Abstract
The coupling of anatomical and functional connectivity at rest suggests that anatomy is essential for wake-typical activity patterns. Here, we study the development of this coupling from wakefulness to deep sleep. Globally, similarity between whole-brain anatomical and functional connectivity networks increased during deep sleep. Regionally, we found differential coupling: during sleep, functional connectivity of primary cortices resembled more the underlying anatomical connectivity, while we observed the opposite in associative cortices. Increased anatomical–functional similarity in sensory areas is consistent with their stereotypical, cross-modal response to the environment during sleep. In distinction, looser coupling—relative to wakeful rest—in higher order integrative cortices suggests that sleep actively disrupts default patterns of functional connectivity in regions essential for the conscious access of information and that anatomical connectivity acts as an anchor for the restoration of their functionality upon awakening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The brain guarantees efficient and flexible behavior by a dynamic functional interplay of neurons fixed in an anatomical network structure. The interaction between structure and function successfully responds to immediate environmental demands as well as chronic challenges including physical damage (Park and Friston 2013). A functional reorganization occurs during the steps from awake conscious awareness to reduced consciousness during deep sleep—including its spontaneous reversal. What are some principles of such adaptive behavior?
Recently, we introduced the hypothesis that the anatomical connectivity of the brain influences functional patterns of activity from wakefulness to sleep (Tagliazucchi et al. 2013b), precluding a more complete breakdown of connectivity during states of deep unconsciousness (Boly et al. 2008). Indeed, it is well established that functional connectivity during wakeful rest can be predicted by the underlying anatomical connections (Hagmann et al. 2008; Greicius et al. 2009). Simple dynamical models unfolding on realistic anatomical connectivity can produce connectivity patterns with high resemblance to empirical observations (Honey et al. 2009; Haimovici et al. 2013; Deco et al. 2014; Stam et al. 2015). Modifications of this anatomical–functional interplay from wakefulness to deep sleep remain to be demonstrated empirically, as well as its relationship with the loss and recovery of conscious awareness we experience daily.
It is becoming increasingly apparent that the difference between anatomical and functional connectivity is crucially important to understand the context-sensitive and dynamic aspect of self-organised neuronal fluctuations. We used sleep to study state-dependent divergence between anatomical and functional connectivity. In other words, we measured the difference between anatomical and functional connectivity as a function of different brain states (depth of sleep as measured electrophysiologically). We hoped to see a global loss (sparsification) of functional connectivity during sleep converging towards the sparse connectivity embodied by anatomical connections. This would be consistent with a loss of (de-synchronised) complex dynamics associated with conscious wakefulness.
For the assessment of this hypothesis in human subjects, we took advantage of the combination of three in vivo non-invasive imaging techniques. Blood-oxygen-level dependent (BOLD) functional magnetic resonance imaging (fMRI) facilitated the study of inter-regionally correlated brain activity throughout the brain. Simultaneous electroencephalography (EEG) was used for the detection of different stages of sleep (AASM 2007). Diffusion weighted imaging (diffusion tensor imaging, DTI) allowed the identification of anatomical connections in the brain enabling the in vivo examination of structural networks. Based on this data, we studied the development of anatomical–functional similarity from wakefulness to deep sleep in 15 healthy participants exhibiting sufficient amounts of wakefulness and all NREM sleep stages. We first analyzed this similarity on a global basis; we then moved towards a regional characterization aiming to evaluate system-specific behavior.
Materials and methods
DTI connectivity network
To construct the DTI network with 401 nodes, we obtained diffusion tensor imaging data from 56 healthy subjects (mean age 32 years, 18 female) and performed whole brain deterministic tractography. First, we divided cortical and sub-cortical grey matter into 401 even-sized regions of interest following the method presented in Zalesky et al. (2010), consisting of the homogenous subdivision of regions in the AAL template (Tzourio-Mazoyer et al. 2002), thus resulting in a less coarser parcellation that still respects anatomical landmarks. Afterwards, we constructed a network by creating a link between two regions in this parcellation if the number of streamlines between them was significantly greater than zero across the group of subjects (p < 0.01, rank sum test, FDR-corrected). Further details on DTI acquisition, preprocessing and network construction can be found in Crossley et al. (2014).
Participants and sleep statistics
71 healthy and non-sleep deprived participants (written informed consent, approval by the local ethics committee, participants not suffering from psychiatric or neurological conditions, or sleep disturbances) took part in the study. Afterwards a sub-sample of 15 subjects was selected based on the presence of at least 100 fMRI volumes (208 s) of wakefulness and each sleep stage, thus allowing for paired comparisons. The total time spent in each sleep stage within this subset was as follows (mean ± SD): Wake = 11.8 ± 6.2 min, N1 sleep = 8.5 ± 2.6 min, N2 sleep = 14.5 ± 5.9 min, N3 sleep = 17 ± 8.3 min. The mean sleep epoch durations were as follows: Wake = 3.9 ± 2.5 min, N1 sleep = 2.8 ± 2.7 min, N2 sleep = 4.3 ± 2.3 min, N3 sleep = 10.4 ± 7.5 min. For each sleep stage, fMRI data were concatenated along all sleep epochs for the functional connectivity analyses.
Within this sub-sample, 9 female and 6 male participants were included (23 ± 3 years, all right handed, not taking medications including stimulants and sedatives). The day of the study all participants reported a waking time between 5:00 AM and 11:00 AM (median: 8:00 AM). The day before the study, participants reported a sleeping time between 10:00 PM and 2:00 AM (median: 11:30 PM). These values remained similar throughout the 6 days prior to the experiment, as reported by the participants.
