Abstract
Background
MALAT1 was discovered as a prognostic marker for lung cancer metastasis and has been found upregulated in many types of tumor, but its transcriptional regulation mechanism in tumors remains unclear.
Methods
A deletion analysis of MALAT1 promoter region was performed to find the cis elements that were critical for the transcriptional activation of MALAT1 gene. Reporter gene assays were employed to analyze the effect of Sp1 on the promoter activity of MALAT1 gene. The binding activity of Sp1 with the promoter of MALAT1 gene was examined by EMSA and ChIP assay. Effects of Sp1 on regulation of MALAT1 were analyzed by RNA interference in vitro and in vivo mouse model.
Results
By means of luciferase assay, Sp1 was found to activate the promoter of the human MALAT1 gene. The binding of Sp1 to this region was also detected by electrophoretic mobility shift and chromatin immunoprecipitation assays. Sp1 knockdown also decreased the MALAT1 and inhibited A549 lung cancer cells’ growth and invasion in vitro. Furthermore, knockdown of Sp1 also mimicked the inhibition of MALAT1 in A549 lung cancer cells’ growth and metastasis in vivo.
Conclusions
Taken together, our data suggest that upregulation of MALAT1 was mediated by the transcription factor Sp1 in A549 lung cancer cells, and Sp1 could be therapeutic target for cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) was originally identified in 2003 as a prognostic parameter for patient survival of stage I lung adenocarcinoma or squamous cell carcinoma patients (Ji et al. 2003). A later study identified MALAT1 as a noncoding transcript enriched in the nucleus of human primary fibroblasts or transformed lymphoblasts (Hutchinson et al. 2007). MALAT1 was one of the first ncRNAs that was associated with lung cancer. Recent work established a critical regulatory function of this lnc RNA in lung cancer metastasis and cell migration (Gutschner et al. 2013a, b; Tano et al. 2010).
Since its discovery in non-small cell lung cancer (NSCLC), overexpressed MALAT1 was found in many solid tumors such as lung cancer, cervical cancer, and HCC (Schmidt et al. 2011; Guo et al. 2010; Li et al. 2014). The association between MALAT1 expression and tumor metastasis prompted us to investigate the exact mechanism of MALAT1 upregulation. It was reported that the posterior pituitary hormone oxytocin could increase the level of MALAT1 in human SK-N-SH neuroblastoma cells; such responsiveness of MALAT1 was regulated by the binding of cyclic AMP-responsive element-binding protein (CREB) to the promoter (Koshimizu et al. 2010). It was also reported that MALAT1 transcription was regulated by Wnt/β-catenin signaling in endometrioid endometrial carcinoma (EEC) (Zhao et al. 2014).
Specificity protein 1 (Sp1) is a transcription factor that is ubiquitously expressed in various cells and tissues. Sp1 recognizes GC-rich regions and binds to DNA through three C2H2-type zinc fingers in the C-terminal domain (Li and Davie 2010). The relative expression of Sp1 in cancer cells has been shown to be higher than that of adjacent normal cells in several tumors, including gastric tumors, breast cancers, and lung cancers (Safe and Abdelrahim 2005; Deacon et al. 2012; Wang et al. 2003; Zannetti et al. 2000). It was also reported that 65 % of lung cancer patients have been shown to have a higher level of Sp1 in tumor tissues (Lin et al. 2010). There are growing evidences showing that Sp1 protein plays a critical role in the growth and metastasis of many tumor types through regulating gene transcription related to growth and proliferation (Black et al. 2001; Wang et al. 2014; Kong et al. 2014). Moreover, Sp1 protein is also very likely to be one of the potential targets for cancer chemotherapy (Jiang et al. 2014; Safe et al. 2014).
In the current study, we found a Sp1-binding site in the promoter region of the MALAT1 gene. Both silencing the Sp1 gene and mutation of the binding site decreased the transcription of MALAT1. EMSA and ChIP assay confirmed the association between the Sp1 protein and MALAT1 promoter. To the best of our knowledge, this is the first evidence to associate the Sp1 protein with the high level of MALAT1 expression in tumor.
Materials and methods
Plasmids and reagent
Plasmid pCMV-Sp1 was purchased from Addgene (Cambridge, MA, USA). Mithramycin A was from Sigma (St. Louis, MI, USA). Plasmid pCMV-Flag-Sp1 was constructed by inserted Sp1 cDNA fragment into pCMV-N-Flag (Beyotime, China).
Cell lines and transfection
Human lung cancer cell lines (A549), human cervical cancer cell line (HeLa), human liver hepatocellular carcinoma cell line (HepG2), normal cells human lung fibroblast cell line (MRC5), and human mammary epithelial cells (MCF10A) were purchased from ATCC. Human liver cells (HL-7702) were bought from the Shanghai Institute for Biological Sciences, Chinese Academy of Sciences. Cell transfection was performed with Lipofectamine transfection reagent (Invitrogen, Waltham, MA, USA). Unless otherwise indicated, cells were harvested at 48 h after transfection.
Generation of MALAT1 promoter-luciferase constructs
A 445-bp fragment (nucleotides from −432 to +13) of human MALAT1 promoter was prepared by PCR amplification of human genomic DNA of MRC5 cells using a sense primer containing Kpn1 restriction site and an antisense primer containing a BglII restriction site. Primers were synthesized on the basis of the reported genomic sequence for human MALAT1, forward 5′-CGGGG TACCT GGGAA AGGAA GACCT AGACT GA-3′ and reverse 5′-GAAGA TCTCC CAGTC CTTTA CAGAA GTCTC G-3′. Following digestion with restriction enzymes, the MALAT1 promoter fragment was directionally cloned into the pGL4.17-basic firefly luciferase expression vector (Promega) to generate a “full-length” MALAT1 reporter construct. Reporter genes containing sequentially truncated fragments (−319/+13, −288/+13, −247/+13, −199/+13) of the MALAT1 promoter region were prepared in a similar manner using different sense primers containing Kpn1 restriction sites and the same antisense primer that was used for the full-length MALAT1 reporter construction. To prepare mutated promoters, the putative site #1 Sp1-binding sequence between nucleotide position −268 to −266 GCG was mutated to TTT and named p-288m/+13; the putative site #2 Sp1-binding site between nucleotide position −307 to −305 GCC was mutated to AAA and named P-319m/+13. The mutation was created from wild-type MALAT1 promoter plasmid p-288/+13 or p-319/+13 by PCR using Mut Express™ II Fast Mutagenesis Kit (Vazyme, China). All of the constructs were verified by sequencing.
