Abstract
Purpose
In non-metastatic neuroblastoma (NB), the identification of the cases that require more intensive treatment is still difficult. Minimal disease (MD) and minimal residual disease (MRD) detection in outcome prediction seems to be important in advanced neuroblastoma, but there are not many studies focused on patients with non-metastatic disease. The aim of this study was to determine whether the presence of MD detected at diagnosis could be associated with bad prognosis.
Procedures
Quantitative reverse transcriptase–polymerase chain reaction QRT–PCR was performed on peripheral blood (PB) and bone marrow (BM) samples from patients with non-metastatic NB at diagnosis for tyrosine hydroxylase (TH) and doublecortin (DCX) mRNAs detection.
Results
The frequencies of detecting MD in our series of 102 patients with non-metastatic NB were as follows: 6.2% (5/81) PB samples and 10.6% (10/94) BM samples. Overall survival was similar for patients who expressed or not the MD biomarkers at diagnosis. However, patients with MD detected in PB showed lower EFS than patients with negative PB (P = 0.038).
Conclusions
Minimal disease detection in PB seems to be useful for predicting relapse probabilities in patients with non-metastatic NB. The stages 1 and 2 patients with neuroblastoma showed high survival rates, and MD was detected in a small number of patients probably being non-contributory for predicting patient outcome. For stage 3 patients with NB, MD detection by QRT–PCR in PB at diagnosis could be useful for predicting outcome and for early and sensitive detection of relapsing disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the most frequent solid tumours in early childhood is neuroblastoma (NB). NB is remarkable for its clinical and genetic heterogeneity. Prognosis fundamentally depends on age at diagnosis (Brodeur and Maris 2002; London et al. 2005), histology (Shimada et al. 1999), tumour stage (Brodeur et al. 1988) and MYCN oncogene amplification status (Brodeur et al. 1984; Seeger et al. 1985). Other molecular parameters such as tumour cell DNA content (Look et al. 1991), gain of chromosome arm 17q (Caron 1995), deletion of chromosome arm 1p (Christiansen and Lampert 1998) and recently 11q (Cohn et al. 2009; Spitz et al. 2006) have shown to be predictive of patient outcome.
Localized NB patients with MYCN amplification (MYCNA) (stages 2 and 3) are included in high-risk protocol and receive an intensive treatment schedule, including induction chemotherapy, surgery, myeloablative chemotherapy followed by autologous stem cell reinfusion, radiotherapy and biological-based therapy with 13-cis-retinoic acid. Nevertheless, high-risk patients have frequently fatal outcome.
On the other hand, patients with localized disease and without MYCNA are generally treated only with surgery (stages 1–2) or with standard–dose chemotherapy followed by surgery (stage 3) showing a good outcome. High 5-year overall survival (OS) and event-free survival (EFS) have been observed in these patients (99 ± 1% and 90 ± 3%, respectively) (Cotterill et al. 2000). Separating patients by stage, OS and EFS are around 95% for stage 1, 86% for stage 2 and 65% for stage 3 patients (Rubie et al. 2011). Despite the good prognosis, some of these patients present metastatic relapses and consequently die.
Solid tumours continuously shed cells into the circulation, and in NB, this minimal disease (MD) can be detected in peripheral blood (PB) and bone marrow (BM) (Moss and Sanders 1990; Swartz et al. 1999). Molecular detection of MD has been considered an important prognostic factor in metastatic NB. Quantitative reverse transcriptase–polymerase chain reaction (QRT–PCR) technique for detecting tumour cells in PB and BM is an extremely sensitive method that makes possible the detection of one tumoral cell in 106 haematopoietic cells (Cheung and Cheung 2001; Lambooy et al. 2003).
Many studies have been performed to identify suitable targets for MD and minimal residual disease (MRD) detection in NB. A recent European study concluded that tyrosine hydroxylase (TH), doublecortin (DCX) and PHOX2B genes are the best candidates for circulating NB cell detection (Viprey et al. 2008). In our group, we routinely perform the MRD analysis in high-risk patients with NB using TH and DCX biomarkers because of their complementarities in tumoral cell detection (Oltra et al. 2005).
