Summary
The nonsteroidal anti-inflammatory drug (NSAID), tolfenamic acid (TA) is emerging as a new anti-cancer agent. TA induces the degradation of specific Specificity protein (Sp) transcription factors, Sp1, Sp3 and Sp4 which are associated with tumor growth and metastasis. In this study we have evaluated the effect of TA on lung cancer using both in vitro and in vivo models. TA in a dose dependent manner inhibited proliferation and cell viability of two different lung cancer cells, A549 and CRL5803. TA treatment for 48 h significantly decreased the expression of Sp1, Sp3 and Sp4. The hepatocyte growth factor receptor, c-Met is overexpressed in a variety of cancers including lung cancer and Sp proteins mediate the regulation of c-Met. TA diminished the expression of c-Met protein and modulates its downstream signaling pathway. Furthermore, TA treatment significantly increased the number of apoptotic cells and pro-apoptotic markers c-PARP and Bax confirming the activation of apoptotic pathways. In vivo studies using the orthotopic mice model for lung cancer showed that TA (25 mg/kg/2 days and 50 mg/kg/2 days) resulted in a dose dependent decrease in tumor formation. The immunohistochemical staining of lung tissue showed high expression of Sp1, Sp3, Sp4, c-Met and phospho Met in control group and a dose dependent decrease in TA treated groups. The crucial findings of this study support that targeting c-Met with a potent inhibitor of Sp proteins is a robust strategy for the implications in lung cancer treatment and TA can serve as a therapeutic agent for this devastating disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the leading cause of cancer related deaths among men and women in the world. It is responsible for one third of deaths among cancer patients in the United States [1, 2] and National Institutes of Cancer (NCI) estimates that 219,440 new cases and 159,390 deaths will occur in 2009 in the United States alone. Lung cancer has a dismal prognosis with 5-year relative survival rates of only 13.6% for men and 17.5% for women which warrants for the development of improved therapies for the treatment of this disease [3].
The main types of lung cancer are small cell lung carcinoma (SCLC) and non-small cell lung carcinoma (NSLC) and majority (75–85%) of lung cancer cases are NSCLC. Scientists and clinicians are continuously investigating for the inactivated tumor suppressor genes and over-expressed growth promoting oncogenes to identify the hallmarks of lung cancer and such hallmarks are essential for the implications in early detection, prevention, and treatment of the disease [4]. It is well documented that several genes including K-ras, p53 and EGFR are associated with the pathogenesis of lung cancer [5–7]. Due to the potential role of EGFR pathway in the pathogenesis of NSCLC, small molecule inhibitors, gefitinib, erlotinib, and cetuximab have been tested and the results showed a limited response rate [8–10]. Now the focus has been shifted to investigate other receptors in tyrosine kinase family with potential role in the pathogenesis of lung cancer [10, 11]. Hepatocyte growth factor (HGF) receptor c-Met, a member of tyrosine kinase receptor family is normally expressed by epithelial cells and has been found to be overexpressed and amplified in a variety of human tumor tissues including lung cancer [12–16]. c-Met/HGF signaling plays key role in growth, motility, invasion, metastasis, angiogenesis, wound healing, and tissue regeneration [10, 17] and its aberrant expression is associated with various cancers. c-Met is functionally expressed in NSCLC [10] and higher levels of HGF is reported in aggressive disease and poor prognosis in lung cancer [18, 19]. It is well established that specificity protein (Sp) family of transcription factors mediate the constitutive expression of c-Met gene since Sp1 and Sp3 bind to the promoter region of c-Met and functionally activate its transcription [20–22].
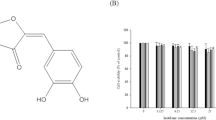
Sp1, a member of Sp protein family is overexpressed in a variety of cancers including gastric, colorectal, pancreatic, epidermal, thyroid, breast and lung cancers. Previous studies from our laboratory showed that Sp proteins mediate a number of genes involved in cell proliferation, survival and angiogenesis and suggested that targeting Sp protein transcription factors can serve as a powerful strategy for the development of anti-cancer agents [23–27]. Our studies on pancreatic cancer and esophageal cancer models have identified a nonsteroidal anti-inflammatory drug (NSAID), tolfenamic acid (TA) with such property to repress specific Sp proteins [22–24, 28–31]. Studies with TA and other structurally related analogs showed that TA is the most potent inhibitor of specific Sp proteins (Sp1, Sp3 and Sp4) expression [28]. TA is currently in Phase I clinical Trial at M. D. Anderson Cancer Center Orlando to test on upper gastrointestinal cancer patients.
In this study, we show that Sp proteins and c-Met are overexpressed in lung cancer cells and tumors in nude mice and TA decreases the lung cancer cell survival, represses Sp proteins, c-Met and key candidates in its downstream signaling. The current pre-clinical data suggest that TA can serve as a potential therapeutic agent for the treatment of lung cancer.