Dual-luciferase assay for promoter activity
Dual-luciferase assay was performed as described previously with some modifications (Li et al. 2013). In brief, cells were plated on 24-well plates, cultured overnight, and transfected using Lipofectamine transfection agent (Invitrogen, Waltham, MA, USA) according to the manufacturer’s protocol. For normalization of luciferase activity, the pRL-TK control vector encoding Renilla luciferase was used for co-transfection together with pGL4.17 plasmids. In some experiments, the pGL4.51-control vector was used in transfection as a positive control of promoter activity. This pGL4.51-control vector contains CMV promoter resulting in strong expression of luciferase gene in many types of mammalian cells. For all experiments, cells were cultured for 48 h after transfection and lysed with the passive lysis buffer (Promega). Lysates were analyzed using Dual-Luciferase Reporter Assay System kit (Promega). Luminescence was measured on luminometer (Turner Biosystems Instrument, Sunnyvale, CA, USA). All experiments were performed at least three times.
Cell migration assay
Wound-healing assay was performed as previously described (Li et al. 2013). Briefly, the cell layer that reached confluence was scratched by a 200 μl pipette tip and cultured at 37 °C. The average extent of wound closure was quantified. The percent of wounded area filled through cell migration was then calculated as follows: [(mean wound width − mean remaining width/mean wound width] × 100 (%). Transwell assay was performed as previously described (Li et al. 2013). Cells were counted on four random fields per well.
Western blot
Samples were separated by SDS-PAGE then transferred to membrane (Millipore). Membranes were blocked in 5 % powdered milk in Tris-buffered saline containing 0.05 % Tween 20 for 1 h at room temperature and probed with the anti-Sp1 (ab13370) or anti-GAPDH (ab37168) primary antibody at 4 °C overnight. After washing three times, the membrane was then incubated with horseradish peroxidase (HRP)-labeled secondary antibody for 2 h at room temperature. The membrane was then developed by using the enhanced chemiluminescent (ECL) detection systems.
Electrophoretic mobility shift assay (EMSA)
Nuclear extracts were prepared from cells, following the previously described procedures (Ishida et al. 2002). Synthetic complementary oligonucleotides containing the Sp1-binding site of MALAT1 promoter were 5′-biotinylated and annealed. The wild-type sequences of the oligonucleotides used are 5′-CAGGC GTTAG GGCGG GGCGC GCGTG C-3′, the mutant sequences are 5′-GGCAC AGGCG TTAΔ6GG CGCGC GTGCG C-3′. DNA–protein-binding assays were carried out and detected using horseradish peroxidase-conjugated streptavidin (LightShift™ chemiluminescent EMSA kit) according to the manufacturer’s instructions.
Chromatin immunoprecipitation assay
ChIP assays were performed by following the chromatin immunoprecipitation (ChIP) assay kit protocol (Millipore catalo# 17-295) (Parakati and DiMario 2013). Approximately 5 × 106 cells were cross-linked with 1 % formaldehyde and collected in lysis buffer. DNA was then sheared to an approximate length of 200–1000 bp. Sheared DNA (50 μl) was incubated with 5 μg of anti-Sp1 ChIP-grade antibody or normal rabbit IgG followed by immunoprecipitation with 20 μl of protein A agarose beads during an overnight incubation at 4 °C with rotation. Enriched DNA was extracted from the DNA/antibody/protein A bead complexes by proteinase K digestion and purified with spin columns. Finally, precipitated MALAT1 DNA was amplified using the following forward and reverse primers: (5′-GGAAG TTGGG CAGCA GCTCC ACG-3′) and (5′-CCACT GGTTC TAACC GGCTC TAG-3′). Unsheared DNA was used as an input control to analyze the immunoprecipitation results. For mithramycin A treatment, the cells were incubated for 12 h in culture medium containing 0.2 μM mithramycin A before cross-linking with 1 % formaldehyde and collection for chromatin immunoprecipitation.
For chromatin immunoprecipitation experiments using wild-type and mutated MALAT1 promoter DNA constructs, 8 μg of pCMV-Flag-Sp1 and 4 μg of wild-type p-288/+13-luc or mutated p-288m/+13-luc were transfected into A549 cells. Cell culture and chromatin immunoprecipitation from A549 cells was carried out as described above. Precipitated DNA was amplified for 30 cycles using the MALAT1 promoter-specific forward primer (GAACA GGCAC AGGCG TTAGG) and a luciferase gene-specific reverse primer (AATGT TTTTG GCATC TTCCA). DNA was resolved in a 1 % agarose gel.
Animals
Athymic nude mice (6–8 weeks of age) were obtained from Shanghai Laboratory Animal Center (Shanghai, China) and housed under germfree conditions. Animal welfare and experimental procedures were performed strictly in accordance with high standard animal welfare and other related ethical regulations approved by Southeast University.
Subcutaneous tumor model
A549 cells (5 × 105 cells in 20 μl) were injected subcutaneously into the dorsal flanks of mice. Tumor volume was monitored by measuring the two maximum perpendicular tumor diameters with calipers every 3 days and calculated using the formula: length × width2 × 0.52. When tumors reached a size of approximately 5 × 5 mm, the mice were arbitrarily assigned to different groups. 10 μg RNAi expression plasmids complexed with in vivo-jetPEI™ (Polyplus-Transfection Inc., New York, NY, USA) were injected into tumor for each animal per injection, and the injection were repeated every 3 days for a total of four times. To evaluate Sp1 and MALAT1 expression level after four times RNAi expression plasmids injections, mice were killed on the 12th day after the first treatment (3 days after the final treatment), and tumors were isolated and RT-PCR and Western blot analysis were performed, whereas for tumor growth assay, the growth of solid tumors were continued monitored by measuring tumor size every 3 days for a 31-day period.