The TH gene codes for the first enzyme in the catecholamine synthesis pathway, which is produced by NB cells (Burchill et al. 2001). The DCX gene codes for a microtubule-associated protein that interacts with and regulates the microtubule cytoskeleton. It is expressed specifically in migrating neurons from the central and peripheral nervous system (Gleeson et al. 1999).
The aim of the present study was to analyse the TH and DCX expression at diagnosis in PB and BM from patients with non-metastatic NB to associate the MD presence with outcome.
Materials and methods
Patients and samples
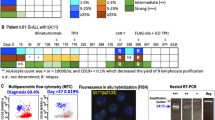
The present study consists of 102 patients with NB from 34 cooperative Spanish hospitals. NB staging was established according to International Neuroblastoma Staging System (INSS) criteria. Samples of fresh or frozen tumour were referred to the Spanish reference centre for pathology and molecular biology NB studies. Samples were centrally reviewed and classified according to the International Neuroblastoma Pathology Committee (INPC) system (Shimada et al. 1999; Burgues et al. 2006). Biological studies included status of MYCN and 1p, both studied by FISH according to ENQUA guidelines (Noguera et al. 2003; Ambros et al. 2003). Patients were included in different National and European studies (LNESG-I and II, INES, EUNS, HR-NBL1 and others). Two BM aspirations and 2 biopsies were performed on routine at diagnosis in all NB cases. All BM were previously analysed by cytological and histological screening and were negative at diagnosis. The MD studies were centrally performed at the Spanish NB MRD reference laboratory, and for these purpose, 2 ml of PB and 0.5 ml of BM were collected in EDTA Vacutainer tubes (BD, UK). All patients had at least one sample, and most of them had both PB and BM. Patients’ samples were considered positive when both markers were detected. Informed consent for samples and data management was obtained in all cases from patients’ parents.
Total RNA extraction
Ficoll gradient centrifugation was used to isolate mononuclear cells from BM and PB specimens, as recommended by the manufacturer (Lymphoprep AXIS-SHIELD PoC AS).
Once purified, mononuclear cells were lysed in RLT (RNasy kit; Qiagen) and immediately stored at −80°C. Total RNA was isolated with a RNasy kit (Qiagen), following the manufacturer’s recommendations.
Retro-transcription (RT)
Total RNA was reverse transcribed in a final volume of 40 μL, following the manufacturer’s guidelines (GeneampGold RNA PCR Core Kit, Applied Biosystems). The reaction mixture was incubated at 25°C for 15 min and at 42°C for 30 min.
Real-time quantitative PCR
Real-time quantitative PCR of DCX, TH and GAPD mRNA was achieved by means of the ABI Prism 7,000 Sequence Detection System (Applied Biosystems). TaqMan MGB probes and primers were used to quantify the DCX, TH and GAPDH expression. All expression assays have FAM reporter dye at the 5’ end and a non-fluorescent quencher at the 3′ end of the probe. GAPDH gene was used as an endogenous reference to control the difference in RNA extraction and cDNA synthesis.
In each 25 μL 96-Well Optical Reaction Plate, 1.5 μL of cDNA template was added to a PCR reaction mix (Applied Biosystems) containing 5 mmol/L MgCl2; 200 mmol/L of each of dATP, dCTP, dGTP and dUTP; 0.05 U/mL AmpliTaq Gold DNA polymerase; and 0.01 U/mL AmpErase UNG to prevent the PCR product from carrying over, as well as a passive reference dye, ROX. This reference dye provides an internal reference to which the reporter dye signal can be normalized, compensating for differences between wells and experiments caused by pipetting errors or by instrument variability; 1.25 μL of DCX, TH or GAPD Assays-on-Demand mixture was also included in the mixture (The TaqMan MGB probes and primers have a concentration of 18 nM for each primer and 5 nM for the probe). The assay references were as follows: Hs00167057_m1, Hs00165941_m1 and Hs99999905_m1, respectively. Each plate was covered with an Optical Adhesive Cover (Applied Biosystems). Samples were always run in duplicate PCR experiments. The initial PCR started using the ABI Prism 7,000 Sequence Detector with a 50°C, 2-minute step to optimize UNG activity, followed by a 95°C, 10-minute step to activate AmpliTaq Gold DNA polymerase and UNG deactivation. Then, 50 2-step cycles were performed, one at 95°C for 15 s and one at 60°C for 1 min. The entire PCR took 2 h and 11 min to complete, with no post-PCR handling. The amount of both DCX and TH was measured in reference to the housekeeping GAPDH gene, also studied in duplicate by obtaining the ΔCT values in the following way: The mean CT for the marker gene was subtracted by the mean CT of GAPDH (ΔCT = CT [Marker]-CT [GAPDH]).