Materials and methods
Cell lines and chemicals
A549 and CRL5803 cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA). F12K with phenol red, 100X antibiotic/antimycotic solution and tolfenamic acid were purchased from Sigma Chemical Co. (St. Louis, MO). Rabbit IgG, antibodies for Sp1, Sp3, Sp4, β-actin and c-Met proteins were obtained from Santa Cruz Biotechnology (Santa Cruz, CA), p-Akt, Akt, Bax, p-ERK1/2, ERK1/2, and c-PARP were obtained from Cell Signaling Technology Inc., (Danvers, MA), and p-Met was purchased from BioSource International Inc., (Camarillo, CA). Colorimetric TUNEL staining kits were purchased from Promega (Madison, WI).
Orthotopic implantation of tumor cells
Male athymic nude mice (NCI-nu) were purchased from the Animal Production Area of the National Cancer Institute Frederick Cancer Research and Development Center (Frederick, MD). Mice were housed and maintained under specific pathogen-free conditions in facilities approved by the American Association for Accreditation of Laboratory Animal Care and in accordance with current regulations and standards of the United States Department of Agriculture, United States Department of Health and Human Services, and National Institutes of Health. Mice were used in accordance with institutional guidelines when they were 8–12 weeks of age. To produce tumors, A549 cells were harvested from subconfluent cultures by a brief exposure to 0.25% trypsin and 0.02% EDTA. Trypsinization was stopped with medium containing 10% fetal bovine serum, and the cells were washed once in serum-free medium and resuspended in HBSS. Only suspensions consisting of single cells with >90% viability were used for the injections. Injection of cells into the right lung was performed as described previously [29]. Mice were killed when moribund (4–5 weeks after injection). The size and weight of the primary lung tumors were recorded. Histopathologic studies confirmed the nature of the disease. For immunohistochemistry (IHC) and histological staining procedures, tumor tissue were fixed in formalin and embedded in paraffin.
Treatment of lung tumors growing in the nude mice
Fourteen days after implantation of tumor cells into the lung of each mouse, 5 mice were killed to confirm the presence of tumor lesions. Tumor-bearing mice were randomized and divided into 3 groups (10 per group). First group (control) was treated with vehicle group-2 and group-3 were treated with 25 mg/kg TA and 50 mg/kg TA on every other day respectively. We have also maintained an additional group (normal) which is injected with saline in lungs to compare the changes in lung weight due to tumor formation. Treatments were continued for 4 weeks and the mice were sacrificed on day 35 and subjected to necropsy.
Histological studies
Mice were sacrificed and body weights were recorded. Lungs were excised, measured, and weighed. For IHC and H&E staining procedures, lung tissues were fixed in formalin and embedded in paraffin. For immunohistochemistry and histologic staining, paraffin embedded tissues were used for identification of c-Met, p-Met, Sp1, Sp3, Sp4 and H&E. Sections (4–6 μm thick) were mounted on positively charged Superfrost slides (Fischer Scientific, Co., Houston, TX) and dried overnight. Sections were deparaffinized in xylene, treated with a graded series of alcohol [100%, 95%, and 80% ethanol (vol/vol) in double distilled water], and rehydrated in PBS (pH 7.5). Antigen retrieval occurred by placing slides in 97°C 0.1 M citrate buffer (pH 6.0) for 20 min. Slides were then washed with PBS that contained 0.1% Triton X. Endogenous peroxidase was blocked with 3% hydrogen peroxide in PBS. Nonspecific binding was blocked with 5% normal horse serum, 1% normal goat serum, and 0.1% Triton X. The slides were incubated at 4°C overnight in a moist chamber with a polyclonal antibody (Santa Cruz Biotechnology, CA, 1:25 dilution). After incubation for 1 h at room temperature with a peroxidase-conjugated rabbit IgG secondary (Santa Cruz Biotechnology, CA, 1:250 dilution), a positive reaction was visualized by incubating the slides with stable 3,3′-diaminobenzidine (Invitrogen Corp, Carlsbad, CA) for 8–10 min. The sections were rinsed with distilled water, counter-stained with Gill’s hematoxylin (Sigma) for 1 min, and mounted with Crystal Mount (Fischer Scientific, Co., Houston, TX). Control samples exposed to secondary antibody alone showed no specific staining.
TUNEL assay
Sections from paraffin-embedded tumor tissues were used for TUNEL staining which was carried out using DeadEnd Colorimetric TUNEL System (Promega, Madison, WI). Paraffin-embedded sections (4–6 μM thick) were processed per manufacture protocol as decribed in our previous publications [28, 29, 31]. Briefly, sections were deparaffinized in xylene and then treated with a graded series of alcohol [100, 95, 85, 70 and 50% ethanol (v/v) in double distilled H20] and rehydrated in PBS (pH 7.5). Tissues were then treated with Proteinase K solution for permeabilization and then refixed with 4% paraformaldehyde solution. Slides were treated with rTdT reaction mix, incubated at 37°C for 1 h., and reactions were terminated by immersing the slides in 2 X SSC solutions for 15 min at room temperature. After blocking endogenous peroxidases activity (by 0.3% hydrogen peroxide) slides were washed with PBS and then incubated with Streptavidin HRP solution for 30 min at room temperature. After washing, slides were incubated with DAB (substrate) solution until a light brown background appears (10 min) and then rinsed several times in deionized water. After mounting, slides were observed using light microscope.
