Abstract
Phosphoglycerate mutase (PGM) is a widely distributed glycolytic enzyme. Two known distinct classes of PGM enzymes were identified, a cofactor-dependent one (dPGM) and a cofactor-independent one (iPGM). A complementary DNA (cDNA) encoding a PGM was cloned from a Clonorchis sinensis cDNA library by large-scale sequencing. This new cDNA contains 955 bp with a putative open reading frame of 256 amino acids, which has a high homology with dPGMs from a number of species. The putative peptide was produced in E. coli and was purified to electrophoretic homogeneity. Enzymatic assays showed that the product of this gene could catalyze the conversion of 3-phosphoglycerate to 2-phosphoglycerate when the cofactor was present and the enzyme activities could be inhibited by vanadate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphoglycerate mutase (PGM, EC5.4.2.1) is an important enzyme that catalyzes the interconversion of 3-phosphoglycerate (3-PGA) and 2-phosphoglycerate (2-PGA) in the glycolytic and gluconeogenic pathways. There exist two types of PGMs according to the requirement for 2,3-bisphosphoglycerate (2,3-BPGA) for their enzyme activities. The PGMs requiring 2,3-BPGA for catalysis are termed cofactor-dependent PGMs (dPGMs). The one that do not require 2,3-BPGA for catalysis are termed cofactor-independent PGMs (iPGMs; Fothergill-Gilmore and Watson 1989).
The dPGM enzyme is composed of about 250 amino acids and distributes in all vertebrates, most invertebrates, some fungi, and bacteria. Their amino acid sequences showed a high degree of conservation across the species. In contract, iPGM is comprised of approximately 500 amino acids. It is present in all plants, algae, and some invertebrates, fungi, and bacteria, predominantly Gram-positive bacteria (Jedrzejas 2000). There is no significant amino acid sequence similarity between dPGMs and iPGMs (Grana et al. 1995). Sequence analysis has revealed that dPGM belongs to an enzyme family that includes acid phosphatase, fructose 2,6-bisphosphatase, and phytase, while iPGM belongs to the superfamily of alkaline phosphatase (Galperin et al. 1998; Jedrzejas 2000; Galperin and Jedrzejas 2001).
These two types of PGM enzymes are also different in their catalytic mechanisms and three-dimensional structure. dPGMs need 2,3-BPGA as a cofactor to be able to transfer phosphate groups from enzyme to substrate, while iPGMs do not need the cofactor, they transfer the phosphate group by themselves. The crystal structure of dPGM was first reported for Saccharomyces cerevisiae (Campbell et al. 1974), and later, a refined, high resolution one was reported (Rigden et al. 1998). It revealed great extensive structural similarity with rat prostatic acid phosphatase (Lindqvist et al. 1993) and rat liver fructose-2, 6-bisphosphatase (Lee et al. 1996). The three enzymes all have two active-site histidines and operate with phosphohistidines intermediates. The structure of the first iPGM was obtained from Bacillus stearothermophilus (Jedrzejas et al. 2000a). It contains two distinctly separated domains having the phosphatase and the phosphotranferase activity, respectively. The phosphatase domain has been shown to have a high similarity to E. coli alkaline phosphatase (Sowadski et al. 1985).
In this work, we report the cloning and characterization of a cDNA encoding a PGM from Clonorchis sinensis, which is one of the most important trematode that causes human clonorchiasis in China, Korea, Japan, and Southeast Asia. Sequence analysis indicated it was a dPGM. Enzymatic assay showed that it had the activities of dPGM, and vanadate could inhibit the enzyme activities. This work is the first report of a PGM in C. sinensis.
Materials and methods
Chemicals
SMART™ cDNA Library Constrction Kit was purchased from Clontech; QIAwell plasmid purification system and Ni-NTA HisTrap resin were purchased from Qiagen. Adenosine diphosphate (ADP) and nicotinamide adenine dinucleotide (reduced form) (NADH) were from Amersco and 3-PGA, 2,3-BPGA, enolase, pyruvate kinase, and lactate dehydrogenase were purchased from Sigma. All kinds of restriction endonucleases were obtained from New England Biolabs.
cDNA library construction and sequencing of the cDNA insert
The collection of adult C. sinensis worms was according to the described methods (Yang et al. 2006). The construction of C. sinensis cDNA library and the large-scale sequencing of cDNA inserts were carried out as described (Song et al. 2004; Zheng et al. 2005; Yang et al. 2006).