Statistical analyses
Mean values comparison was performed by the Student’s t test. For EFS analysis, time to event was defined as the time from diagnosis until the time of first occurrence of relapse, progression or death. For OS, time to event was defined as time until death or until last contact if the patient was alive. Survival analysis was performed to compare OS and EFS between groups using the Kaplan–Meier method, and differences between groups were tested using log-rank tests. These statistical analyses were run using SPSS, version 12.0.
Results
Samples from 102 NB patients with non-metastatic disease at diagnosis were classified as follows: 38 stage 1, 11 stage 2 and 53 stage 3. Stages 1–2 patients were treated only with surgery followed by a complete re-evaluation according to the LNESG-I or II protocol. Stage 3 patients also received chemotherapy according to the INES or EUNS study. Stages 2 and 3 patients with MYCNA received high-dose chemotherapy, myeloablative therapy with haematopoietic stem cell rescue, local radiotherapy and 13-cis retinoic therapy in conformity with the HR-NBL1 study. Clinical, biological and follow-up data are summarized in Table 1. Favourable histology was found in 54 patients (twenty-six stage 1, four stage 2 and twenty-four stage 3). Twenty-eight patients presented unfavourable histology (two stage 1, three stage 2 and twenty-three stage 3). INPC classification was not informed in 20 cases. MYCN amplification was detected in 16 patients (two stage 2 and fourteen stage 3). Deleted 1p chromosome was observed in 17 patients (two stage 1, one stage 2 and fourteen stage 3), while imbalanced 1p chromosome was found in 6 patients (three stage 1, one stage 2 and two stage 3). Relapses were observed in one stage 1 and three stage 2 (one local and two local and metastatic). However, all of them are currently alive. With respect to the stage 3 patients with NB, four patients presented disease progression and seven relapsed (three presented metastatic relapses, one presented local relapse and one presented local and metastatic relapse; relapse location was not informed in two patients). Thirteen stage 3 patients died, eleven of them due to disease progression, one due to surgical complications and one died 21 days after diagnosis because the patient’s parents refused any treatment (Table 1).
The expression of TH and DCX was analysed in 81 PB and 94 BM samples collected at diagnosis from the 102 patients included in the study. We detected positive PB samples in 5 patients (one stage 1 (3%), none stage 2 (0%) and four stage 3 (9%)). In addition, 10 patients presented positive BM samples (two stage 1 (6%), none stage 2 (0%) and eight stage 3 patients (16%)). The number and percentages of positive PB and BM samples depending on the TH and DCX expression and separating patients according to the stage are detailed in Table 2.
OS and EFS were similar for patients expressing or not the MD markers in BM. This analysis was also done excluding the patient who died due to surgical complications and the untreated case, and the same results were obtained. When we separated patients by stage, there were no differences in OS or in EFS for stages 1–2 and stage 3 patients according to the detection of MD in BM or not (Fig 1).
Significant results were found in survival analysis when we studied MD detection in PB. The analysis shows that non-metastatic NB patients with MD detected in PB presented lower EFS than patients with negative PB (P = 0.038). OS was similar for patients expressing, or not, the MD markers in PB. We did the analysis also separating patients by stage. Neither did we find significant differences in OS or in EFS in stages 1–2 depending on MD detection or not. In stage 3 patients, however, we found that cases with MD detected in PB showed lower EFS than patients without MD in PB (Fig. 2). These results were very close to be significant (P = 0.057).
We studied the prognostic impact of unfavourable histology, age at diagnosis, MYCNA and 1p deletion in our series. Except for age at diagnosis, survival analyses revealed that patients with those risk factors showed significant lower OS and EFS than patients with favourable histology, no MYCNA and undeleted 1p chromosome, as expected (Fig 3). Patients older than 18 months showed a tendency of bad outcome in EFS but the differences were not statistically significant (P = 0.08).