Morphometric analysis
Lung specimens collected at the time of harvesting were stored in formalin, embedded in paraffin and sections were prepared. These sections were subjected to Hematoxillin and eosin staining and processed for histopathological analysis. Morphometric analysis was performed using Nikon Eclipse E400 microscope, under a Nikon plan Fluor 10X/0.30 DIC L (∞/0.17 WD16.0) objective. Images were captured with Leica DFC 300 FX camera attached to the microscope. LEICA Application Suite Version 2.8.1 software was used for the morphometric measurements. The morphometric analysis of tumor area was performed according to the procedure described previously [32]. Length and width of normal and malignant tissue were measured for each field (Supplementary data-Figure-I) and the data were analyzed using Graph Pad prism V5.0 statistical software and Excel office professional.
Immunoblotting
Protein expression was evaluated by Western blot analysis as described in our previous published work [28, 29, 31]. Cell lysates were prepared using lysis buffer [20 mmol/L Tris-HCl (pH 7.4), 150 mmol/L Sodium Chloride, 1 mmol/L EDTA, 1% Triton X-100, 0.1% SDS, 1 mmol/L sodium orthovanadate, 1 mmol/L phenylmethylsulfonyl fluoride, 1μmol/L leupeptin, and 1μg/mL aprotinin]. After centrifugation of the lysate at 15,000× g for 20 min, the supernatants were recovered, and protein was quantified by the Bradford protein assay using a reagent kit from Bio-Rad Laboratories. Protein samples (20–60 μg) were size-separated by electrophoresis on SDS-polyacrylamide gels under nonreducing conditions. Separated proteins were electroblotted onto nitrocellulose membranes. The blot was blocked by incubating in blocking buffer [5% skim milk, 10 mmol/L Tris (pH 7.5), 10 mmol/L sodium chloride, and 0.1% Tween 20] for 1 h at 20°C and then incubated with the primary antibody overnight at 4°C. Incubation with a horseradish peroxidase-conjugated anti-mouse or rabbit secondary antibody was then carried out at 20°C for 4 h. Antibody-bound proteins were detected by the Enhanced Chemiluminescence Western blotting analysis system (Perkin-Elmer Life and Analytical Sciences).
Reporter assays
For reporter assays, A549 and CRL 5803 cells (5 × 104) were seeded in 12-well plate in 1 ml DMEM supplemented with 5% fetal bovine serum. After 24 h, cells were transfected with variable lengths (0.9 kb-full length; 0.7 kb; 0.2 kb and 0.1 kb) of c- Met luc constructs (500 ng) and 5–6 h post-transfection cells were treated with DMSO or TA (50 µM). 24 h later (post treatment), cells were harvested and lysed and luciferase activity (relative to β-galactosidase activity) was measured by dual luciferase reporter assay system following manufacturer’s protocol (Promega Corporation, WI). Cells transfection was carried out using Lipofectamine 2000 (Invitrogen, CA) following the manufacturer’s protocol.
Statistical analysis
Statistical significance was determined by analysis of variance and Scheffe’s test, and the levels of probability are noted. The results of cell culture studies are expressed as mean ± SD.
Results
TA inhibits proliferation, cell viability and induces apoptosis in A549 and CRL5803 cells
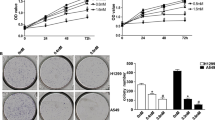
The effect of TA (25, 36, 50 µM) on cell proliferation (Fig. 1a & b) and cell viability (Fig. 1c & d) was tested on A549 and CRL5803 human lung cancer cells. TA decreased both cell proliferation and viability in a dose dependant manner. The IC50 values for cell proliferation and cell viability were 53.41 µM/77.5 µM and 50.42 µM/53.36 µM at 2 days and 36.5 µM/48.07 µM and 38.52 µM/44.42 µM at 6 days respectively for A549 and CRL5803. Since TA inhibited the cell survival in a dose dependent manner, we have also evaluated the effect of TA on inducing programmed cell death-apoptosis. TA significantly increased the number of apoptotic cells both in A549 and CRL5803 cells following the treatment for 48 h and this increase was also dose dependent (Fig. 2a & b). The apoptosis-inducing effect of TA was further confirmed by examining the expression of selected pro-apoptotic markers, c-PARP and Bax. The results showed that TA (50 µM; 48 h) significantly increased the expression of both c-PARP and Bax in A549 and CRL5803 cells. Upregulation of Bax and PARP, the major executioners of mitochondrial apoptotic pathway suggested the TA induced intrinsic apoptotic pathways.
