Abstract
Purpose
Thyroid cancer is the most common endocrine malignancy worldwide. The molecular mechanisms underlying thyroid tumorigenesis remain unclear. Some studies suggested that the ZCCHC12 gene correlates with certain diseases. However, the function of ZCCHC12 in thyroid cancer has yet to be determined. This study investigated the role of the ZCCHC12 gene in papillary thyroid cancer (PTC).
Methods
We conducted a comprehensive analysis of massively parallel whole-transcriptome resequencing of matched PTC tumors and normal tissues in 19 patients. Results showed that the expression of ZCCHC12 was significantly upregulated in thyroid cancer. qRT-PCR was performed to confirm previous results. The functions of the ZCCHC12 gene in PTC cell lines (TPC1 and BCPAP) transfected with small interfering RNA were determined through cell colony formation assay, migration assay, and invasion assay.
Results
The ZCCHC12 gene was remarkably upregulated in primary PTC tumors in both validated cohort (T:N = 1.80 ± 2.58:0.23 ± 0.50, P < 0.01) and TCGA cohort (T:N = 7.63 ± 3.25:1.55 ± 1.71, P < 0.01). We also achieved area under the curve (AUC of ROC) of 87.9% for the validated cohort while 91.4% for the TCGA cohort to classify PTC tumors and normal tissues. ZCCHC12 overexpression correlated with lymph node metastasis in both cohorts (P < 0.05). In in vitro experiments, ZCCHC12 downregulation significantly inhibited the colony formation, migration, and invasion of PTC cells.
Conclusion
Our study indicated that ZCCHC12 gene has important biological functions and acts as a metastasis-related oncogene in PTC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thyroid cancer is the most common endocrine malignancy worldwide, with 64,300 newly estimated diagnosed cases and 1980 estimated deaths in the USA in 2015 (Siegel et al. 2015). In China, the number of newly estimated diagnosed cases and deaths was 90,000 and 6800, respectively (Chen et al. 2016). Papillary thyroid cancer (PTC), characterized by diverse biological behavior, accounts for approximately 80% of thyroid cancer cases (Morris et al. 2016). The tumorigenesis and development of thyroid cancer are mainly driven by genomic variations, including the activation of tumor promoter genes and the silencing of tumor suppressor genes (Xing 2005, 2010, 2013). It was confirmed that BRAF mutation could accelerate PTC tumorigenesis and progression by aberrantly activating the MAP kinase pathway (Xing 2005). Mutations in the TERT (Liu et al. 2013), RAS (Liu et al. 2008), PIK3CA (Garcia-Rostan et al. 2005), and TP53 (Donghi et al. 1993) genes also play an important role in thyroid cancer initiation and development. Although genetic research has achieved much progress, the mechanisms underlying PTC tumorigenesis remain unclear. Thus, the molecular mechanisms underlying PTC tumorigenesis should be elucidated to identify potential therapeutic targets and novel diagnostic and prognostic biomarkers.
The mouse zinc finger CCHC domain containing 12 gene (ZCCHC12), also known as Smad-interacting zinc finger protein 1 (Sizn1), is localized on chromosome X. This gene serves as a transcriptional coactivator in the bone morphogenic protein (BMP) signaling pathway (Cho et al. 2008b). The human ZCCHC12 gene shares 74% sequence identity with the mouse ZCCHC12 gene. Variants of the human ZCCHC12 gene have been associated with some diseases (Cho et al. 2008a, 2009; Li et al. 2012), but the role of ZCCHC12 gene in the initiation and progression of thyroid carcinoma has yet to be clarified. Therefore, investigation of the biological functions of the human ZCCHC12 gene is crucial to understand PTC tumorigenesis and progression.
In our previous study, we performed the whole-transcriptome resequencing of 19 pairs of PTC tumors and adjacent normal tissues and found that human ZCCHC12 expression is remarkably upregulated in PTC tumors. A systematic literature review revealed that the function of human ZCCHC12 in PTC tumorigenesis is still under exploration. Thus, we compared the ZCCCH12 expression in PTC tumors and in normal tissues on a large scale. We also analyzed the relationship between ZCCHC12 expression and clinical features, as well as examined the function of ZCCHC12 in PTC cell lines by using small interfering RNA (si-RNA). The current study aimed to investigate the role of the ZCCHC12 gene in PTC.