EEG-fMRI acquisition
Scanning sessions took place during the evening (the experimental sessions started at 7:00 PM, functional MRI scanning started between 8:00 PM and 9:00 PM). We instructed subjects to close their eyes and lie still and relaxed. EEG via a cap (modified BrainCapMR, Easycap, Herrsching, Germany) was recorded during fMRI acquisition (1505 volumes of T2*-weighted echo planar images, TR/TE = 2080/30 ms, matrix 64 × 64, voxel size 3 × 3 × 2 mm3, distance factor 50 %; FOV 192 mm2) at 3 T (Siemens Trio, Erlangen, Germany) with an optimized polysomnographic setting [chin and tibial EMG, ECG, EOG recorded bipolarly (sampling rate 5 kHz, low pass filter 1 kHz), 30 EEG channels recorded with FCz as the reference (sampling rate 5 kHz, low pass filter 250 Hz), and pulse oxymetry, respiration recorded via sensors from the Trio (sampling rate 50 Hz)] and MR scanner compatible devices (BrainAmp MR+, BrainAmp ExG; Brain Products, Gilching, Germany). In order to reduce EEG artifacts due to vibrations, the scanner helium pump was turned off during the experiments.
MRI and pulse artifact correction were performed based on the average artifact subtraction (AAS) method (Allen et al. 1998) as implemented in Vision Analyzer2 (Brain Products, Germany) followed by objective (CBC parameters, Vision Analyzer) ICA-based rejection of residual artifact-laden components after AAS resulting in EEG with a sampling rate of 250 Hz. EEG was sleep staged by an expert following AASM criteria (AASM 2007).
fMRI data pre-processing
Using Statistical Parametric Mapping (SPM8, www.fil.ion.ucl.ac.uk/spm) EPI data were realigned, normalized (MNI space) and spatially smoothed (Gaussian kernel, 8 mm3 full width at half maximum). Data were re-sampled to 4 × 4 × 4 mm resolution to facilitate further processing. Cardiac, respiratory [both estimated using the RETROICOR method, (Glover et al. 2000)] and head motion realignment time series (3 head displacement parameters, 3 head rotation parameters and their first three derivatives, resulting in a total of 24 motion realignment regressors) were partialled out via linear regression. Finally, data were band-pass filtered in the range 0.01–0.1 Hz (Cordes et al. 2001) using a sixth-order Butterworth filter. The frame-wise displacement (Power et al. 2012) and the variance explained by the RETROICOR regressors were not found to be significantly different between wakefulness and NREM sleep stages (see the supplementary information for details).
Functional network construction
We constructed functional networks by extracting average BOLD signals from regions of interest and computing the linear correlation values among all pairs of signals. This procedure results in the correlation matrix \( C_{ij} \), given by,
where \( x_{i,j} \) are BOLD signals at ROIs i and j, \( \sigma_{i,j} \) are the standard deviations of the signals and <, > is temporal averaging. We defined our functional adjacency matrix based on positive correlations; since we did not regress the global fMRI signal, we precluded strong negative functional connectivity (Murphy et al. 2009). In this work, the ROIs were those used in the parcellation leading to the construction of the DTI network.
For comparisons with anatomical connectivity networks, the correlation matrices \( C_{ij} \) were thresholded to yield binary adjacency matrices \( A_{ij} \) such that \( A_{ij} = 1 \) if \( C_{ij} > \rho \) and \( A_{ij} = 0 \) otherwise. The parameter \( \rho \) was chosen to fix the ratio of present connections in the network (given by \( \mathop \sum \nolimits_{i > j} A_{ij} \)) to the total possible number of connections. This ratio is termed link density. It is important to fix the link density when comparing two networks (or in this case, their similarity with a third network, i.e. the anatomical connectivity network) since otherwise differences could arise because the means of the respective \( C_{ij} \) are different (and therefore the number of non-zero entries in \( A_{ij} \)) and not because connections are re-organized.
Since the link density is a free parameter, we repeated the analyses for a range of link densities (between 0.01 and 0.2 in steps of 0.01 for the global network similarity between anatomy and function). When comparing functional networks to their anatomical counterparts, the chosen link density ranges always included the link density of the DTI network.
Similarity measures between functional and anatomical connectivity
We applied two different metrics to measure the similarity between functional and anatomical networks. For binary adjacency matrices we computed the linear correlation (\( R \)) between them. \( R > 0 \) is obtained if the matrices are correlated (i.e. the presence of a link in one network predicts the presence of the same link in the other network and vice versa). \( R < 0 \) is obtained if the matrices are anti-correlated (i.e. the presence of a link in one network predicts the absence of the same link in the other network and vice versa). For binary vectors, the linear correlation is formally equivalent to the Phi coefficient \( \phi \), which is understood in terms of a contingency table. For two binary signals \( x_{i} \) and \( y_{i} \), there are four possible combinations: \( (x_{i} = 0,y_{i} = 0) \), \( (x_{i} = 1,y_{i} = 0) \), \( (x_{i} = 0,y_{i} = 1) \), \( (x_{i} = 1,y_{i} = 1) \). If \( N_{00} \), \( N_{10} \), \( N_{01} \), \( N_{11} \) denote the number of times each of these four possibilities occur then,
The second similarity metric between functional and anatomical networks was based on comparing the modular decomposition of networks. Intuitively, a module is a group of nodes having denser within-group connections compared to the connections with other modules. A possible way to obtain the modular structure of a network is to maximize the modularity \( Q \), which estimates the difference between the number of intra-module links and the expected number (for the same partition) in a random network of the same size (Newman and Girvan 2004). Given a partition of the nodes into subsets, \( Q \) is defined as,
In this equation the sum comprises all nodes in the network, \( L \) equals the total number of links (\( \mathop \sum \nolimits_{i > j} A_{ij} \)), \( k_{i} = \mathop \sum \nolimits_{i} A_{ij} \) is the degree (number of links attached to each node) and \( \delta \left( {i,j} \right) = 1 \) if nodes \( i \) and \( j \) belong to the same subset of the partition, and 0 otherwise. We applied a modularity maximization algorithm by Newman (2006) based on the eigenspectrum of the modularity matrix, as implemented in the Brain Connectivity Toolbox (Rubinov and Sporns 2010).