Bioinformatics analysis of CsPGM gene
DNA and the deduced protein sequence were analyzed using the BLASTN and BLASTP at NCBI Web Server (http://www.ncbi.nlm.nih.gov/blast). The open reading frame (ORF) was predicted by ORF finder program (http://www.ncbi.nlm.nih.gov/gorf/gorf.html). Sequence alignment was carried out using GeneDoc software.
Expression of CsPGM cDNA in E.coli
Primers were designed according to the putative ORF of C. sinensis phosphoglycerate mutase (CsPGM). The sense primer was 5′GGAATTCCATATGTACAAAACAAACTATATGG3′, and the antisense primer was 5′CCGCTCGAGCTTCTTTTTACCCTGATCGGC3′ with an NdeI site and an XhoI site incorporated, respectively. Polymerase chain reaction (PCR) was carried for 30 cycles at 94°C for 30 s, 55°C for 30 s, and 72°C for 45 s. The reaction was continued for 10 min at 72°C after the last cycle. The amplified PCR fragment was digested with NdeI/XhoI and cloned into the expression vector pET24b (+). The ligation mixture was transformed into E. coli BL21 (DE3). After being cultured overnight in Luria–Bertani (LB) plate containing 50 μg/ml kanamycin, plasmids were isolated and sequenced to confirm the correct insertion of the cDNA fragment. After sequencing, a correct transformant was picked up and cultured at 37°C in LB media containing 50 μg/ml kanamycin. The bacterial cells were induced by final concentration of 1 mM isopropylthiogalactoside (IPTG) when they grew to OD600 = 0.4 ∼ 0.6 and continuously cultured for 3 h.
Purification of recombinant CsPGM
A single transformant was inoculated into 2 ml LB media containing 50 μg/ml kanamycin and cultured overnight. Then, the overnight culture was diluted into 200 ml LB containing 50 μg/ml kanamycin and grew at 37°C for 3 h. The bacterial cells were induced with 1 mM IPTG for 3 h before harvesting at 5,000 g for 10 min at 4°C. The cell pellets were resuspended in phosphate-buffered saline (140 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, and 1.8 mM KH2PO4) containing 0.5 mM phenylmethylsulphonyl fluoride and 2 mM β-mercaptoethanol for sonication with 1-s work and 1-s pause in between for 10 min. The homogenate was centrifuged at 12,000×g 4°C for 15 min, and the supernatant was collected and loaded onto the Ni–NTA HisTrap resin. CsPGM–his fusion protein was eluted with 50 and 150 mM imidazole and analyzed with sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). The purified protein was dialyzed with 30 mM Tris–HCl pH 7.0. Protein concentration was determined using the Bradford method (Bradford 1976).
Enzymatic activities assay of recombinant CsPGM
The CsPGM activity was assayed in the forward reaction from 3-PGA to 2-PGA by measuring the decrease of NADH in a standard enzyme-coupled assay (Fraser et al. 1999). The reaction mixture contained 30 mM Tris–HCl pH 7.0, 20 mM KCl, 5 mM MgCl2, 1 mM ADP, 0.15 mM NADH, 10 mM 3-PGA, 0.1 mM 2,3-BPGA, 2 U enolase, 2 U pyruvate kinase, and 2 U lactate dehydrogenase. Reactions were performed for 5 min with data collected at 15-s intervals at wave of 340 nm. The enzyme reactions were measured with a U-3000 spectrophotometer (Hatachi, Japan). The optimal temperature and pH of the CsPGM enzyme were determined by incubating the standard reaction mixture at various temperatures and in buffer of various pH values. Kinetic parameters were determined at 25°C in Tris–HCl buffer (pH 7.0) by varying the concentration of 3-PGA (0.1 ∼ 5 mM) with 0.1 μg recombinant CsPGM. The results were analyzed by double reciprocal Lineweaver–Burk plot.