Materials and methods
Patients and samples
With the approval of the Ethnic Committee of the First Affiliated Hospital of Wenzhou Medical University, a total of 47 primary PTCs and matched non-cancerous thyroid tissues were obtained at the time of initial surgery. Samples were snap-frozen in liquid nitrogen immediately after resection and then stored at −80 °C before RNA extraction. All samples were reviewed retrospectively to confirm the histological diagnosis and to ensure abundant cancer content of the tumor by two pathologists. Informed consent for the scientific use of biological material was obtained from each patient. The normalized mRNA expression counts were obtained via the TCGA portal and are expressed as RNA-Seq by transcripts per kilobase million (TPM) values. We downloaded TCGA clinical data in ‘Biotab’ format from the TCGA portal.
RNA extraction and real-time reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA from the tissues and cells were isolated by TRIzol reagent according to manufacturer’s protocol (Life Technologies, Carlsbad, CA), and the cDNA was prepared using ReverTra Ace® qPCR RT Kit (Toyobo, Osaka, Japan). Real-time reverse transcription polymerase chain reaction (qRT-PCR) was performed using Thunderbird SYBR qPCR Mix (Toyobo, Osaka, Japan) in the Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA). The primer sequences for PCR are as follows: ZCCHC12, forward CTGGGGAGAAGTCTCGGTC and reverse GGCCCCTAAGGGTTTTCATCA. Each sample was in triplicate. GAPDH was used as an internal control.
Cell lines and cell culture
Human thyroid cancer cell lines TPC1 and BCPAP were kindly provided by Professor Mingzhao Xing of the Johns Hopkins University School of Medicine, Baltimore, MA, USA. TPC-1 and BCPAP were cultured at 37 °C with 5% CO2 in RPMI 1640 with 10% fetal bovine serum (FBS) (Life Technologies, Carlsbad, CA) and 1× MEM nonessential amino acids +1 × sodium pyruvate.
RNA interference
Small interfering RNA (si-RNA) targeting ZCCHC12 and control si-RNA (si-NC) were provided by Gene Pharma (Shanghai, China) for si-RNA-mediated gene knockdown. The sequences of si-RNA are as follows: ZCCHC12, forward 5′-GGAUGAAGUGCUGGUCAUUTT-3′, reverse 5′-AAUGACCAGCACUUCAUCCTT-3′. Cells were transfected at 50% confluence using Lipo iMAX (Invitrogen, Grand Island, NY), with a final si-RNA concentration of 100 nM (TPC-1) or 50 nM (BCPAP), respectively. Cells were harvested 48 h after transfection for subsequent RNA expression analysis. All knockdown experiments were performed in three replicates.
Cell colony formation assay
Colony formation assay was performed using monolayer culture. The indicated cells were seeded into six-well plates at 3 × 103 cells/well (TPC1), 5 × 103 cells/well (BCPAP). The medium was refreshed every 2 days. After 8–14 days of culture, surviving colonies were fixated with 4% PFA (paraformaldehyde, Sigma, USA) for 30 min and stained with 0.01% crystal violet for 30 min, and then, those colonies were counted. All colony formation assays were performed in three replicates.
Transwell migration and invasion assay
Migration and invasion assays were performed using transwell chambers. For migration assays, we used transwell cell culture chambers (Corning Costar Corp, Cambridge, MA, USA). The two transfectant cells (5 × 104 cells) were seeded with 10% FBS medium onto the top chamber, and the bottom chamber was filled with same concentration medium. For invasion assays, we used BioCoat™ Matrigel Invasion chamber 24-Well Plate 8.0 Micron (Corning, NY, USA). Cells (10 × 104 for TPC1,15 × 104 for BCPAP) were seeded with 10% FBS medium onto the top chamber, and the bottom chamber was filled with 20% FBS medium. After 24 h of seeding, the indicated cells and the bottom membranes were fixated with 4% PFA (Sigma) for 30 min, stained with 0.01% crystal violet for 30 min, and then photographed under a microscope at a magnification of 200×.
