Abstract
Serrated adenocarcinoma (SAC) is a recently defined subtype of colorectal carcinoma (CRC). However, in cases where an adjacent serrated adenoma is absent and the differentiation is poor, the diagnosis of SAC can be challenging. BRAF V600E mutation is a characteristic molecular change for the serrated route, but the utility of the newly described BRAF V600E-specific immunohistochemistry in the recognition of SAC is unclear. In this study, we conducted immunohistochemical determination of BRAF V600E mutation and correlated the results to BRAF mutation status and the histological features of SAC in a cohort of 147 CRC patients. There were 13 (8.8 %) BRAF-mutated CRCs confirmed by DNA sequencing. The sensitivity of immunohistochemistry in detecting BRAF V600E mutation was 100 % (13/13) and the specificity was 99.3 % (133/134). Three evaluators independently analyzed the immunohistochemical sections and the correlation between all the evaluators was perfect (κ = 1). In histologic examination, 33 (22.4 %) of the CRCs were classified as SACs. Twelve of 13 (92.3 %) BRAF-mutated CRCs were evaluated to represent serrated type growth pattern. One of 13 (7.7 %) showed poor differentiation not enabling convincing classification. In conclusion, we found immunohistochemistry to be accurate in the detection of the BRAF V600E mutation, with potential applications in the recognition of the BRAF-mutated SACs. Especially in cases where the adjacent adenoma is absent and the tumor is poorly differentiated, BRAF immunohistochemistry could be utilized as an aid to detect SACs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the Western world, colorectal cancer (CRC) is among the three most common cancer types [1]. There are at least three distinct molecular pathways leading to CRC. About 60–80 % are associated with early APC mutations and chromosomal instability [2]. Microsatellite instability (MSI) characterizes the rest of CRCs. Majority of MSI cancers are sporadic, resulting mainly from silencing of MLH1 by promoter methylation [3]. About 3 % of all CRCs are associated with Lynch syndrome, a result of germline mutation of DNA mismatch repair (MMR) [4].
Serrated adenocarcinoma (SAC) is a recently defined subtype of CRC arising via the serrated pathway and representing at least 10 % of all CRCs. Its occurrence is more frequent in women and the majority of them are located either in the cecum and ascending colon or in the rectum [5, 6]. Current diagnostic criteria for SAC are based on the recognition of the adjacent serrated polyp (hyperplastic polyp, HP; sessile serrated adenoma, SSA; or traditional serrated adenoma, TSA) next to the carcinoma or the defined histological criteria included in the World Health Organization (WHO) classification [7, 8]. However, in cases where the adjacent adenoma is absent, and the differentiation is poor, the recognition of the SAC can be challenging.
SACs differ from the conventional adenocarcinomas (CC) also on the molecular basis. In conventional adenomas and carcinomas, MSI occurs infrequently, while in serrated polyps and carcinomas, it is a common phenomenon [5, 9, 10] mostly resulting from the hypermethylation of the CpG islands in the promoter area of the MMR gene MLH1 [11, 12]. Besides MSI, oncogenic mutation in BRAF is characteristic to the serrated pathway and is present in the vast majority of the sporadic MSI CRCs [13–15].
BRAF is one of the direct downstream effectors of KRAS and an important participant of the mitogen-activated protein kinase-extracellular signal-regulated kinase (MAPK-ERK) pathway [16]. MAPK-ERK mediates the cellular response to extracellular signals regulating cell growth, differentiation, and apoptosis. In BRAF-mutated CRCs, somatic BRAF mutation is an early phenomenon in carcinogenesis detected already in aberrant crypt foci [8, 17]. The most common BRAF mutation is V600E, accounting for nearly all of the oncogenic BRAF mutations in CRC. This mutation is strongly associated with DNA methylation abnormalities and MSI [18, 19].
Recently, immunohistochemical analysis was reported being an accurate and rapid method for detecting the presence of BRAF V600E mutation in patients with metastatic melanoma [20], and its feasibility was soon established also in CRC [21–23]. Although BRAF V600E mutation has been proposed to be specific for SACs and its precursors [13, 24], the utility of BRAF V600E-specific immunohistochemistry (VE1 IHC) in the detection of SAC is unclear.