Inhibition of the CsPGM’s activity
Vanadate inhibition experiments were carried out in glycolytic direction by incubation of sodium metavanadate and recombinant CsPGM for 10 min at room temperature in buffer pH 7.0, then added them to the reaction mixture and started the assay. Each concentration of vanadate was measured three times under saturating substrate concentrations (5 mM 3-PGA).
Results
Cloning and sequence analysis of CsPGM gene
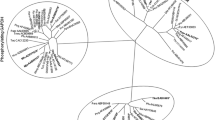
CsPGM gene was cloned from the adult C. sinensis cDNA library by large-scale sequencing. The cDNA is 955 bp in length with a putative ORF of 256 amino acids (Fig. 1). Bioinformatics analysis revealed that CsPGM shares a high degree of homology (58% identity and 73% similarity) with human B-form PGM. It still has a high homology with dPGMs from other species (Fig. 2).
Nucleotide sequence and the deduced amino acid sequence of the Clonorchis sinensis PGM cDNA. The putative translation start codon is in bold letters, and the stop codon is indicated by an asterisk. The polyadenylation signal at the 3′-end is underlined. The nucleotide sequence reported in this paper has been submitted to the GenBank database with accession number AY796059
Sequence alignment of Clonorchis sinensis PGM and PGMs from other organisms. Conserved amino acids are shaded. Conserved residues that constitute the substrate binding site (asterisk) and the catalytic histidine (inverted filled triangle) are indicated. The accession numbers of the aligned proteins are as follows: AY796059 (Clonorchis sinensis), AAH53356 (human brain), AAH73741 (human muscle), NP_059024 (rat), NP_061358 (mouse), S50326 (Drosophila melanogaster), CAA41595 (Saccharomyces cerevisiae), and NP_415276 (Escherichia coli)
Expression and purification of CsPGM protein
The cDNA of the putative CsPGM ORF was amplified by PCR and cloned into the expression vector pET24b(+). The recombinant vector was named pET-CsPGM. This vector has a hexahistidine tag at the end of multiclone sites; therefore, Ni-NTA HisTrap resin was used to purify CsPGM–his fusion protein. As judged from SDS-PAGE analysis, CsPGM protein was purified to apparent homogeneity (Fig. 3) with 150 mM imidazole elution. The apparent molecular mass of the protein was about 29.8 kDa. The concentration of the purified protein was 4 mg/ml, and it was used for enzymatic activity assay.
Expression and purification of recombinant CsPGM. Proteins were resolved on 12% SDS-PAGE and stained with Coomassie blue. Lane 1 Protein marker; lane 2 total cell protein sample before induced by IPTG; lane 3 total cell protein sample after induced by 1 mM IPTG for 3 h; lane 4 elution fraction with 50 mM imidazole; lane 5 elution fraction with 150 mM imidazole
Enzymatic activity assay of CsPGM protein
The activity of the purified recombinant CsPGM was detected in glycolytic direction (conversation of 3-PGA to 2-PGA). Based on the bioinformatics analysis, CsPGM belongs to dPGM. The CsPGM showed PGM activity when the enzymatic assay system contained 2,3-BPGA. The optimal temperature of this enzyme was 45°C, and its optimal pH value was 7.0. The Km value for the substrate 3-PGA was 9.76 × 10-4 M, which had the same magnitude with the E. coli dPGM (Fraser et al. 1999).
Inhibition effect of vanadate on CsPGM’s activity
Vanadate is known to be a potent inhibitor of cofactor-dependent PGMs but does not inhibit cofactor-independent PGMs (Carreras et al. 1980). Our inhibition experiments showed that vanadate could strongly inhibit CsPGM’s activity when 1 μM inhibitor was used. The inhibition effect was gradually increased according to the concentration increasing of vanadate. By fitting the data for dPGM inhibition with an equation for reversible, competitive inhibition, the inhibitory constant of 15.7 nM was determined.
Discussion
Studies showed that PGM is not only an important enzyme in the glycolytic and gluconeogenic pathways, it also plays essential roles in other ways. When reducing the activities of PGM by RNA interference in C. elegans, it led to multiple developmental defects such as embryonic lethality, larval lethality, and abnormal body morphology (Zhang et al. 2004). Deletion of the iPGM gene in a spore-forming bacterium, Bacillus subtilis, resulted in extremely slow growth and an inability to produce spores (Leyva-Vazquez and Setlow 1994). Inactivation of the iPGM locus by a transposon insertion in the tomato bacterial pathogen Pseudomonas syringae resulted in a mutant strain that could not grow or infect tomatoes (Morris et al. 1995).