Effects on survival of age at diagnosis, histology, MYCN amplification and 1p chromosome status of the 102 patients with non-metastatic neuroblastoma. <18 months: patients younger than 18 months, >18 months: patients older than 18 months, FH favourable histology, UH unfavourable histology, MYCNA MYCN amplification, MYCN not A MYCN not amplified, normal 1p: normal 1p chromosome, imbal. 1p: imbalanced 1p chromosome, del.1p: deleted 1p chromosome
The presence of MD at diagnosis and the previously mentioned risk factors were also analysed in our series. Out of 5 patients with MD detected in PB, 3 did not have any of the studied risk factors. Two cases showed deleted 1p chromosome and MYCN amplification, and one of them, older than 18 months, presented unfavourable histology. Regarding the 10 patients with MD detected in BM, 8 were younger than 18 months and presented normal MYCN status. MYCNA was detected in 2 patients. Also, 2 showed deleted 1p chromosome and imbalance 1p chromosome. Furthermore, five patients had unfavourable histology, and one patient presented the 4 studied risk factors. We could not establish an association between MD detection and the presence or not of the above-mentioned risk factors, due to the small number of localized NB patients with MD at diagnosis.
Discussion
Regarding localized NB, the identification of cases that require special attention is still difficult. MYCN amplification, deletion of chromosome arm 1p and recently 11q are well documented as recognized biological risk factors, but there are cases without such risk factors with a bad outcome. MD and MRD detection in outcome prediction seems to be important in advanced NB, but there are not many studies focused on patients with non-metastatic disease.
In the present study, we analysed the impact of the MD detection in PB and BM from localized patients with NB at diagnosis using QRT–PCR. All BM were previously analysed by cytological screening and were negative at diagnosis. However, this technique has limited sensitivity and is unable to reliably detect MD (Cheung et al. 1997). We detected MD in PB samples from 5 patients (one stage 1 (3%), none stage 2 (0%) and four stage 3 patients (9%)). In addition, 10 patients presented positive BM samples (two stage 1 (6%), none stage 2 (0%) and eight stage 3 patients (16%)) Positive samples were considered when both MD markers were detected, taking into account that TH may be also present in some T-lymphocytes (Cosentino et al. 2007) and lead to false positivity.
The low relapse rates observed in our series fit well with those expected in patients with localized disease (De Bernardi et al. 2008). According to our results, TH and DCX mRNA detection in PB of children with localized disease at diagnosis predicts relapse probabilities, reflecting the propensity of dissemination via bloodstream. When we divided our series into stages 1–2 and stage 3 groups, there was a tendency of bad outcome in stage 3 patients with the MD biomarkers detected in PB. This situation is very close to be statistically significant, so we considered that increasing the number of analysed patients will clarify that MD detection by QRT–PCR in PB from stage 3 patients is helpful to predict patient outcome.
Concerning the stages 1–2 cases, we only studied the relapse risk based on the MD detection. Some groups have found in long-term follow-up of children with stages 1 and 2 NB high survival rates and a small proportion with local recurrence (Perez et al. 2000; De Bernardi et al. 2008). We reached the same conclusion, and even though some of them have relapsed, all are currently alive. MD was detected in a small percentage of our stages 1 and 2 patients with NB, and no correlation between MD detection and outcome was found, suggesting that for this group MD studies are non-contributory for predicting patient outcome.
Positive results previously obtained by immunocytochemistry (IC) in BM and PB samples at diagnosis from stages 1, 2 and 3 patients correlated with unfavourable outcome (Corrias et al. 2004; Corrias et al. 2008). With respect to stage 4 patients, TH mRNA detected in PB at diagnosis has been described as a poor prognostic indicator (Gleeson et al. 1999; Reynolds 2004). In addition, high transcript concentration of TH and other MRD markers in PB and BM at diagnosis have been associated with poor prognosis in patients with NB. In particular, TH in BM above median indicated worse outcome for a homogenous cohort with HR-NB (Träger et al. 2008).