Tolfenamic acid inhibits cell proliferation and cell viability of lung cancer cells. Lung cancer cells a A549 and b CRL5803 cells were treated with DMSO or 25, 36, 50 µM TA and cells were counted at 2, 4 and 6 days after the treatment as described in the materials and methods section. Cell viability test was also performed on these cells using CellTiter Glo kit (Promega). Lung cancer cells c A549 and d CRL5803 were seeded on 96 well plates. After 24 h cells were treated with DMSO or 25, 36, 50 µM of TA. Cell viability was estimated at 2, 4 and 6 days after the treatment. Data shown is mean ± S.E.M. of at least three separate determinations for each group
TA induced apoptosis in A549 and CRL5803 cells. The effect of TA on cell apoptosis was examined in lung cancer cells, A549 and CRL5803. Cells were treated with DMSO or increasing (25, 36, 50 µM) doses of TA. 48 h following the treatment the number of apoptotic cells was counted as described in the methods section. Results showed a significant increase in the number of apoptotic cells both in a A549 and b CRL5803 cells following the treatment of TA. Data shown is mean ± SD (n = 4) and asterisks indicate significant difference from corresponding controls (*: p < 0.05; **: p < 0.005; ***: p < 0.001). c Lung cancer cells were treated with DMSO or TA (50 µM) for 48 h and whole cell lystaes were prepared and expression of pro-apoptotic markers c-PARP and Bax were evaluated through Western blot analysis. The data were obtained from at least three different determinations and representative gels were shown in the figure (c)
TA represses Sp1, Sp3 and Sp4 transcription factors in lung cancer cells
Sp1, Sp3 and Sp4 are the sequence specific transcription factors that recognize and bind to GC rich sequences [33]. Since large number of genes including the associated with tumor growth and metastasis contain GC-box in promoter region, Sp proteins play critical role in the transcriptional activation of these genes [34, 35]. TA is a known potent inhibitor of Sp proteins as evinced by the studies on pancreatic cancer and esophageal cancer models and it is very crucial to test for such response in lung cancer model. We examined the concentration and time dependent effects of TA on the expression of Sp1, Sp3 and Sp4 in lung cancer cells, A549 and CRL5803. As shown in Fig. 3, TA (50 µM; 48 h) decreased the expression of Sp1, Sp3 and Sp4 proteins in both cell lines and these results are in consistent with our published the results on pancreatic cancer cells.
Tolfenamic acid induces the degradation of Sp proteins and other key proteins associated with tumor growth: Lung cancer cells A549 and CRL5803 were treated with DMSO or 50 µm of TA for 48 h. Whole cell lysates were prepared and the expression of various proteins including Sp1, Sp3, Sp4, c-Met, ERK1/2, and Akt were evaluated through Western blot analysis. The data were obtained from at least three different determinations and representative gels were shown in the figure
TA represses expression of Sp-dependent genes including c-Met
Compounds such as TA, curcumin and betulinic acid that decrease Sp proteins also downregulate expression of several Sp-dependent genes and proteins. The hepatocyte growth factor receptor c-Met is overexpressed in a variety of cancers including lung cancer and it is clear that Sp1 and Sp3 mediate the expression of c-Met. TA (50 µM; 48 h) decreased the expression of c-Met and its phosphorylated form (p-Met) in both cell lines studied (Fig. 3; Supplementary data Figure-II). TA-induced inhibition of c-Met and p-Met expression suggests that it can produce beneficial effect on c-Met downstream signaling pathway. We have screened the expression of a few candidates associated with the downstream signaling of c-Met and found that TA (50 µM; 48 h) significantly decreased the phosphorylation of ERK1/2 and Akt (Fig. 3).
TA decreased transcriptional activity of c-Met in lung cancer cells
The efficacy of TA on the transcriptional activity of c-Met was evaluated in lung cancer cells, A549 and CRL5803 using different c-Met promoter constructs. Full length and shorter promoter constructs have been used to identify functional Sp1 sites that might be responsible for the inhibitory effects of tolfenamic acid. Cells (5 × 104) were seeded into 12 well plates and transfected with full length (0.9 kb) and three shorter (0.7 kb, 0.2 kb and 0.1 kb) c-Met luc constructs. After 5–6 h post transfection, cells were treated with TA (50 µM) for 24 h and then dual luciferase assays were performed as per the manufacturer’s instructions. Luciferase reporter assays showed that tolfenamic acid significantly (p < 0.05) decreased c-met transcriptional activity in both cell lines, A549 (Fig. 4a) and CRL5803 (Fig. 4b). These promoter analysis studies have shown that basal promoter activity of c-Met decreased in the shorter constructs compare to the full length promoter and this might be in part to the loss of the upstream GC rich and other regulatory sites. However, promoter inhibitory effect of tolfenamic acid in the full length construct was also maintained in the shorter constructs which suggest that the effect of tolfenamic acid is mainly medicated through proximal Sp1 sites.
TA decreased transcriptional activity of c-Met in lung cancer cells. A549 and CRL5803 cells were transfected with c-Met constructs with variable lengths (0.9 kb-full length; 0.7 kb, 0.2 kb and 0.1 kb). Following 24 h treatment with TA (50 µM), cells were harvested and dual luciferase assays were performed. Data shown were the mean ± SD of at least three different determinations and the bars maerked with ‘*’ are significantly different from corresponding controls (p < 0.05)
In vivo carcinogenicity studies and immunohistochemistry of lung tumors harvested from orthotopic mice
Morphometric analysis
Athymic nude mice bearing A549 cells in the lung were treated with vehicle or two doses (25 mg/kg/2 days or 50 mg/kg/2 days) of TA for 4 weeks and the tumor tissues were subjected to histopathological and histochemical analysis. The results showed that TA significantly diminished the tumor growth as evinced by the parameters tested (Fig. 5). TA significantly reduced the increase in lung weight and this response was dose dependent. The median lung weight of 50 mg/kg/2 days dose group (mice injected with saline). Morphometric analysis of tumor area showed a significant/dose dependent decrease and this decrease was around 80% in the group received 50 mg/kg/ TA on every other day.
