf324 mRNAs. Biochim Biophys Acta 1307:259–262
Kubo T, Mikami T, Kinoshita T (1993) The sugar beet mitochondrial genome contains an ORF sharing sequence homology with the gene for the 30 kDa subunit of bovine mitochondrial complex I. Mol Gen Genet 241:479–481
Kubo T, Nishizawa S, Sugawara A, Itchoda N, Estiati A, Mikami T (2000a) The complete nucleotide sequence of the mitochondrial genome of sugar beet (Beta vulgaris L.) reveals a novel gene for tRNACys (GCA). Nucleic Acids Res 28:2571–2576
Kubo T, Yamamoto MP, Mikami T (2000b) The nad4L-orf25 gene cluster is conserved and expressed in sugar beet mitochondria. Theor Appl Genet 100:214–220
Mower JP (2005) PREP-Mt: predictive RNA editor for plant mitochondrial genes. BMC Bioinformatics 6:96
Mundel C, Schuster W (1996) Loss of RNA editing of rps1 sequences in Oenothera mitochondria. Curr Genet 30:455–460
Neuwirt J, Takenaka M, van der Merwe JA, Brennicke A (2005) An in vitro RNA editing system from cauliflower mitochondria: editing site recognition parameters can vary in different plant species. RNA 11:1563–1570
Nishizawa S, Kubo T, Mikami T (2000) Variable number of tandem repeat loci in the mitochondrial genomes of beets. Curr Genet 37:34–38
Notsu Y, Masood S, Nishikawa T, Kubo N, Akiduki G, Nakazono M, Hirai A, Kadowaki K (2002) The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: frequent DNA sequence acquisition and loss during the evolution of flowering plants. Mol Genet Genomics 268:434–445
Oda K, Yamato K, Ohta E, Nakamura Y, Takemura M, Nozato N, Akashi K, Kanegae T, Ogura Y, Kohchi T, Ohyama K (1992) Gene organization deduced from the complete sequence of liverwort Marchantia polymorpha mitochondrial DNA: a primitive form of plant mitochondrial genome. J Mol Biol 223:1–7
Ohyama K, Oda K, Ohta E, Takemura M (1993) Gene organization and evolution of introns of a liverwort, Marchantia polymorpha, mitochondrial genome. In: Brennicke A, Kuck U (eds) Plant mitochondria. Verlag Chemie, Weinheim, pp 115–129
Onodera Y, Yamamoto MP, Kubo T, Mikami T (1999) Heterogeneity of the atp6 presequences in normal and different sources of male-sterile cytoplasms of sugar beet. J Plant Physiol 155:656–660
Quinones V, Zanlungo S, Moenne A, Gomez I, Holuigue L, Litvak S, Jordana X (1996) The rpl5-rps14-cob gene arrangement in Solanum tuberosum: rps14 is a transcribed and unedited pseudogene. Plant Mol Biol 31:937–943
Sandoval P, Leon G, Gomez I, Carmona R, Figueroa P, Holuigue L, Araya A, Jordana X (2004) Transfer of RPS14 and RPL5 from the mitochondrion to the nucleus in grasses. Gene 324:139–147
Satoh M, Kubo T, Nishizawa S, Estiati A, Itchoda N, Mikami T (2004) The cytoplasmic male-sterile type and normal type mitochondrial genomes of sugar beet share the same complement of genes of known function but differ in the content of expressed ORFs. Mol Genet Genomics 272:247–256
Schuster W, Brennicke A (1991) RNA editing makes mistakes in plant mitochondria: editing loses sense in transcripts of a rps19 pseudogene and in creating stop codons in coxI and rps3 mRNAs of Oenothera. Nucleic Acids Res 19:6923–6928
Schuster W, Ternes R, Knoop V, Hiesel R, Wissinger B, Brennicke A (1991) Distribution of RNA editing sites in Oenothera mitochondrial mRNAs and rRNAs. Curr Genet 20:397–404
Schuster W, Wissinger B, Unseld M, Brennicke A (1990) Transcripts of the NADH-dehydrogenase subunit 3 gene are differentially edited in Oenothera mitochondria. EMBO J 9:263–269
Senda M, Mikami T, Kinoshita T (1993) The sugar beet mitochondrial gene for the ATPase alpha-subunit: sequence, transcription and rearrangements in cytoplasmic male-sterile plants. Curr Genet 24:164–170
Siqueira SF, Dias SM, Hardouin P, Pereira FR, Lejeune B, de Souza AP (2002) Transcription of succinate dehydrogenase subunit 4 (sdh4) gene in potato: detection of extensive RNA editing and co-transcription with cytochrome oxidase subunit III (cox3) gene. Curr Genet 41:282–289
Steinhauser S, Beckert S, Capesius I, Malek O, Knoop V (1999) Plant mitochondrial RNA editing. J Mol Evol 48:303–312
Sugiyama Y, Watase Y, Nagase M, Makita N, Yagura S, Hirai A, Sugiura M (2005) The complete nucleotide sequence and multipartite organization of the tobacco mitochondrial genome: comparative analysis of mitochondrial genomes in higher plants. Mol Genet Genomics 272:603–615
Sunkel S, Brennicke A, Knoop V (1994) RNA editing of a conserved reading frame in plant mitochondria increases its similarity to two overlapping reading frames in Escherichia coli. Mol Gen Genet 242:65–72
Sutton CA, Conklin PL, Pruitt KD, Hanson MR (1991) Editing of pre-mRNAs can occur before cis- and trans-splicing in Petunia mitochondria. Mol Cell Biol 11:4274–4277
Thomson MC, Macfarlane JL, Beagley CT, Wolstenholme DR (1994) RNA editing of mat-r transcripts in maize and soybean increases similarity of the encoded protein to fungal and bryophyte group II intron maturases: evidence that mat-r encodes a functional protein. Nucleic Acids Res 22:5745–5752
Turmel M, Otis C, Lemieux C (2003) The mitochondrial genome of Chara vulgaris: insights into the mitochondrial DNA architecture of the last common ancestor of green algae and land plants. Plant Cell 15:1888–1903
Wilson RK, Hanson MR (1996) Preferential RNA editing at specific sites within transcripts of two plant mitochondrial genes does not depend on transcriptional context or nuclear genotype. Curr Genet 30:502–508
Yang AJ, Mulligan RM (1991) RNA editing intermediates of cox2 transcripts in maize mitochondria. Mol Cell Biol 11:4278–4281
Acknowledgments
We wish to thank the National Agricultural Research Center for Hokkaido Region (Hokkaido, Japan) for providing the seeds used in this study. This research was supported by NIH Grant R01-GM-70612 to JDP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Herrmann
Nucleotide sequence data generated for this study are available in the DDBJ/EMBL/GenBank databases under accession numbers DQ381444–DQ381465.
An erratum to this article can be found at http://dx.doi.org/10.1007/s00438-006-0160-6
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mower, J.P., Palmer, J.D. Patterns of partial RNA editing in mitochondrial genes of Beta vulgaris . Mol Genet Genomics 276, 285–293 (2006). https://doi.org/10.1007/s00438-006-0139-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-006-0139-3