Once we identified the modular structure of anatomical and functional networks, we compared them using the corrected-for-chance Rand Index (Rand 1971), a measure of modular decomposition (or clustering) similarity ranging between 0 (no agreement in the modular structure) and 1 (complete agreement).
Similarity between functional and anatomical connectivity neighborhoods
For a given link density we define the connectivity neighborhood of node \( i \) as \( n_{j} = A_{ij} \). According to this definition, the jth entry of \( n_{j} \) is 1 if nodes i and j share a direct connection in the network and is zero otherwise. This definition is motivated by the concept of a “connectivity fingerprint” associated with each brain area, introduced by Passingham and colleagues (Passingham et al. 2002).
We obtained the connectivity neighborhood for each node in the anatomical network as well as in the individual functional connectivity networks across all conditions, and did this for a range of local link densities of the connectivity of each node with the network. To estimate the similarity between anatomical and functional connectivity neighborhoods we computed the Hamming distance between the respective binary vectors \( n_{i} \), normalized by their length. The Hamming distance is defined as the number of substitution of symbols (in this case 0 or 1) needed to transform one sequence into another and vice versa. In this case, the Hamming distance equals twice the number of connections that must be re-wired to transform one connectivity neighborhood into the other (since re-wiring a link can be achieved by two symbol substitutions). We demonstrate a schematic of this procedure in Fig. S5. Thus, high Hamming distances imply a stronger departure from the underlying anatomical connectivity.
Statistical testing
We performed paired t test comparisons between wakefulness and all NREM sleep stages. In the analysis of regional function-anatomy differences results were reported at p < 0.05 FDR, corrected for multiple comparisons separately for each contrast (Storey 2002).
Visualization
3D renderings of network nodes and surface data were performed using the BrainNet software (Xia et al. 2013).
Results
Throughout the following sub-sections we present results derived from metrics of global and local similarity of functional connectivity with the underlying anatomical network. We first establish the preservation of global anatomical–functional similarity during light sleep [N1 sleep (AASM 2007)] and the increase of global anatomical–functional similarity in deeper sleep stages (N2, N3) compared to wakefulness. We then move from whole-brain networks to a finer (local) characterization of the regions involved and demonstrate differential regional behavior.
Similarity between global functional and anatomical connectivity networks
The correlation matrices between the BOLD signals from all 401 regions of interest are shown in Fig. 1 for wakefulness and all sleep stages. The binary adjacency matrix of the anatomical connectivity network is shown below the diagonal of the wakefulness correlation matrix. These matrices display in the intersection of the ith row and jth column whether a connection between the ith and jth nodes exists (adjacency matrix) or the functional connectivity between the associated BOLD time series (correlation matrix). The significant functional connectivity differences between wakefulness and sleep (and whether they occur in the presence or absence of anatomical connectivity) are presented below the diagonal of the remaining matrices in Fig. 1.
Functional connectivity changes during sleep in relation to anatomical connectivity. Average correlation (R) matrices between the BOLD signal time series of all 401 regions of interest. Results are presented for wakefulness and all sleep stages. The adjacency matrix of the anatomical connectivity network is shown below the diagonal of the wakefulness correlation matrix. Significant differences between sleep and wakefulness are shown below the diagonal of the other matrices (p < 0.05, paired t test, false discovery rate (FDR) controlled for multiple comparisons) and color-coded depending on the presence of underlying anatomical connectivity
Based on this information, we first studied the similarity between whole-brain anatomical and functional connectivity networks across wakefulness and all NREM sleep stages. Two (potentially simultaneous) scenarios will increase global similarity between functional connectivity and the underlying anatomical backbone: firstly, a decrease of functional connectivity between regions not directly connected anatomically and secondly an increase in functional connectivity between regions sharing a direct anatomical link, as illustrated in Fig. 2a. We tested for this effect by calculating the probability of finding decreased functional connectivity during sleep conditional to the lack of anatomical connectivity; P(decrease|unconnected) = 0.023 and 0.154 for N2 and N3 sleep, respectively. This was indeed 28 and 18 % (N2 and N3 sleep, respectively) greater than that of finding decreased functional connectivity conditional to the presence of an anatomical connection; P(decrease|connected) = 0.016 and 0.125 for N2 and N3 sleep, respectively; see Fig. 2a. In addition to this effect of pruning the functional network towards anatomical links, the probability of sleep increasing functional connectivity given anatomical links was P(increase|connected) = 0.0108 and 0.0078 (for N2 and N3 sleep), 118 and 192 % higher than that of finding connectivity increases in the absence of anatomical links, P(increase|unconnected) = 0.0052 and 0.0012 for N2 and N3 sleep; see Fig. 2a.