In this study, we conducted VE1 IHC on 147 CRCs and correlated the results on the BRAF mutational status. Our specific point of interest was to find out whether VE1 IHC can be utilized as an aid to detect poorly differentiated or undifferentiated SACs.
Materials and methods
Patients
All newly diagnosed CRC patients operated surgically in Oulu University Hospital between the years 2006 and 2010 were introduced in this study. One hundred forty-seven patients were both eligible for the study and had signed informed consent to participate [25]. The Ethical Committee of Oulu University Hospital accepted the study design (58/2005, 184/2009). Clinical records and a questionnaire were used in the collection of the clinical details of the patients and controls. The patients with T3 or T4 rectal tumors (n = 32) received preoperative radiotherapy or chemoradiotherapy [25].
Histological analysis
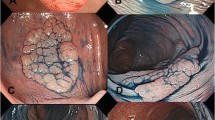
The surgical samples were fixed in 10 % formalin and embedded in paraffin. Five-micrometer sections cut from the embedded specimens were stained with hematoxylin-eosin (H & E). Histological type of the tumors, WHO grade, and TNM stage were evaluated from the H & E sections. The SACs were detected by the established criteria [7, 8, 26], including epithelial serrations, clear or eosinophilic cytoplasm, abundant cytoplasm, vesicular nuclei, distinct nucleoli, intracellular and extracellular mucin production, and absence or scarceness of necrosis (Table 1). In mucinous SACs, cell balls and papillary rods were considered to support the diagnosis of SAC (Fig. 1). The histological type of the CRC was evaluated without the knowledge of the BRAF mutation status.
Representative images of serrated adenocarcinoma histology. a BRAF V600E-mutated CRC displaying characteristic features of colorectal serrated adenocarcinoma, including epithelial serration, eosinophilic cytoplasm, and well-preserved polarity. b A traditional serrated adenoma (TSA) adjacent to the cancer
Tissue microarray construction
A tissue microarray (TMA) was utilized in the immunohistochemical analyses in this study. The H & E slides were used to mark the desired tumor locations. Depending on the size of the tumor, a total of one to four (median 3) cores of 3.0 mm diameter were manually sampled for each case yielding an overall tumor area of 7.1–28.3 mm2. One to three (median 2) of these cores were acquired from invasive front of the tumor containing the point of deepest invasion and the rest were from intratumoral locations.
DNA extraction and BRAF mutation analysis
Ten-micrometer-thick sections were cut from paraffin-embedded tumor samples. The sections were placed on a glass slide and the tumor area was scraped with a surgical blade. DNA was extracted by standard phenol chloroform extraction.
Mutation analysis of BRAF V600E was done by direct sequencing of PCR-amplified DNA. For PCR procedure, a Phusion High-Fidelity PCR kit (New England Biolabs, UK) was used and PCR reaction conditions were selected according to the kit’s guidelines using 25 ng of template DNA. The primers used in the analysis were 5′-AAACTCTTCATAATGCTTGCTCTG-3′ (forward) and 5′-GGCCAAAAATTTAATCAGTGGA-3′ (reverse) for BRAF V600E. In PCR amplification, a PTC-200 thermal cycler (MJ Research, Waltham, MA, USA) was used. PCR conditions are available on request. Five microliters of PCR products were enzymatically purified before sequencing using a 2.5-μl mixture of exonuclease I (Exo 1, Fermentas #EN0581) and shrimp alkaline phosphatase (SAP, Fermentas #EN0511) containing 10 U Exo 1 and 2 U SAP.
The DNA sequencing was performed in both directions by ABI 3130 xl Genetic Analyzer (Applied Biosystems, Foster City, CA, USA), and the sequencing data was analyzed with Chromas 1.6 sequencing analysis software (Technelysium Pty, Halensvale, Australia). Mutations were ensured by repeating the PCR and sequencing procedures.
Immunohistochemistry
Immunohistochemical staining of BRAF V600E was performed with BRAF V600E-specific monoclonal antibody (VE1; Spring Bioscience, Pleasanton, CA) at a dilution of 1:2,000 with OptiView Amplification using Ventana BenchMark XT immunostainer (Ventana Medical Systems, Tucson, AZ) [23].