Morphometric analysis to elucidate the effect of Tolfenamic acid on tumor growth in orthotopic lung cancer mice model. Mice injected with saline (normal group) or A549 cells (control and TA treated groups) in lungs as described in the methods section. Animals were treated with vehicle (normal/control groups) or TA (25 mg/kg/2 days or 50 mg/kg/2 days) and the lungs were harvested and weights were recorded (a). The Lung tissues sections were analyzed to determine the percent of tumor area. The length and width of normal tissue (N) and tumor tissue (T) were measured (b) and percent tumor area was calculated (c) as described in the methods section. Data represents Mean ± SEM and the bars marked with ‘*’ are significantly different from control group (p: <0.0001)
Immuno staining
TUNEL staining was performed on sections prepared from tumors of the control and TA-treated groups. The results revealed a marked increase (dose dependent) in apoptosis in TA treated groups (Fig. 6a) and these results are correlated with downregulation of Sp proteins and c-Met in tumors from treated vs. control animals. These results demonstrate that TA is highly effective in inhibiting lung tumors growth in nude mice and interestingly, these tumors shows a better response to TA treatment when compared with our previously reported results on pancreatic tumors. Tumors tissue sections were then stained to determine the expression of Sp1, Sp3, Sp4, c-Met and p-Met. Control group showed intense staining reflecting high expression of Sp and c-Met proteins compared to low expression in TA treated groups (Fig. 6b). These in vivo data further confirm the efficacy of TA in decreasing c-Met expression by targeting selected Sp proteins.
Immunohistochemical analysis showing the effect of TA in lung tissue sections from the orthotopic lung cancer mice. Mice were injected with A549 cells in lungs, treated with TA (25 mg/kg/2 days or 50 mg/kg/2 days), and lung tissues were harvested. The tissue sections were prepared and subjected to immunostaining for (A) apoptosis (TUNEL) and the expression of Sp proteins (Sp1, Sp3, Sp4), c-Met and p-Met as described in the methods section. TUNEL assay revealed that TA treatment increased the number of apoptotic cells in a dose dependent manner (a). IHC studies showed that Sp proteins, c-Met and its phosphorylated forms were highly expressed in control group and dose dependently reduced due to TA treatment (b)
Discussion
The aberrant expression of hepatocyte growth factor receptor c-Met is associated with several cancers including lung cancer and the studies from our and others laboratories showed that targeting critical candidates in the tyrosine kinas receptors family such as c-Met is a very promising strategy for the implications in cancer treatment [10–16, 22, 36]. c-Met is functionally expressed in NSCLC [10] and higher levels of HGF are reported in aggressive disease and poor prognosis in lung cancer [18, 19]. It is well established that specificity protein (Sp) family of transcription factors mediate the constitutive expression of c-met gene since Sp1 and Sp3 bind to the promoter region of c-met and functionally activate its transcription [20, 21]. Previous studies from our laboratory showed that NSAID, TA inhibited the tumor growth and metastasis of pancreatic cancer and esophageal cancer [22, 28, 31]. The underlying mechanism of its action is not fully evaluated; however the anticancer activity of TA is primarily attributed to its efficacy on the degradation of specific Sp proteins Sp1, Sp3 and Sp4. In this study we investigated the anti-cancer activity of TA in lung cancer model, targeting specific Sp transcription factors and c-Met with the potential implications for the treatment of this disease.
As shown in Fig. 1, TA significantly decreased the cell proliferation and viability in A549 and CRL5803 cells. In consistent with these results the number of apoptotic cells was also increased with TA dose increase (Fig. 2). These results showed that TA treatment caused cell growth arrest and steering cells to undergo apoptosis. To further confirm the initiation of cell apoptosis and potential underlying mechanism, we have tested the expression of Bax and cleaved- poly-ADP-ribose polymerase (PARP) in A549 and CRL5803 cells. The results showed a significant increase both in Bax and c-PARP expressions following TA (50 µM) treatment for 48 h. (Fig. 2). The increase in Bax can decrease the mitochondrial potential [37] which will make the cells bioenergytically deficient and subsequently leading to cell death. In a typical apoptotic program, the alterations in Bax could result the release of cytochrome-c from mitochondria and the accumulation of cytochrome-c in cytosol interacts with Apaf-1 and procaspase-9 leading to the formation of apoptosome which could cleaves and induces the inactivation of PARP [38–42].