Deep sleep increases the similarity between global functional and anatomical connectivity networks. a Illustration of how the selective breakdown of functional connections, which are not associated with structural links, and the strengthening of those, which are associated with such leads to increased similarity of anatomical and functional connectivity networks during deep sleep. b Probability of finding functional connectivity (FC) decreases and increases (sleep vs. wakefulness) given the presence or absence of anatomical connectivity. c Similarity (Pearson’s correlation coefficient between binary adjacency matrices) between functional and anatomical connectivity matrices as a function of link density, plotted for N1, N2 and N3 sleep. Error bars are ±SEM. We observed significant increases only for N2 and N3 versus wakefulness
To further quantify these observations we introduced a metric of network similarity given by the correlation coefficient between binarized functional and anatomical adjacency matrices and analyzed link densities of the functional connectivity network ranging from 0.01 to 0.2 in steps of 0.01. This range included the (fixed) density of the anatomical network. Results are plotted in Fig. 2c. Consistent with our previous observations, we observed an increase in global functional–anatomical connectivity coupling relative to wakefulness only in the two deepest sleep stages (N2 and N3) but not in N1 sleep. This result held for all link densities under consideration.
Similarity between the modular structure of functional and anatomical networks
Both anatomical and functional networks are modular, with groups of nodes having dense connections between them and sparser connections with other, similarly well-connected groups of nodes. We applied a module detection algorithm to functional and anatomical networks and studied the similarity of the resulting modular structure across wakefulness and all NREM sleep stages. To compare the overlap between different partitions of a network into modules we employed the corrected-for-chance Rand Index, which reaches the maximum value of 1 for equal partitions.
Figure 3a shows the modular decompositions of average functional connectivity networks during wakefulness and all sleep stages. In this visualization, the adjacency matrices were thresholded to match the link density of the anatomical connectivity network.
The modular organization of functional connectivity networks is reshaped towards that of anatomical connectivity networks during deep sleep. a The modular structure of average functional connectivity matrices obtained from wakefulness, N1, N2 and N3 sleep. Different colors of nodes and links code for module membership (inter-modular links are shown in grey). b Similarity between the modular decomposition of functional and anatomical connectivity networks (quantified by the corrected-for-chance Rand Index) as a function of link density. Results are plotted separately for wake vs. N1, N2 and N3 sleep. Error bars are ±SEM
The Rand index between the modular structure of functional and anatomical networks is shown in Fig. 3b as a function of link density for wakefulness vs. all NREM sleep stages. We found an increased Rand Index in N2 and N3 sleep (i.e. better structure–function match) compared to wakefulness; we observed this for almost all the explored link densities. Differences were present in N1 sleep vs. wakefulness, too, also in the form of increased similarity between functional and anatomical connectivity networks but with a lower likelihood and for fewer link density values.
Similarity between local functional and anatomical connectivity
We can obtain a more detailed picture of the brain regions associated with increased or decreased function-anatomy similarity by studying the local connectivity of single nodes in the network. For a given link density within the neighborhood, the connectivity neighborhood of a node is defined as a binary vector representing those nodes directly connected with it. This local connectivity description of a single node can be obtained both in terms of anatomical and functional connections. Then, the similarity between the functional and anatomical connectivity neighborhoods can be measured by computing the Hamming distance between both binary vectors: the larger the distance, the less similar the functional neighborhood to the anatomical neighborhood. The Hamming distance is the number of symbol substitutions required to transform one vector into the other. In this case, it equals twice the number of connections that must be re-wired to transform one connectivity neighborhood into another (we show a schematic of the procedure followed to compute the Hamming distance in Fig. S5).
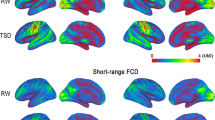
We present the results of this analysis in Fig. 4 for wakefulness vs. N2 sleep and in Fig. 5 for wakefulness vs. N3 sleep. Statistical comparisons (at a level of p < 0.05, paired t test, FDR-controlled for multiple comparisons) did not reveal any significant difference for wakefulness vs. N1 sleep, but widespread changes appeared in comparison to N2 sleep (Fig. 4) and N3 sleep (Fig. 5). To visualize the anatomical areas where these differences were located, we mapped the nodes to the brain parcellation serving as the basis for the anatomical connectivity network. Figures 4a and 5a report increases in the anatomical–functional distances, Figs. 4b and 5b decreases (for any link density in both cases). Panel D of Figs. 4 and 5 shows a summary of AAL regions (Tzourio-Mazoyer et al. 2002) where changes were observed, ranked by percentage of nodes within each region presenting statistically significant differences. We found decreases in the anatomical–functional Hamming distance, i.e. increased similarity in nodes comprising the sensorimotor, visual and auditory networks. Interestingly, we also revealed a set of regions with increased Hamming distance (i.e. decreased similarity); these regions did not reside inside specific RSN but comprised mostly midline, inferior temporal, frontal and parietal areas. However, they overlapped with the default mode and executive control networks in N3 sleep. The patterns of nodal average Euclidean distance to all other nodes in the network (Fig. S2) do not resemble the results in Figs. 4 and 5, suggesting that the region-specific distribution of nodal distances does not drive these results.