Statistical analyses
The statistical analyses were carried out using statistical analysis software SPSS Statistics 20 (IBM, Chicago, IL). Cross-tabulation was used to present the associations between the categorical variables. The chi-square test was used to estimate the statistical significance. p value under 0.05 was considered statistically significant. The kappa coefficient (κ) was used to measure inter-rater agreement.
Results
Immunohistochemistry
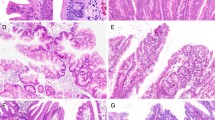
After an overview of the immunohistochemical sections, we decided to evaluate them on a two-tiered scale, 0 denoting negative and 1 defined as positive staining. The prerequisite for the positivity was diffuse distribution in the tumor cells. The sections were independently evaluated by three researchers. Immunohistochemistry using BRAF V600E mutation-specific antibody VE1 was successful in all 147 cases, and an excellent correlation existed in detecting V600E mutation by immunohistochemistry and by sequencing a corresponding area of BRAF exon 15 (Table 2). DNA sequencing revealed 13 (8.8 %) BRAF V600E-mutated cases, and VE1 antibody stained all of these cancers. All or almost all tumor cells showed diffuse and strong cytoplasmic positivity, whereas stromal cells did not show any VE1 staining (Fig. 2a, b). Other mutations of BRAF, such as BRAFV600K, were not detected in the sequencing procedure.
Representative images of BRAF immunohistochemistry. a, b Two examples of CRC with BRAF V600E mutation detected by DNA sequence analysis showing strong immunoreaction against BRAF in epithelial cells while stromal cells are negative. c CRC with no evidence of BRAF V600E mutation in DNA sequence analysis
According to the genetic analyses, 134 of the CRC cases were carrying wild-type BRAF protein. One hundred thirty-three genetically wild-type CRCs (99.3 %) were also negative in VE1 staining when only one genetically BRAF wild-type case was VE1 positive in immunohistochemistry. This particular case remained VE1 positive when the immunohistochemistry was repeated on whole sections, while the repeated sequencing showed BRAF V600E mutation once and wild-type BRAF twice. This case was histologically poorly differentiated and all tumor areas contained a high number of nontumor cells. Thus, the high number of nonneoplastic cells might have been the source of false-negative results in PCR-based sequencing. Thus, the IHC proved better in the detection of BRAF V600E mutation, and the sensitivity of sequencing in detecting BRAF V600E mutation was 100 % (13/13) and the specificity was 99.3 % (133/134). The correlation between all three evaluators was perfect (κ = 1).
Correlation between BRAF mutation and serrated morphology
In histologic examination, 33 (22.4 %) of the CRCs were classified as SAC and 114 (77.6 %) were classified as CCs. Sixteen (48.5 %) of the SACs had an adjacent serrated adenoma, and of these, ten (62.5 %) were TSAs, five (31.3 %) were SSAs, and one (6.3 %) harbored features of SSA and TSA. All adenomas contained cytological dysplasia.
Twelve of 13 (92.3 %) genetically tested BRAF-mutated CRCs were evaluated to represent serrated type growth pattern and five of them (38.5 %) had an adjacent serrated polyp. The only BRAF-mutated case classified as CC showed poor differentiation and also had some features of SAC, including eosinophilic cytoplasm, easily discernible vesicular nuclei, distinct nucleoli, and scarcity of necrosis. Twenty (14.9 %) of the genetically wild-type BRAF CRCs were classified as SACs.
Discussion
To our knowledge, this was the first study to investigate the feasibility of the VE1 IHC in detecting the BRAF V600E-mutated SACs. We found an excellent correlation between the VE1 IHC and mutation analysis; 12 of 13 BRAF-mutated CRCs were classified as SACs and 1 of 13 was poorly differentiated but had some features of SAC histology when all H & E slides were carefully examined. Around one third of SACs have been reported to carry BRAF V600E mutation, which means that VE1 was not sensitive but highly specific for detecting SAC [24, 27].
Several studies have reported of the high accuracy of VE1 mutation-specific IHC in CRC [21, 22] which is in concordance with our findings. So far, the studies have suggested that VE1 IHC may have potential clinical significance in the exclusion of Lynch syndrome from the MSI-H CRCs [28–30] based on the concept of Lynch syndrome rarely harboring the BRAF mutation [31].