It is well established that Sp1, Sp3 and Sp4 are associated with tumor growth and these Sp proteins are highly expressed in tumor tissues and TA induced the degradation of Sp1, Sp3 and Sp4 and inhibit the expression of several Sp-dependent genes in pancreatic and esophageal cancer cells [22, 28]. In the current study we have evaluated the effect of TA on Sp proteins and other key genes associated with tumor progression in A549 and CRL5803 cells (Fig. 3) and found that TA decreased the expression of Sp1, Sp3 and Sp4. TA also inhibited the expression of c-Met and p-Met both in A549 and CRL5803 cells. We have also evaluated the effect of TA on the transcriptional activity of c-Met in lung cancer cells, A549 and CRL5803 using c-Met constructs with variable lengths (Fig. 4a & b). These results revealed that TA significantly inhibited the transcriptional activity of c-Met and this inhibitory effect was seen in full length as well as in shorter promoter constructs suggesting the importance of proximal Sp sites in medicating such effect of tolfenamic acid. Since TA modulated the activity and expression of c-Met, it is also important to investigate the response on key candidates in its downstream signaling pathway so we have studied Akt and ERK1/2 proteins (both constitutive and phosphorylated forms) in A549 and CRL5803 cells. TA significantly reduced the phosphorylation status of Akt and ERK1/2. These data demonstrate that TA repressed the expression of Sp proteins and c-Met and so interfere with c-Met survival signaling pathway. These in vitro results suggest that targeting Sp proteins for degradation in lung cancer cells/tumors with TA represent an alternate approach for inhibiting c-Met protein or signaling pathway.
The in vivo anticarcinogenic activity of TA was investigated in orthotopic mice model bearing A549 cells in lung and results showed significant decrease in tumor size and incidences of tumor formation. TA treatment (25 mg/kg/2 days and 50 mg/kg/2 days) decreases (Fig. 5) tumor size and weight (as indicated from whole lung). In addition, TUNEL staining in tumor sections from TA treated mice were significantly increased reflecting the activation of apoptosis compared ot he control group (Fig. 6a). In consistent with in vitro findings, in vivo results also confirmed that TA treatment decreased Sp1, Sp3, Sp4, c-Met and p-Met staining in tumor sections in a dose dependant manner (Fig. 6b). These results demonstrate that TA is a new potential anticancer agent for the treatment of lung cancer and these findings complements our previous studies on pancreatic and esophageal cancer models [22, 28, 31]. The mechanism of action of TA as a tumor growth inhibitor is due, in part, to the repression of Sp transcription factors and Sp-dependent genes, many of which are overexpressed in tumors and cancer cell lines [43].
Recent studies have proposed the association of Sp1 transcription factor in lung cancer [35, 44–47]; however, in this study we show for the first time that targeting selective Specificity proteins (Sp1, Sp3 and Sp4) by tolfenamic acid could be a promising approach to inhibit lung cancer cells/tumor growth using both in vitro and in vivo models. Furthermore, we report for the first time that tolfenamic acid decreases c-Met expression through degradation of Sp proteins in lung cancer cells and tumors in nude mice bearing A549 cells in lung. This is particularly relevant for the potential efficacy of TA for treatment of lung cancer since both c-Met and p-Met are overexpressed in lung cancer cells and tumors and it could be a potential target for the treatment of lung cancer and such investigations will be critical for the development of TA and related compounds as a novel class of mechanism based drugs for the treatment of lung cancer.
References
Jemal A, Siegel R, Ward E, Murray T, Xu J, Smigal C, Thun MJ (2006) Cancer statistics, 2006. CA Cancer J Clin 56:106–130
Bey EA, Bentle MS, Reinicke KE, Dong Y, Yang CR, Girard L, Minna JD, Bornmann WG, Gao J, Boothman DA (2007) An NQO1- and PARP-1-mediated cell death pathway induced in non-small-cell lung cancer cells by beta-lapachone. Proc Natl Acad Sci USA 104:11832–11837
Albert JM, Cao C, Kim KW, Willey CD, Geng L, Xiao D, Wang H, Sandler A, Johnson DH, Colevas AD, Low J, Rothenberg ML, Lu B (2007) Inhibition of poly(ADP-ribose) polymerase enhances cell death and improves tumor growth delay in irradiated lung cancer models. Clin Cancer Res 13:3033–3042
Fong KM, Sekido Y, Gazdar AF, Minna JD (2003) Lung cancer. 9: molecular biology of lung cancer: clinical implications. Thorax 58:892–900
Devereux TR, Taylor JA, Barrett JC (1996) Molecular mechanisms of lung cancer. Interaction of environmental and genetic factors. Giles F. Filley Lecture. Chest 109:14S–19S
Herbst RS, Heymach JV, Lippman SM (2008) Lung cancer. N Engl J Med 359:1367–1380
Aviel-Ronen S, Blackhall FH, Shepherd FA, Tsao MS (2006) K-ras mutations in non-small-cell lung carcinoma: a review. Clin Lung Cancer 8:30–38
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, Haber DA (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350:2129–2139
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, Meyerson M (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304:1497–1500