Experimental lung metastases model
Experimental lung metastases model was performed as described previously with some modifications (Lo et al. 2011). Briefly, A549 cells transfected with RNAi expression plasmids or the control vector were harvested from six-well plates, washed with phosphate-buffered saline (PBS), and resuspended at 2 × 107 cells/ml. Suspended cells (0.1 ml) were injected into the tail veins of mice. Seven weeks after the injection, animals were euthanized and the lungs were removed and photographed with a digital camera. Visible tumors on the lung surface were counted. Lungs were fixed and stained with hematoxylin and eosin (H&E) for further pathological confirmation.
RNAi experiments
Pre-miRNA expression cassettes were designed to target the Sp1 gene or MALAT1 by using the BLOCK-iT Pol II miR RNAi Expression vector kits and manufacturer’s instructions (Invitrogen, Waltham, MA, USA) (Fu et al. 2009). For Sp1 knockdown, a double-stranded oligonucleotide predicted to form a microRNA (miRNA) was synthesized by using the following primer pairs: forward, 5-TGCTG GGCAT AGCAG CAATG ATGTT GGTTT TGGCC ACTGA CTGAC CAACA TCAGC TGCTA TGCC-3; reverse, 5-CCTGG GCATA GCAGC TGATG TTGGT CAGTC AGTGG CCAAA ACCAA CATCA TTGCT GCTAT GCCC-3), The primers used For MALAT1 knockdown: forward, 5-TGCTG GCGTC ATGGA TTTCA AGGTC TGTTT TGGCC ACTGA CTGAC AGACC TTGAT CCATG ACGC-3, reverse, 5-CCTGG CGTCA TGGAT CAAGG TCTGT CAGTC AGTGG CCAAA ACAGA CCTTG AAATC CATGA CGCC-3. Double-stranded oligos were then cloned into pcDNA6.2-GW/EmGFPmiR expression vectors (Invitrogen, Waltham, MA, USA). BLOCK-iT™ Pol II miR-luc validated miRNA control vector (Invitrogen, Waltham, MA, USA) was used as RNAi control. These plasmids were transfected into A549 cells using Lipofectamine 2000 (Invitrogen, Waltham, MA, USA). The cells were processed for immunoblot analyses 48 h after transfection.
Cell proliferation assay
The effects of Sp1- or MALAT1-specific miRNA on A549 cell proliferation were assessed by the MTT assay. Briefly, A549 cells in the exponential growth phase were seeded at a final density of 6 × 104 cells/ml into a 96-well plate. After 24 h posttransfection, the cell viability was determined by the colorimetric MTT [3-(4, 5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide] assay at wavelength 490 nm by TECAN Safire Fluorescence Absorbance and Luminescence Reader (Vienna, VA, USA). The cell viability was calculated according to the formula: \({\text{Cell}}\;{\text{viability }}\left( \% \right) = {\text{average A49}}0 \;{\text{nm of treated group}}/{\text{average A49}}0\; {\text{nm of control group}} \times 100 \;\% .\) All assays were performed in triplicate and independently repeated twice.
Statistical methods
Data are presented as mean ± SD of triplicate experiments. The statistical analysis involving two groups was performed by means of Student’s t test. P < 0.05 was considered to be significant.
Results
Identification of the MALAT1 gene promoter
The longest MALAT1 transcript from human is 8708 nt in length [GenBank: NR_002819.2], and the transcriptional start site (TSS) was determined by 5′-RACE (Hutchinson et al. 2007). In order to identify the region crucial for MALAT1 promoter activity, a reporter gene assay was performed using pGL4.17-basic vector containing various lengths of the MALAT1 promoter region (Fig. 1a, b). The construct p-319/+13-luc and p-288/+13-luc exhibited the similar promoter activity (Fig. 1c). The promoter activities in further deleted segments were decreased dramatically. These results indicated that the sequence between −319 and +247 is critical for basal MALAT1 gene transcription.
Identification of the region(s) required for the transcriptional regulation of MALAT1 gene. a Nucleotide sequence of the promoter region of MALAT1 gene. The predicted binding sites of Sp1 are underlined. +1 indicates the position of the transcription initiation site of MALAT1 gene. b The schematic diagram of the luciferase reporter constructs containing the indicated genomic fragments of MALAT1 gene. The positions relative to the transcriptional initiation site of MALAT1 gene (+1) are indicated. c The results of the luciferase reporter assays. A549, HeLa, or HepG2 cells were transiently co-transfected in triplicate in 24-well plates with the indicated luciferase reporter constructs together with the Renilla luciferase reporter plasmid (pRL-TK). Forty-eight hours after transfection, firefly and Renilla luciferase activities were measured by dual-luciferase assay system. Data obtained from a representative of at least three independent experiments are shown as fold induction compared to the activity of cells transfected with the empty pGL4.17-basic luciferase reporter vector. The results are presented as mean and SD of triplicates from a representative experiment. Asterisk represents statistically significant difference with P < 0.01 between pGL4.17-basic vector and MALAT1 promoter construct
The proximal Sp1-binding site plays an important role in the regulation of MALAT1 transcription.
Through computational analysis, two binding sites for Sp1 were predicted in this region (Fig. 1a, b). To determine whether Sp1 could contribute to the transcriptional regulation of MALAT1 gene, mutations were introduced into either Sp1 site in the p-319/+13-luc and p-288/+13-luc luciferase reporter construct by site-directed mutagenesis (Fig. 2a, left panel). A549, HeLa, or HepG2 cells were transiently co-transfected with the indicated luciferase reporter constructs along with pRL-TK. Forty-eight hours after transfection, cells were lysed and their luciferase activities were examined. As shown in the right panel of Fig. 2a, mutation of the distal Sp1-binding site had a negligible effect on the promoter activity of MALAT1 gene, and disruption of the proximal Sp1-binding site resulted in a significant reduction in the promoter activity of MALAT1 gene. Consistent with these results, overexpression of Sp1 failed to stimulate the MALAT1 promoter activity devoid of the proximal Sp1-binding site, but Sp1 can still activate the MALAT1 promoter activity devoid of the distal Sp1-binding site (Fig. 2b, c). These results strongly suggest that Sp1 plays an important role in the transcriptional regulation of MALAT1 gene through the proximal Sp1-binding site.