It is worth mentioning the fact that only the MD detection in PB and not in BM samples showed this putative influence on survival. The use of PB and not BM samples for the study is an additional advantage, considering that patient will not be exposed to an invasive technique. Many evidences have shown that the presence of circulating tumour cells in the PB of patients with solid tumours correlates with clinical outcome (Steen et al. 2008; Bozionellou et al. 2004). However, some results are in conflict, and for this reason, the biological significance of circulating tumour cells and the clinical relevance of their detection are still unclear. These cells may generate metastatic deposits, depending on the expression of the appropriate molecules, or die by apoptotic mechanisms. Tumours present genetically heterogeneous cell subpopulations with different metastatic potential, so the presence of these circulating tumour cells is necessary, but not sufficient for the metastasis development (Mocellin et al. 2006).
On the other hand, we also confirm in our series that localized NB patients with unfavourable histology, MYCNA or 1p deletion showed significant lower OS and EFS than patients without those risk factors. Due to the heterogeneity of this neuroblastic tumour, only the combination of clinical and biological factors can explain the tumour behaviour.
In conclusion, MD detection in PB at diagnosis seems to be useful for predicting relapse probabilities in patients with non-metastatic NB. Stages 1 and 2 patients with NB showed high survival rates, and the MD markers were detected in a small number of patients probably being non-contributory for predicting patient outcome. For stage 3 patients with NB, MD detection by QRT–PCR in PB at diagnosis could be useful for predicting outcome and for early and sensitive detection of relapsing disease. QRT–PCR is a sensitive, easy and not expensive method for the detection of MD. However, in order to use a uniform methodology for analysis and reporting, we recommend performing the study of MD in PB from stage 3 patients with NB at diagnosis in a reference laboratory and to include this into an international study protocol. In this group of patients, the study of MD in PB at different time points of evolution/treatment could be another interesting challenge.
References
Ambros IM, Benard J, Boavida M, Bown N, Caron H, Combaret V, Couturier J, Darnfors C, Delattre O, Freeman-Edward J, Gambini C, Gross N, Hattinger CM, Luegmayr A, Lunec J, Martinsson T, Mazzocco K, Navarro S, Noguera R, O’Neill S, Potschger U, Rumpler S, Speleman F, Tonini GP, Valent A, Van Roy N, Amann G, De Bernardi B, Kogner P, Ladenstein R, Michon J, Pearson AD, Ambros PF (2003) Quality assessment of genetic markers used for therapy stratification. J Clin Oncol 21(11):2077–2084
Bozionellou V, Mavroudis D, Perraki M et al (2004) Trastuzumab administration can effectively target chemotherapy-resistant cytokeratin-19 messenger RNA-positive tumor cells in the peripheral blood and bone marrow of patients with breast cancer. Clin Cancer Res 10(24):8185–8194
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM (1984) Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science 224:1121–1124
Brodeur GM, Seeger RC, Barrett A, Berthold F, Castleberry RP, D’Angio G, De Bernardi B, Evans AE, Favrot M, Freeman AI, Haase G, Hartmann O, Hayes FA, Helson L, Kemshead J, Lampert F, Ninane J, Ohkawa H, Philip T, Pinkerton CR, Pritchard J, Sawada T, Siegel S, Smith EI, Tsuchida Y, Voute PA (1988) International criteria for diagnosis, staging, and response to treatment in patients with neuroblastoma. J Clin Oncol 6:1874–1881
Brodeur GM, Maris JM (2002) Neuroblastoma. In: Pizzo PA, Poplack DG (eds) Principles and practice of paediatric oncology, 4th edn. JB. Lippincott Co., Philadelphia, pp 895–938