In this study, all BRAF-mutated CRCs showed diffuse and strong VE1 staining and VE1 staining (sensitivity 100 %). Only one case was evaluated VE1 IHC positive in the absence of mutation on initial DNA sequencing. Thus, the specificity was 99.3 %. The discrepancy between staining and mutation analysis in the single case might have been caused by the high number of nontumor nuclei and thus enrichment of normal over mutated BRAF allele. This would lead to the nonrepeated sequencing result and conflicting interpretation of the mutation status.
So far, only one study has examined the feasibility of the novel VE1 IHC for detecting the polyps of the serrated route [32]. In that study Mesteri et al. found 141 of 194 (72.7 %) serrated lesions to show the positive, cytoplasmic staining for the BRAF V600E antibody: 100 % of SSAs/SSPs, 94 % of TSAs and 62 % of HPs while all conventional adenomas were negative for VE1. This is in concordance with the earlier reports based on mutation analyses and highlights the importance of early BRAF mutation in the serrated route of CRC [24].
Recently published studies have also indicated that the BRAF-mutated CRCs favor the proximal location, female gender, and older patient age [8]. These clinicopathological characteristics are typical of SACs supporting the concept of the specificity of BRAF mutation to SAC. Our results are convergent with these findings since ten (76.9 %) of the BRAF-mutated CRCs were female and the patients’ mean age was 71, the youngest being 52 and the oldest 87. Interestingly, the distribution of the cases differed from expected as nine (69.2 %) of the cases were localized in the distal colon, and the remaining four cases (30.8 %) were proximal.
To our knowledge, this is the first study testing the utility of VE1 IHC on the detection of SACs. Twelve of 13 BRAF-mutated CRCs were evaluated as SACs prior to the stratification with the BRAF mutation status. One of 13 BRAF-mutated CRC showed poor differentiation which made the precise classification demanding without consideration of BRAF mutation status. However, this case showed also some features of SAC, and the existence of BRAF mutation in addition to SAC features suggests the serrated origin of this case also. Taken together, our results are in concordance with earlier reports of the specificity of BRAF mutation to SACs [14, 24, 33], suggesting that BRAF immunohistochemistry can be utilized as an aid to detect poorly differentiated SACs harboring BRAF mutations.
The identification of BRAF mutation may also be important when choosing the treatment. Recent studies have linked the BRAF-mutated CRCs with the suboptimal response with anti-EGFR monoclonal antibodies (MoAbs), which has already been detected with the KRAS mutation [34–36]. This is reasonable when considering the constant activation of the MAPK pathway by either KRAS or BRAF mutations independently of the epidermal growth factor receptor (EGFR) [17, 37].
In addition to selecting patients not benefiting the anti-EGFR treatment, VE1 IHC could be used to select patients to treat with V600E-mutated BRAF inhibitors vemurafenib (PLX4032) and dabrafenib. In V600E-mutated metastatic melanoma cases, these BRAF inhibitors induce tumor regression in a high proportion of patients and vemurafenib improves overall survival as compared with standard of care chemotherapy [38]. So far, the CRC treatment experiments with BRAF inhibitors have shown disappointingly limited therapeutic potential in CRC, but in the future, the combination of BRAF inhibitors with other treatment strategies and/or PI3k/mTOR inhibition could lead to better response rates in CRC [39].
In conclusion, we proved VE1 IHC to be accurate in the detection of the BRAF V600E mutation, with potential applications in the recognition of the BRAF-mutated SACs. Especially in cases where the adjacent adenoma is absent and the tumor is poorly differentiated, VE1 IHC could be utilized as an aid to detect SACs. In the future, VE1 IHC might also help to identify patients who would benefit from anti-EGFR MoAbs and V600E-mutated BRAF inhibitor therapy.