Ma PC, Jagadeeswaran R, Jagadeesh S, Tretiakova MS, Nallasura V, Fox EA, Hansen M, Schaefer E, Naoki K, Lader A, Richards W, Sugarbaker D, Husain AN, Christensen JG, Salgia R (2005) Functional expression and mutations of c-Met and its therapeutic inhibition with SU11274 and small interfering RNA in non-small cell lung cancer. Cancer Res 65:1479–1488
Hahn O, Salgia R (2005) Novel therapies in lung cancer. Hematol Oncol Clin North Am 19:343–367 vii
Natali PG, Prat M, Nicotra MR, Bigotti A, Olivero M, Comoglio PM, Di Renzo MF (1996) Overexpression of the met/HGF receptor in renal cell carcinomas. Int J Cancer 69:212–217
Olivero M, Rizzo M, Madeddu R, Casadio C, Pennacchietti S, Nicotra MR, Prat M, Maggi G, Arena N, Natali PG, Comoglio PM, Di Renzo MF (1996) Overexpression and activation of hepatocyte growth factor/scatter factor in human non-small-cell lung carcinomas. Br J Cancer 74:1862–1868
Hellman A, Zlotorynski E, Scherer SW, Cheung J, Vincent JB, Smith DI, Trakhtenbrot L, Kerem B (2002) A role for common fragile site induction in amplification of human oncogenes. Cancer Cell 1:89–97
Di Renzo MF, Olivero M, Katsaros D, Crepaldi T, Gaglia P, Zola P, Sismondi P, Comoglio PM (1994) Overexpression of the Met/HGF receptor in ovarian cancer. Int J Cancer 58:658–662
Maulik G, Kijima T, Ma PC, Ghosh SK, Lin J, Shapiro GI, Schaefer E, Tibaldi E, Johnson BE, Salgia R (2002) Modulation of the c-Met/hepatocyte growth factor pathway in small cell lung cancer. Clin Cancer Res 8:620–627
Ma PC, Maulik G, Christensen J, Salgia R (2003) c-Met: structure, functions and potential for therapeutic inhibition. Cancer Metastasis Rev 22:309–325
Siegfried JM, Weissfeld LA, Luketich JD, Weyant RJ, Gubish CT, Landreneau RJ (1998) The clinical significance of hepatocyte growth factor for non-small cell lung cancer. Ann Thorac Surg 66:1915–1918
Bharti A, Ma PC, Maulik G, Singh R, Khan E, Skarin AT, Salgia R (2004) Haptoglobin alpha-subunit and hepatocyte growth factor can potentially serve as serum tumor biomarkers in small cell lung cancer. Anticancer Res 24:1031–1038
Zhang X, Li Y, Dai C, Yang J, Mundel P, Liu Y (2003) Sp1 and Sp3 transcription factors synergistically regulate HGF receptor gene expression in kidney. Am J Physiol Renal Physiol 284:F82–94
Zhang X, Liu Y (2003) Suppression of HGF receptor gene expression by oxidative stress is mediated through the interplay between Sp1 and Egr-1. Am J Physiol Renal Physiol 284:F1216–1225
Papineni S, Chintharlapalli S, Abdelrahim M, Lee SO, Burghardt R, Abudayyeh A, Baker C, Herrera L, Safe S (2009) Tolfenamic acid inhibits esophageal cancer through repression of specificity proteins and c-Met. Carcinogenesis 30:1193–1201
Abdelrahim M, Safe S (2005) Cyclooxygenase-2 inhibitors decrease vascular endothelial growth factor expression in colon cancer cells by enhanced degradation of Sp1 and Sp4 proteins. Mol Pharmacol 68:317–329
Abdelrahim M, Smith R 3rd, Burghardt R, Safe S (2004) Role of Sp proteins in regulation of vascular endothelial growth factor expression and proliferation of pancreatic cancer cells. Cancer Res 64:6740–6749
Abdelrahim M, Samudio I, Smith R 3rd, Burghardt R, Safe S (2002) Small inhibitory RNA duplexes for Sp1 mRNA block basal and estrogen-induced gene expression and cell cycle progression in MCF-7 breast cancer cells. J Biol Chem 277:28815–28822
Hong J, Samudio I, Liu S, Abdelrahim M, Safe S (2004) Peroxisome proliferator-activated receptor gamma-dependent activation of p21 in Panc-28 pancreatic cancer cells involves Sp1 and Sp4 proteins. Endocrinology 145:5774–5785
Khan S, Abdelrahim M, Samudio I, Safe S (2003) Estrogen receptor/Sp1 complexes are required for induction of cad gene expression by 17beta-estradiol in breast cancer cells. Endocrinology 144:2325–2335
Abdelrahim M, Baker CH, Abbruzzese JL, Safe S (2006) Tolfenamic acid and pancreatic cancer growth, angiogenesis, and Sp protein degradation. J Natl Cancer Inst 98:855–868
Abdelrahim M, Baker CH, Abbruzzese JL, Sheikh-Hamad D, Liu S, Cho SD, Yoon K, Safe S (2007) Regulation of vascular endothelial growth factor receptor-1 expression by specificity proteins 1, 3, and 4 in pancreatic cancer cells. Cancer Res 67:3286–3294
Abdelrahim M, Liu S, Safe S (2005) Induction of endoplasmic reticulum-induced stress genes in Panc-1 pancreatic cancer cells is dependent on Sp proteins. J Biol Chem 280:16508–16513
Konduri S, Colon J, Baker CH, Safe S, Abbruzzese JL, Abudayyeh A, Basha MR, Abdelrahim M (2009) Tolfenamic acid enhances pancreatic cancer cell and tumor response to radiation therapy by inhibiting survivin protein expression. Mol Cancer Ther 8:533–542
Lee JC, Krochak R, Blouin A, Kanterakis S, Chatterjee S, Arguiri E, Vachani A, Solomides CC, Cengel KA, Christofidou-Solomidou M (2009) Dietary flaxseed prevents radiation-induced oxidative lung damage, inflammation and fibrosis in a mouse model of thoracic radiation injury. Cancer Biol Ther 8:47–53
Gidoni D, Dynan WS, Tjian R (1984) Multiple specific contacts between a mammalian transcription factor and its cognate promoters. Nature 312:409–413
Suske G (1999) The Sp-family of transcription factors. Gene 238:291–300
Cui J, Meng X, Gao X, Tan G (2009) Curcumin decreases the expression of Pokemon by suppressing the binding activity of the Sp1 protein in human lung cancer cells. Mol Biol Rep [Epub ahead of print].