Burchill SA, Lewis IJ, Abrams KR et al (2001) Circulating neuroblastoma cells detected by reverse transcriptase polymerase chain reaction for tyrosine hydroxylase mRNA are an independent poor prognostic indicator in stage 4 neuroblastoma in children over 1 year. J Clin Oncol 19:1795–1801
Burgues O, Navarro S, Noguera R, Pellín A, Ruiz A, Castel V, Llombart-Bosch A (2006) Prognostic value of the international neuroblastoma pathology classification in neuroblastoma (schwannian stroma-poor) and comparison with other prognostic factors: a study of 182 cases from the spanish neuroblastoma registry. Virchows Arch 449(4):410–420
Caron H (1995) Allelic loss of chromosome 1 and additional chromosome 17 material are both unfavourable prognostic markers in neuroblastoma. Med Pediatr Oncol 24:215–222
Cheung IY, Cheung NK (2001) Quantitation of marrow disease in neuroblastoma by real-time reverse transcription-PCR. Clin Cancer Res 7:1698–1705
Cheung NKV, Heller G, Kushner BH et al (1997) Detection of metastatic neuroblastoma in bone marrow: when is routine histology intensive? J Clin Oncol 15(8):2807–2817
Christiansen H, Lampert F (1998) Tumour karyotype discriminates between good and bad prognostic outcome in neuroblastoma. Br J Cancer 57:121–126
Cohn SL, Pearson AD, London WB, Monclair T, Ambros PF, Brodeur GM, Faldum A, Hero B, Iehara T, Machin D, Mosseri V, Simon T, Garaventa A, Castel V, Matthay KK (2009) The international neuroblastoma risk group (INRG) classification system: an INRG task force report. J Clin Oncol 27:289–297
Corrias MV, Faulkner LB, Pistorio A, Rosanda C, Callea F, Lo Piccolo MS, Scaruffi P, Marchi C, Lacitignola L, Occhino M, Gambini C, Tonini GP, Haupt R, De Bernardi B, Pistoia V, Garaventa A (2004) Detection of neuroblastoma cells in bone marrow and peripheral blood by different techniques: accuracy and relationship with clinical features of patients. Clin Cancer Res 10:7978–7985
Corrias MV, Parodi S, Haupt R, Lacitignola L, Negri F, Sementa AR, Dau D, Scuderi F, Carlini B, Bianchi M, Casale F, Faulkner L, Garaventa A (2008) Detection of GD2-positive cells in bone marrow samples and survival of patients with localised neuroblastoma. Br J Cancer 98(2):263–269
Cosentino M, Fietta AM, Ferrari M, Rasini E, Bombelli R, Carcano E, Saporiti F, Meloni F, Marino F, Lecchini S (2007) Human CD4_CD25_ regulatory T cells selectively express tyrosine hydroxylase and contain endogenous catecholamines subserving an autocrine/paracrine inhibitory functional loop. Blood 109:632–642
Cotterill SJ, Pearson AD, Pritchard J, Foot AB, Roald B, Kohler JA, Imeson J (2000) Clinical prognostic factors in 1,277 patients with neuroblastoma: results of the European neuroblastoma study group ‘survey’ 1982–1992. Eur J Cancer 36:901–908
De Bernardi B, Mosseri V, Rubie H, Castel V, Foot A, Ladenstein R, Laureys G, Beck-Popovic M, De Lacerda A et al (2008) Treatment of localised resectable neuroblastoma. results of the LNESG I study by the SIOP Europe neuroblastoma group. Br J Cancer 99:1027–1033
Gleeson JG, Lin PT, Flanagan LA et al (1999) Doublecortin is a microtubuleassociated protein and is expressed widely by migrating neurons. Neuron 23:257–271
Lambooy LH, Gidding CE, van den Heuvel LP, Hulsbergen-van de Kaa CA, Ligtenberg M, Bökkerink JP, De Abreu RA (2003) Real-time analysis of TH gene expression: a sensitive and semiquantitative marker for minimal residual disease detection of neuroblastoma. Clin Cancer Res 9:812–819
London WB, Castleberry RP, Matthay KK, Look AT, Seeger RC, Shimada H, Thorner P, Brodeur G, Maris JM, Reynolds CP, Cohn SL (2005) Evidence for an age cut-off greater than 365 days for neuroblastoma risk group stratification in the children’s oncology group. J Clin Oncol 23(27):6459–6465