The proximal Sp1-binding site plays an important role in the regulation of MALAT1 transcription. a MALAT1 promoter activity was decreased in the absence of proximal Sp1-binding site. Site-directed mutations were introduced into the parental p-319/+13-luc and p-288/+13-luc luciferase reporter constructs to disrupt the indicated Sp1-binding sites (left panel). Right panel shows the results of luciferase reporter assay. A549, HeLa, or HepG2 cells were transiently co-transfected with the indicated luciferase reporter constructs together with pRL-TK. Forty-eight hours after transfection, luciferase activities were measured as described in Fig. 1c. Asterisk represents statistically significant difference with P < 0.05 between p-288/+13-luc MALAT1 promoter construct WT and the construct of p-288m/+13-luc with mutant proximal SP1 sites. Ns represents no significant difference between p-319/+13-luc and p-319m/+13-luc. b, c Overexpression of Sp1-activated MALAT1 promoter activity through proximal Sp1-binding site. Stimulatory effect of Sp1 overexpression on MALAT1 promoter activation was prevented in the absence of proximal Sp1-binding site. Human A549, HeLa, and HepG2 cells were co-transfected with 1 μg Sp1 expressing plasmid, or empty vector along with 1 μg wild-type p-319/+13-luc or mutated p-319m/+13-luc (b) or 1 μg wild-type p-288/+13-luc or mutated p-288m/+13-luc (c), and Renilla expressing plasmid. Luciferase activity of cells co-transfected with empty vector and wild-type p-319/+13-luc (b) or p-288/+13-luc (c) was arbitrarily set at 1. Asterisk represents statistically significant difference with P < 0.01 between MALAT1 promoter construct (p-319/+13-luc, p-319m/+13-luc or p-288/+13-luc) with and without plasmid expression of SP1. Ns represents no significant difference between p-288m/+13-luc with and without plasmid expression of SP1
The direct binding activity of Sp1 was further confirmed by EMSA and ChIP assay
To determine whether Sp1 binds to the MALAT1 promoter region via the proximal Sp1-binding site, EMSA was performed. As shown in Fig. 3a, DNA–protein complex was detected when the nuclear extracts of A549 cells were incubated with the double-stranded oligonucleotide probe containing the proximal Sp1-binding site. This DNA–protein complex was prevented in competition experiments, and the specificity was confirmed by super-shift assay with Sp1 antibody.
Binding of Sp1 transcription factor on MALAT1 promoter. a EMSA analysis. The −279 to −254 ntDNA probe bound to the nuclear extracts of A549 cells, and this binding was blocked by excess unlabeled consensus probe. Sp1 antibody supershifted the binding band. b ChIP assay was used to confirm the Sp1-binding activity. Formaldehyde-cross-linked chromatin was isolated from A549 cells, sonicated, and immunoprecipitated with anti-Sp1. b-1 Chromatin was prepared from nontransfected A549 cells and immunoprecipitated using the Sp1 antibody. Endogenous MALAT1 DNA was amplified using MALAT1 promoter-specific forward and reverse primers. Lane 1, input chromatin prior to immunoprecipitation. Lane 2, immunoprecipitation with normal rabbit IgG. Lane 3, immunoprecipitation with the Sp1 antibody. Lane 4–6, the group of Sp1 inhibitor mithramycin A treatment. Lane 7, amplification of genomic DNA (gDNA) containing the MALAT1 promoter sequence. Lane 8, immunoprecipitation without antibody (no Ab). b-2 Chromatin was prepared from A549 transfected with pCMV-Flag-Sp1, which expresses Sp1 coupled to the FLAG epitope tag. A549 were also transfected with the wild-type MALAT1 promoter coupled to the luciferase reporter gene (p-288/+13-luc) or the MALAT1 promoter-luciferase construct containing the mutation of the Sp-binding site (p-288m/+13-luc). Chromatin from transfected cells was immunoprecipitated with the FLAG epitope tag antibody. DNA containing either the wild-type or the mutated MALAT1-luciferase sequence was amplified using the MALAT1 promoter-specific forward primer and a luciferase gene-specific reverse primer. Lane 1, input chromatin prior to immunoprecipitation. Lane 2, immunoprecipitation with normal rabbit IgG. Lane 3, immunoprecipitation without antibody. Lane 4, immunoprecipitation with the FLAG antibody. Lane 5, amplification of plasmid DNA containing either the wild-type or the mutated MALAT1 promoter sequence
To examine whether Sp1 could bind to the MALAT1 promoter region in cells, we performed chromatin immunoprecipitation (ChIP) assays. Cross-linked chromatin was prepared from A549 cells and immunoprecipitated with control IgG or anti-Sp1; the immunoprecipitated genomic DNA was purified and amplified by PCR using the primers specific for the MALAT1. As shown in Fig. 3b, ChIP assays demonstrated that Sp1 is recruited onto the MALAT1 promoter. We also examined the possible effect of Sp1 inhibitor mithramycin A (MA) on the transcription of MALAT1 gene. MA is a cell-permeable agent that binds to GC-rich DNA sequence thereby blocking the binding of transcription factors such as Sp1- to GC-specific regions of DNA. A549 cells were treated with MA, and the ChIP assay result showed that MA treatment decreased the binding of Sp1 to MALAT1 promoter region significantly. To further determine whether Sp1 interacted specifically with the proximal Sp-binding site as suggested by the electromobility shift assays (Fig. 3a) and endogenous chromatin immunoprecipitation (Fig. 3b-1), pCMV-Flag-Sp1 and wild type p-288/+13-luc or mutated p-288m+13-luc were transfected into A549 cells. Similar to the procedure of endogenous MALAT1 chromatin immunoprecipitation, exogenous MALAT1 chromatin was immunoprecipitated by the FLAG antibody. PCR was performed using the MALAT1 promoter specific forward primer and a luciferase gene specific reverse primer to distinguish amplification of transfected DNA versus endogenous MALAT1 promoter sequence (Fig. 3b-2). The result showed that exogenous wild type MALAT1 promoter chromatin was immunoprecipitated by the FLAG antibody and amplified. However, no PCR product of MALAT1 promoter with the mutated Sp1 binding site was detected. These results indicate that Sp1 interacts with the proximal Sp-binding site within the MALAT1 promoter. Taken together, these data demonstrates that Sp1 directly binds to the promoter of MALAT1.