Differential coupling between functional and DTI connectivity networks in sensorimotor regions and frontal, midline, parietal and associative cortices during N2 sleep vs. wakefulness. a Rendering of the nodes for which a significant (p < 0.05, paired t test, FDR corrected for multiple comparisons) increase (red) in the Hamming distance between anatomical and functional connectivity neighborhoods was found. The outlines of five canonical RSN from Beckmann et al. (2005) are superimposed onto the renderings. b The same information as in panel B but for decreases (blue) in the Hamming distance between anatomical and functional connectivity neighborhoods. c AAL nodes (see Table S1 and Fig. S10 for region locations) ranked for the percentage of the nodes they contain with a Hamming distance above the threshold of statistical significance. Vertical bars are color-coded for the RSN membership (DMN and executive control for wake <sleep and sensorimotor RSN for wake >sleep; in both cases grey for none) of the corresponding AAL region (see Table S1 for abbreviations), as determined by the position of its center of mass
Differential coupling between functional and DTI connectivity networks in sensorimotor regions and frontal, midline, parietal and associative cortices during N3 sleep vs. wakefulness. a Rendering of the nodes for which a significant (p < 0.05, paired t test, FDR corrected for multiple comparisons) increase (red) in the Hamming distance between anatomical and functional connectivity neighborhoods was found. The outlines of five canonical RSN from Beckmann et al. (2005) are superimposed onto the renderings. b The same information as in panel B but for decreases (blue) in the Hamming distance between anatomical and functional connectivity neighborhoods. c AAL nodes ranked for the percentage of the nodes they contain with a Hamming distance above the threshold of statistical significance (same as in Fig. 4c)
Result robustness against subcortical influence and motion-induced variance
Extensive changes in subcortical neuromodulatory drive to cortical sites might induce artifactual changes in functional connectivity during sleep (Lee and Dan 2012). We excluded the possibility that an interaction between sleep stage and such subcortically modulated connectivity induced shifts in anatomical–functional coupling (Fig. S3). For this, we reproduced our core results computing functional connectivity via partial correlations including left and right thalamus (from the AAL template) and brainstem [MNI coordinates (0, −22, −34)] time series as partial regressors.
In addition, we assessed whether our main results exhibited correlations with individual frame-wise displacement (FD) values, as well as with the variance explained by the RETROICOR regressors. We present the results in Figs. S6 and S7 of the supplementary information. Not any significant correlations between FD values and anatomical–functional coupling (for all link densities) and anatomical–functional neighborhood similarities were present. For the variance explained by the RETROICOR regressors, few and isolated nodes presented correlations with their anatomical–functional neighborhood similarities; however, these were not widespread enough to explain the patterns presented in Figs. 4 and 5 and comprised different anatomical nodes.
Finally, the main results were reproduced after erasing volumes associated with relatively large (>0.4 mm) framewise displacement values (Power et al. 2012). Results are shown in Fig. S4. There were not any significant differences between wakefulness and all NREM sleep stages in terms of the number of erased volumes (see supplementary methods section).
Discussion
We investigated the relationship between the brain’s anatomical connectivity and functional connectivity in the context of different degrees of conscious awareness. For this purpose we adopted deep NREM sleep as a model of a reversible and physiological unconscious state and examined the similarity between functional and anatomical connectivity networks across all NREM sleep stages vs. wakefulness. Globally, we found a correspondence between anatomical and functional connectivity during wakefulness, which on average increased with deepening sleep. Supporting the connection of our results to diminished conscious awareness, differences were predominantly observed during deep (N2 and N3) sleep, but not during light (N1) sleep, the latter being characterized by richer conscious experiences (see Tagliazucchi et al. 2013a for a discussion). These results are in line with a recent study in monkeys also showing that structure–function similarity was higher in the state of reduced conscious awareness (anesthesia) than during wakeful rest (Barttfeld et al. 2015). Our study reaches further in that we identify differential regional behavior: compared to wakefulness, primary auditory, sensory, motor and visual cortices showed an increased functional–structural coupling during the sleep stages N2 and N3 while deeper midline and frontal-parietal regions exhibited a dissociation of function from structure.
Our observations link longstanding views on development and consciousness (Herrick 1893) with modern concepts of structure–function interaction (Park and Friston 2013) and their relationship with brain dynamics (Chialvo 2010; Haimovici et al. 2013).
Developmental considerations on structure and function
The human brain distinguishes itself from that of close relatives like macaques or chimpanzees in its unique size and in its costly brain wiring (Bullmore and Sporns 2012; Hofman 2014; Rilling 2014). Another unique characteristic is its degree of gyrification, with high levels of connectivity suggested as one driving force of increased cortical folding (Van Essen 1997; Zilles et al. 2013). The regionally differential behavior of the structural–functional interplay we observed in primary cortices versus higher cerebral associative cortices parallels development. Phylogenetically, the associative parietal cortex has developed later than the primary motor, sensory and prefrontal cortices, and is estimated to have expanded by an order of magnitude in the human lineage compared to the macaque. In contrast, early sensory areas expanded far less. Ontogenetically, fiber tracts in the regions of high evolutionary expansion continue to grow into the third decade of life, and this postnatal human maturation of connectivity may further cortical folding (Kaas et al. 2013; Van Essen 2013; Zilles et al. 2013). In humans and the great apes, the early sensory and motor areas are heavily myelinated, in contrast to lateral temporal, parietal, and prefrontal regions (Van Essen 2013). Hence, primary cortices are phylogenetically old, have fast conducting fiber tracts and conclude their maturation prenatally. Such features reflect the evolution of a long-preserved efficient system with a finite number of degrees of freedom. At the same time, the mentioned properties contrast those of brain regions supporting cognitive functioning with a much less constrained functional repertoire which cannot be hard-wired (Bullmore and Sporns 2012).