Statistical analysis
The Mann–Whitney U test was used to evaluate the expression difference between PTC tumors and adjacent normal tissues in the validated cohort and the TCGA cohort. Chi-square test or Fisher’s exact test was used to evaluate the relationship between clinicopathological characteristics and ZCCHC12 expression. Logistic regression analysis was performed to assess the lymph node metastatic risk of ZCCHC12. All P values were two-sided, and a P value of <0.05 was considered statistically significant. All statistical analyses were performed using SPSS version 22.0 (Chicago, IL). GraphPad Prism 5 (GraphPad Software, La Jolla, CA, USA) was used for graphs.
Results
ZCCHC12 gene was overexpressed in PTC
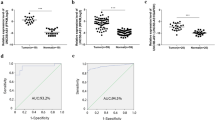
To confirm the data of whole-transcriptome resequencing, we first detected the mRNA expression of ZCCHC12 in 47 pairs of PTC tumor samples and matched adjacent non-cancerous tissues by qRT-PCR. As shown in Fig. 1a, the expression of ZCCHC12 was significantly upregulated in the tumor samples compared with matched adjacent non-cancerous tissues (T:N = 1.80 ± 2.58:0.23 ± 0.50, P < 0.01). This finding was further confirmed by The Cancer Genome Atlas (TCGA) dataset, which presented that the mRNA expression level of ZCCHC12 was remarkably higher in PTC than in the normal control (T:N = 7.63 ± 3.25:1.55 ± 1.71, P < 0.01) (Fig. 1b). A receiver operator characteristic (ROC) curve was plotted to determine the diagnostic value of ZCCHC12 expression. We achieved sensitivity and specificity rates of 93.62 and 74.47% for the validated cohort (AUC: 87.9%) while 86.53 and 96.61% for the TCGA cohort (AUC: 91.4%) to discriminate PTC tumors from normal tissues (Fig. 1c, d).
ZCCHC12 expression was significantly upregulated in thyroid cancer samples compared with normal thyroid tissues in both validated cohort and TCGA cohort. Notes a In the validated cohort, ZCCHC12 expression was examined by qRT-PCR in 47 paired PTC samples and adjacent normal tissues (Mann–Whitney U test, ***P < 0.01). b The TCGA cohort contained 490 tumor samples and 59 normal tissue samples. TPM was represented for ZCCHC12 expression. Data were represented as log2 fold changes. Analysis was done using the Mann–Whitney U test (***P < 0.01). All data were median ± interquartile ranges of ZCCHC12 expression. c ROC curve for ZCCHC12 expression to diagnose PTC in the validated cohort. The area under the ROC curve (AUC) was 87.9%, with 93.62% sensitivity and 74.47% specificity. d ROC curve for ZCCHC12 expression to diagnose PTC in the TCGA cohort. The AUC was 91.4%, with 86.53% sensitivity and 96.61% specificity
Relationship between ZCCHC12 expression and clinicopathologic features
To determine whether or not ZCCHC12 expression is associated with the tumorigenesis and progression of PTC, we investigated the relationship of this gene with clinicopathologic features. We divided PTC patients into low- and high-ZCCHC12-expression groups on the basis of the median value. In the TCGA cohort, results revealed that age (P < 0.001), lymph node metastasis (P < 0.001), and histological type (P = 0.002) were significantly related to high ZCCHC12 expression (Table 1). Meanwhile, the results from the validated cohort were consistent with the previous finding that high ZCCH12 expression corresponds to more lymph node metastasis (P = 0.022) (Table 2). However, no significant associations were found between ZCCH12 expression and gender, tumor size, and clinical stage in both cohorts (P > 0.05).