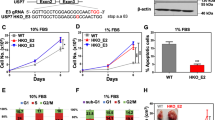
Knockdown of Sp1 reduces the MALAT1 expression and associated metastatic capacity of A549 lung cancer cells
A previous study has shown that A549 cells express higher Sp1 protein levels as compared to normal bronchial epithelial cells (Chen et al. 2011). This was also confirmed by our experiments. As shown in Fig. 4a-1, A549 cells have higher Sp1 expression level than MRC5 cells, a normal lung cell line. Meanwhile, MALAT1 expression was also increased in metastasis A549 cells than that in MRC5 (Fig. 4a-2). Since Sp1 plays a critical role in MALAT1 transcription, both Sp1 RNAi and MALAT1 RNAi resulted in a significant decrease of MALAT1 mRNA level in A549 cells, as determined by RT-PCR analysis (Fig. 4b). To evaluate whether Sp1 is responsible for A549 cell metastasis phenotype, wound-healing experiments were performed. The results are shown in Fig. 4c, and RNAi-mediated downregulation of Sp1 or MALAT1 in A549 cells resulted in fewer A549 cells to migrate into wound area as compared with cells transfected with control RNAi plasmid. Similarly, knockdown of Sp1 and knockdown of MALAT1 in A549 cells also resulted in significant reduction in migration of A549 cells in transwell migration experiments (Fig. 4d).
Sp1-mediated transcriptional regulation of MALAT1 plays a critical role in A549 cells’ migration and proliferation. a MALAT1 expression level correlated positively with Sp1 protein level in human lung adenocarcinoma cell line A549 and human lung normal cell line MRC5. a-1, a-2 Elevated expression of Sp1 protein and MALAT1 expression in A549 compared with MRC5. Sp1 protein of each cell line was determined by Western blotting using the anti-Sp1 antibody (a-1). A-2 MALAT1 expression was analyzed by RT-PCR. GAPDH was used as an internal control. b Sp1-specific miRNA (Sp1 RNAi) reduced the endogenous Sp1 mRNA and protein levels in A549 cells (b-1, b-2). Knockdown of Sp1 by Sp1-specific miRNA or Knockdown of MALAT1 by MALAT1-specific miRNA in A549 cells led to a reduction in MALAT1 mRNA (b-3, b-4). Cells were transfected with Sp1 or MALAT1 miRNA expression plasmids. After 48 h, total RNA was collected for assessment of transcript levels by RT-PCR. Western blot analysis was performed to determine the Sp1 protein levels. Transfection with miR-luc control plasmid was performed as control RNAi. GAPDH was used as an internal control. c Wound-healing assay was used to detect A549 cell motility changes after silencing MALAT1 by Sp1 RNAi and MALAT1 RNAi. The A549 cells were transfected with Sp1 or MALAT1 miR RNAi expression plasmid or control RNAi plasmid, then wound-healing scratch motility assays were performed in the fibronectin-coated plates in the presence of serum. Cell migration was assessed at 0 and 48 h. Representative images are shown (c-1). c-2 Statistical analysis of wound closure. Gap size at 0 h was set to 100 % and percentage of closed wound was calculated after 48 h after image analysis (n = 3; ***P < 0.01). d Transwell assays demonstrated knockdown of Sp1-suppressed A549 cell invasiveness. After A549 cells were transfected with control or MALAT1 miRNA or Sp1 miRNA expression plasmid for 24 h, cells were collected and resuspended in culture medium at a density of 1 × 106 cells/ml. Hundred microliters of the cell suspension was plated into the upper wells of transwell inserts containing 8-μm pore polycarbonate membranes pre-coated with fibronectin (10 μg/ml) on the under surface. The cells were allowed to migrate for 20 h at 37 °C, then cells on upper wells were removed gently and those migrated to the under surface were fixed and stained. Representative membranes stained with Giemsa are shown (d-1), the arrows indicated the migrant cells. Scale bar 100 μm. Quantitative analysis of the number of the cells migrated to the down side of the membrane (d-2). Data are mean ± SD of three independent experiments. ***P < 0.01 versus control. e Proliferation assay showed reduced proliferation in Sp1 or MALAT1-downregulated A549 cells. A549 cell viability after transfection with Sp1- or MALAT1-specific miRNA expression plasmids or control plasmids 24 h was measured by MTT assay. ***P < 0.01 versus control
Effect of downregulated MALAT1 expression on the proliferation of A549 lung cancer cells
To investigate the possible function of MALAT1 on the growth of A549 cells, MTT assay was adopted to further evaluate the anti proliferation effect of Sp1 or MALAT1 RNAi. As shown in Fig. 4e, Sp1- or MALAT1-specific miRNA significantly suppressed A549 cell proliferation.
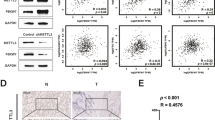
The correlation between the Sp1 and MALAT1 in liver and breast cancer cell lines
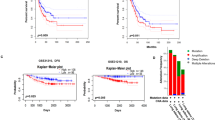
The expression of MALAT1 correlates with tumor development, progression, or survival in lung, liver, and breast cancer. Overexpression of MALAT1 was found in these solid tumors (Schmidt et al. 2011; Gutschner et al. 2013a; Li et al. 2014). So we further tested whether the Sp1 was also correlated with the MALAT1 level in liver and breast cancer cells. As shown in Fig. 5, Sp1 protein level was highly expressed in breast cancer cells MCF-7 and liver cancer cells HepG2; MALAT1 level was also increased in these tumor cells compared with the normal cell.