Look AT, Hayes FA, Shuster JJ, Douglass EC, Castleberry RP, Bowman LC, Smith EI, Brodeur GM (1991) Clinical relevance of tumour cells ploidy and N-myc gene amplification in childhood neuroblastoma: a paediatric oncology group study. J Clin Oncol 9:581–591
Mocellin S, Keilholz U, Rossi CR, Nitti D (2006) Circulating tumour cells: the ‘leukemic phase’ of solid cancers. Trends Mol Med 12:130–139
Moss J, Sanders DG (1990) Detection of neuroblastoma cells in blood. J Clin Oncol 8:736–740
Noguera R, Canete A, Pellin A, Ruiz A, Tasso M, Navarro S, Castel V, Llombart-Bosch A (2003) MYCN gain and MYCN amplification in a stage 4S neuroblastoma. Cancer Genet Cytogenet 140(2):157–161
Oltra S, Martinez F, Orellana C, Grau E, Fernandez JM, Canete A, Castel V (2005) The doublecortin gene, a new molecular marker to detect minimal residual disease in neuroblastoma. Diagn Mol Pathol 14(1):53–57
Perez C, Matthay K, Atkinson J, Seeger R, Shimada H, Haase G, Stram D, Gerbing R, Lukens J (2000) Biologic variables in the outcome of stages I and II neuroblastoma treated with surgery as primary therapy: a children’s cancer group study. J Clin Oncol 18(1):18–26
Reynolds CP (2004) Detection and treatment of minimal residual disease in high-risk neuroblastoma. Pediatr Transplant 8(Suppl 5):56–66
Rubie H, De Bernardi B, Gerrard M, Canete A, Ladenstein R, Couturier J, Ambros P, Munzer C, Pearson AD, Garaventa A, Brock P, Castel V, Valteau-Couanet D, Holmes K, Di Cataldo A, Brichard B, Mosseri V, Marquez C, Plantaz D, Boni L, Michon J (2011) Excellent outcome with reduced treatment in infants with nonmetastatic and unresectable neuroblastoma without MYCN amplification: results of the prospective INES 99.1. J Clin Oncol 29(4):449–455
Seeger RC, Brodeur GM, Sather H, Dalton A, Siegel SE, Wong KY, Hammond D (1985) Association of multiple copies of the N-myc oncogene with rapid progression of neuroblastomas. N Engl J Med 313:1111–1116
Shimada H, Ambros IM, Dehner LP, Hata J, Joshi VV, Roald B (1999) Terminology and morphologic criteria of neuroblastic tumors: recommendations by the international neuroblastoma pathology committee. Cancer 86(2):349–363
Spitz R, Hero B, Simon T, Berthold F (2006) Loss in chromosome 11q identifies tumors with increased risk for metastatic relapses in localized and 4S neuroblastoma. Clin Cancer Res 12(11):3368–3373
Steen S, Nemunaitis J, Fisher T, Kuhn J (2008) Circulating tumor cells in melanoma: a review of the literature and description of a novel technique. Bayl Univ Med Cent 21(2):127–132
Swartz MA, Kristensen CA, Melder RJ, Roberge S, Calautti E, Fukumura D et al (1999) Cells shed from tumours show reduced clonogenicity, resistance to apoptosis, and in vivo tumorigenicity. Br J Cancer 81:756–759
Träger C, Vernby A, Kullman A, Ora I, Kogner P, Kågedal B (2008) mRNAs of tyrosine hydroxylase and dopa decarboxylase but not of GD2 synthase are specific for neuroblastoma minimal disease and predicts outcome for children with high-risk disease when measured at diagnosis. Int J Cancer 123:2849–2855
Viprey VF, Lastowska MA, Corrias MV, Swerts K, Jackson MS, Burchill SA (2008) Minimal disease monitoring by QRT-PCR: guidelines for identification and systematic validation of molecular markers prior to evaluation in prospective clinical trials. J Pathol 216:245–252
Acknowledgments
We thank Désirée Ramal for her help in the data management and all the Spanish collaborating hospitals who register their patients. This work was supported in part by: FIS PS09/02323, FIS PS02/0315 and Cancercare Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yáñez, Y., Grau, E., Oltra, S. et al. Minimal disease detection in peripheral blood and bone marrow from patients with non-metastatic neuroblastoma. J Cancer Res Clin Oncol 137, 1263–1272 (2011). https://doi.org/10.1007/s00432-011-0997-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-011-0997-x