High ZCCHC12 expression aggravates the lymph node metastatic risk of PTC patients
The relationship between ZCCHC12 expression and lymph node metastasis was then further analyzed. Univariate logistic regression analysis showed that the significant variables for lymph node metastasis were age (OR 0.660, 95% CI 0.486–0.896, P = 0.008), histological type (OR 0.731, 95% CI 0.546–0.946, P = 0.017), tumor size (OR 1.956, 95% CI 1.626–2.353, P < 0.001), and ZCCHC12 expression (OR 2.071, 95% CI 1.488–2.884, P < 0.001) (Table 3). Multivariate logistic analysis using the four mentioned parameters also revealed that tumor size (OR 2.128, 95% CI 1.747–2.591, P < 0.001) and ZCCHC12 expression (OR 1.992, 95% CI 1.392–2.851, P < 0.001) were positively correlated with more lymph node metastasis, whereas age (OR 0.611, 95% CI 0.435–0.857, P = 0.004) and histological type (OR 0.715, 95% CI 0.543–0.942, P = 0.017) were negatively associated with lymph node metastasis (Table 4). In summary, ZCCHC12 overexpression could aggravate the risk of lymph node metastasis in PTC patients.
Downregulation of ZCCHC12 expression inhibits thyroid cancer cell colony formation
Given that the ZCCH12 gene is commonly overexpressed in PTC, we hypothesized that this gene performs an oncogenic role in thyroid cancer initiation and progression. Thus, we tested the growth-inhibitory effect by silencing ZCCHC12 expression in TPC1 and BCPAP cells using targeted si-RNA. qRT-PCR was performed to confirm the efficiency of ZCCHC12 knockdown by targeted si-RNA (Fig. 2a). Results revealed that ZCCHC12 knockdown effectively inhibited the colony formation of thyroid cancer cells compared with the control (P < 0.01) (Fig. 2b, c).
ZCCHC12 knockdown in two papillary thyroid cancer cell lines (TPC1 and BCPAP) and colony formation assay. Notes a ZCCHC12 knockdown was checked by qRT-PCR. ZCCHC12 mRNA expression was significantly downregulated in TPC1 and BCPAP cells transfected with targeted si-RNA compared with NC (Student’s t test, ***P < 0.01). b Colony formation assay. TPC1 and BCPAP cells transfected with targeted si-RNA and NC cells were seeded in six-well plates at proper density. After 8–14 days of culture, surviving colonies were fixated with 4% PFA and stained with 0.01% crystal violet. c Relative quantification of the colony number. The columns represent the mean colony number from at least three independent experiments, and the little vertical bars on the top of the columns represent SD. ***P < 0.01 in comparison with NC using Student’s t test
ZCCHC12 knockdown inhibits thyroid cancer cells migration and invasion
Considering that the clinical feature analysis revealed an association between high ZCCHC12 expression and lymph node metastasis, we examined the effect of ZCCHC12 on the migration and invasion of thyroid cancer cells. Our data revealed that ZCCHC12 knockdown could dramatically compromise the cell migration and invasion of thyroid cancer cells compared with the control groups (P < 0.01) (Fig. 3). The in vitro data are consistent with our previous clinical feature analysis results, suggesting that ZCCHC12 is a vital gene for the metastasis of human thyroid cancers.
Downregulation of ZCCHC12 inhibits papillary thyroid cancer cell lines (TPC1 and BCPAP) cell migration and invasion. Notes a Migration assay. Migrating cell number was much less in TPC1 and BCPAP cells transfected with targeted si-RNA compared with NC. b Relative quantification of the migrating cell number. The columns represent the mean migrating cell number from at least three independent experiments, and the little vertical bars on the top of the columns represent SD. ***P < 0.01 in comparison with NC using Student’s t test. c Invasion assay. Invading cell number was much less in TPC1 and BCPAP cells transfected with targeted si-RNA compared with NC. d Relative quantification of the invading cell number. The columns represent the mean invading cell number from at least three independent experiments, and the little vertical bars on the top of the columns represent SD. ***P < 0.01 in comparison with NC using Student’s t test
Discussion
Thyroid cancer is a highly heterogeneous disease that can be classified into several histological types depending on molecular etiology (Lloyd et al. 2011). Patients with PTC, being the most common type of thyroid carcinomas, have relatively better prognosis with lower mortality rate than patients with the other types (Xing 2013). However, PTC could exhibit different biological behavior in tumorigenesis because of genomic variations (Landa et al. 2016; Lloyd et al. 2011). Thus, making treatment decisions heavily relies on the information provided by current clinical and pathologic parameters is insufficient to personalize therapeutic regimen and evaluate the risk for each PTC patient (American Thyroid Association Guidelines Taskforce on Thyroid et al. 2009). It is urgent for us to find novel molecular biomarkers that can predict the progression and metastasis status of PTCs in clinical practice.