MALAT1 expression level correlated positively with Sp1 protein level in human breast cancer cell MCF-7 and liver cancer cell HepG2. Elevated expression of Sp1 protein and MALAT1 expression in MCF-7 and HepG2 compared with normal cells MCF10A and HL-7702. Sp1 protein of each cell line was determined by Western blotting using the anti-Sp1 antibody (a). b MALAT1 expression was analyzed by RT-PCR. GAPDH was used as an internal control
Downregulation of MALAT1 or Sp1 expression by RNAi reduces A549 lung cancer cell growth in vivo
It was reported that MALAT1 promotes tumor growth in vivo (Schmidt et al. 2011). Next, we postulated that delivery of miRNA expression plasmid against Sp1 or MALAT1 into tumor in vivo will delay the tumor growth similarly. As expected, when jetPEI-complexed Sp1 or MALAT1 miRNA expression plasmids were injected into tumors of nude mice, the solid tumor growth of groups of MALAT1 RNAi as well as Sp1 RNAi was greatly suppressed as compared with other control groups (Fig. 6a). Thirty-one days after the first treatment, animals were killed. It was reported that this mouse model was used to study tumor growth, and it is not an established metastasis model (Schmidt et al. 2011), and we still discovered one tumor metastasis in a right axillary lymph node of one mouse of PBS-treated group. Besides, no other metastasis was found on examination of the other lymph nodes or lungs. This result was consistent with that previously reported (Schmidt et al. 2011).
Knockdown of Sp1 or MALAT1-suppressed tumor growth in nude mice subcutaneous tumor model. a A549 cells were injected subcutaneously into the dorsal flanks of athymic nude mice. When tumors reached a size of approximately 5 × 5 mm, mice were intratumorally injected with in vivo-jetPEI™-complexed Sp1- or MALAT1-specific miRNA expression plasmids. Control groups were intratumorally injected with miR-luc control plasmid or PBS. Effect of intratumoral injection of miRNA expression plasmids on the primary tumor growth was shown. Values were expressed as the mean ± SD from eight animals. *P < 0.05 as compared to the control groups. b Expression of Sp1 and MALAT1 in tumors after RNAi treatment. After four times intratumoral injection of Sp1- or MALAT1-specific miRNA expression plasmids, tumors were isolated on day 12 (3 days after the fourth treatment) and subjected to RT-PCR (b-1) and Western blotting (b-2), PCR primers were specific for human Sp1, MALAT1, and GAPDH (internal control)
To examine whether the Sp1-specific miRNA treatment for tumor could decrease the MALAT1 expression in vivo, after four times treatment, tumor were taken from mice and analyzed with RT-PCR and Western blotting. The result revealed that both groups receiving either Sp1- or MALAT1-specific miRNA expression plasmids treatment showed downregulated expressions of MALAT1 (Fig. 6b).
Knockdown MALAT1 or Sp1 by RNAi suppress A549 cell metastasis in vivo
To test the tumor suppressor activity of MALAT1 or Sp1 RNAi on experimental lung metastases, A549 cells transfected with MALAT1 or Sp1 miRNA expression plasmids were injected into nude mice. After 7 weeks, animals were killed for lung tissue examination. Metastatic nodules on the surface of lungs were counted. MALAT1 or Sp1 knockdown group had significantly fewer tumor nodules in the lung compared to control groups (Fig. 7a, b). This difference was further confirmed through H&E staining of lung sections (Fig. 7c).
Knockdown of Sp1 or MALAT1 suppressed tumor metastasis in nude mice tumor model. a Experimental lung metastasis was established in nude mice by injecting Sp1 or MALAT1 knockdown A549 cells into nude mice. Seven weeks after tail vein injection, formation of lung tumor nodules was analyzed. The lungs were resected and fixed. MALAT1 or Sp1 RNAi resulted in a reduction in the number of metastatic nodules compared with those in the control group. Representative pictures with the tumor nodules on lung surfaces were shown. b Statistical analysis. Values were expressed as the mean ± SD from eight animals. ***P < 0.01 as compared to the control groups. c H&E-stained lung sections showed micrometastases of MALAT1 or Sp1 knockdown tumor cells, whereas extensive metastases were found in control groups
Discussion
With the innovations in RNA-seq technologies, large numbers of lnc RNAs in mammals are being identified and characterized (Nielsen et al. 2014). Although the function of most lncRNAs remains unknown, many of them have been suggested to play crucial roles in various biological processes ranging from epigenetic gene regulation, transcriptional control, to posttranscriptional regulation during normal development and diseases, including cancers (Batista and Chang 2013; Wapinski and Chang 2011). This kind of ncRNAs is similar to mRNAs in gene structure and is associated with influencing the recruitment of chromatin modifier factors to specific chromatin sites and providing architectural support to the hierarchical subnuclear organization and keeping proteins tethered to specific cellular compartments (Yang et al. 2014; Singh and Prasanth 2013). MALAT1 is one of the first found cancer-associated lncRNAs, which is also referred to as nuclear-enriched abundant transcript 2 (NEAT2) (Ji et al. 2003). Dysregulation of this evolutionarily conserved long noncoding RNA has been linked to many human cancer types. However, the molecular mechanism of MALAT1 action is currently under debate: One recent study proposed a role for MALAT1 in cell cycle progression through its involvement in regulating E2F1 activity (Yang et al. 2011b), another report suggested that MALAT1 positively regulates the expression of the oncogenic transcription factor BMYB (Tripathi et al. 2013), whereas others provide evidence that MALAT1 regulates alternative splicing by controlling the activity of the SR protein family of splicing factors (Tripathi et al. 2010). In addition, the molecular mechanism of the upregulation of MALAT1 in cancer was not clear yet. To investigate the possible transcriptional regulation for MALAT1, we cloned the promoter of MALAT1 and performed dual-luciferase assay. We found two potential Sp1-binding sites in the core promoter of MALAT1, and further mutation analysis excluded the role of distal binding site. The proximal Sp1-binding site was confirmed by the reporter assays and EMSA analysis (Figs. 2, 3). ChIP assay further confirmed the binding activity of Sp1 in vivo (Fig. 3). We also constructed the miR RNAi expression vector to knockdown Sp1 or MALAT1. RNAi-mediated knockdown of endogenous Sp1 resulted in a significant downregulation of MALAT1 gene and decreased the migration and proliferation of A549 cells (Fig. 4). Finally, knockdown of Sp1 gene also mimicked the A549 growth inhibition and invasion inhibition as that induced by MALAT1 RNAi in nude mice model (Figs. 6, 7).