Herrera LJ, El-Hefnawy T, Queiroz de Oliveira PE, Raja S, Finkelstein S, Gooding W, Luketich JD, Godfrey TE, Hughes SJ (2005) The HGF receptor c-Met is overexpressed in esophageal adenocarcinoma. Neoplasia 7:75–84
Wong BS, Hsiao YC, Lin TW, Chen KS, Chen PN, Kuo WH, Chu SC, Hsieh YS (2009) The in vitro and in vivo apoptotic effects of Mahonia oiwakensis on human lung cancer cells. Chem Biol Interact 180:165–174
Daniel PT, Wieder T, Sturm I, Schulze-Osthoff K (2001) The kiss of death: promises and failures of death receptors and ligands in cancer therapy. Leukemia 15:1022–1032
Gogvadze V, Orrenius S, Zhivotovsky B (2006) Multiple pathways of cytochrome c release from mitochondria in apoptosis. Biochim Biophys Acta 1757:639–647
Lee HJ, Lee HJ, Lee EO, Ko SG, Bae HS, Kim CH, Ahn KS, Lu J, Kim SH (2008) Mitochondria-cytochrome C-caspase-9 cascade mediates isorhamnetin-induced apoptosis. Cancer Lett 270:342–353
Lee JH, Lee YH, Lee HJ, Lee HJ, Lee EO, Ahn KS, Shim BS, Bae H, Choi SH, Ahn KS, Baek NI, Kim DK, Kim SH (2009) Caspase and mitogen activated protein kinase pathways are involved in Solanum lyratum herba induced apoptosis. J Ethnopharmacol 123:121–127
Lee KB, Kim KR, Huh TL, Lee YM (2008) Proton induces apoptosis of hypoxic tumor cells by the p53-dependent and p38/JNK MAPK signaling pathways. Int J Oncol 33:1247–1256
Safe S, Abdelrahim M (2005) Sp transcription factor family and its role in cancer. Eur J Cancer 41:2438–2448
Kang Y, Hong JA, Chen GA, Nguyen DM, Schrump DS (2007) Dynamic transcriptional regulatory complexes including BORIS, CTCF and Sp1 modulate NY-ESO-1 expression in lung cancer cells. Oncogene 26:4394–4403
Zheng Y, Ritzenthaler JD, Sun X, Roman J, Han S (2009) Prostaglandin E2 stimulates human lung carcinoma cell growth through induction of integrin-linked kinase: the involvement of EP4 and Sp1. Cancer Res 69:896–904
Tsou JH, Chang KY, Wang WC, Tseng JT, Su WC, Hung LY, Chang WC, Chen BK (2008) Nucleolin regulates c-Jun/Sp1-dependent transcriptional activation of cPLA2alpha in phorbol ester-treated non-small cell lung cancer A549 cells. Nucleic Acids Res 36:217–227
Sun X, Ritzenthaler JD, Zhong X, Zheng Y, Roman J, Han S (2009) Nicotine stimulates PPARbeta/delta expression in human lung carcinoma cells through activation of PI3K/mTOR and suppression of AP-2alpha. Cancer Res 69:6445–6453
Acknowledgements
Authors thank M. D. Anderson Cancer Center Orlando’s Cancer Research Institute for providing necessary financial and technical assistance. The assistance of Donna Schade and Beth Isley is greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jimmie Colon and Md. Riyaz Basha have equal contributions.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PPT 609 kb)
Rights and permissions
About this article
Cite this article
Colon, J., Basha, M.R., Madero-Visbal, R. et al. Tolfenamic acid decreases c-Met expression through Sp proteins degradation and inhibits lung cancer cells growth and tumor formation in orthotopic mice. Invest New Drugs 29, 41–51 (2011). https://doi.org/10.1007/s10637-009-9331-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-009-9331-8