Only few studies have reported that the abnormal expression of the human ZCCHC12 gene accounts for different types of diseases. Cho et al. (2008a) found that different sequence variants in the human ZCCHC12 gene could lead to non-syndromic, X-linked mental retardation. Furthermore, ZCCHC12 could reportedly localize to promyelocytic leukemia protein nuclear bodies (PML-NBs) and participate in SUMO attachment (Cho et al. 2009). Li et al. (2009) found that the ZCCHC12 gene is notably expressed in the human brain and mouse embryonic brain and serves as a positive modulator of activator protein-1 (AP-1). Meanwhile, Li et al. (2012) confirmed that the mRNA expression of ZCCHC12 is preferentially increased in PTC tumors compared with nodular goiters, implicating the ZCCHC12 gene as a potential diagnostic molecular marker of PTC. However, the role of ZCCHC12 in PTC tumorigenesis remains unclear.
In the present study, we reported and validated that ZCCHC12 gene expression is significantly upregulated in PTC compared with adjacent non-cancerous tissues in 47 PTC patients. The abnormal expression of ZCCHC12 was also confirmed in the TCGA cohort. Clinicopathologic feature analysis revealed that ZCCHC12 gene expression positively correlated with lymph node metastasis in both the validated and TCGA cohorts. Interestingly, high ZCCHC12 expression positively correlated with the classical type of PTC in the TCGA cohort, indicating that ZCCHC12 is involved in the subtype transformation of PTC, which could increase the prognostic risk of patients (Shi et al. 2016). Moreover, logistic regression analysis indicated that high ZCCHC12 expression could aggravate the risk of lymph node metastasis in PTC patients. Loss-of-function analysis showed that ZCCHC12 knockdown can inhibit the cell colony formation of PTC cells in vitro. Moreover, withdrawal of ZCCHC12 significantly impaired the migration and invasion of PTC cells, which was consistent with our previous findings. Taken together, our study demonstrated that ZCCHC12 is involved in PTC progression and acts as a metastasis-related oncogene. To the best of our knowledge, this study is the first to report the function of ZCCHC12 in the progression of thyroid cancer.
Our study still contains some limitations. Given the lack of data on disease-free survival and overall survival, the prognostic value of ZCCHC12 must be discussed upon complete data collection. Furthermore, the mechanisms of action of ZCCHC12 in PTC tumorigenesis, especially in tumor metastasis, remain unclear and warrant further investigation.
In conclusion, ZCCHC12 is generally overexpressed in primary PTC. ZCCHC12 knockdown inhibits thyroid tumorigenesis by impairing cell colony formation, metastasis, and invasion. The results of this study provide a new potential marker for the diagnosis and treatment of thyroid cancer.