Sp1 is a well-known member of the Krüppel-like factor (KLF) family of transcriptional regulators. The Sp/KLF transcription factor family comprises more than 20 members that play common or distinctive physiological roles in transcription. These proteins contain highly conserved C2H2 zinc finger motifs in their carboxyl terminal and bind to GC-rich sites (Li and Davie 2010; Black et al. 2001). For example, Sp1 activates FGFR1 promoter in proliferating myoblasts,while KLF10 repress FGFR1 transcription by binding to the same Sp site in the FGFR1 promoter (Parakati and DiMario 2013). Other reports demonstrated that Sp1 and Sp3 selectively bind to different Sp sites on the same promoter (Li and Davie 2010). Indeed, regulation of transcription from “Sp1 sites” can be highly complex. Differential binding of Sp/KLF factors to different Sp1 sites and differences in the activity or expression of individual family members may contribute to the gene transcription fine-tuned. Our preliminary experiments showed that Sp1 can regulate lncRNA MALAT1. It was also reported that in liver cancer cells, another lnc RNA lnc-HEIH was regulated by Sp1 (Yang et al. 2011a). As both Sp1 and MALAT1 are known to promote a metastasis phenotype in tumor cells, it is likely that MALAT1 and lnc-HEIH act as a downstream effector of Sp1 in accelerating tumor progression. Many types of tumors have a much higher level of Sp1 transcription factor than normal cells (Fig. 5), and this trend is associated with metastasis progress (Lou et al. 2005). Our findings are useful in understanding MALAT1 gene regulation in tumors. However, there is still much to learn that whether other potential Sp/KLF family members are also involved in regulating MALAT1 gene expression. For example, some cancer cells express not only Sp1 but also Sp3 and Sp4, and Sp3 and Sp4 are also associated with tumor growth and metastasis (Colon et al. 2011; Jutooru et al. 2014; Nair et al. 2013, 2014). Further investigations are also required to test the signaling pathways of Sp/KLF transcription factors that are involved in MALAT1 regulation in relation to cell growth and tumorigenesis.
Abbreviations
- MALAT1:
-
Metastasis-associated lung adenocarcinoma transcript 1
- EMSA:
-
Electrophoretic mobility shift assay
- ChIP:
-
Chromatin immunoprecipitation assay
- lnc RNA:
-
Long noncoding RNA
- ncRNA:
-
Noncoding RNA
References
Batista PJ, Chang HY (2013) Long noncoding RNAs: cellular address codes in development and disease. Cell 152:1298–1307
Black AR, Black JD, Azizkhan-Clifford J (2001) Sp1 and kruppellike factor family of transcription factors in cell growth regulation and cancer. J Cell Physiol 188:143–160
Chen Y, Wang X, Li W, Zhang H, Zhao C, Li Y, Wang Z, Chen C (2011) Sp1 upregulates survivin expression in adenocarcinoma of lung cell line A549. Anat Rec (Hoboken) 294:774–780
Colon J, Basha MR, Madero-Visbal R, Konduri S, Baker CH, Herrera LJ, Safe S, Sheikh-Hamad D, Abudayyeh A, Alvarado B, Abdelrahim M (2011) Tolfenamic acid decreases c-Met expression through Sp proteins degradation and inhibits lung cancer cells growth and tumor formation in orthotopic mice. Invest New Drugs 29:41–51
Deacon K, Onion D, Kumari R, Watson SA, Knox AJ (2012) Elevated SP-1 transcription factor expression and activity drives basal and hypoxia-induced vascular endothelial growth factor (VEGF) expression in non-small cell lung cancer. J Biol Chem 287:39967–39981
Fu T, Li P, Wang H, He Y, Luo D, Zhang A, Tong W, Zhang L, Liu B, Hu C (2009) c-Rel is a transcriptional repressor of EPHB2 in colorectal cancer. J Pathol 219:103–113
Guo F, Li Y, Liu Y, Wang J, Li Y, Li G (2010) Inhibition of metastasis-associated lung adenocarcinoma transcript 1 in CaSki human cervical cancer cells suppresses cell proliferation and invasion. Acta Biochim Biophys Sin (Shanghai) 42:224–229
Gutschner T, Hämmerle M, Diederichs S (2013a) MALAT1—a paradigm for long noncoding RNA function in cancer. J Mol Med 91:791–801
Gutschner T, Hämmerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zörnig M, MacLeod AR, Spector DL, Diederichs S (2013b) The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res 73:1180–1189
Hutchinson JN, Ensminger AW, Clemson CM, Lynch CR, Lawrence JB, Chess A (2007) A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8:39
Ishida A, Fujita N, Kitazawa R, Tsuruo T (2002) Transforming growth factor-β induces expression of receptor activator of NF-κ B ligand in vascular endothelial cells derived from bone. J Biol Chem 277:26217–26224
Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, Müller-Tidow C (2003) MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22:8031–8041
Jiang J, Lv X, Fan L, Huang G, Zhan Y, Wang M, Lu H (2014) MicroRNA-27b suppresses growth and invasion of NSCLC cells by targeting Sp1. Tumour Biol 35:10019–10023
Jutooru I, Guthrie AS, Chadalapaka G, Pathi S, Kim K, Burghardt R, Jin UH, Safe S (2014) Mechanism of action of phenethylisothiocyanate and other reactive oxygen species-inducing anticancer agents. Mol Cell Biol 34:2382–2395
Kong LM, Liao CG, Zhang Y, Xu J, Li Y, Huang W, Zhang Y, Bian H, Chen ZN (2014) A regulatory loop involving miR-22, Sp1, and c-Myc modulates CD147 expression in breast cancer invasion and metastasis. Cancer Res 74:3764–3778
Koshimizu TA, Fujiwara Y, Sakai N, Shibata K, Tsuchiya H (2010) Oxytocin stimulates expression of a noncoding RNA tumor marker in a human neuroblastoma cell line. Life Sci 86:455–460