Phosphoglycerate mutases are divided into two classes based on their requirement for the cofactor 2,3-diphosphoglycerate. The cofactor-dependent and cofactor-independent PGMs have different catalytic mechanism. dPGM needs 2,3-BPGA as a cofactor and catalyzes the intermolecular transfer of the phosphate group between the monophosphoglycerates and the cofactor through a phosphohistidine intermediate (Rigden et al. 2002; Jedrzejas et al. 2000a). In contrast, iPGM does not require 2,3-BPGA as a cofactor; it catalyzes the intramolecular transfer of the phosphate group on monophosphoglycerates through a phosphoserine intermediate (Jedrzejas et al. 2000b). These two types of PGMs have their own different active sites amino acid. Our enzymatic assay suggested that CsPGM is a dPGM, and sequences analysis indicated that the histidines lie in position 17, and 190 might be its active site.
Vanadate is a potent inhibitor of the dPGM and does not inactivate iPGM (Fraser et al. 1999). It is often used to discriminate the dPGM from that of the structurally unrelated iPGM (Jedrzejas et al. 2000b). In our results of inhibition experiments, vanadate could inhibit the CsPGM’s activity distinctly. It is supposed that vanadate influences the spatial structure of PGM when it binds to the enzyme, as the studies of circular dichroism spectroscopy revealed; when vanadate is added to the CsPGM solution, the secondary structure of the protein had changed (data not shown). The structure of E.coli dPGM complexed with vanadate has shown that the inhibitor is present in the active site of the PGM (Bond et al. 2002). We supposed that vanadate probably binds to CsPGM in the histidine active site and alters its conformation, thus preventing the binding of substrate to CsPGM, resulting in the inhibition.
The whole-genome sequence has predicted that C. elegans has only iPGM, and E. coli has both iPGM and dPGM (Zhang et al. 2004; Fraser et al. 1999). In our work, a new cDNA encoding a PGM was cloned from C. sinensis by a cDNA library construction and large-scale sequencing. Sequence analysis and enzymatic assay revealed that CsPGM belongs to the dPGM family. To illustrate this, CsPGM’s catalytic mechanism, crystal structure, and mutagenesis experiments should be done, and these works are under way.
References
Bond CS, White MF, Hunter WN (2002) Mechanistic implications for Escherichia coli cofactor-dependent phosphoglycerate mutase based on the high-resolution crystal structure of a vanadate complex. J Mol Biol 316:1071–1081
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Campbell JW, Watson HC, Hodgson GI (1974) Structure of yeast phosphoglycerate mutase. Nature 250:301–303
Carreras J, Bartrons R, Grisolia S (1980) Vanadate inhibits 2,3-bisphosphoglycerate dependent phosphoglycerate mutases but does not affect the 2,3-bisphosphoglycerate independent phosphoglycerate mutases. Biochem Biophys Res Commun 96:1267–1273
Fothergill-Gilmore LA, Watson HC (1989) The phosphoglycerate mutases. Adv Enzymol Relat Areas Mol Biol 62:227–313
Fraser HI, Kvaratskhelia M, White MF (1999) The two analogous phosphoglycerate mutases of Escherichia coli. FEBS Lett 455:344–348
Galperin MY, Jedrzejas MJ (2001) Conserved core structure and active site residues in alkaline phosphatase superfamily enzymes. Proteins 45:318–324
Galperin MY, Bairoch A, Koonin EV (1998) A superfamily of metalloenzymes unifies phosphopentomutase and cofactor-independent phosphoglycerate mutase with alkaline phosphatases and sulfatases. Protein Sci 7:1829–1835
Grana X, Perez de la Ossa P, Broceno C, Stocker M, Garriga J, Puigdomenech P, Climent F (1995) 2,3-Bisphosphoglycerate-independent phosphoglycerate mutase is conserved among different phylogenic kingdoms. Comp Biochem Physiol B Biochem Mol Biol 112:287–293