We found that functional connectivity in sensory(-motor) cortices during wakefulness is less bound to anatomy than during sleep, which might reflect the integration of primary sensory input and the planning of motor output during wakefulness. This could augment the range of the sensory-motor operations beyond those already mapped out anatomically, thus driving functional connectivity away from the basic structural connections per se providing mere essential functionality. Vice versa, with ceasing conscious awareness, higher order processing is reduced including frontal-parietal feedback to primary cortices (Boly et al. 2011), and functional activity falls back towards the anatomical web like we observed during sleep. Although hardwiring also exists for the associative cortices (Greicius et al. 2009), it is phylogenetically and ontogenetically more dynamic than that of the sensory-motor systems. This diversity might allow for human individuality and the variety of subjective conscious experiences shaping further throughout one’s lifetime, ranging from deep unconsciousness to their expansion in the psychedelic state (Tagliazucchi et al. 2014). Still, only a functional structure superseding, i.e. departing from, the underlying anatomy guarantees conscious processing (Tagliazucchi et al. 2013a). However, why then, did we observe a further deviation of functional from structural connectivity in associative cortices during deep sleep compared to wakefulness?
Conscious by default?
Let us assume that the high similarity of the functional to the structural connectivity represents a rather basic, or “default” state of a system’s activity as we argued in the context of the primary sensory-motor regions. Accordingly, the functional connectivity of the associative midline, inferior temporal, frontal and parietal cortices being closer to structural connectivity during wakefulness than during sleep would suggest that for these regions the default state is wakefulness, i.e. conscious awareness. This in turn would imply reduced conscious awareness as an actively induced condition—even during sleep—during which the system is driven beyond the connectivity repertoire inherent to anatomy. We previously demonstrated activity in a most similar set of regions mediating sleep protection versus arousal during K-complexes (KC), an electrophysiological hallmark of deeper sleep (Jahnke et al. 2012). Connectivity analysis revealed primary sensory cortex as the first region influenced during KC and that midline regions activated in association with the sleep protecting part of the KC. We concluded that KC-associated activations in sensory areas suggest the existence of low level information processing in sleep during KC interacting with the anterior insula reflecting periodical monitoring of the environment with basic information processing facilitating either an arousal (to environmental threat) or sleep protection (Jahnke et al. 2012). This would be one example of how an active process originating in the primary sensory cortices drives activity in higher order brain regions during sleep.
The concept of an active process inhibiting conscious awareness could explain how consciousness can be regained spontaneously: suspension of this active process would yield a fallback of connectivity towards the underlying anatomical network, i.e. a state of connectivity closer to that observed during wakefulness. In the context of disease, if the condition having led to the reduction in consciousness ceases to exist, the natural state of wakeful awareness can be regained unless co-existing brain injury induced structural damage in the anatomical network subserving consciousness to a degree impossible to compensate for.
Brain dynamics and the dissociation of function from structure
Our results fit with modeling studies suggesting that during NREM sleep different cortical areas adopt different dynamical regimes: In a system below its critical point perturbations die down quickly, while above the critical point they expand in an uncontrolled fashion. Only at or near criticality, a controlled and efficient one to one propagation occurs, which is also when structural and functional connectivity match best (Deco et al. 2014) and the reactivity or susceptibility to perturbations is maximized. Using realistic anatomical connectivity (Hagmann et al. 2008), Haimovici and colleagues (Haimovici et al. 2013) modelled brain activity and found that near the system’s critical point functional connectivity achieved maximal concordance with empirical data, arising in sets of brain regions described as “resting state” (or intrinsic connectivity) networks (Beckmann et al. 2005). Accordingly, high structure–function similarity in sensory networks during deep sleep suggests operation of the sensory networks closer to the critical point. In contrast, during wakefulness, lower structure–function correlations are consistent with a subcritical state of the sensory networks when input load is high, such that self-dampening activity could reduce the risk of over-excitation. Activity in associative midline, frontal and parietal regions shows an inverse behavior, i.e. high structure–function coupling during conscious wakefulness suggestive of optimal exploration of the underlying anatomical connectivity near the critical point (Deco and Kringelbach 2014; Deco et al. 2014; Stam et al. 2015) and departure from criticality during deep sleep when consciousness is strongly reduced. Since cortical activity is non-vanishing during sleep—indeed, increased activity levels have been reported (Gabbott and Rolls 2013)—this departure could be towards a super-critical state when system excitation is not self-limited. Future modeling efforts should study the link between different states of consciousness and the dynamical regime of cortical activity (critical and sub/supercritical).
Relating functional changes to individual anatomy
Across scales and species past efforts mainly focused either on network function (Kandel et al. 2013) or structure (DeFelipe 2010). However, cognition and behavior result from dynamic neuronal interactions on anatomically fixed connectivity (Park and Friston 2013). The repertoire of functional networks is interlinked with the structural architecture of connections but with a dynamic divergence (Honey et al. 2009; Skudlarski et al. 2008). Our results extend these observations by revealing system-specific regional variations of the structure–function relationship and call for a reconsideration of the standard methodology of resting state analyses: an assessment of the re-organization of functional connectivity relative to the backbone of anatomical connections.