Li L, Davie JR (2010) The role of Sp1 and Sp3 in normal and cancer cell biology. Ann Anat 192:275–283
Li S, Huang X, Zhang D, Huang Q, Pei G, Wang L, Jiang W, Hu Q, Tan R, Hua ZC (2013) Requirement of PEA3 for transcriptional activation of FAK gene in tumor metastasis. PLoS ONE 8:e79336
Li G, Zhang H, Wan X, Yang X, Zhu C, Wang A, He L, Miao R, Chen S, Zhao H (2014) Long noncoding RNA plays a key role in metastasis and prognosis of hepatocellular carcinoma. Biomed Res Int 2014:780521
Lin RK, Wu CY, Chang JW, Juan LJ, Hsu HS, Chen CY, Lu YY, Tang YA, Yang YC, Yang PC, Wang YC (2010) Dysregulation of p53/Sp1 control leads to DNA methyltransferase-1 overexpression in lung cancer. Cancer Res 70:5807–5817
Lo FY, Tan YH, Cheng HC, Salgia R, Wang YC (2011) An E3 ubiquitin ligase: c-Cbl: a new therapeutic target of lung cancer. Cancer 117:5344–5350
Lou Z, O’Reilly S, Liang H, Maher VM, Sleight SD, McCormick JJ (2005) Down-regulation of over-expressed Spl protein in human fibrosarcoma cell lines inhibits tumor formation. Cancer Res 65:1007–1017
Nair V, Pathi S, Jutooru I, Sreevalsan S, Basha R, Abdelrahim M, Samudio I, Safe S (2013) Metformin inhibits pancreatic cancer cell and tumor growth and downregulates Sp transcription factors. Carcinogenesis 34:2870–2879
Nair V, Sreevalsan S, Basha R, Abdelrahim M, Abudayyeh A, Rodrigues Hoffman A, Safe S (2014) Mechanism of metformin-dependent inhibition of mammalian target of rapamycin (mTOR) and Ras activity in pancreatic cancer: role of specificity protein (Sp) transcription factors. J Biol Chem 289:27692–27701
Nielsen MM, Tehler D, Vang S, Sudzina F, Hedegaard J, Nordentoft I, Orntoft TF, Lund AH, Pedersen JS (2014) Identification of expressed and conserved human noncoding RNAs. RNA 20:236–251
Parakati R, DiMario JX (2013) Repression of myoblast proliferation and fibroblast growth factor receptor 1 promoter activity by KLF10 protein. J Biol Chem 288(19):13876–13884
Safe S, Abdelrahim M (2005) Sp transcription factor family and its role in cancer. Eur J Cancer 41:2438–2448
Safe S, Imanirad P, Sreevalsan S, Nair V, Jutooru I (2014) Transcription factor Sp1, also known as specificity protein 1 as a therapeutic target. Expert Opin Ther Targets 18:759–769
Schmidt LH, Spieker T, Koschmieder S, Schäffers S, Humberg J, Jungen D, Bulk E, Hascher A, Wittmer D, Marra A, Hillejan L, Wiebe K, Berdel WE, Wiewrodt R, Muller-Tidow C (2011) The long noncoding MALAT-1 RNA indicates a poor prognosis in non-small cell lung cancer and induces migration and tumor growth. J Thorac Oncol 6:1984–1992
Singh DK, Prasanth KV (2013) Functional insights into the role of nuclear-retained long noncoding RNAs in gene expression control in mammalian cells. Chromosome Res 21:695–711
Tano K, Mizuno R, Okada T, Rakwal R, Shibato J, Masuo Y, Ijiri K, Akimitsu N (2010) MALAT-1 enhances cell motility of lung adenocarcinoma cells by influencing the expression of motility-related genes. FEBS Lett 584:4575–4580
Tripathi V, Ellis JD, Shen Z, Song DY, Pan Q, Watt AT, Freier SM, Bennett CF, Sharma A, Bubulya PA, Blencowe BJ, Prasanth SG, Prasanth KV (2010) The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 39:925–938
Tripathi V, Shen Z, Chakraborty A, Giri S, Freier SM, Wu X, Zhang Y, Gorospe M, Prasanth SG, Lal A, Prasanth KV (2013) Long noncoding RNA MALAT1 controls cell cycle progression by regulating the expression of oncogenic transcription factor B-MYB. PLoS Genet 9:e1003368
Wang L, Wei D, Huang S, Peng Z, Le X, Wu TT, Yao J, Ajani J, Xie K (2003) Transcription factor Sp1 expression is a significant predictor of survival in human gastric cancer. Clin Cancer Res 9:6371–6380
Wang H, Gutierrez-Uzquiza A, Garg R, Barrio-Real L, Abera MB, Lopez-Haber C, Rosemblit C, Lu H, Abba M, Kazanietz MG (2014) Transcriptional regulation of oncogenic protein kinase Cϵ (PKCϵ) by STAT1 and Sp1 proteins. J Biol Chem 289:19823–19838
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–361
Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, Shang JL, Gao CF, Zhang FR, Wang F, Sun SH (2011a) Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology 54:1679–1689
Yang L, Lin C, Liu W, Zhang J, Ohgi KA, Grinstein JD, Dorrestein PC, Rosenfeld MG (2011b) ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147:773–788
Yang G, Lu X, Yuan L (2014) LncRNA: a link between RNA and cancer. Biochim Biophys Acta 1839:1097–1109
Zannetti A, Del Vecchio S, Carriero MV, Fonti R, Franco P, Botti G, D’Aiuto G, Stoppelli MP, Salvatore M (2000) Coordinate up-regulation of Sp1 DNA-binding activity and urokinase receptor expression in breast carcinoma. Cancer Res 60:1546–1551
Zhao Y, Yang Y, Trovik J, Sun K, Zhou L, Jiang P, Lau TS, Hoivik EA, Salvesen HB, Sun H, Wang H (2014) A novel wnt regulatory axis in endometrioid endometrial cancer. Cancer Res 74:5103–5117
Acknowledgments
This work was supported by Grant from the Chinese National Nature Science Foundation (31070706) and by the Fund from Nanjing Health Bureau (ykk10082).
Conflict of interest
The authors declared that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Shufeng Li, Qiwei Wang and Qian Qiang have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, S., Wang, Q., Qiang, Q. et al. Sp1-mediated transcriptional regulation of MALAT1 plays a critical role in tumor. J Cancer Res Clin Oncol 141, 1909–1920 (2015). https://doi.org/10.1007/s00432-015-1951-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-015-1951-0