Abbreviations
- CC:
-
Conventional adenocarcinoma
- CRC:
-
Colorectal cancer
- EGFR:
-
Epidermal growth factor receptor
- HP:
-
Hyperplastic polyp
- IHC:
-
Immunohistochemistry
- MAPK-ERK:
-
Mitogen-activated protein kinase-extracellular signal-regulated kinase
- MMR:
-
Mismatch repair
- MoAbs:
-
Monoclonal antibodies
- MSI:
-
Microsatellite instability
- SAC:
-
Serrated adenocarcinoma
- SSA:
-
Sessile serrated adenoma
- TMA:
-
Tissue microarray
- TSA:
-
Traditional serrated adenoma
References
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ (2009) Cancer statistics, 2009. CA Cancer J Clin 59:225–249
Worthley DL, Leggett BA (2010) Colorectal cancer: molecular features and clinical opportunities. Clin Biochem Rev 31:31–38
Boland CR, Goel A (2010) Microsatellite instability in colorectal cancer. Gastroenterology 138:2073–2087.e3
Aaltonen LA, Salovaara R, Kristo P, Canzian F, Hemminki A, Peltomäki P, Chadwick RB, Kääriäinen H, Eskelinen M, Järvinen H, Mecklin JP, de la Chapelle A (1998) Incidence of hereditary nonpolyposis colorectal cancer and the feasibility of molecular screening for the disease. N Engl J Med 338:1481–1487
Mäkinen MJ, George SM, Jernvall P, Mäkelä J, Vihko P, Karttunen TJ (2001) Colorectal carcinoma associated with serrated adenoma—prevalence, histological features, and prognosis. J Pathol 193:286–294
Garcia-Solano J, Perez-Guillermo M, Conesa-Zamora P, Acosta-Ortega J, Trujillo-Santos J, Cerezuela-Fuentes P, Makinen MJ (2010) Clinicopathologic study of 85 colorectal serrated adenocarcinomas: further insights into the full recognition of a new subset of colorectal carcinoma. Hum Pathol 41(10):1359–1368
Tuppurainen K, Mäkinen JM, Junttila O, Liakka A, Kyllönen AP, Tuominen H, Karttunen TJ, Mäkinen MJ (2005) Morphology and microsatellite instability in sporadic serrated and non-serrated colorectal cancer. J Pathol 207:285–294
Mäkinen MJ (2007) Colorectal serrated adenocarcinoma. Histopathology 50:131–150
Iino H, Jass JR, Simms LA, Young J, Leggett B, Ajioka Y, Watanabe H (1999) DNA microsatellite instability in hyperplastic polyps, serrated adenomas, and mixed polyps: a mild mutator pathway for colorectal cancer? J Clin Pathol 52:5–9
Jass JR, Iino H, Ruszkiewicz A, Painter D, Solomon MJ, Koorey DJ, Cohn D, Furlong KL, Walsh MD, Palazzo J, Edmonston TB, Fishel R, Young J, Leggett BA (2000) Neoplastic progression occurs through mutator pathways in hyperplastic polyposis of the colorectum. Gut 47:43–49
McGivern A, Wynter CV, Whitehall VL, Kambara T, Spring KJ, Walsh MD, Barker MA, Arnold S, Simms LA, Leggett BA, Young J, Jass JR (2004) Promoter hypermethylation frequency and BRAF mutations distinguish hereditary non-polyposis colon cancer from sporadic MSI-H colon cancer. Fam Cancer 3:101–107
Poynter JN, Siegmund KD, Weisenberger DJ, Long TI, Thibodeau SN, Lindor N, Young J, Jenkins MA, Hopper JL, Baron JA, Buchanan D, Casey G, Levine AJ, Le Marchand L, Gallinger S, Bapat B, Potter JD, Newcomb PA, Haile RW, Laird PW, Colon Cancer Family Registry Investigators (2008) Molecular characterization of MSI-H colorectal cancer by MLHI promoter methylation, immunohistochemistry, and mismatch repair germline mutation screening. Cancer Epidemiol Biomarkers Prev 17:3208–3215
Kambara T, Simms LA, Whitehall VL, Spring KJ, Wynter CV, Walsh MD, Barker MA, Arnold S, McGivern A, Matsubara N, Tanaka N, Higuchi T, Young J, Jass JR, Leggett BA (2004) BRAF mutation is associated with DNA methylation in serrated polyps and cancers of the colorectum. Gut 53:1137–1144