Just like the structure–function interaction gives rise to cognition and behavior, an alteration in the liaison can yield pathology. Given our data and the empirical and modelling studies discussed above (Barttfeld et al. 2015; Haimovici et al. 2013; Honey et al. 2009; Park and Friston 2013; Skudlarski et al. 2008) it is conceivable that a system at rest behaves functionally normal, but when perturbed reacts in a pathological manner if structural connectivity is not normal. It follows that in-born, degenerative or traumatic brain injury immediately affecting structure at the same time will affect function. A crucial step is hence to understand when and how the structure-dependent function compensates for the structural pathology (plasticity) and when it does not. Such knowledge will promote the development of beneficial behavioral, pharmacological, neuromodulatory or even surgical interventions.
Limitations
Diffusion imaging may be prone to inaccuracies when estimating long-range connections (Reveley et al. 2015), in particular, interhemispheric connections between homologous brain regions (Messé et al. 2014). However, we first observed (Fig. S8) that homotopic functional connectivity increased (relative to wakefulness) in primary sensory areas (N1, N2 and N3 sleep) and decreased in higher-level associative cortices (N3 sleep). This strongly suggests that underestimation of homotopic anatomical connections does not drive our results, since in this case the opposite effect is expected. Second, following Messé et al. (2014), we repeated all computations adding anatomical connections between homologous brain regions, and our results persisted (Fig. S9).
We used DTI from a different cohort of subjects to investigate the relationship between functional and anatomical connectivity. While this approach did not allow us to study the effects of variability at a single subject basis, the employed network was derived from a relatively large number of healthy participants and thus we take it as a representative average anatomical network.
Conclusion
Going from conscious awareness to deep sleep, cerebral structure–function coupling behaves reciprocally in phylogenetically distinguishable brain systems facilitating the biologically advantageous regaining of consciousness. Our identification of state- and region-dependent structural–functional interplay in the brain implies that future studies of higher brain function and pathology need to co-assess functional and anatomical network properties since otherwise we must miss the full picture of our complex brain.
References
AASM (2007) The AASM manual for the scoring of sleep and associated events- rules, terminology and technical specifications. American Academy of Sleep Medicine, Chicago
Allen PJ, Polizzi G, Krakow K, Fish DR, Lemieux L (1998) Identification of EEG events in the MR scanner: the problem of pulse artifact and a method for its subtraction. Neuroimage 8:229–239
Barttfeld P, Uhrig L, Sitt JD, Sigman M, Jarraya B, Dehaene S (2015) Signature of consciousness in the dynamics of resting-state brain activity. Proc Natl Acad Sci USA 112:887–892
Beckmann CF, DeLuca M, Devlin JT, Smith SM (2005) Investigations into resting-state connectivity using independent component analysis. Phil Trans Roy Soc B 360:1001–1013
Boly M, Phillips C, Tshibanda L, Vanhaudenhuyse A, Schabus M, Dang-Vu TT, Moonen G, Hustinx R, Maquet P, Laureys S (2008) Intrinsic brain activity in altered states of consciousness: how conscious is the default mode of brain function? Ann N Y Acad Sci 1129:119–129
Boly M, Garrido MI, Gosseries O, Bruno MA, Boveroux P, Schnakers C, Massimini M, Litvak V, Laureys S, Friston K (2011) Preserved feedforward but impaired top-down processes in the vegetative state. Science 332:858–862
Bullmore E, Sporns O (2012) The economy of brain network organization. Nat Rev Neurosci 13:336–349
Chialvo DR (2010) Emergent complex neural dynamics. Nat Phys 6:744–750
Cordes D, Haughton VM, Arfanakis K, Carew JD, Turski PA, Moritz CH, Quigley MA, Meyerand ME (2001) Frequencies contributing to functional connectivity in the cerebral cortex in “resting-state” data. AJNR Am J Neuroradiol 22:1326–1333
Crossley NA, Mechelli A, Scott J, Carletti F, Fox PT, McGuire P, Bullmore ET (2014) The hubs of the human connectome are generally implicated in the anatomy of brain disorders. Brain 137:2382–2395
Deco G, Kringelbach ML (2014) Great expectations: using whole-brain computational connectomics for understanding neuropsychiatric disorders. Neuron 84:892–905
Deco G, McIntosh AR, Shen K, Hutchison RM, Menon RS, Everling S, Hagmann P, Jirsa VK (2014) Identification of optimal structural connectivity using functional connectivity and neural modeling. J Neurosci 34:7910–7916
DeFelipe J (2010) From the connectome to the synaptome: an epic love story. Science 330:1198–1201
Gabbott PL, Rolls ET (2013) Increased neuronal firing in resting and sleep in areas of the macaque medial prefrontal cortex. Eur J Neurosci 37:1737–1746
Glover GH, Li TQ, Ress D (2000) Image-based method for retrospective correction of physiological motion effects in fMRI: RETROICOR. Magn Reson Med 44:162–167
Greicius MD, Supekar K, Menon V, Dougherty RF (2009) Resting-state functional connectivity reflects structural connectivity in the default mode network. Cereb Cortex 19:72–78
Hagmann P, Cammoun L, Gigandet X, Meuli R, Honey CJ, Wedeen VJ, Sporns O (2008) Mapping the structural core of human cerebral cortex. PLoS Biol 6:e159