References
American Thyroid Association Guidelines Taskforce on Thyroid N et al (2009) Revised American Thyroid Association management guidelines for patients with thyroid nodules and differentiated thyroid cancer. Thyroid Off J Am Thyroid Assoc 19:1167–1214. doi:10.1089/thy.2009.0110
Chen W et al (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66:115–132. doi:10.3322/caac.21338
Cho G et al (2008a) Evidence that SIZN1 is a candidate X-linked mental retardation gene. Am J Med Genet Part A 146A:2644–2650. doi:10.1002/ajmg.a.32472
Cho G, Lim Y, Zand D, Golden JA (2008b) Sizn1 is a novel protein that functions as a transcriptional coactivator of bone morphogenic protein signaling. Mol Cell Biol 28:1565–1572. doi:10.1128/MCB.01038-07
Cho G, Lim Y, Golden JA (2009) SUMO interaction motifs in Sizn1 are required for promyelocytic leukemia protein nuclear body localization and for transcriptional activation. J Biol Chem 284:19592–19600. doi:10.1074/jbc.M109.010181
Donghi R, Longoni A, Pilotti S, Michieli P, Della Porta G, Pierotti MA (1993) Gene p53 mutations are restricted to poorly differentiated and undifferentiated carcinomas of the thyroid gland. J Clin Investig 91:1753–1760. doi:10.1172/JCI116385
Garcia-Rostan G et al (2005) Mutation of the PIK3CA gene in anaplastic thyroid cancer. Cancer Res 65:10199–10207. doi:10.1158/0008-5472.CAN-04-4259
Landa I et al (2016) Genomic and transcriptomic hallmarks of poorly differentiated and anaplastic thyroid cancers. J Clin Investig 126:1052–1066. doi:10.1172/JCI85271
Li H et al (2009) Human ZCCHC12 activates AP-1 and CREB signaling as a transcriptional co-activator. Acta Biochim Biophys Sin 41:535–544
Li QL, Chen FJ, Lai R, Guo ZM, Luo R, Yang AK (2012) ZCCHC12, a potential molecular marker of papillary thyroid carcinoma: a preliminary study. Med Oncol 29:1409–1417. doi:10.1007/s12032-011-0018-6
Liu Z et al (2008) Highly prevalent genetic alterations in receptor tyrosine kinases and phosphatidylinositol 3-kinase/Akt and mitogen-activated protein kinase pathways in anaplastic and follicular thyroid cancers. J Clin Endocrinol Metab 93:3106–3116. doi:10.1210/jc.2008-0273
Liu X et al (2013) Highly prevalent TERT promoter mutations in aggressive thyroid cancers. Endocr Relat Cancer 20:603–610. doi:10.1530/ERC-13-0210
Lloyd RV, Buehler D, Khanafshar E (2011) Papillary thyroid carcinoma variants. Head Neck Pathol 5:51–56. doi:10.1007/s12105-010-0236-9
Morris LG, Tuttle RM, Davies L (2016) Changing trends in the incidence of thyroid cancer in the United States. JAMA Otolaryngol Head Neck Surg 142:709–711. doi:10.1001/jamaoto.2016.0230
Shi X et al (2016) Differential clinicopathological risk and prognosis of major papillary thyroid cancer variants. J Clin Endocrinol Metab 101:264–274. doi:10.1210/jc.2015-2917
Siegel RL, Miller KD, Jemal A (2015) Cancer statistics, 2015. CA Cancer J Clin 65:5–29. doi:10.3322/caac.21254
Xing M (2005) BRAF mutation in thyroid cancer. Endocr Relat Cancer 12:245–262. doi:10.1677/erc.1.0978
Xing M (2010) Genetic alterations in the phosphatidylinositol-3 kinase/Akt pathway in thyroid cancer. Thyroid Off J Am Thyroid Assoc 20:697–706. doi:10.1089/thy.2010.1646
Xing M (2013) Molecular pathogenesis and mechanisms of thyroid cancer. Nat Rev Cancer 13:184–199. doi:10.1038/nrc3431
Acknowledgements
This work was supported by a Grant from the Major Science and Technology Projects of Zhejiang Province (2015C03052), Wenzhou Science and Technology Planning Project (Y20160126), and Scientific Research Incubator Project of The First Affiliated Hospital of Wenzhou Medical University (No. FHY2014018).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The author declare that they no conflicts of interest.
Ethical approval
Ethical approval for this study was obtained from the Ethnic Committee of the First Affiliated Hospital of Wenzhou Medical University.
Informed consent
Written informed consent was obtained from each individual participant.
Rights and permissions
About this article
Cite this article
Wang, O., Zheng, Z., Wang, Q. et al. ZCCHC12, a novel oncogene in papillary thyroid cancer. J Cancer Res Clin Oncol 143, 1679–1686 (2017). https://doi.org/10.1007/s00432-017-2414-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-017-2414-6