O’Brien MJ, Yang S, Mack C, Xu H, Huang CS, Mulcahy E, Amorosino M, Farraye FA (2006) Comparison of microsatellite instability, CpG island methylation phenotype, BRAF and KRAS status in serrated polyps and traditional adenomas indicates separate pathways to distinct colorectal carcinoma end points. Am J Surg Pathol 30:1491–1501
Patil DT, Shadrach BL, Rybicki LA, Leach BH, Pai RK (2012) Proximal colon cancers and the serrated pathway: a systematic analysis of precursor histology and BRAF mutation status. Mod Pathol 25:1423–1431
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA (2002) Mutations of the BRAF gene in human cancer. Nature 417:949–954
Schlessinger J (2002) Ligand-induced, receptor-mediated dimerization and activation of EGF receptor. Cell 110:669–672
Nagasaka T, Sasamoto H, Notohara K, Cullings HM, Takeda M, Kimura K, Kambara T, MacPhee DG, Young J, Leggett BA, Jass JR, Tanaka N, Matsubara N (2004) Colorectal cancer with mutation in BRAF, KRAS, and wild-type with respect to both oncogenes showing different patterns of DNA methylation. J Clin Oncol 22:4584–4594
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, Kang GH, Widschwendter M, Weener D, Buchanan D, Koh H, Simms L, Barker M, Leggett B, Levine J, Kim M, French AJ, Thibodeau SN, Jass J, Haile R, Laird PW (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38:787–793
Long GV, Wilmott JS, Capper D, Preusser M, Zhang YE, Thompson JF, Kefford RF, von Deimling A, Scolyer RA (2013) Immunohistochemistry is highly sensitive and specific for the detection of V600E BRAF mutation in melanoma. Am J Surg Pathol 37:61–65
Sinicrope FA, Smyrk TC, Tougeron D, Thibodeau SN, Singh S, Muranyi A, Shanmugam K, Grogan TM, Alberts SR, Shi Q (2013) Mutation-specific antibody detects mutant BRAF(V600E) protein expression in human colon carcinomas. Cancer 119:2765–2770
Affolter K, Samowitz W, Tripp S, Bronner MP (2013) BRAF V600E mutation detection by immunohistochemistry in colorectal carcinoma. Genes Chromosomes Cancer 52:748–752
Thiel A, Heinonen M, Kantonen J, Gylling A, Lahtinen L, Korhonen M, Kytölä S, Mecklin JP, Orpana A, Peltomäki P, Ristimäki A (2013) BRAF mutation in sporadic colorectal cancer and Lynch syndrome. Virchows Arch 463(5):613–621
Stefanius K, Ylitalo L, Tuomisto A, Kuivila R, Kantola T, Sirnio P, Karttunen TJ, Mäkinen MJ (2011) Frequent mutations of KRAS in addition to BRAF in colorectal serrated adenocarcinoma. Histopathology 58:679–692
Kantola T, Klintrup K, Väyrynen JP, Vornanen J, Bloigu R, Karhu T, Herzig K, Näpänkangas J, Mäkelä J, Karttunen TJ, Tuomisto A, Mäkinen MJ (2012) Stage-dependent alterations of the serum cytokine pattern in colorectal carcinoma. Br J Cancer 107:1729–1736
Hamilton SR, Bosman FT, Boffetta P, Ilyas M, Morreau H, Nakamura S, Quirke P, Riboli E, Sobin LH (2010) Carcinoma of the colon and rectum. In: Bosman FT, Carneiro F, Hruban RH, Theise ND (eds) WHO classification of tumours of the digestive system, 4th edn. IARC, Lyon, pp 134–146
Garcia-Solano J, Conesa-Zamora P, Carbonell P, Trujillo-Santos J, Torres-Moreno DD, Pagan-Gomez I, Rodriguez-Braun E, Perez-Guillermo M (2012) Colorectal serrated adenocarcinoma shows a different profile of oncogene mutations, MSI status and DNA repair protein expression compared to conventional and sporadic MSI-H carcinomas. Int J Cancer 131:1790–1799
Capper D, Voigt A, Bozukova G, Ahadova A, Kickingereder P, von Deimling A, von Knebel DM, Kloor M (2013) BRAF V600E-specific immunohistochemistry for the exclusion of Lynch syndrome in MSI-H colorectal cancer. Int J Cancer 133:1624–1630