Haimovici A, Tagliazucchi E, Balenzuela P, Chialvo DR (2013) Brain organization into resting state networks emerges at criticality on a model of the human connectome. Phys Rev Lett 110:178101
Herrick CL (1893) The Evolution of Consciousness and of the Cortex. Science 21:351–352
Hofman MA (2014) Evolution of the human brain: when bigger is better. Front Neuroanat 8:15
Honey CJ, Sporns O, Cammoun L, Gigandet X, Thiran JP, Meuli R, Hagmann P (2009) Predicting human resting-state functional connectivity from structural connectivity. Proc Natl Acad Sci USA 106:2035–2040
Jahnke K, von Wegner F, Morzelewski A, Borisov S, Maischein M, Steinmetz H, Laufs H (2012) To wake or not to wake? The two-sided nature of the human K-complex. Neuroimage 59(2):1631–1638. doi:10.1016/j.neuroimage.2011.09.013
Kaas JH, Gharbawie OA, Stepniewska I (2013) Cortical networks for ethologically relevant behaviors in primates. Am J Primatol 75:407–414
Kandel ER, Markram H, Matthews PM, Yuste R, Koch C (2013) Neuroscience thinks big (and collaboratively). Nat Rev Neurosci 14:659–664
Lee SH, Dan Y (2012) Neuromodulation of brain states. Neuron 76(1):209–222
Messé A, Rudrauf D, Benali H, Marrelec G (2014) Relating structure and function in the human brain: relative contributions of anatomy, stationary dynamics, and non-stationarities. PLoS Comp Biol 10(3):e1003530
Murphy K, Birn RM, Handwerker DA, Jones TB, Bandettini PA (2009) The impact of global signal regression on resting state correlations: are anti-correlated networks introduced? Neuroimage 44(3):893–905
Newman ME (2006) Modularity and community structure in networks. Proc Natl Acad Sci USA 103:8577–8582
Newman ME, Girvan M (2004) Finding and evaluating community structure in networks. Phys Rev E 69:026113
Park HJ, Friston K (2013) Structural and functional brain networks: from connections to cognition. Science 342:1238411
Passingham RE, Stephan KE, Kotter R (2002) The anatomical basis of functional localization in the cortex. Nat Rev Neurosci 3:606–616
Power JD, Barnes KA, Snyder AZ, Schlaggar BL, Petersen SE (2012) Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59(3):2142–2154
Rand WM (1971) Objective criteria for the evaluation of clustering methods. JASA 66:846–850
Reveley C, Seth AK, Pierpaoli C, Silva AC, Yu D, Saunders RC, Leopold D, Ye FQ (2015) Superficial white matter fiber systems impede detection of long-range cortical connections in diffusion MR tractography. Proc Natl Acad Sci USA 112(21):E2820–E2828
Rilling JK (2014) Comparative primate neuroimaging: insights into human brain evolution. Trends Cogn Sci 18:46–55
Rubinov M, Sporns O (2010) Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52:1059–1069
Skudlarski P, Jagannathan K, Calhoun VD, Hampson M, Skudlarska BA, Pearlson G (2008) Measuring brain connectivity: diffusion tensor imaging validates resting state temporal correlations. Neuroimage 43:554–561
Stam CJ, van Straaten ECW, Van Dellen E, Tewarie P, Gong G, Hillebrand A, Meier J, Van Mieghem P (2015) The relation between structural and functional connectivity patterns in complex brain networks. Int J Psychophysiol. doi:10.1016/j.ijpsycho.2015.02.011
Storey JD (2002) A direct approach to false discovery rates. J Roy Stat Soc B 64(3):479–498
Tagliazucchi E, Behrens M, Laufs H (2013a) Sleep neuroimaging and models of consciousness. Front Psychol 4:256
Tagliazucchi E, von Wegner F, Morzelewski A, Brodbeck V, Jahnke K, Laufs H (2013b) Breakdown of long-range temporal dependence in default mode and attention networks during deep sleep. Proc Natl Acad Sci USA 110:15419–15424
Tagliazucchi E, Carhart-Harris R, Leech R, Nutt D, Chialvo DR (2014) Enhanced repertoire of brain dynamical states during the psychedelic experience. Hum Brain Mapp 35:5442–5456
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, Mazoyer B, Joliot M (2002) Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15:273–289
Van Essen DC (1997) A tension-based theory of morphogenesis and compact wiring in the central nervous system. Nature 385:313–318
Van Essen DC (2013) Cartography and connectomes. Neuron 80:775–790
Xia M, Wang J, He Y (2013) BrainNet Viewer: a network visualization tool for human brain connectomics. PLoS One 8(7):e68910
Zalesky A, Fornito A, Harding IH, Cocchi L, Yücel M, Pantelis C, Bullmore ET (2010) Whole-brain anatomical networks: does the choice of nodes matter? Neuroimage 50:970–983
Zilles K, Palomero-Gallagher N, Amunts K (2013) Development of cortical folding during evolution and ontogeny. Trends Neurosci 36:275–284
Acknowledgments
This work was supported by the Bundesministerium für Bildung und Forschung (grant number 01 EV 0703) and the LOEWE Neuronale Koordination Forschungsschwerpunkt Frankfurt (NeFF). We are indebted to Helmuth Steinmetz and Günther Deuschl for their patronage; Astrid Morzelewski for data acquisition and sleep scoring together with Kolja Jahnke; Sandra Anti, Ralf Deichmann and Steffen Volz for extensive MRI support; Thomas Sattler for excellent IT infrastructure maintenance; and our volunteers for participation in the study. We thank an anonymous reviewer and Olaf Sporns for most constructive comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tagliazucchi, E., Crossley, N., Bullmore, E.T. et al. Deep sleep divides the cortex into opposite modes of anatomical–functional coupling. Brain Struct Funct 221, 4221–4234 (2016). https://doi.org/10.1007/s00429-015-1162-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00429-015-1162-0