Jedrzejas MJ (2000) Structure, function, and evolution of phosphoglycerate mutases: comparison with fructose-2,6-bisphosphatase, acid phosphatase, and alkaline phosphatase. Prog Biophys Mol Biol 73:263–287
Jedrzejas MJ, Chander M, Setlow P, Krishnasamy G (2000a) Structure and mechanism of action of a novel phosphoglycerate mutase from Bacillus stearothermophilu. EMBO J 19:1419–1431
Jedrzejas MJ, Chander M, Setlow P, Krishnasamy G (2000b) Mechanism of catalysis of the cofactor-independent phosphoglycerate mutase from Bacillus stearothermophilus. Crystal structure of the complex with 2-phosphoglycerate. J Biol Chem 275:23146–23153
Lee YH, Ogata C, Pflugrath JW, Levitt DG, Sarma R, Banaszak LJ, Pilkis SJ (1996) Crystal structure of the rat liver fructose-2,6-bisphosphatase based on selenomethionine multiwavelength anomalous dispersion phases. Biochemistry 35:6010–6019
Leyva-Vazquez MA, Setlow P (1994) Cloning and nucleotide sequences of the genes encoding triose phosphate isomerase, phosphoglycerate mutase, and enolase from Bacillus subtilis. J Bacteriol 176:3903–3910
Lindqvist Y, Schneider G, Vihko P (1993) Three-dimensional structure of rat acid phosphatase in complex with L(+)-tartrate. J Biol Chem 268:20744–20746
Morris VL, Jackson DP, Grattan M, Ainsworth T, Cuppels DA (1995) Isolation and sequence analysis of the Pseudomonas syringae pv. tomato gene encoding a 2,3-diphosphoglycerate-independent phosphoglyceromutase. J Bacteriol 177:1727–1733
Rigden DJ, Alexeev D, Phillips SE, Fothergill-Gilmore LA (1998) The 2.3 Å X-ray crystal structure of S. cerevisiae phosphoglycerate mutase. J Mol Biol 276:449–459
Rigden DJ, Mello LV, Setlow P, Jedrzejas MJ (2002) Structure and mechanism of action of a cofactor-dependent phosphoglycerate mutase homolog from Bacillus stearothermophilus with broad specificity phosphatase activity. J Mol Biol 315:1129–1143
Song L, Chen S, Yu X, Wu Z, Xu J, Yang G, Zheng N, Hu X, Guo L, Dai J, Xu J, Ji C, Gu S, Ying K (2004) Molecular cloning and characterization of cDNA encoding a ubiquitin-conjugating enzyme from Clonorchis sinensis. Parasitol Res 94:227–232
Sowadski JM, Handschumacher MD, Murthy HM, Foster BA, Wyckoff HW (1985) Refined structure of alkaline phosphatase from Escherichia coli at 2.8 Å resolution. J Mol Biol 186:417–433
Yang G, Jing C, Zhu P, Hu X, Xu J, Wu Z, Yu X (2006) Molecular cloning and characterization of a novel lactate dehydrogenase gene from Clonorchis sinensis. Parasitol Res 99:55–64
Zhang Y, Foster JM, Kumar S, Fougere M, Carlow CK (2004) Cofactor-independent phosphoglycerate mutase has an essential role in Caenorhabditis elegans and is conserved in parasitic nematodes. J Biol Chem 279:37185–37190
Zheng N, Xu J, Wu Z, Chen J, Hu X, Song L, Yang G, Ji C, Chen S, Gu S, Ying K, Yu X (2005) Clonorchis sinensis: molecular cloning and functional expression of novel cytosolic malate dehydrogenase. Exp Parasitol 109:220–227
Acknowledgment
This research was supported by grants from the Natural Science Foundation of Guangdong Province, China (No.2002B31005) and the Key Program of Science and Technology Department of Guangdong Province, China (No.200223-E4022). The experiments comply with the current laws of the country in which the experiments were performed.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Song, L., Xu, Z. & Yu, X. Molecular cloning and characterization of a phosphoglycerate mutase gene from Clonorchis sinensis . Parasitol Res 101, 709–714 (2007). https://doi.org/10.1007/s00436-007-0540-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-007-0540-9