Toon CW, Walsh MD, Chou A, Capper D, Clarkson A, Sioson L, Clarke S, Mead S, Walters RJ, Clendenning M, Rosty C, Young JP, Win AK, Hopper JL, Crook A, von Deimling A, Jenkins MA, Buchanan DD, Gill AJ (2013) BRAFV600e immunohistochemistry facilitates universal screening of colorectal cancers for Lynch syndrome. Am J Surg Pathol 37:1592–1602
Jin M, Hampel H, Zhou X, Schunemann L, Yearsley M, Frankel WL (2013) BRAF V600E mutation analysis simplifies the testing algorithm for Lynch syndrome. Am J Clin Pathol 140:177–183
Parsons MT, Buchanan DD, Thompson B, Young JP, Spurdle AB (2012) Correlation of tumour BRAF mutations and MLH1 methylation with germline mismatch repair (MMR) gene mutation status: a literature review assessing utility of tumour features for MMR variant classification. J Med Genet 49:151–157
Mesteri I, Bayer G, Meyer J, Capper D, Schoppmann SF, von Deimling A, Birner P (2014) Improved molecular classification of serrated lesions of the colon by immunohistochemical detection of BRAF V600E. Mod Pathol 27(1):135–144
Spring KJ, Zhao ZZ, Karamatic R, Walsh MD, Whitehall VL, Pike T, Simms LA, Young J, James M, Montgomery GW, Appleyard M, Hewett D, Togashi K, Jass JR, Leggett BA (2006) High prevalence of sessile serrated adenomas with BRAF mutations: a prospective study of patients undergoing colonoscopy. Gastroenterology 131:1400–1407
Benvenuti S, Sartore-Bianchi A, Di Nicolantonio F, Zanon C, Moroni M, Veronese S, Siena S, Bardelli A (2007) Oncogenic activation of the RAS/RAF signaling pathway impairs the response of metastatic colorectal cancers to anti-epidermal growth factor receptor antibody therapies. Cancer Res 67:2643–2648
Lievre A, Blons H, Laurent-Puig P (2010) Oncogenic mutations as predictive factors in colorectal cancer. Oncogene 29:3033–3043
Mao C, Liao RY, Qiu LX, Wang XW, Ding H, Chen Q (2011) BRAF V600E mutation and resistance to anti-EGFR monoclonal antibodies in patients with metastatic colorectal cancer: a meta-analysis. Mol Biol Rep 38:2219–2223
Peyssonnaux C, Eychene A (2001) The Raf/MEK/ERK pathway: new concepts of activation. Biol Cell 93:53–62
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, Jouary T, Schadendorf D, Ribas A, O’Day SJ, Sosman JA, Kirkwood JM, Eggermont AM, Dreno B, Nolop K, Li J, Nelson B, Hou J, Lee RJ, Flaherty KT, McArthur GA, BRIM-3 Study Group (2011) Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 364:2507–2516
Coffee EM, Faber AC, Roper J, Sinnamon MJ, Goel G, Keung L, Wang WV, Vecchione L, de Vriendt V, Weinstein BJ, Bronson RT, Tejpar S, Xavier RJ, Engelman JA, Martin ES, Hung KE (2013) Concomitant BRAF and PI3K/mTOR blockade is required for effective treatment of BRAF(V600E) colorectal cancer. Clin Cancer Res 19:2688–2698
Acknowledgments
The authors wish to express their gratitude to Ms Riitta Vuento for her excellent assistance in the preparation of study material. This study was supported by grants from the Academy of Finland, Emil Aaltonen Foundation, Finnish Cancer Society, Finnish Medical Foundation, Northern Finland Cancer Foundation, and Oulu University Scholarship Foundation.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sajanti, S., Sirniö, P., Väyrynen, J.P. et al. VE1 immunohistochemistry accurately detects BRAF V600E mutations in colorectal carcinoma and can be utilized in the detection of poorly differentiated colorectal serrated adenocarcinoma. Virchows Arch 464, 637–643 (2014). https://doi.org/10.1007/s00428-014-1555-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-014-1555-0